exam 3 biology
1/18
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
19 Terms
study of describing, naming/ classifying organisms
taxonomy
scientific name parts
1st genus: capitalized n itlicuzed (underlined when written)
2nd specific epithet
carl linnaeus
made up scientific name. formal way of introducing species
category vs taxon
rank or lvl of taxonomic classification ex:fam or genus
tacon: a named group of organisms ex: urus
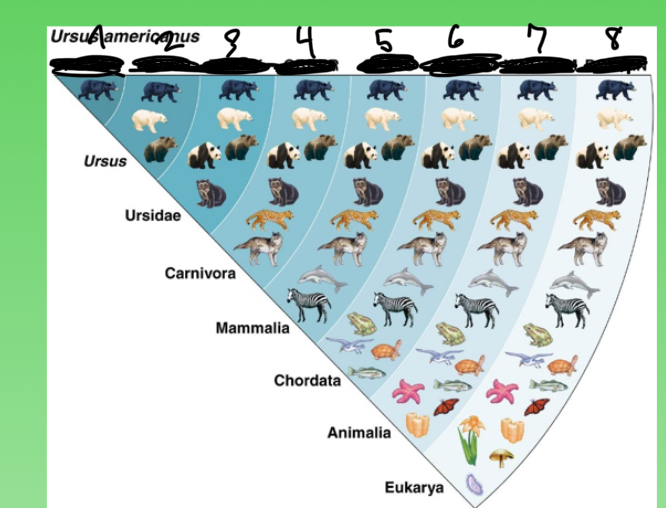
name the categories 1-7
1 species
2 genus
3 fam
4 order
5 class
6 pylum
7 kingdom
8 domain
what is the tree of life and what does it show us
its a model that shows evolution relationships between all living organism's on earth
it shows us how different species r related through common ancestors
what r the 3 domains in tree of life
bacteria, archea, n protist.
what r the 4 kingdoms in tree of life
protist, fungi, plantae, animalia
systemics
studyof diversity n life n its evolutionary rls
what is phylogeny
evo history of a group of organisms. presented as a phylogenetic tree
the phylogentic tree is made up of what three things
root: oldest point of time in tree, common ancestor
line: lineages, species
nodes: last common ancestor ofthe branches above
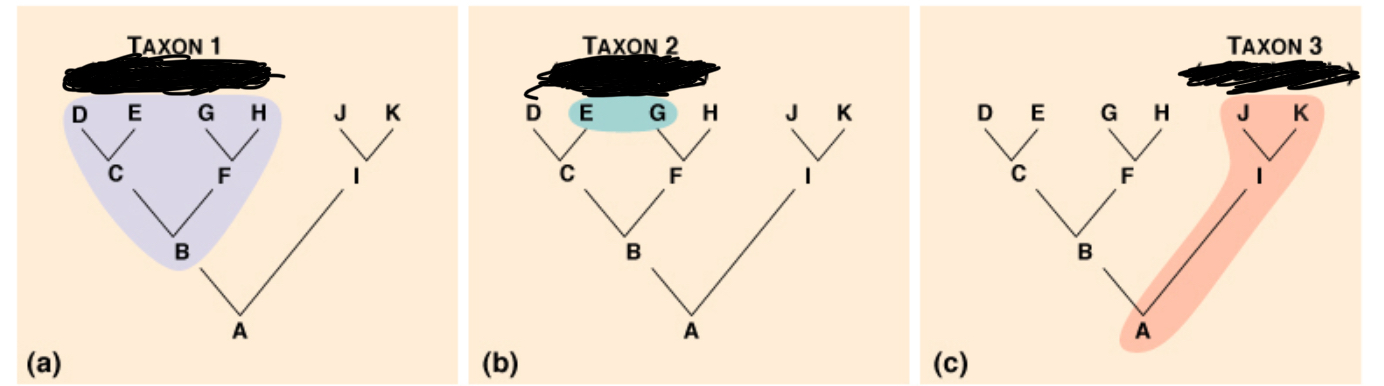
Label a- c n why each label is that
a monophyletic has to do w a comon ancestor n all of its descendants (known as clades)
b polyphletic unrelated orhanism grouped by common trait lack a common ancestor
c paraphyletic contains common anecestor but not all descendants
what is homoplasy
individuals can look similar but not b closely related at all n thats bc of convergent evo
convergent evolution resulting structures r…
analogus sturcture that evolved diff in species
homology va homoplasy
plasy: results FROM convergent evo. makes them analogus structures, similar func but not development path, NOT ON PHYLOGENTIC TREE
logy: common ancestry, same developmental path n position not necessarily function, results IN homolgues sturctures, ON PHYLOGENTIC TREE
polymerase chain reaction
where u make a bunch of copies of dna from a small sample (molecular data)
advantages of moleculear data
can compare distantly related species that dont look similar and one who arent related but look similar so u dont have to go based off looks.
can factorslf environmental phentotype affect on species
cladistics
ased on shared n derived characters states which results in cladograms