oxidative phosphoryaltion
1/18
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced |
|---|
No study sessions yet.
19 Terms
five oligomeric complexes
I NADH-ubiquinone (Q) oxidoreductase
II succinate-ubiquinone (Q) oxidoreductase
III ubiquinol- cytochrome c reductase
IV cytochrome c oxidase
V ATP synthase
ubiquinone
oxidised form of coenzyme q
aliphatic chain
electrons are captured from the head
coenzyme q
reduced one electron at a time
semiquinone is a free radical from reduction of coenzyme q
reduced to dihydroquinone
which is then oxidised one electron at a time
complex i - enzyme and modules
NADH-ubiquinone reductase
largest of complexes
nadh binds to dehydrogenase n module
ubiquinone found in q module
H+ move across the membrane arm P module
protons and electron movement with complex i
two electrons from NADH transferred - oxidation to NAD+
4 protons move from mitochondrial matrix to the inter membrane space
against the conc gradient, nadh giving electrons to ubiquinone releases energy to allow this
FMN acts as intermediate
complex ii properties and enzyme
only enzyme common to citric acid cycle and respiratory chain
does not move protons
succinate Q reductase
electron movement and products from complex ii step
FADH2 accepts the two electrons
ubiquinone → ubiquinol
succinate → fumarate releases two electrons to reduce FAD to FADH2 then ubiquinone to ubiquinol
cytochrome c
electron carrier between complexes III and IV
alpha helical haem protein, iron atom in the centre, does not bind oxygen
small, highly soluble protein from the intermembrane space protein, associated to inner mitochondrial membrane
complex iii enzyme
ubiquinol-cytochrome c oxidoreductase
electron and protons transfer in complex iii
4 H+ are translocated, two from the matrix and two from QH2 using free eenrgy
electrons are transferred from ubiquinol (QH2) to two molecules of cytochrome C - Q cycle
iron atom can change oxidative state in cytochrome C which allows the transfer of electrons
complex iv function
receives electrons from cytochrome C carrier, one at a time
iron atoms and copper atoms are both reduced and oxidised as electrons flow to oxygen
catalyses the reduction of oxygen and water
two more hydrogen ions are translocated using free energy
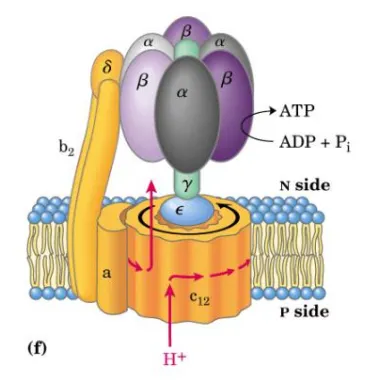
complex v enzyme, location and structure
atp synthase
found at the tips of the cristae
knob and stalk structure
F1 - catalytic subunits (faces the matrix)
F0 - proton channel (embedded in the inner membrane)
proton movement for complex v
use the proton gradient energy for the synthesis of ATP
protons flow back into the matrix via ATP synthase
through rotation of proton channel and move up the stable ring
F1 subunits
3a, 3beta, 1 gamma, 1δ, 1ε
alpha and beta alternate like orange segments
does not move
gamma is main component of central axle and allows the rotation
δ is in peripheral stalk
each beta subunit has active site for ATP synthesis
rotational catalysis conformations for each beta unit
open state - available to bind ADP and Pi
loose state - active site closes loosely on ADP and Pi
tight state - converts ADP and Pi into ATP
what drives the conformational changes of beta subunits for f1
flow of protons drive F0 and gamma rotation
this forces cyclic conformational changes to each beta subunit
number of protons per ATP
one full rotation of ATP synthase - 3 molecules of ATP
number of subunits in c ring of F0 = number of H+ needed for one full rotation
number of H+ per ATP
oxidative phosphorylation ~3H+/ATP
what transports atp, adp and pi
adenine nucleotide translocate (antiporter)
then pi enters through symport mechanism with H+
atp synthase stochiometry
P:O ratio = molecules of ADP phosphorylated / atoms of oxygen reduced
4H+ per ATP synthesised
one H+ needed for transport of Pi across inner mitochondrial membran e
NADH: 2 e–, 10 H+ transported, ~2.5 ATP/O FADH2: 2 e–, 6 H+ transported, ~1.5 ATP/O