Micro Unit 3 Exam
1/95
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
96 Terms
Catabolism
Breaks down large molecules into precursor metabolites and reducing molecules needed in anabolism. Also, generates energy needed for motility and transport and polymerization.
Anabolism
Polymerizes molecules by assembling precursor metabolites from catabolism through energy from catabolism.
catabolism produces the metabolites…
Pyruvate
Glyceraldehyde-3-P
Oxaloacetate
Phosphoenolpyruvate
Fructose-6-P
Glucose-6-P
Catabolism produces energy in form of the nucleotide triphosphates…
ATP
GTP
Catabolism produces the Reducing Power (RP) electron donors…
NADH
NADPH
FADH2
Anabolism produces…
building blocks
amino acids
sugars
nucleotides
large biomolecules
carbs
proteins
lipids
nucleic acids
supramolecular systems
Basic cell metabolism
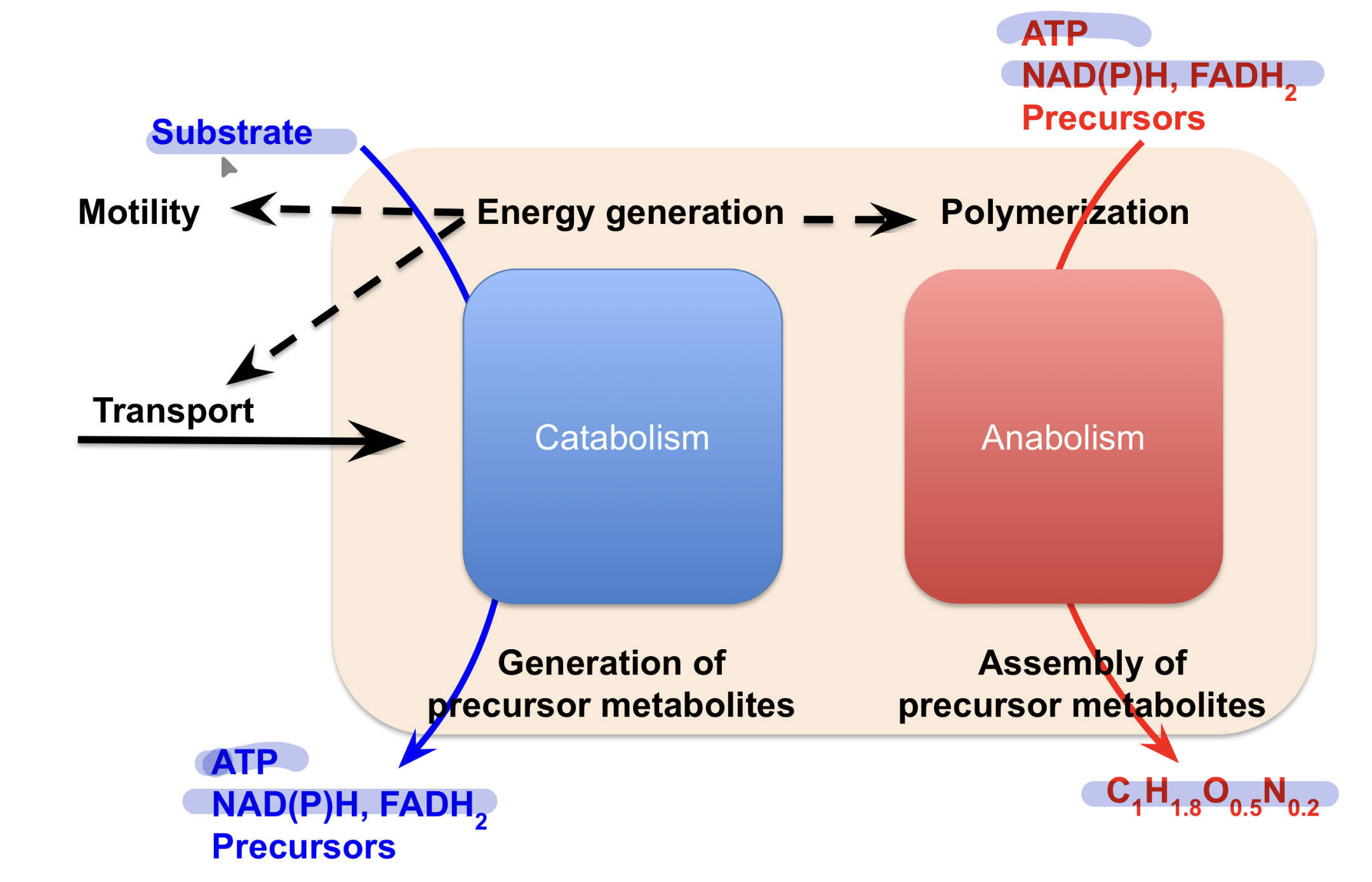
Cell energy cycle
Catabolic processes produce ATP from ADP + Pi needed for chemical, transport, and mechanical work where ATP is hydrolyzed back to ADP +Pi
Standard Reduction Potential (E’0)
ability to gain electrons (higher E’0 = better acceptor)
Energy sources (ex. to make ATP)
Light
Chemicals (ex. glucose)
Light energy source
Phototroph
Chemical energy source
Chemotroph
Electrons sources (ex. to make RP)
Inorganic (ex. water)
Organic (ex. glucose)
Inorganic electron source
Lithotroph
Organic electron source
Organotroph
Carbon sources (ex. to make precursor metabolites)
CO2
Organic (ex. glucose)
Both = mixotroph
CO2 carbon source
Autotroph
Organic carbon source
Heterotroph
What is name of organisms that do aerobic respiration, anaerobic respiration, and fermentation?
Chemoorganoheterotroph
What is name of organisms that use inorganic chemicals?
Chemolithoautotroph
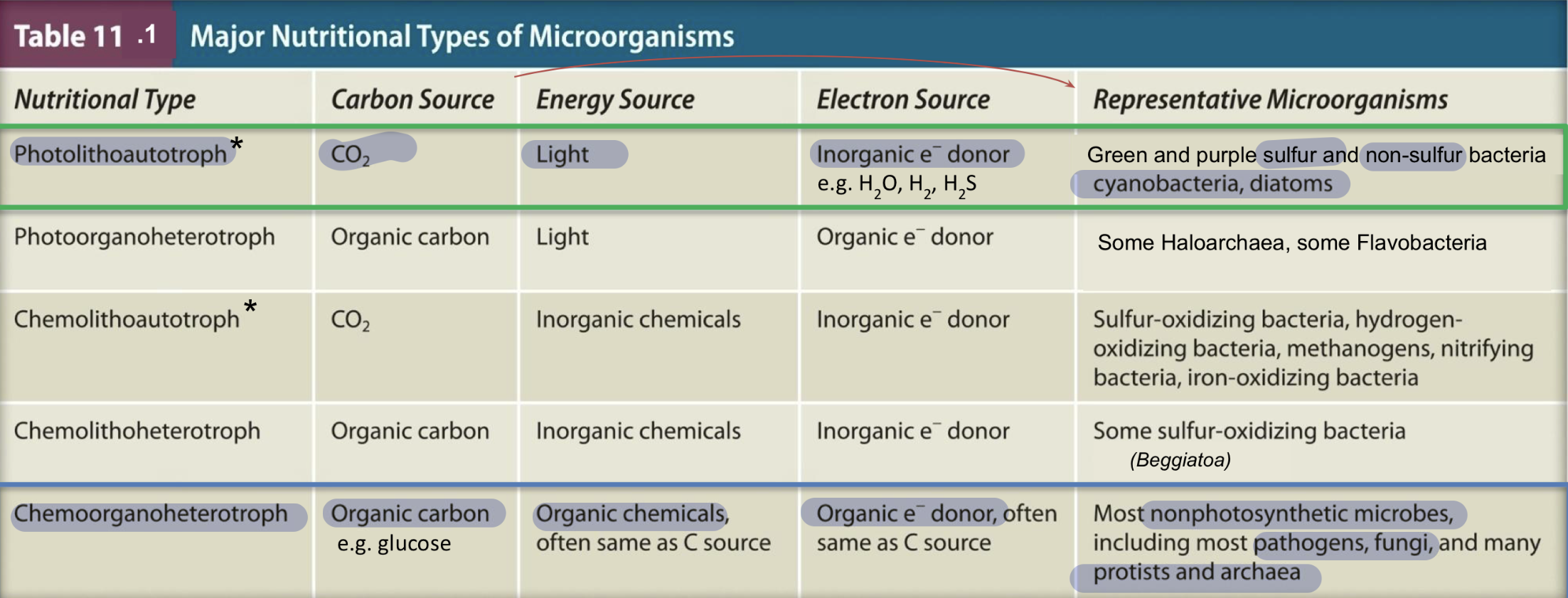

Summary of nutritional types of microorganisms
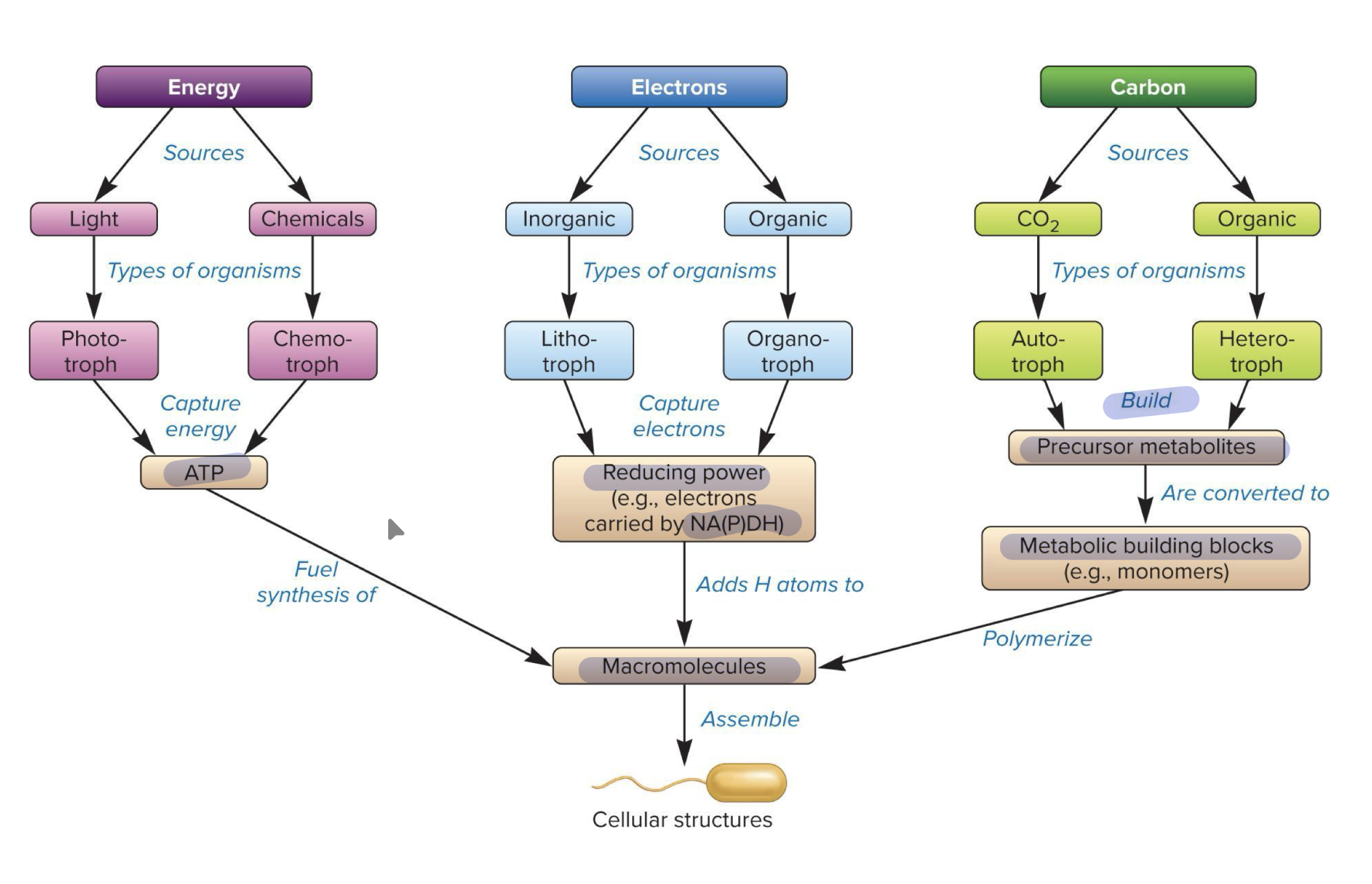
Bacteria and Archaea have huge metabolic flexibility
found everywhere
many can change nutrition types based on environment (ex. sulfuric acid)
5 types of metabolism
aerobic respiration
anaerobic respiration
fermentation
chemolithotrophy
photosynthesis
Chemoorganotropic fueling rxns (energy from organic chemicals) (can be done by hetero or auto ex. cyanobacteria)
aerobic respiration
anaerobic respiration
fermentation
Aerobic respiration
(starts w/ glucose or other molecules that can be funneled in)
1) glycolytic pathway
Glucose to pyruvate
3 routes
Glycolysis (E, RP, PM)
Entner-Doudoroff (ED) pathway (E, RP, PM)
Pentose Phosphate Pathway (PPP) (only RP, PM)
2) TCA pathway
3) ETC
Glycolysis
universal across all tree of life
all cells except bacteria that rely only on ED or PPP
2 stages
6C phase: Glucose (C6) splits into 2 glyceraldehyde 3-phosphate (C3)
3C phase: Oxidation of each glyceraldehyde 3-phosphate into pyruvate
Produces 2 NADH molecules
Produces 2 net ATP
initial invest 2 ATPs
Produce 4 total ATP by sub level phosphorylation
produces precursor metabolites for anabolism through some intermediates and pyruvate
F6P —> NAM, NAG
PLs
amino acids
Entner-Duodoroff (ED) Pathway
specific to some bacteria
starts with glucose
Produces 1 pyruvate and 1 Glyceraldehyde 3-phosphate (later continues to reg glycolysis)
Only 1 net ATP (uses 1 and then produces 2 in glycolysis)
Advantage = growth on aldonic acids (ex. E.coli can switch to ED and grow on gluconate (intermediate of ED) when no glucose)
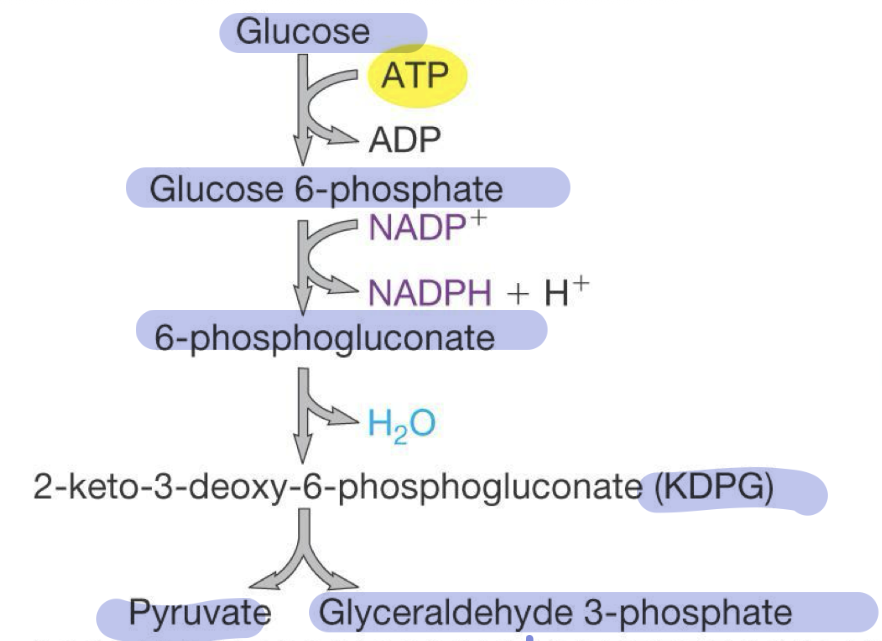
Pentose Phosphate Pathway (PPP)
bacteria and eukaryotes
No ATP produced
Glucose 6-phophate converted to
variety of 3C-7C sugars (like G3P for glycolysis) (sugars used for anabolic of amino acids)
pentose precursors for nucleic acids
Produces NADPH —> for anabolism (not ETC)
TCA cycle (tricarboxylic acid)
Pre-step: oxidation pf pyruvate —> acetyl —> acetyl-CoA
produces 1CO2 and 1NADH
Achieved by pyruvate dehydrogenase complex (uses coenzyme A)
Citric acid cycle: oxidation of acetyl-CoA to CO2
produces 2CO2 and 1GTP (or ATP)
Produces RP: 3NADH and 1FADH2
achieved by 8 diff enzymes
2 carbons from acetyl-COA eventually leave as 2CO2 —- each initial glucose needs 3 runs of TCA to fully oxidize
produces precursor metabolites
Electron Transport Chain and Oxidative Phosphorylation
NADH and FADH2 generated from glycolysis and TCA flow down ETC to O2 (final e- acceptor) —> reduced to H2O
energy from flow of electrons down proteins pumps protons —> proton gradient —> ATP synthase
Why cells go through so much to form H2O from O2?
ETC allows energy to release gradually —> more efficient and more ATP
Eukaryotic ETC
4 multiprotein complexes, coenzyme Q, cytochrome c in inner membrane of mitochondria
energy release pumps H+ to mito intermembrane space —> proton gradient
H+ returns to matrix via ATP synthase —> ATP synthesis by proton motive force
Energy used for…
ATP
flagella
transport
Prokaryotic ETC
variable in length and composition
embedded in plasma membrane (sometimes per plasmic space or outer membrane)
energy released pumps H+ to periplasmic space —> proton gradient —> H+ returns to cytoplasm via ATP synthase —> ATP synthesis by proton motive force
Variations in ETC depend on C source
Glucose as C source — chemoorganoheterotrophy uses this aerobic respiration — ex. paracoccus denitrificans
Methanol can be used as C source through methylotrophy
Variations in ETC depend on oxygen availability (E.coli)
2 alternative terminal oxidases in low or high O2 conditions (or fermentation or anaerobic)
High aeration, log phase = normal O2, normal ETC
Low aeration, stationary phase = little O2, diff aerobic respiration
Total ATP yield per glucose is variable
variable ETCs
some proton motive force used for transport or movement (flagella)
some intermediate metabolites channeled to anabolism
Max ATP per glucose in eukaryotes = 32
lower ATP in prokaryotes due to less complexes in ETC —> less passage of electrons —> less chemoosmosis —> less oxidative phosphorylation
Aerobic respiration summary
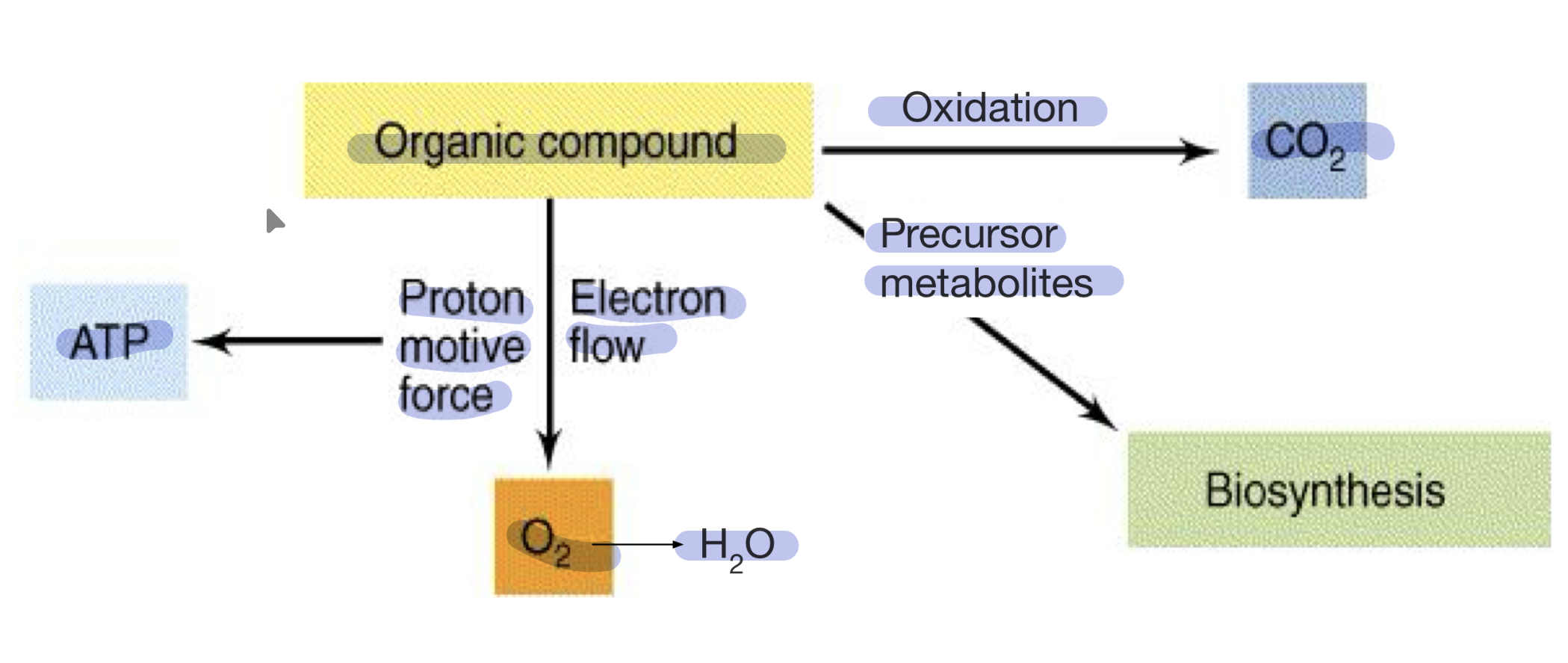
Anaerobic respiration
prokaryotes and protists (only in microbes)
Similarities to aerobic respiration
organic substrate is oxidized (CnH2O —> CO2)
resulting NADH and FADH2 used in ETC
Differences from aerobic respiration
O2 not final electron acceptor (NO3- and SO42- common)
Less ATP produced
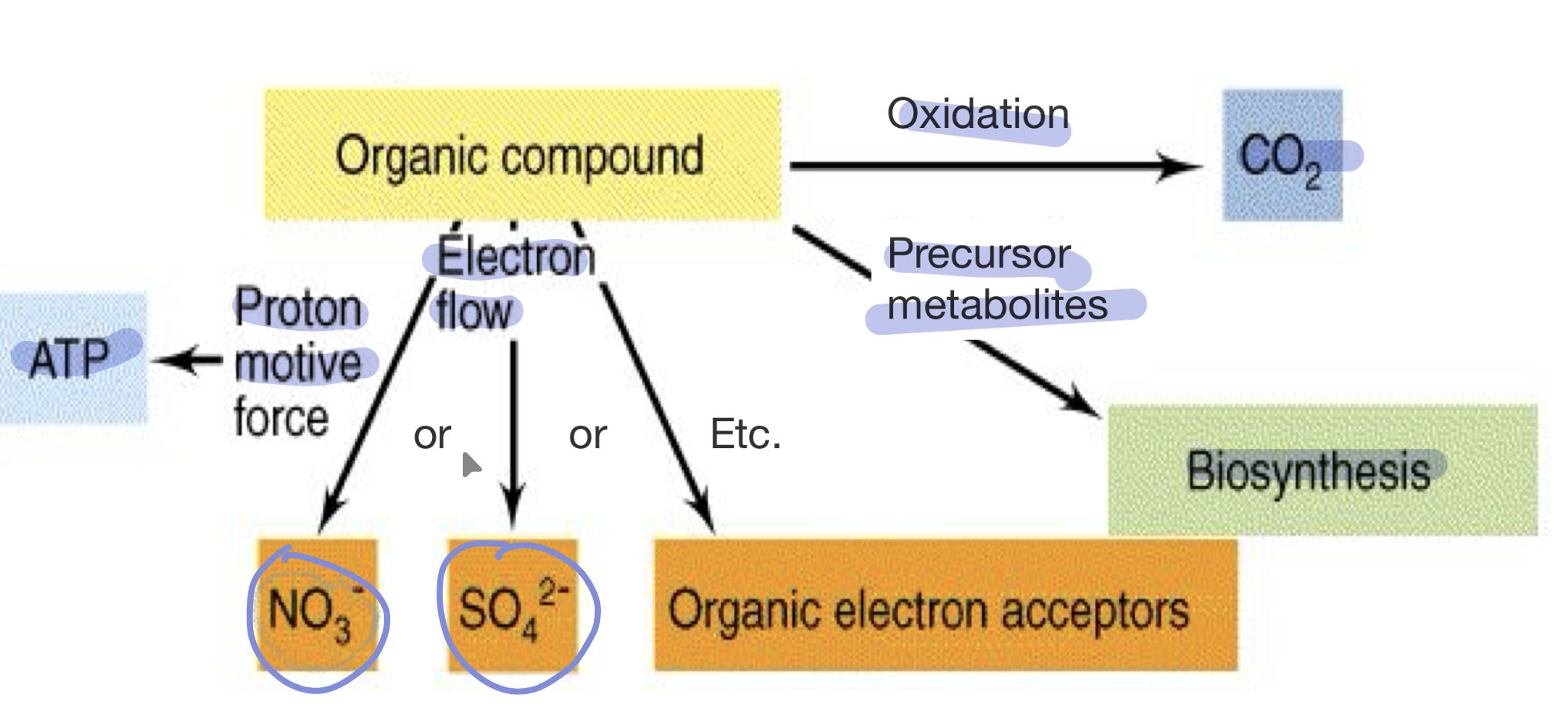
Dissimilatory nitrate reduction
NO3- (final electron acceptor) —> NO2- (reduced product)
Denitrification
NO3- (final electron acceptor) —> NO2-, N2O, N2 (reduced product)
Dissimilatory sulfate reduction
SO42- (final electron acceptor) —> H2S (reduced product)
Methanogenesis (chemolithoautotrohpy)
CO2 (final electron acceptor) —> CH4 (reduced product)
Acetogenesis
CO2 (final electron acceptor) —> acetate (reduced product)
Ferric reduction
Fe3+ (ferric) (final electron acceptor) —> Fe2+ (ferrous) (reduced product)
Anaerobic respiration yields less ATP due to…
smaller diff in reduction potential from NADH/FADH2 to substrates other than O2 → less H+ transport → less ATP
Ranking from most to least ATP: aerobic → ferric reduction → denitrification → dissimilatory nitrate reduction
Difference btw dissimilatory nitrate reduction and denitrification
Dissimilatory nitrate reduction makes N unavailable for cell assimilation
When gas is produced → denitrification
E.coli does not do denitrification
Paracoccus denitrificans using aerobic respiration do denitrification: Nitrate —> nitrite by Nar but then reduced fully to nitrogen by Nir and Nas
Fermentation
In many prokaryotes, some fungi and protists, some animal cells
ETC not used because
there is none (ex. symbiotic bacteria and archaea)
ETC is repressed (some facultative bacteria in low O2, such as S. aureus when overgrowing —> mannitol fermentation)
ETC cannot function due to lack of final electron acceptor (ex. some facultative anaerobes such as E.coli when no O2 or NO3-)
Glycolysis still occurs
NADH produced must be oxidized back to NAD+ (instead of ETC)
Pyruvate produced is reduced to acid (lactic acid) or alcohol (ex. ethanol) — pyruvate acts as endogenous final electron acceptor
many fermentation pathways due to different microbe lineages — named after product
If produces ATP, from sub level phosphorylation
max yield = 2 or 4 ATP
Alcoholic fermentation
produces ethanol and CO2 (ex. yeasts)
Lactate (lactic acid) fermentation
homofermentative = produces mostly lactate
Heterofermentative = produces also ethanol and CO2
Mixed acids fermentation
produces ethanol and complex mix of organic acids
Butanediol fermentation
produces ethanol and butanediol
Is lactate fermentation the same as fermentation of lactose?
Yes —- lactose → galactose → glucose → glycolysis → fermentation → lactic acid
Fermentation of mannitol
No acidity: Mannitol → fructose → Fructose 6-phosphoric acid → glycolysis → fermentation → ethanol
Acidity: Mannitol → fructose → Fructose 6-phosphoric acid → acid products
Challenges of fermentation
regenerates NAD+
If all energy comes from sub level phosphorylation → no proton motive force generated for transport and flagella motility → H+ accumulates in cell → acidification
So, fermentors used ATP synthase in reverse direction: pumps H+ out of cell hydrolyzing ATP and generating proton motive force
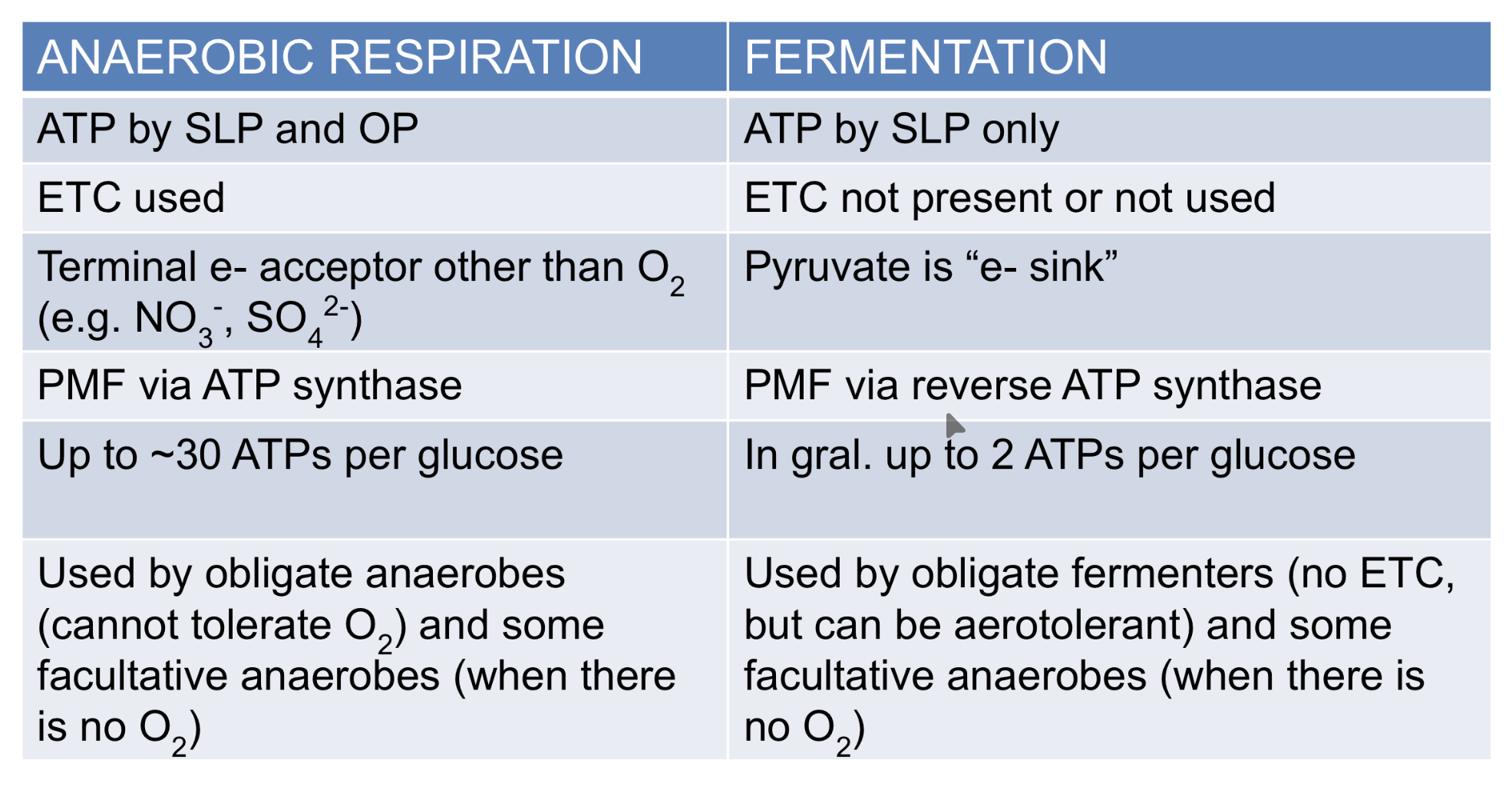
Anaerobic respiration vs fermentation
ATP by…
ETC used or not
Final electron acceptor?
Proton motive force by…
Total ATP generated
Used by…

Chemolithotrophic fueling reactions
Similar to aerobic respiration, but oxidation of an inorganic molecule
Still ETC and ATP by ox phos
Carbon fixation: Carbon source is often CO2 cause most are chemolithoautotrophs and some are chemolithomixotrophs (also use organic C)
Chemolithotrophs energy sources and electrons
energy from oxidation of inorganic electron donor (H2, NH4+/NO2-, SH2, Fe2+) (reduced molecules)
Reduction of electron acceptors
mostly O2 = aerobic respiration
some others = anaerobic respiration
Chemolithotrophs impact on environment
They need to do multiple cycles of rxns to get enough ATP and reductive power → use up a lot of those inorganic molecules from the environment
plays a major role in biogeochemical cycles = N, S, Fe
Chemolithotroph ETC
ETC is similar to aerobic/anaerobic respiration
energy is produced (ATP)
Phototropic fueling rxns
Chlorophyll-based phototrophy
Light → chlorophyll or bacteriochlorophyll → ETC → PMF → photo phosphorylation → ATP
Carbon fixation: Carbon source (often CO2 for photolithoautotrophs) → biosynthesis
Electrons from H2O or H2, SH2
NADP+/NADPH
Rhodopsin-based phototrophy
Light → bacteriorhodopsin → PMF → photo phosphorylation → ATP
Carbon fixation = organic carbon source = photoorganoheterotrophs
electrons from organic source
Chlorophyll- and Bacteriochlorophyll-based phototrophy
Eukaryotes
have chloroplasts
2 photosystems
electron donor = H2O
energy products = ATP + NADPH
carbon source = CO2
aerobic, generates O2
thylakoid membrane
Cyanobacteria
no chloroplasts
2 photosystems
electron donor = H2O
energy products = ATP + NADPH
carbon source = CO2
aerobic, generates O2
thylakoid membrane is PM?
Green and purple sulfur and non-sulfur bacteria, heliobacteria, acidobacteria
bacteriochlorophyll
1 photosystem
electron donors = H2, H2S, S, organic matter
energy product = ATP
Carbon source = organic or CO2
anaerobic, no O2
Plasma membrane invagination
Chlorophyll-based phototrophy (cyanobacteria, algae = phytoplankton)
Light rxns: photosystems and photophospshorylation (similar to ETC ox phos except starts with light)
Photosystem II: light is used to pull electrons from H2O, which is then oxidized to O2
Photosystem I: electrons are boosted by light and the used to produce NADPH
Coupling 2 photosystems is an ETC that leads to a H+ gradient and synthesis of ATP by photophosphorylation
Bacteriochlorophyll-based phototrophy (green and purple sulfur and non-sulfur bacteria, heliobacteria, acidobacteria)
only 1 photosystem
no oxygen produced
no NADPH produced
ATP synthase still makes ATP from proton motive force
Rhodopsin-based phototrophy
rhodopsin captures light
What are the energetic demands of biosynthesis in a microbe?
Anything cell needs to make is done via biosynthesis using C, ATP and reducing power
What are the carbon precursors metabolites, ATP, and reductive power used for?
ex. glycolytic pathways and TCA cycle provides for:
gluconeogenesis
amino acid synthesis
nucleotide synthesis
peptidoglycan synthesis (some F6P diverted to this instead of glycolysis)
Assimilation via carbon fixation
Carbon fixation = taking CO2 from atmosphere and turning it into glucose
Calvin cycle (anabolic pathway)
Half of all organic carbon on earth is produces by microbial photosynthesis
Key enzyme: RubisCO (ribulose bisphosphate carboxylase) = enzyme that captures gas and turns it into organic molecule
Takes place in carboxysomes for prokaryotes
18 ATP and 12 NADPH required per molecule of glucose produced
Reductive TCA cycle
by chemolithoautotrophs and anoxygenic phototrophs (that don’t have rubisco)
TCA runs in reverse
product is acetyl-CoA (→ is then used to make glucose)
Requires ATP and electrons (from chemolithotrophy)
Way to incorporate sulfate in chemolithotrophy
Assimilatory sulfate reduction: takes sulfate and turns it into cysteine
Way to incorporate nitrate in chemolithotrophy
Assimilatory nitrate reduction: reduce nitrate NO3- to Ammonia NH3 then to organic matter
diff from dissimilatory nitrate reduction in anaerobic respiration (nitrate is just final electron acceptor)
uses nitrate reductase (Nas) in the cytoplasm
done in plants, algae, and cyanobacteria
Way to incorporate nitrogen
Nitrogen fixation
only done by nitrogen-fixing microbes
reduction of atmospheric nitrogen (N2) to ammonia (NH3)
catalyzed by enzyme nitrogenase
requires a lot of ATP and electrons
done by cyanobacteria, Rhizobia (live in plant roots), methanogenic archaea, and alga (have nitroplast which is a nitrogen-fixing organelle)
Microbes with different environments are distributed in the environment based on (soil, water, a culture, your body) based on:
temperature
osmotic pressure
pH
oxygen concentration
redox potential (availability of final electron acceptors and donors)
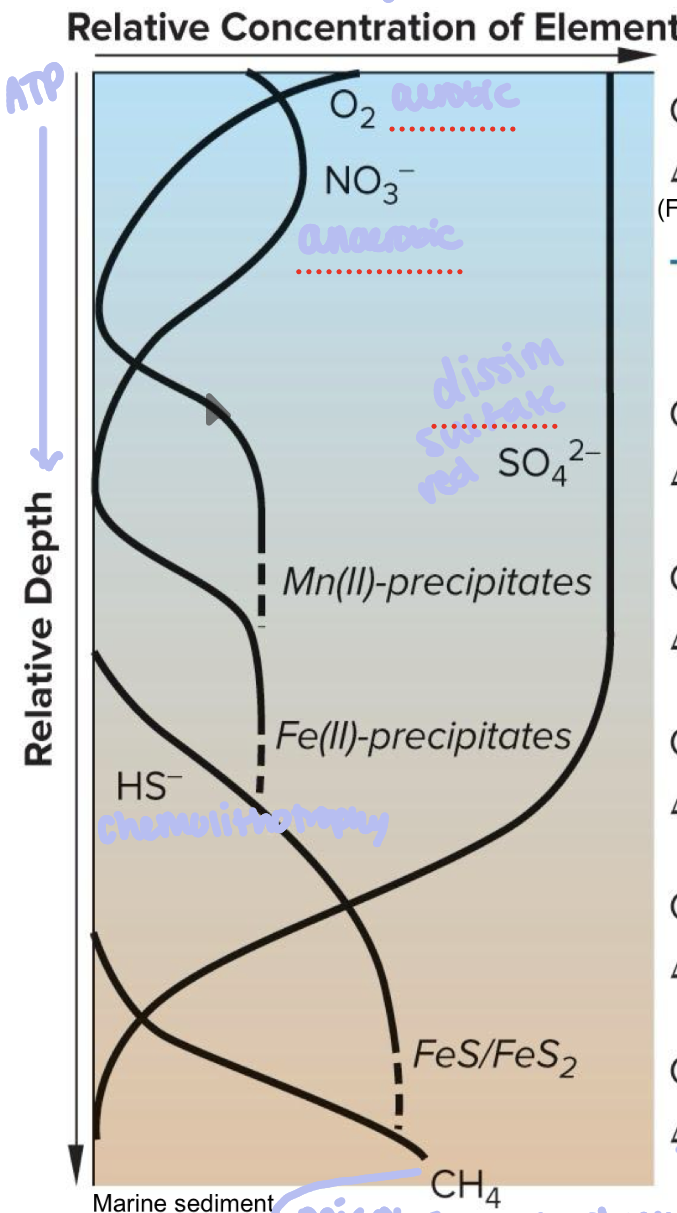
As you go deeper in sediment…
molecule availability changes and so do the microbes based on types of metabolism
towards the top = most O2 = aerobic respiration
NO3- = anaerobic respiration (denitrification)
SO42-= dissimilatory sulfate reduction
Fe(III) = ferric reduction
HS- = chemolithotrophy
ATP produced by each microbe decreases
microbes have adaptations where they don’t need that much ATP or have other mechanisms to get it
What came first, anaerobic or aerobic respiration?
Anaerobic respiration then when cyanobacteria first evolved they produced O2 for earth
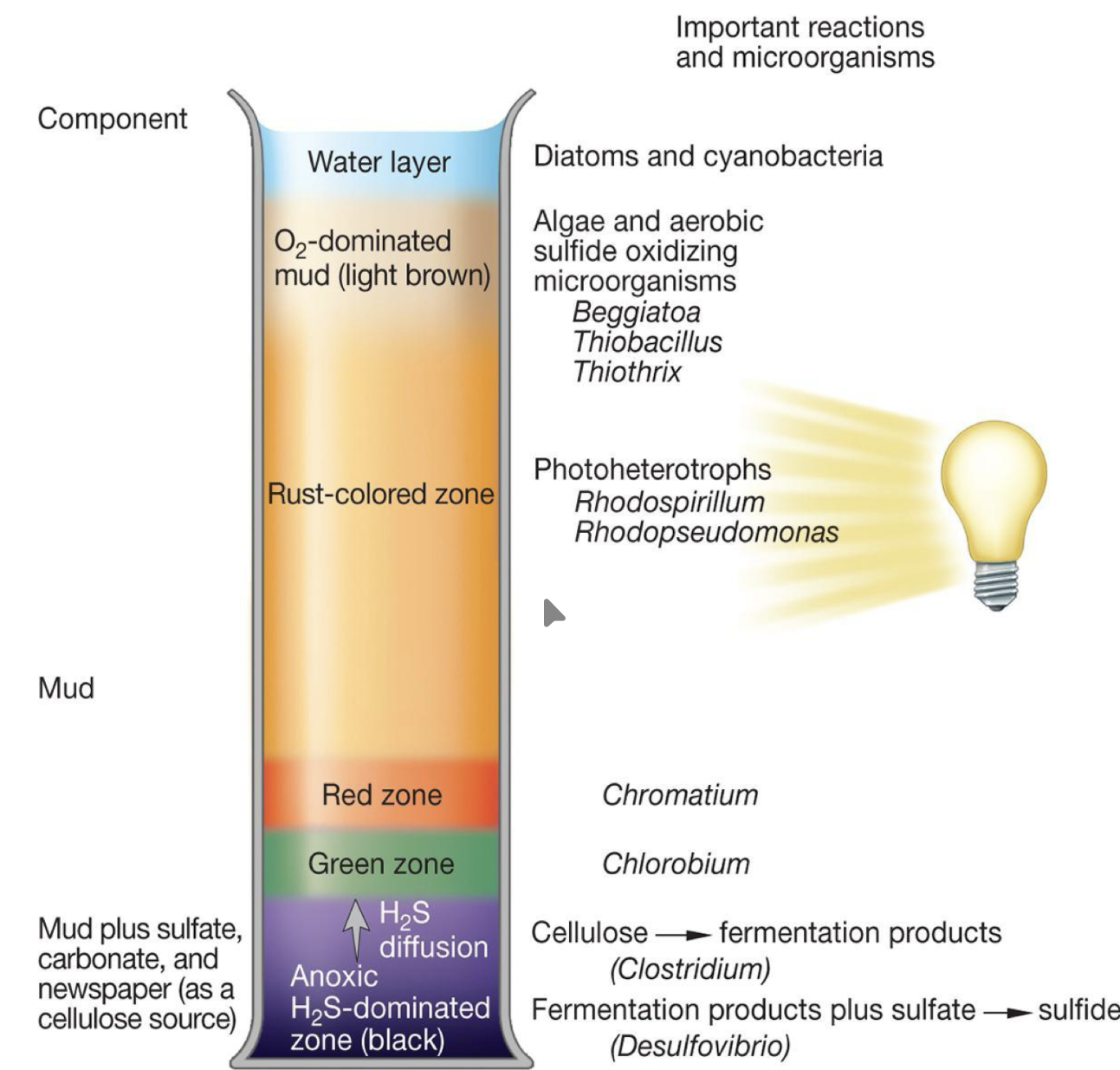
Winogradsky columns
demonstrate diversity of microbial metabolisms with different colors representing different products from different metabolisms
How can sulfur be made
cysteine degradation
anaerobic respiration
Cable bacteria metabolism
have cells that allow electrons to travel up to an oxic zone so that O2 can be final electron acceptor → more ATP
Microbial metabolisms impact on biogeochemical cycles
carbon
nitrogen
Microbial metabolism impact on carbon cycles
CO2 → organic matter: CO2 fixation (photo and chemoautotrophy)
organic matter → CO2: respiration and fermentation, decay and mineralization
CO2 → CH4: methanogenesis (only microbes)
CH4 → CO2: methano and methyl (only microbes)
Methanogenesis and Methanotrophy
Methanogenesis: reduction of CO2 to CH4
Methanotrophy: Oxidation of CH4 to CO2
**Both processes happen in adjacent habitats and sometimes involve metabolic syntrophy or “cross feeding”
Microbial metabolism impact on nitrogen cycles
Driven by microbes —> would stop without microbes
Nitrogen cycle
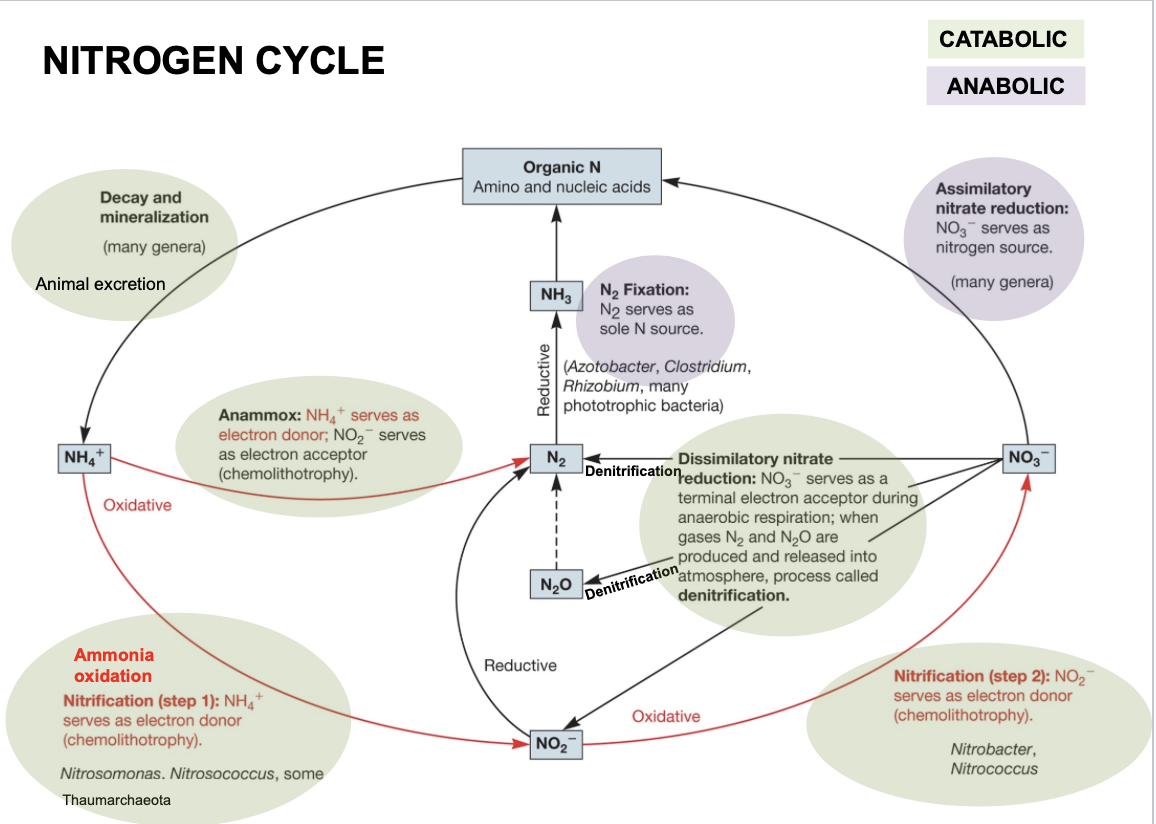
Nitrate (NO3-) → organic N molecules (amino and nucleic acids)
Assimilatory nitrate reduction
NO3- is N source
many microbes
anabolic
reduction
occurs in both aerobic and anaerobic conditions
Organic N molecules (amino and nucleic acids) → ammonium (NH4+)
Decay and Mineralization
many microbes
reduction
catabolic
Animal excretion
Ammonium (NH4+) → Nitrogen gas (N2)
Chemolithotrophy - Anammox
Ammonium (NH4+) is electron donor
Nitrite (NO2-) is electron acceptor
oxidation
catabolic
Ammonium (NH4+) → nitrite (NO2-)
Chemolithotrophy - Ammonia oxidation
nitrification step 1: NH4+ is electron donor
oxidation
catabolic
Nitrite (NO2-) → Nitrate (NO3-)
Chemolithotrophy Nitrification (step 2):
NO2- is electron donor
oxidation
catabolic
Nitrite (NO2-) → Nitrogen gas (N2)
Reductive
Nitrate (NO3-) → Nitrite (NO2-), nitrous oxide (N2O), nitrogen gas (N2)
Anaerobic respiration - Dissimilatory nitrate reduction
reduction
catabolic
chemoorganoheterotrophy
For nitrous oxide (N2O) and nitrogen gas (N2) (because gases) - Denitrification
reductive
catabolic
Nitrogen gas (N2) → NH3 → organic N (amino acids and nucleic acids)
Nitrogen fixation
reductive
anabolic
Rhizonium and many phototrophic bacteria
N2 serves as sole N source
Global impacts of microbes
Microbial metabolisms influence concentrations of …
greenhouse gases (CO2, CH4, N2O) → global warming
N compounds in soil and aquatic environments → eutrophication
Ways microbial metabolism lower CO2
All autotrophic bacteria absorb CO2
Ways microbial metabolism increase methane gas
Microbes in cow stomach have enzymes that help degrade cellulose → these microbes do methanogenesis —> CH4
methanogenic archaea
Permafrost sewing = frozen soil melting because of global warming → wakes up dormant methanogenic microbes in soil → CH4
positive feedback
Eutrophication (increase N in environment)
fertilizer → excess N in soil for microbial metabolism → N2 and N2O released into atmosphere
N pollution in water (waste from rain, sewage, runoff) → phytoplankton bloom → phytoplankton sink because too many of them → microbes decompose phytoplankton using O2 as electron acceptor → hypoxia (low O2) → aquatic life death, poor water quality → affects public health and economy
Why is culturing difficult for characterizing environmental microbes
Most environmental microbes don’t like to live in the lab
Quantification of environmental microbes can be done via…
flow cytometry (single cell count at a time)
fluorescence microscopy (DAPI staining)
Metabarcoding and Metagenomics
Used to study taxonomy and functions of culturable and uncultivable microbes
1) filter water
2) extract DNA
3) PCR
4) sequence
Metabarcoding: sequence 16S or 18S (for protists) rRNA gene of thousands of species (not Sanger)
Metagenomics: sequence all genes of many species without PCR
Activities of microbes can be studied via
Extracting mRNA for specific gene of interest → run quantitative PCR
**mRNA not DNA because want to ensure gene is expressed
Why care about microbes in environment
Recycle C,N, and other nutrients
Interact (+ or -) with climate change, eutrophication and other environmental issues
maintain soil integrity
sustain aquatic food webs
have key interactions with other microbes and non-microbes (ex. eating eachother)
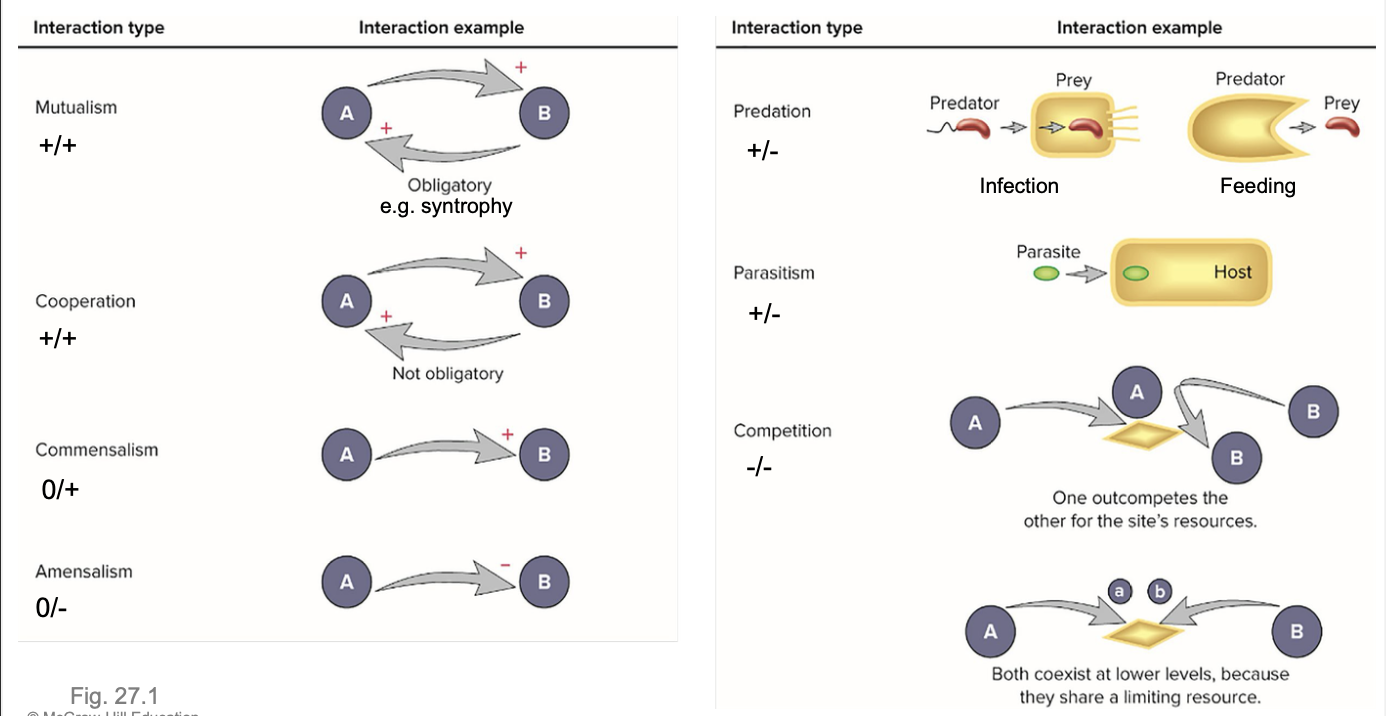
Microbe-Microbe and Microbe-host interactions