Biology of Cancer- Exam 1 (Ch.1-4 and JC 1&2)
1/123
Earn XP
Description and Tags
Exam 1 flashcards for first 4 chapters and 1-2 journal clubs. There will be vocab reperats because I am doing it by chaoter but I will try to have the examples/ implications fo why they are relevant in different ways for the different chapters when posssible. The chapters are not labeled because they are seperated by critical thinking questions.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
124 Terms
Incidence
Total number of new cases in a given time
Prevalence
Total number of cases at a give time
Mortality
Death caused by cancer
Carcinoma
• “-carcinoma” = Epithelial cells ( ~85% of cancers)
Sarcoma
• “-sarcoma” = Connective Tissue
Lymphoma
• “-lymphoma” = Cells of the lymphatic system
Leukemia
• “-leukemia” = Circulating blood cells
Benign
Benign tumors are confined to a certain area in the body
• Benign tumors are abnormally growing cells that do not invade other tissues and remain restricted to a defined area
• Benign tumors are easier to treat (e.g surgery) and generally have
better prognoses
• Exceptions can be tumors in areas that are hard to reach surgically (e.g. the brain)
Malignant
• “Malignant” is an adjective to describe a property of
tumor cells. Malignant tumors have the ability to spread
throughout the body
-Malignant tumor cells break confinement and spread through the body
-Malignant cancers are abnormally growing cells that have gained
additional mutations that allow them to leave their tissue or origin
Metastasis
• “Metastasis” is a process. It says that tumor cells are
spreading through the body to form secondary tumors.
• Malignant tumors are tumors that have the ability to
metastasize
Carcinogenesis
• Carcinogenesis is the multi-step
process by which a healthy cells
transforms into a cancerous cell, and is
caused by:
• Inherited DNA mutations (Nature/Genetics)
• Environmental carcinogens (Environment)
Carcinogens
A carcinogen is any substance or agent that promotes the development of cancer
• Different environmental carcinogens are
related to different cancers
• UV = Skin cancer
• Cigarette smoke = Lung cancer
• Diet/Exercise
• Reproductive life = Breast cancer
• Alcohol = Liver cancer
Mutagens
Mutagens cause abnormal changes to DNA to disrupt normal cellular function
>Mutations to DNA sequence lead to changes in cell protein expression (function)
-Accumulated mutations are passed to new cells via mitosis
*All cancers have an underlying genetic component
Tumor Suppressor Genes
Tumor suppressor genes
• Non-mutated: Suppresses cancerous
pathways (e.g. cell cycle progression)
• Mutated: Fails to suppress cancerous
pathways
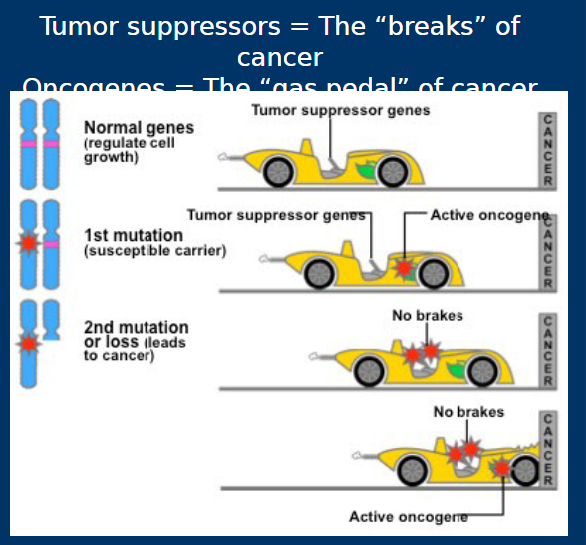
Proto-Oncogenes/Oncogenes
• An oncogene is a gene mutated such that its protein product is produced in higher quantities or has increased activity
• Non-mutated versions known as proto-oncogenes
• Mutated oncogenes promote tumorigenesis and cell cycle progression
Germline mutation
Germline mutations are inherited mutations and increase susceptibility
Somatic mutation
Somatic mutations are acquired mutations
Chemotherapy
Chemotherapy: Use of chemicals designed to kill cancer cells
• Better efficacy against malignant tumors
• Often associated with severe off-target effects
Radiation
Radiation: High energy targeted at tumor leading to death of tumor cells
• Ineffective against malignant tumors
• Potential off-target effects
Efficacy
Efficacy: How well do they treat (effect) the cancer
Safety
Safety: To what extent do they cause off target health effects
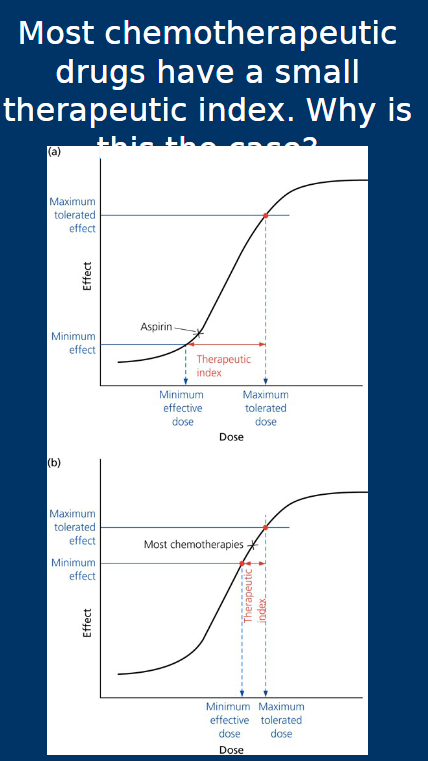
Therapeutic index
The therapeutic index
• Difference between minimum effective dose and maximum tolerated
dose
• Larger the value, safer the drug
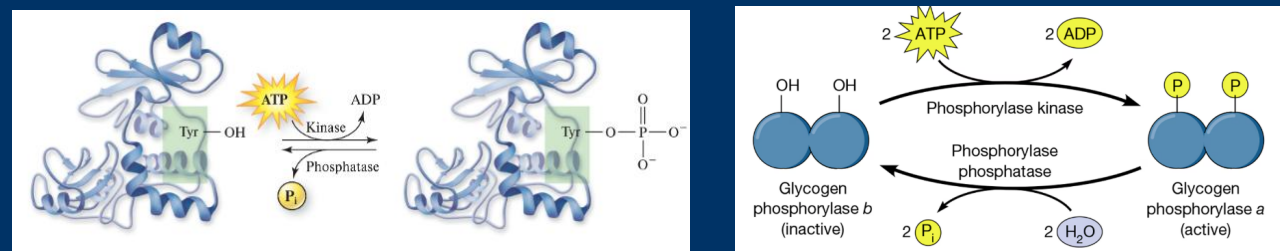
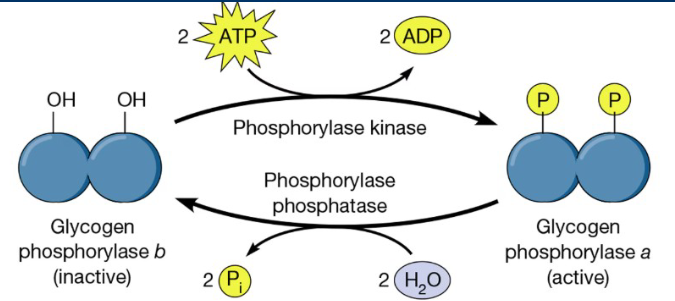
Phosphorylation
Phosphorylation is one of the most common and impactful covalent
modification that affects enzyme activity
• Phosphorylation is the addition of a highly negative phosphate
groups (PO4). The presence of this negative charge can have huge
effects on enzyme activity
• The addition of phosphates are catalyzed by enzymes called kinases
and the removal of phosphates are catalyzed by phosphatases
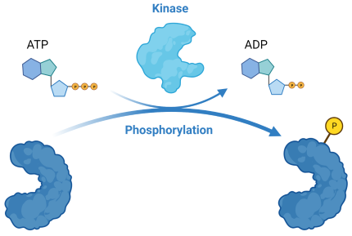
Kinases
Kinases are enzymes that add phosphate groups (phosphorylation)
The addition of phosphates are catalyzed by enzymes called kinases
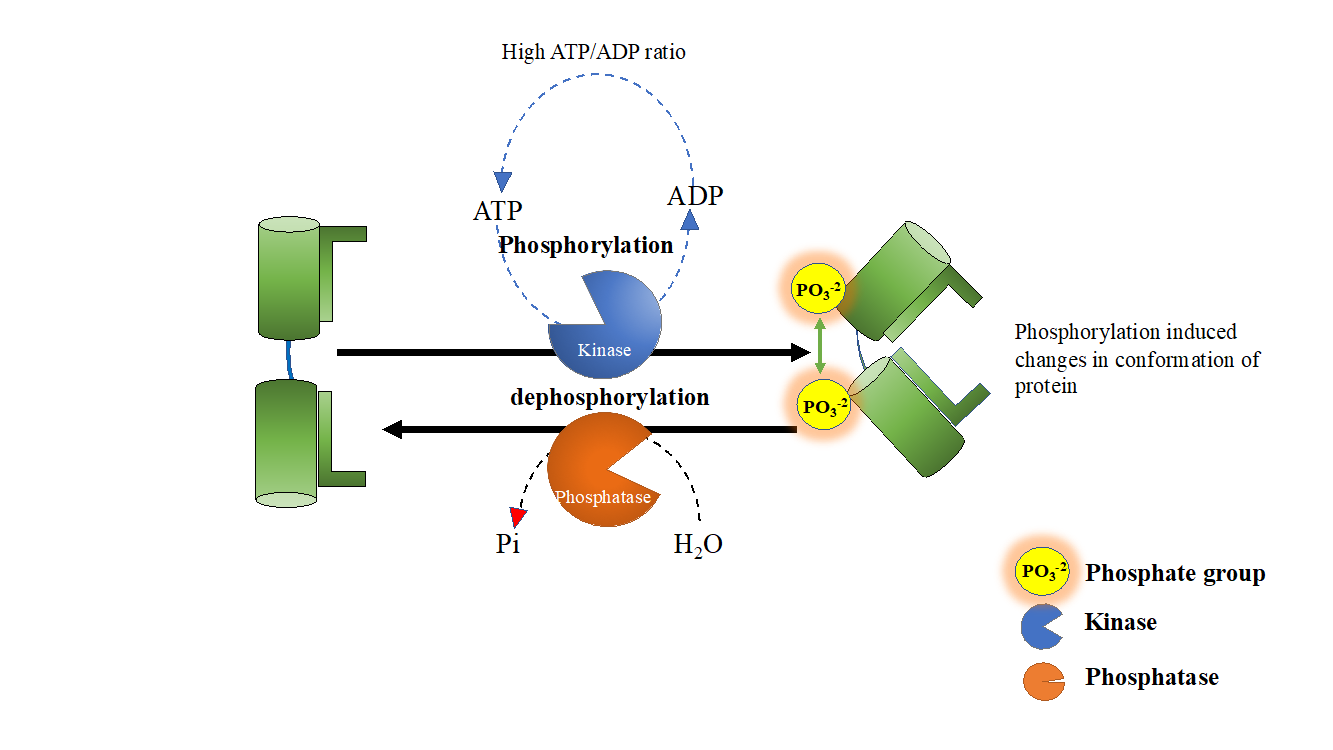
Phosphatases
The addition of phosphates are catalyzed by enzymes called kinases
and the removal of phosphates are catalyzed by phosphatases
Phosphatases are enzymes that remove phosphate groups from molecules (dephosphorylation)
Critical Thinking Questions
1.) Why would a cancer have high incidence but low mortality? Why
would a cancer have low incidence by high mortality?
2.) Why do benign tumors generally have better prognoses than
malignant tumors?
3.) Why is surgery not enough for malignant cancers?
4.) Why can it be difficult to separate environmental and genetic causes of cancer?
1.) High incidence and low mortality it is very common but it is very treatable or potentially slow growing conversely, low incidence but high mortality means the cancer is likely aggressive/ fast growing and not very treatable.
2.) Benign tumors have better prognoses because they don’t metastasize and there for if they are in an optimal location they generally can be removed and there isn’t fear like malignant tumors some cells may have got left behind and it starts growing again
3.) Surgery is not an option for malignant tumors because they metastasize they are not located at just one site in the body rather they spread to different sites and invade healthy cells
4.) Without direct experiment it is difficult to separate environmental and genetic causes of cancer because genes and environment interact so its hard to distinguish the interplay where one starts and the other ends because environmental factors can cause mutations but how do you know what mutations are acquired vs inherited and how genetic predispositions affect how the environment affects you
Critical Thinking Questions
5.) In what ways do the principles of natural selection help explain the
acquisition of tumor resistance to therapeutics?
6.) In what ways would the genetic variability of cancers make cancers
difficult to treat?
7.) Why does it make sense that many inherited cancer susceptibility
genes often function in DNA repair pathways?
5.) Survival and reproduction of the fittest really because some of the cells may have some partial or full resistance and the therapeutics will kill the cells that have no resistance and this acts as a selective pressure meaning the resistant minority repopulates and becomes the majority and now the therapeutics no longer work.
6.) Genetic variability means that cancers would not all react to the same treatment even one cancer can be made up of genetic variable cells so there is no saying it can’t be or become drug resistant which means what works in one may case may not work for another
7.) It make sense that many inherited cancer susceptibility genes often function in DNA repair pathways because these pathways are crucial for removing defects and maintaining genomic stability so them not functioning allows these mutations accumulate which can lead to cancer
> EX: BRCA1/2 (homologous recombination) or MMR genes (mismatch repair) fix errors, and losing them creates genomic instability, allowing cancer-driving mutations to persist and transform normal cells
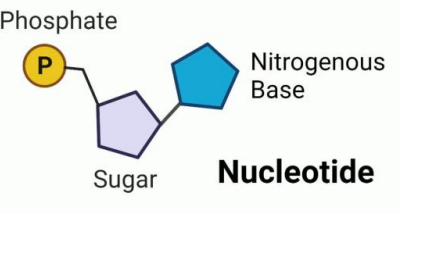
Nucleotide
Nucleotides are organic molecules acting as the fundamental building blocks of DNA and RNA, composed of a nitrogenous base, a five-carbon sugar, and a phosphate group
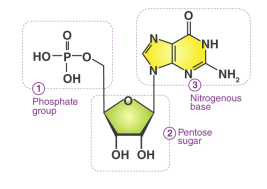
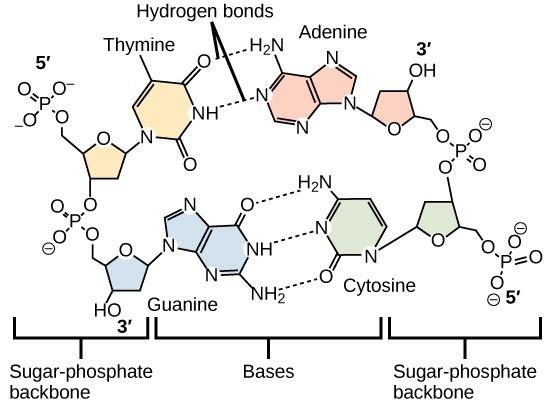
Hydrogen bonds
Hold together complementary base pairs in DNA
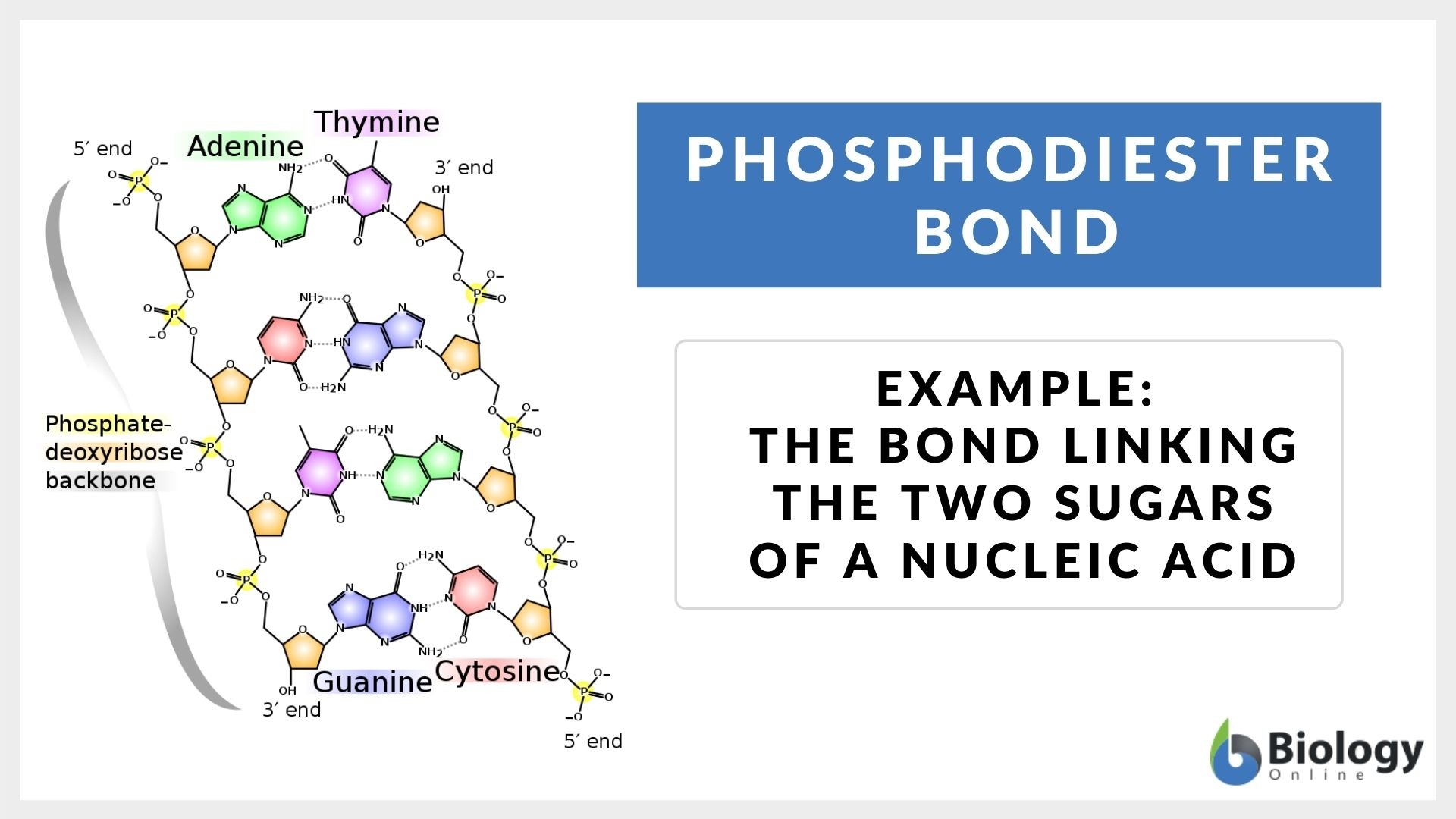
Phosphodiester bonds
Connect adjacent nucleotides through phosphate backbone in DNA
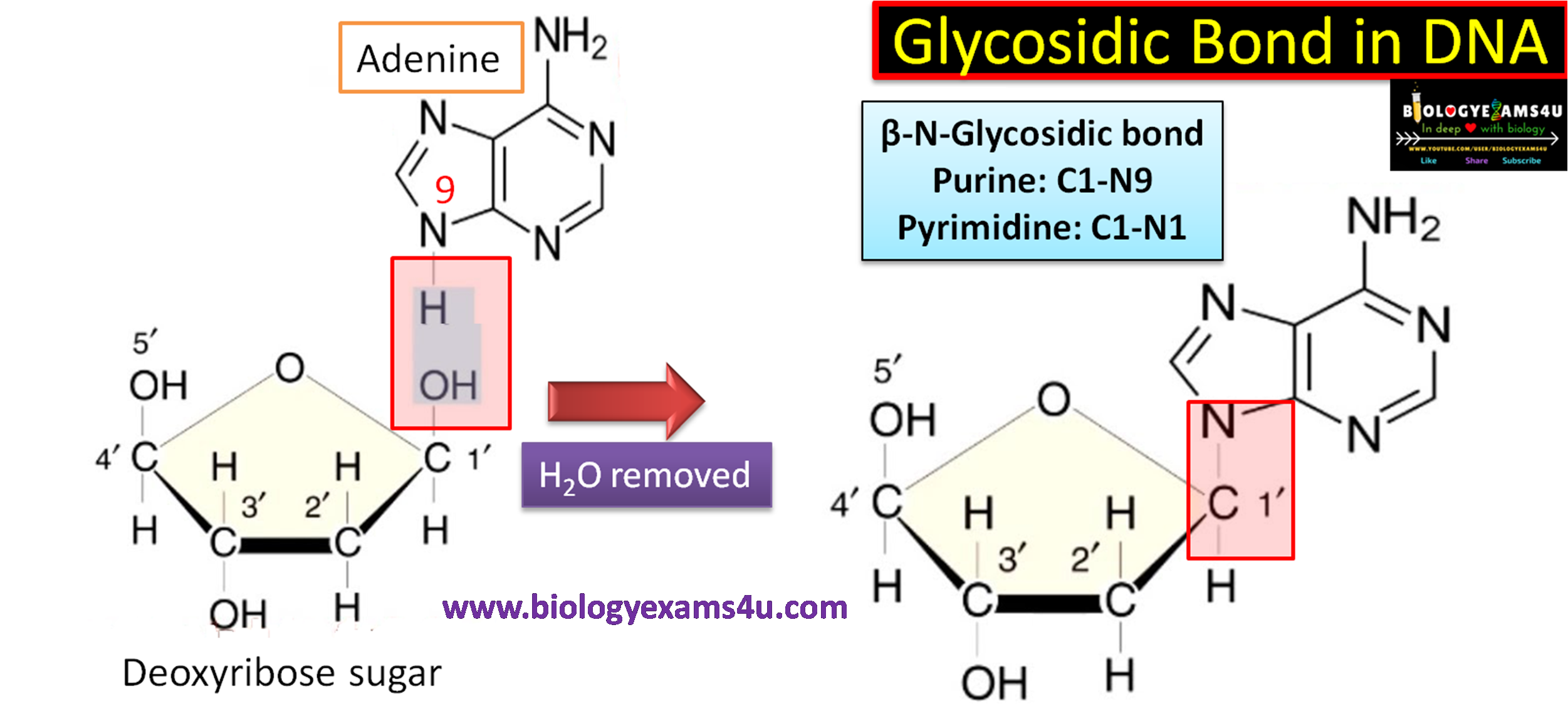
Glycosidic bonds
Connect nitrogenous base to deoxyribose sugar
Chromatin
• DNA strands are organized and compacted
• The organized form of DNA is called chromatin
• Refers to DNA packaged with proteins
• Chromatin is organized at multiple different layers
• Histones are the protein that is most important for DNA organization
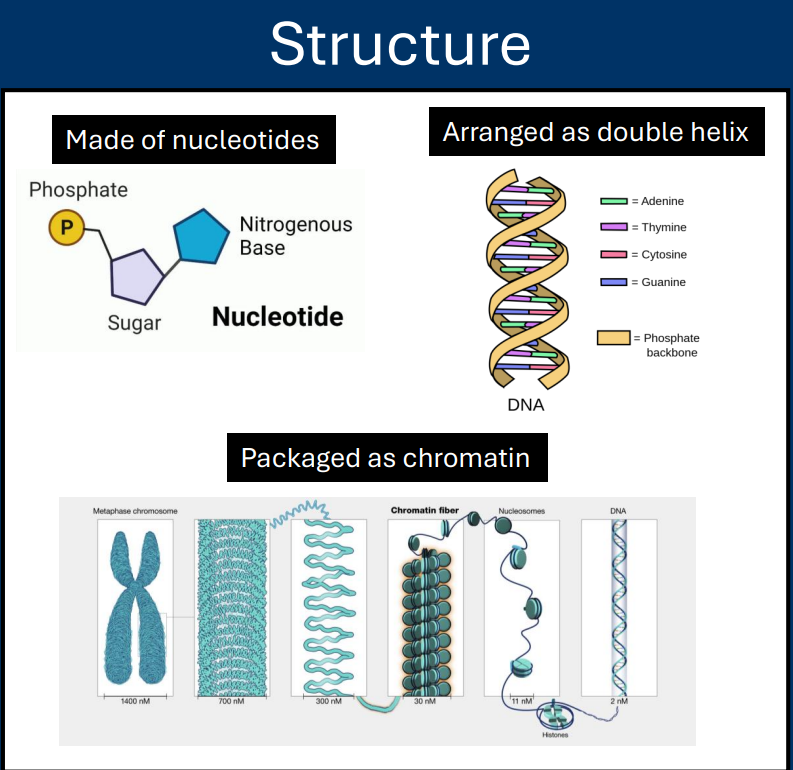
Histones
Histones are the protein that is most important for DNA organization
Each nucleosome includes:
• Histone proteins (positively charged)
• DNA (negatively charged)
• Cells regulate DNA accessibility by altering the charges of histones
Chromosomes
Eukaryotes store DNA in the form of linear chromosomes
Gene
Genes make up a portion of the genome
• A gene is a specific sequence of DNA that encodes for either RNA or Protein
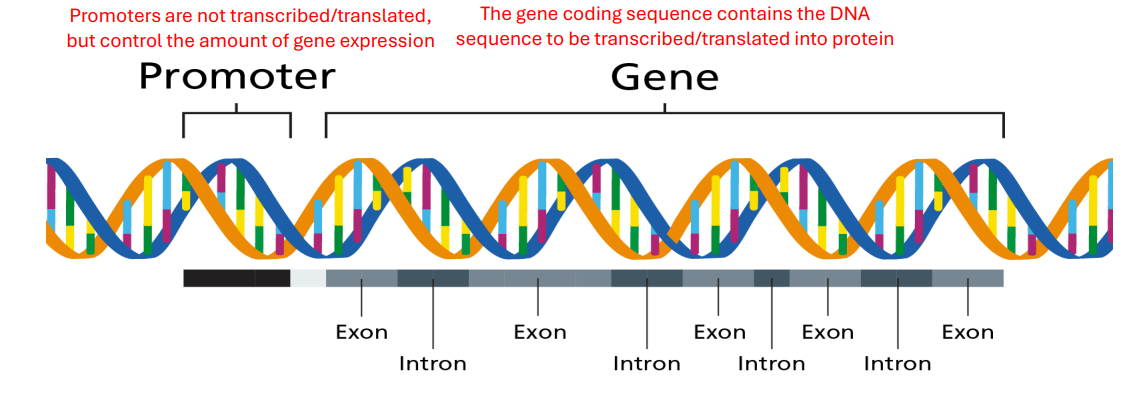
Promoter
Promoters are regions of DNA upstream of a gene that regulate the level of gene expression
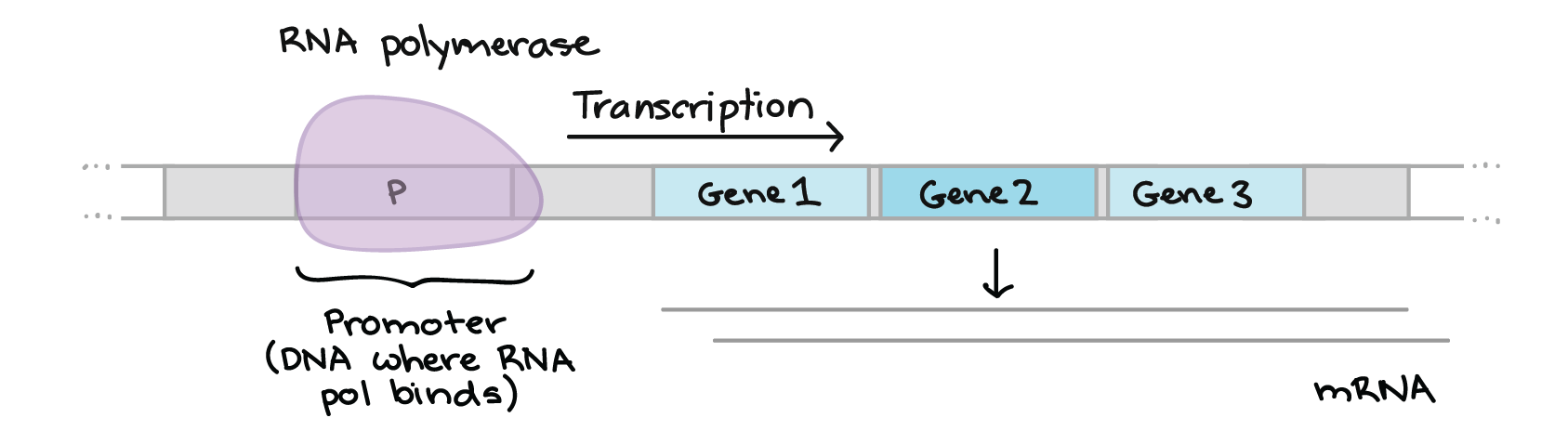
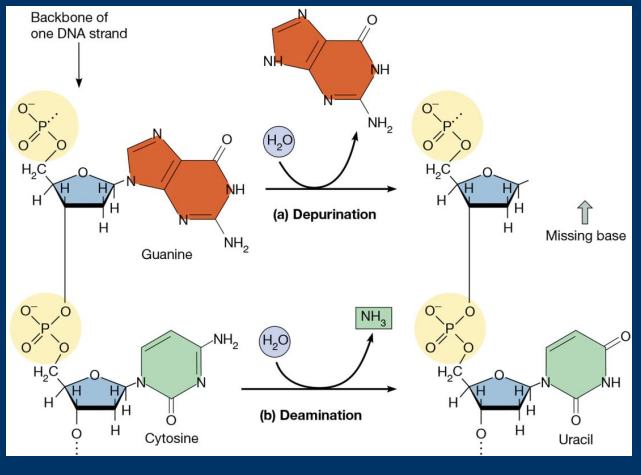
Depurination
• Depurination: Spontaneous loss of purines (Guanine/Adenine)
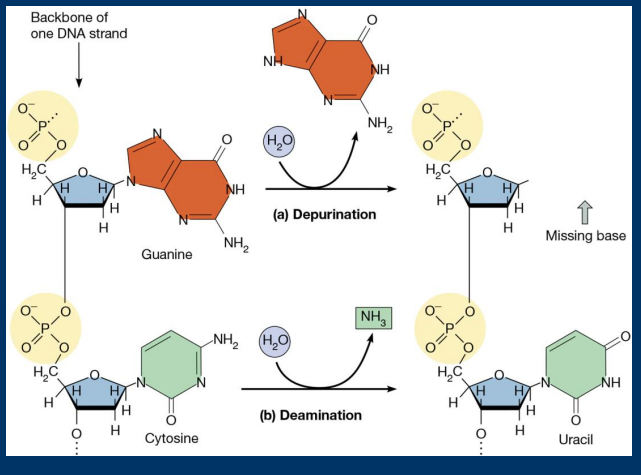
Deamination
• Deamination: Spontaneous loss of an amine groups (converts cytosine to uracil)
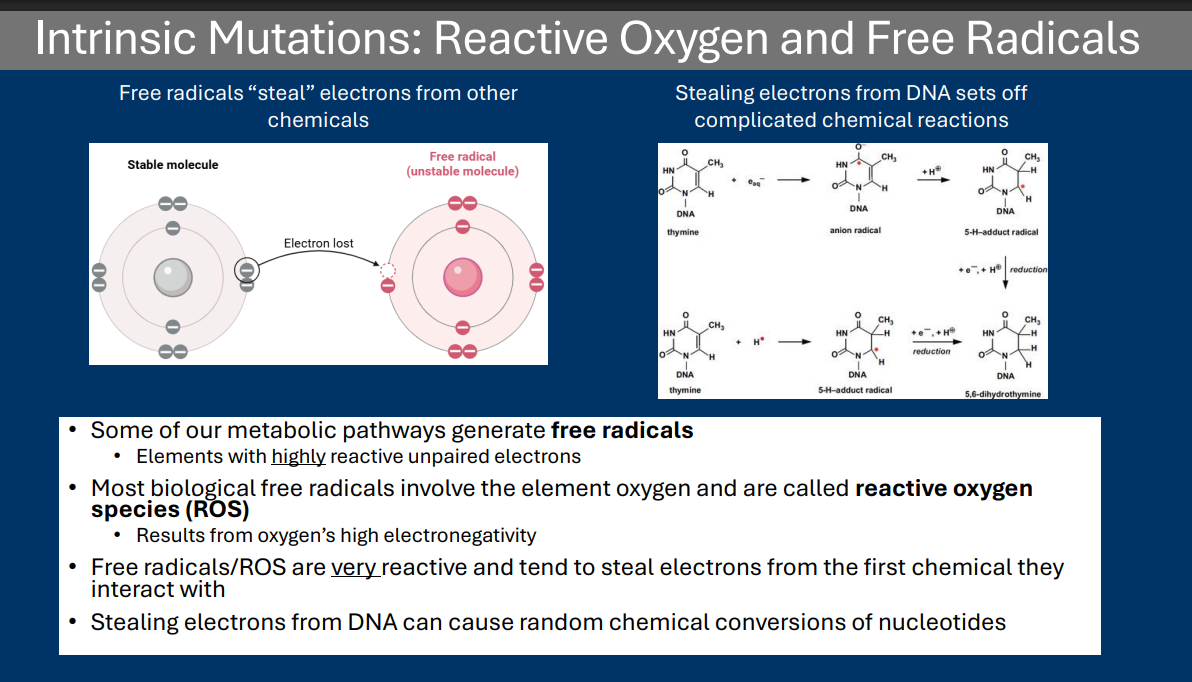
Free Radicals/ROS
• Most biological free radicals involve the element oxygen and are called reactive oxygen species (ROS)
• Results from oxygen’s high electronegativity
• Free radicals/ROS are very reactive and tend to steal electrons from the first chemical they interact with
• Stealing electrons from DNA can cause random chemical conversions of nucleotides
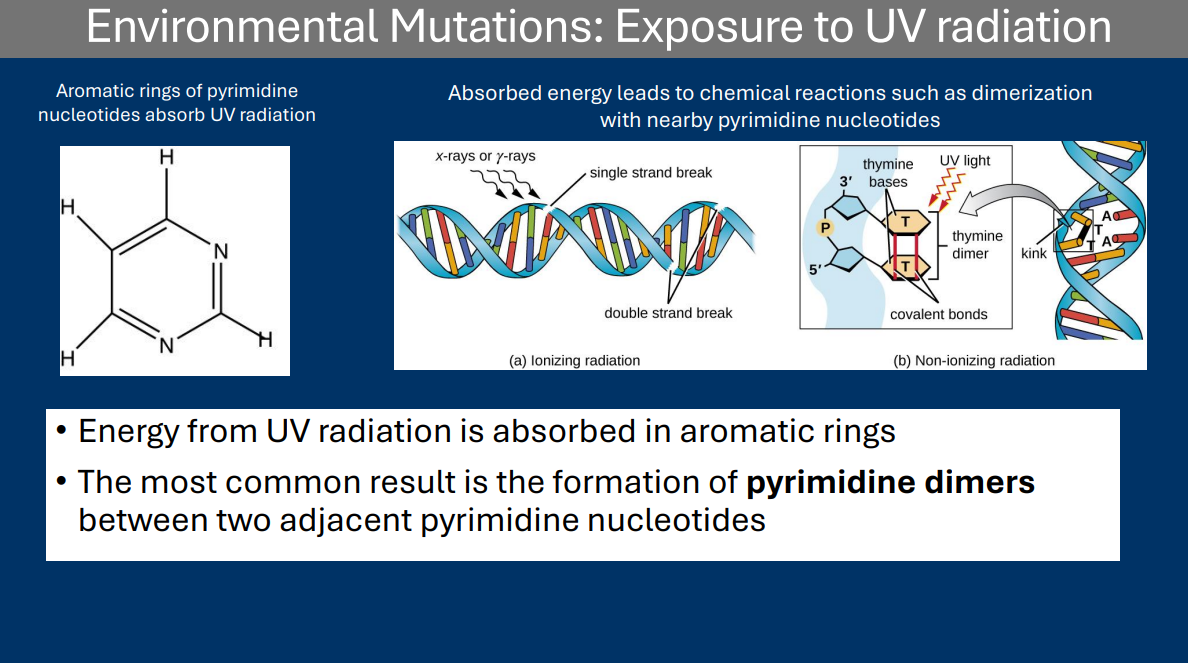
Ultraviolet radiation
• Radiation is a form of energy that can cause chemical reactions in DNA
• Not all forms of radiation have enough energy to cause mutation (e.g. visible light is a form of radiation) • High energy forms of radiation can cause DNA damage
• X-rays can cause damage, but only after long and repeated exposure
• The most common form of cancer-causing radiation are Ultraviolet (UV) radiation
• Classified as UV-A, -B, and -C
• UV-B most commonly associated with carcinogenesis
Pyrimidine Dimers
• Energy from UV radiation is absorbed in aromatic rings
• The most common result is the formation of pyrimidine dimers between two adjacent pyrimidine nucleotides
Hormones/Estrogen
Endogenous hormones can act as carcinogenes
• Our body’s natural chemical hormones can stimulate cell proliferation
• Hormone-related cancers include Breast, Endometrial, Ovarian, Prostate, Testicular, Thyroid
• Estrogen is a hormone that stimulate cell proliferation in breast tissue during pregnancy
• During cancer, estrogen can stimulate cancers cell proliferation
Indel
Indels are mutations that insert or delete a nucleotide
Chromosome Translocation
Chromosomal translocation are major structural rearrangements of chromosomes
Silent Mutation
• Silent mutations have no effect on protein sequence
• Codon redundance
Missense Mutation
Missense mutations affect the identity of a single amino acid
Nonsense Mutation
• Nonsense mutations cause the appearance of a premature STOP codon
Frameshift Mutation
• Frameshift mutations (similar to indels) lead to a change in the open reading frame
• Many frameshift mutations are also nonsense mutations
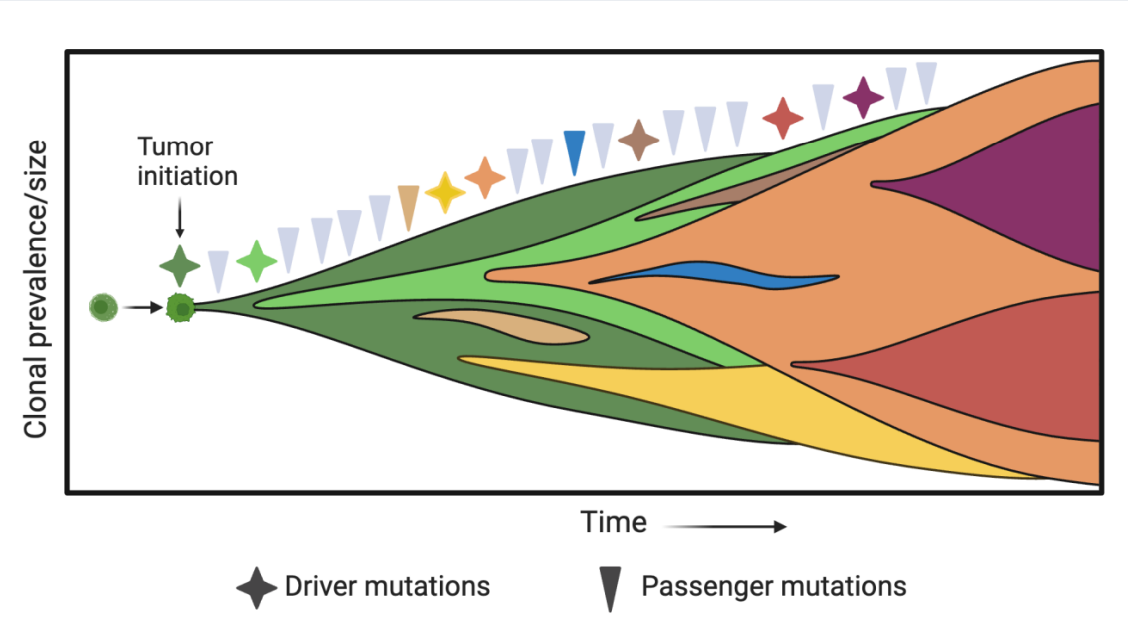
Driver/Passenger Mutation
• Recall that most DNA mutations will have no effect on the development of cancer
• Driver mutations confer a growth advantage (e.g. cause cancer)
• Passenger mutations are acquired during cancer growth but do not affect cancer development
• (confers growth advantage) versus passenger mutations
• ~5-7 driver mutations required for most cancers (may be as high as 20)
•Examples of driver mutations:
• In DNA repair pathways (causes more mutations)
• Heightened cell cycle progression (causes more mutations)
Base Excision Repair
• Base excision repair (BER) corrects small mutations that affect a single base (e.g. deaminations, alkylations)
• Mutated base pairs create small bumps in DNA that are recognized by DNA damage-specific glycosidases that remove mutated base
• Can scan millions of bases per second!
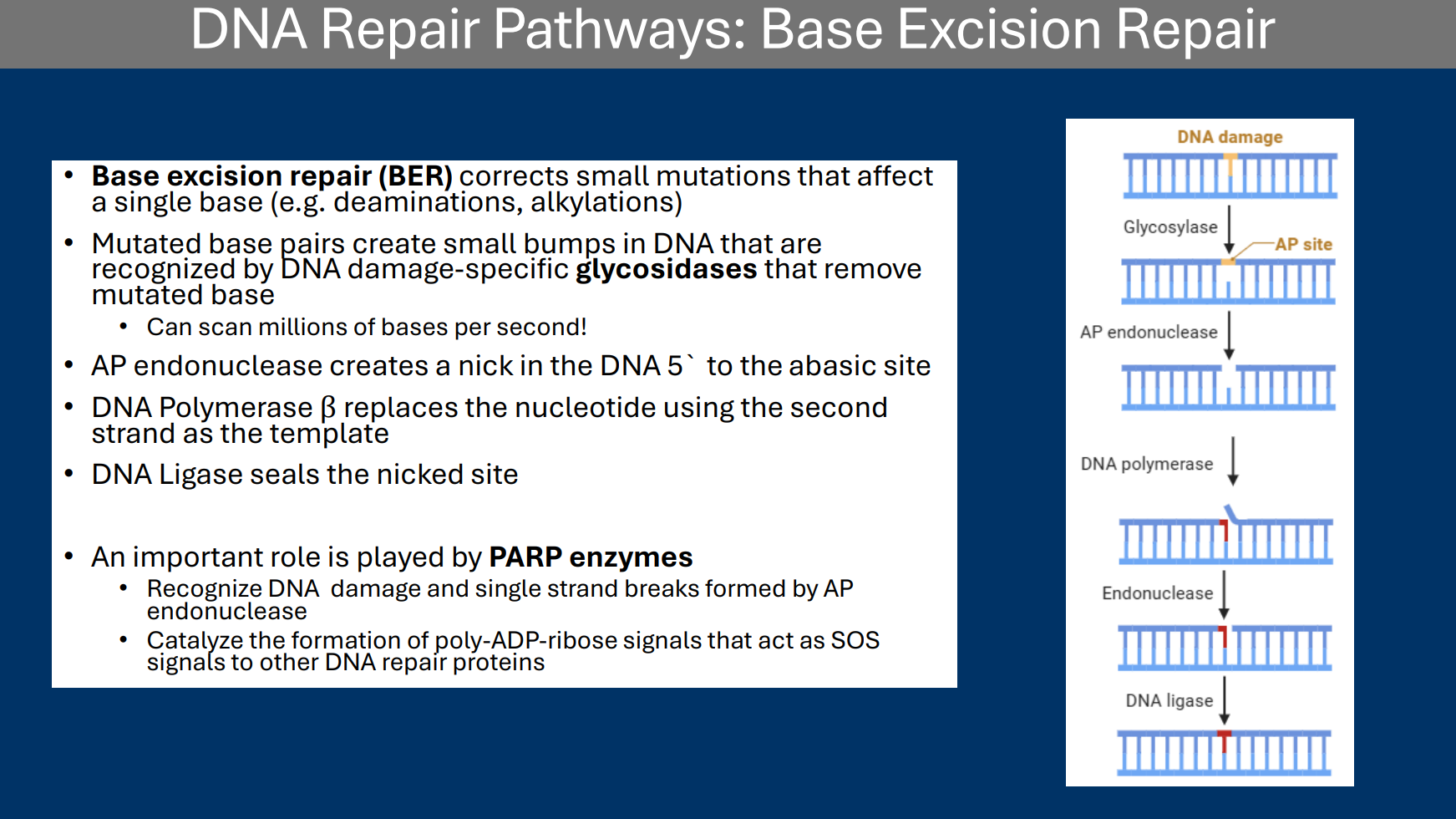

Glycosidases
Enzymes that break down glycosidic bonds
An example is they are used in base excision repair Mutated base pairs create small bumps in DNA that are recognized by DNA damage-specific glycosidases that remove mutated base
PARP enzymes
• An important role is played by PARP enzymes
• Recognize DNA damage and single strand breaks formed by AP endonuclease
• Catalyze the formation of poly-ADP-ribose signals that act as SOS signals to other DNA repair proteins
Nucleotide Excision Repair
• Nucleotide excision repair (NER) is used to correct for larger, helix-distorting lesions (e.g. pyrimidine dimers) and bulky DNA adducts
• During NER the entire nucleotide is removed as well as ~40nt on either side of the mutation
• Accomplished by NER endonuclease
• DNA Polymerase then fills in the removed region using the non-mutated strand
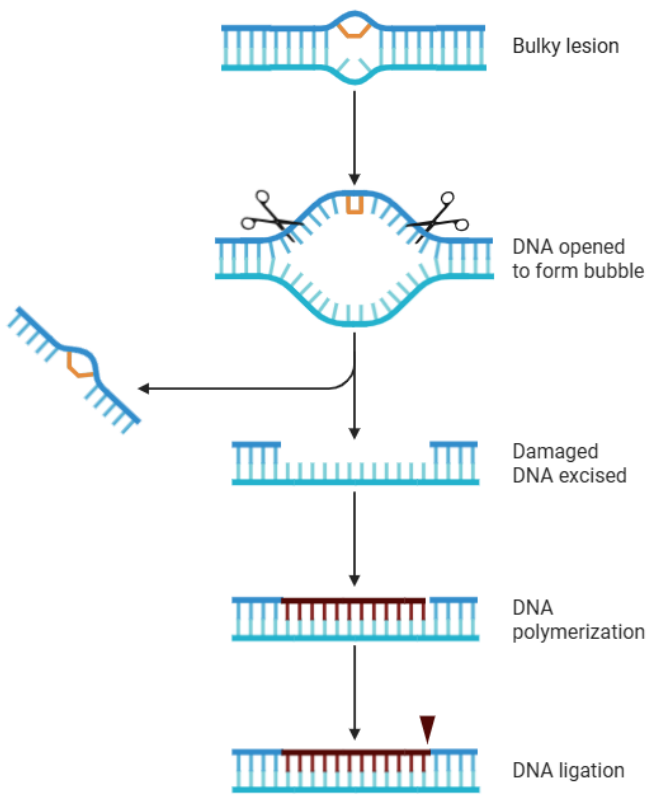
NER Endonucleases
Nucleotide excision repair (NER) endonucleases are specialized enzymes, that execute precise incisions on damaged DNA strands to remove bulky lesions
Mismatch Repair
• Mismatch repair fixes errors caused by DNA Polymerase during DNA synthesis (but missed by proofreading function)
• Base pair mismatches/indels recognized by poor hydrogen bonding
• Regulated by proteins of the hMut family
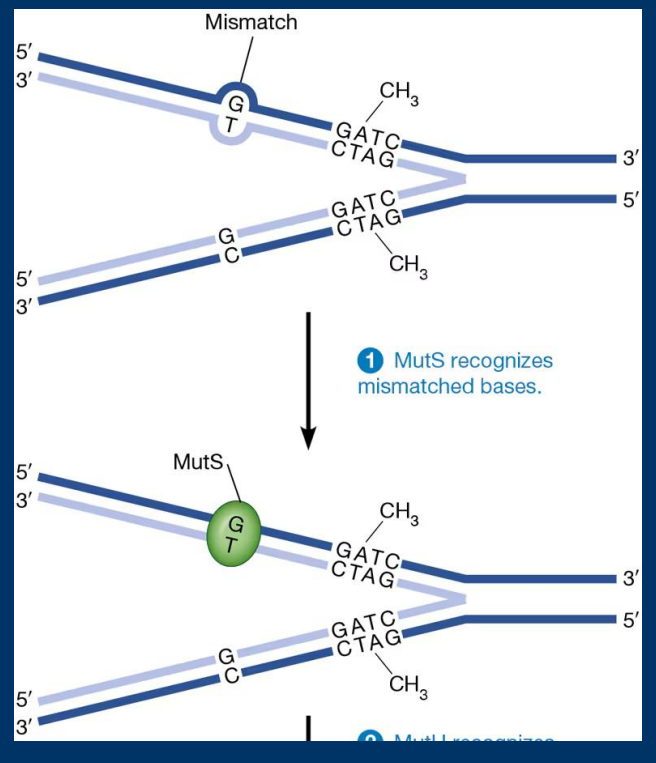
hMut Proteins
They are are essential human DNA repair enzymes that fix mismatched bases and small loops formed during DNA replication. They function in the Mismatch Repair (MMR) pathway by identifying errors detected by MutS complexes, cutting the faulty DNA strand, and initiating its removal to ensure genomic stability.
hMUT (Methylmalonyl-CoA mutase) proteins are essential enzymes, typically human-derived (hMUT), that break down specific amino acids and fats
Double Strand Breaks
• DNA double strand breaks (DSBs) are some of the worse forms of DNA damage
• DSBs mean no template strand is available to help with correction
• Two mechanisms are available to correct DSBs
Homologous Recombination
Homologous recombination uses sister chromatids (formed during DNA synthesis) to direct DNA repair machinery
• Recognition of double-strand break by ATM (ataxia telangiectasia mutated) kinase
• Formation of a Holiday junction with sister chromatids to allow DNA synthesis by DNA polymerase
ATM proteins
ATM (ataxia telangiectasia mutated)
ATM (Ataxia Telangiectasia Mutated) is a vital protein kinase that acts as a "master controller" for detecting and repairing broken DNA strands, particularly double-strand breaks. It resides in the cell nucleus, activating repair pathways, controlling the cell cycle, and triggering cell death if damage is too severe.
Non-homologous End Joining
• Non-homologous end joining (NHEJ) combines together DSBs at random and is therefore more error prone
Critical Thinking Questions
1.) What are the three different types of bonds found in a DNA molecule?
2.)What are the three different sources of DNA mutation discussed in class?
3.) What is the difference between germline and somatic mutations?
4.)What are some examples of intrinsic mutation mechanisms discussed in class?
1.) Hydrogen bonds, phosphodiester bonds, glycosidic bonds
2.)
• Inherited mutations (germline mutations)
• Intrinsic mutations that occur spontaneously during the life of a cell
• Environmental mutagens that occur due to exposure to carcinogens
3.) Germline are inherited (occur in sperm/egg) and in all cells somatic are required and only effect those after it (body cells)
4.)
-Errors by DNA Polymerase
-Spontaneous chemical reactions (Depurination/deamination)
-Reactive oxygen species and free radicals
Critical Thinking Questions
5.) What are some examples of environmental mutation mechanisms discussed in class?
6.) What are the two ways in which viral infection could lead to carcinogenesis?
7.)What are the different ways that mutations can affect DNA sequence?
8.)What are the different ways that mutations can affect protein sequence?
5.)
-Exposure to radiation
-Exposure to UV radiation
-chemical carcinogens
6.)
• Mechanism #1: Virus-encoded proteins may function to increase cell proliferation.
• Mechanisms #2: Incorporation of viral DNA into the genome
7.)
• Transitions convert a purine to a purine or a pyrimidine to a pyrimidine
• Transversions convert a purine to a pyrimidine
• Indels are mutations that insert or delete a nucleotide
• Chromosomal translocation are major structural rearrangements of chromosomes
8.)
• Silent mutations have no effect on protein sequence
• Codon redundance
• Missense mutations affect the identity of a single amino acid
• Nonsense mutations cause the appearance of a premature STOP codon
• Frameshift mutations (similar to indels) lead to a change in the open reading frame
Critical Thinking Questions
9.) What are the four DNA repair pathways discussed in class? What is a protein that is associated with each pathway?
10.) Why do base excision repair pathways require a glycosidase and nucleotide excision repair pathways require an endonuclease?
11.) Why would it be the case that many germline/inherited cancer susceptibility genes play a role in DNA repair?
12.)Why can the induction of DNA damage both cause cancer and be used as a therapeutic for treating cancer?
9.)
• Base excision repair →glycosidases
• Nucleotide excision repair →NER endonuclease
• Mismatch repair →hMut family
•Double stranded breaks →ATM (ataxia telangiectasia mutated) kinase
10.)
Base excision repair (BER) uses a glycosidase because it specifically targets small, non-helix-distorting base damage by cleaving only the damaged base, leaving the sugar-phosphate backbone intact. Nucleotide excision repair (NER) requires an endonuclease to cut out a whole oligonucleotide chunk (containing bulky damage like UV-induced thymine dimers) because these lesions distort the DNA helix, requiring the entire segment to be removed.
11.)
Many germline, inherited cancer susceptibility genes play a role in DNA repair because they act as caretakers of the genome. Inheriting a mutation in these genes leads to deficient repair mechanisms, causing an accumulation of DNA damage and mutations in somatic cells.
12.)
DNA damage acts as a "double-edged sword" because low-level, chronic damage triggers mutations leading to cancer initiation, while high-level, acute damage overwhelms cancer cells' already compromised repair systems, causing cell death. It causes cancer by inducing genomic instability, but treats it by activating apoptosis in rapidly dividing, repair-deficient cells.
Transcription Factors
• Transcription factors (TFs) are proteins that regulate the level of expression for specific genes in the genome
• TFs interact with regulatory DNA sequences and control the production of protein coding DNA sequences
• There are ~3,000 different transcription factors in the human genome
• ~10,000 – 20,000 total genes
• Each transcription factor regulates the expression of many different genes
(Transcription factors determine patterns of gene expression→ transcription factors interact with regulatory DNA sequence and determine how much of a gene is transcribed)
Promoters
Promoters: Found immediately upstream of a gene coding sequence. These sequences are required for gene expression (i.e. they “promote” expression)
-Regulatory regions found in DNA
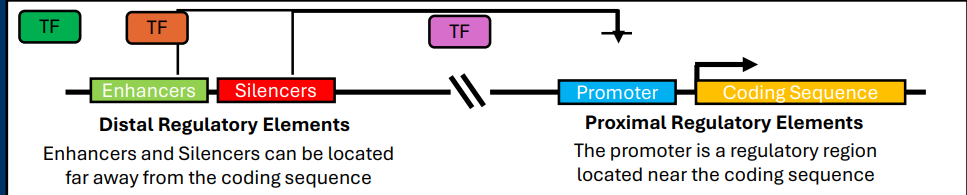
Enhancers
Enhancers: Often located far away from coding sequences in DNA strand. Positively effect (i.e. “enhance”) gene expression
Silencers
Silencers: Located far away from coding sequence in DNA strand. Negatively affect (i.e. “silence”) gene expression
Consensus Sequence
• TFs “recognize” specific sequences of DNA known as a consensus sequence
The presence of a consensus sequence explains why a specific transcription factor regulates some genes but not others
→idealized DNA sequences representing the most common, optimal base at each position to which a specific transcription factor binds
Chromatin Remodeling
*• Chromatin remodeling is the process of regulating DNA accessibility (e.g. euchromatin versus heterochromatin) *
• This is generally accomplished by chromatin remodeling enzymes
• Examples = SWI/SNF or CHD4
• Chromatin remodelers add post-translational modifications to histone tails that affect DNA interaction
• Methylation, Acetylation, Ubiquitination, Phosphorylation
• Know the identity of all four modifications, but we will focus on one: Acetylation
Histone
Histones are essential proteins that package and organize DNA into chromatin within the cell nucleus, acting like spools for the negatively charged DNA to wind around, forming nucleosomes, which compact the DNA and regulate gene activity through chemical modifications on their tails, controlling access for gene expression.
-Histone modifications regulate DNA accessibility
-Histone modifications regulate chromatin remodeling
• Histone proteins have an overall positive charge, and DNA has an overall negative charge the is responsible for histone/DNA interactions
• Chromatin accessibility is regulated by affecting histone/DNA interaction
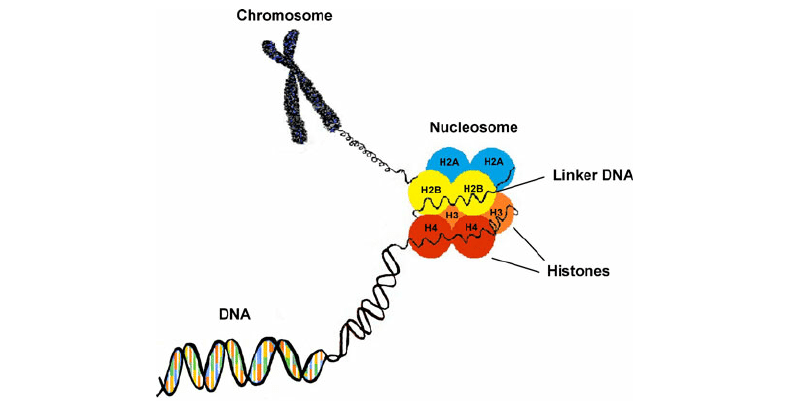
Nucleosome
A nucleosome: Eight positively charged histones wrapped with negative DNA
-The nucleosome is the core of chromatin structure
• The core unit of chromatin regulation is the nucleosome
• Eight histone proteins (called an octamer) wrapped with DNA
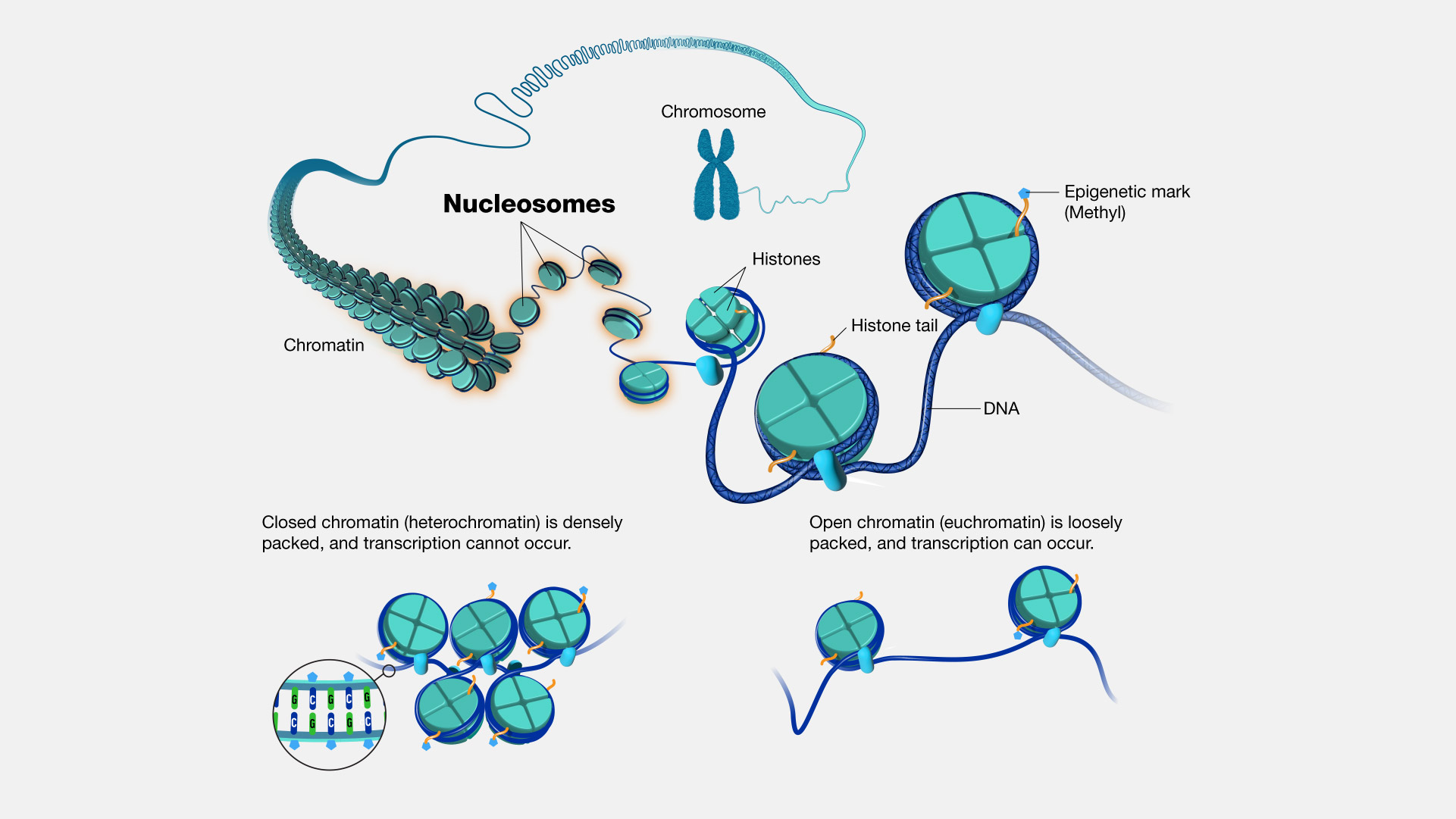
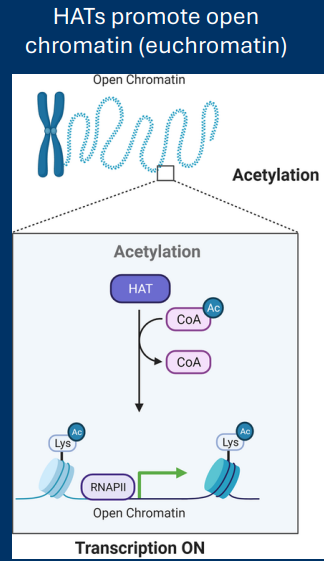
Histone Acetyltransferase
Histone acetyltransferases (HATs)
• Add acetyl groups to histones
• Relax DNA/histone interaction
• Increase gene transcription
-Getting rid of the + = relax
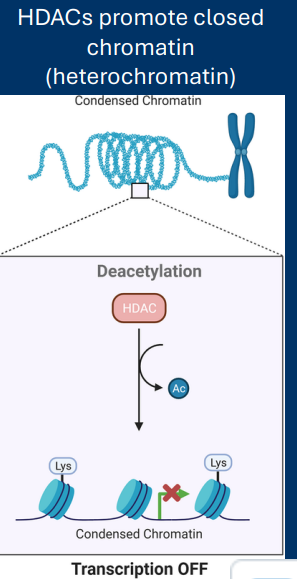
Histone Deacetyltransferase
Histone deacetyl transferases (HDACs)
• Remove acetyl groups from histones
• Increase DNA/histone interaction
• Decrease gene expression
-releases + = condense
->decrease of turn off gene expression
DNA Methylation
• DNA methylation is a common form of epigenetic gene regulation
• Addition of a methyl groups (CH3 ) to cytosine nucleotides by DNA methyltransferase enzymes
• ~3-4% of all cytosines in the genome are methylated
• Most DNA methylation occurs in promoter regions rich in Cytosine and Guanine (known as CpG islands)
• DNA methylation represses gene transcription by preventing transcription factor binding
Hypermethylation/Hypomethylation
• Cancer is associated with a global decrease in DNA methylation (hypomethylation)
• The promoters of many specific cancer related genes are hypermethylated (more than normal)
Ex: in cancer cells you would expect tumor suppressor genes to be hypermethylated because they aren’t suppressing the tumor
and you would expect proto-oncogenes to be hypomethylated because they are on in excess to cause the cancerous phenotype
Telomeres/Telomerase
-The very ends of eukaryotic chromosomes are known as telomeres
→Due to the problems of lagging strand DNA synthesis, telomeres are shortened with each successive round of DNA replication/cell division
• This creates a “countdown” on cell division→ senescence
-Stem cells produce the telomerase enzyme to prevent telomere shrinking during cell division
• Telomerase can enzymatically lengthen telomeres preventing telomere shortening
Critical Thinking Questions
1.) What is the difference between cell type-specific transcriptional regulation and inducible transcriptional regulation?
2.) In what ways are mutations to regulatory sequences different that mutations to coding sequences? In what ways can they be similar?
3.)What are the three different domains that can be found in transcription factor proteins? What is the function of each domain?
1.)
Cell type-specific regulation determines gene expression based on tissue identity using specific transcription factors. Conversely, inducible regulation activates gene expression in response to external stimuli. (Cell type regulation dictates where a gene is active; induction dictates when)
2.)
Mutations to regulatory sequences alter the timing, location, and level of gene expression by disrupting transcription factor binding, affecting gene activity without changing the protein structure. Coding mutations alter the amino acid sequence, often resulting in non-functional or shortened proteins. Both types can significantly impact phenotypic traits and cause disease.
3.)
• DNA-binding domains: Physically interacts with regulatory DNA sequences
• Transcriptional activator domains: Recruits additional proteins to regulate gene expression
• Ligand-binding domains: Allow for stimuli induced activation or repression
Critical Thinking Questions
4.)How could a DNA repair using non-homologous end joining lead to transcriptional misregulation of cancer-related genes?
5.) How can the function of HATs and HDACs be explained in terms of charged interactions between DNA and histones?
6.) In a cancerous cell, how would you expect DNA methylation of a tumor suppressor to be affected? How would you expect DNA methylation of a proto-oncogene to be affected?
7.) Why is the enzyme telomerase normally only expressed in stem cells? What is a mechanism that could lead to its expression in cancerous cells?
4.)NHEJ repairs DNA double-strand breaks in an error-prone manner it directly ligates broken ends , directly causing transcriptional misregulation of cancer-related genes through the introduction of things like chromosomal translocations and just overall introducing pieces that do not go together. It introduces disruptions and potential alterations of regulation sequences.
5.)
(-) and (+)’s attract HATs regulate by adding an acetyl group to the histone tails which blocks the positive charge meaning negative DNA is not as attracted so the state is lax and reduced.
HDACs unblock the (+) (deacetylation) of the histone tails so the (-) DNA is attracted again and is in a tight/condensed state.
6.) in cancer cells you would expect tumor suppressor genes to be hypermethylated because they aren’t suppressing the tumor
and you would expect proto-oncogenes to be hypomethylated because they are on in excess to cause the cancerous phenotype
7.)
Stem cells require high proliferation capacity to regenerate tissues, necessitating telomerase to maintain telomere length. If there is a mutation in the promoter which relieves repression of telomerase it can be created in non stem cells because a non-stem cells has the correct machinery to produce telomerase it just doesn’t it because it does not have the right transcription factor.
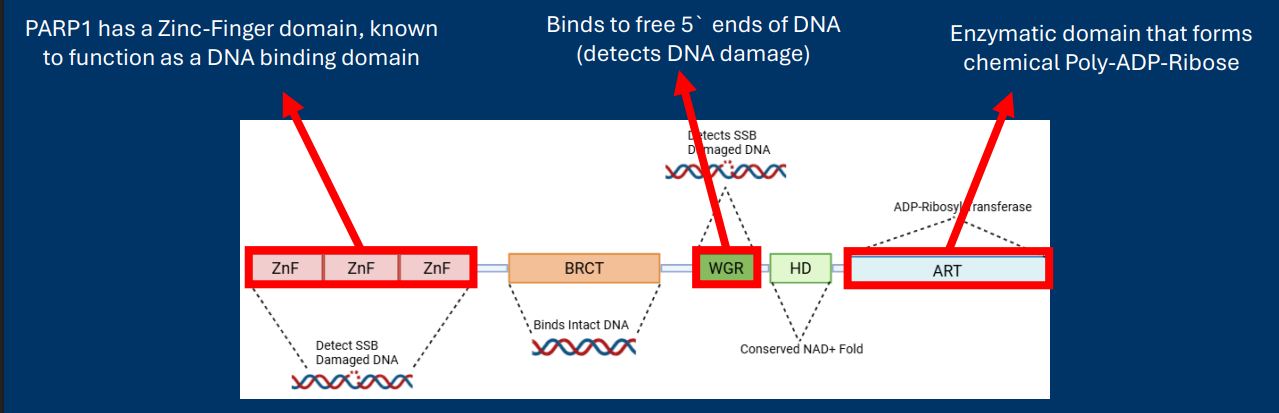
PARP Protein Domain Structure (Journal Club #1)
PARP1 has a Zinc-Finger domain, known to function as a DNA binding domain
WGR Binds to free 5` ends of DNA (detects DNA damage)
ART Enzymatic domain that forms chemical Poly-ADP-Ribose
• A protein domain is a region of primary sequence that can stably fold into a structure with a specific function
• The function of protein domains can often be determined by comparison to other proteins with the same domain
PARP enzymes involved in what DNA Repair Pathways
Base Excision Repair (BER)
(ss strand break/single base damage)
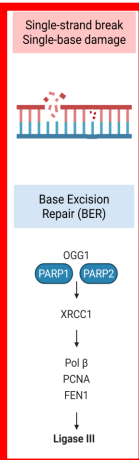
BRAC DNA Repair Pathway
Homologous recombination
(double strand break)
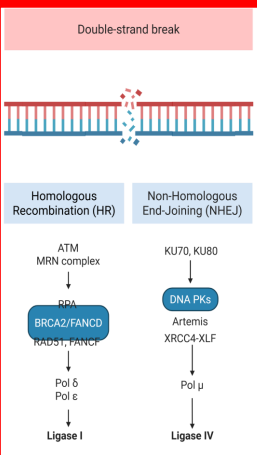
Olaparib is an inhibitor of PARP
Potential mechanism :
It acts by targeting the catalytic PARP domain, specifically blocking the enzyme's ADP-ribosyltransferase (ART) activity, while simultaneously "trapping" the PARP enzyme on the DNA.
synthetic lethality
A synthetic lethal treatment occurs when the simultaneous loss of two pathways leads to cell death
How does Olaparib have a synthetic lethal effect?
• If the PARP system fails, the single strand break will become a double strand break the next time the cell undergoes division
So if olaparib inhibits PARP then it becomes a double stranded break and since there is already a BRAC mutation the cell can repair and it dies
• BRCA mutation carriers already have mutation to one pathway
• This gives the drug specificity towards cancer cells • Olaparib targets the second pathway
• This is an example of targeted therapy
• Therapeutics that are selective for the underlying mechanisms
• Contrast with chemotherapeutics which are less selective
Goals of Phase 1 trials:
Goals of Phase I trials:
• Identify maximum tolerated dose/toxicities
• Determine effective dose
• Preliminary look at drug efficacy is small cohort
maximum tolerated dose
The maximum tolerated dose (MTD) is the highest amount of a drug or treatment. It produces the desired effect without causing unacceptable side effects or toxicity. This is determined during Phase I clinical trials.

How does molecular weight effect where a band would be in gel electropheresis?
molecules are separated by size, with higher molecular weight (HMW) bands located near the top (closer to the wells/cathode) and lower molecular weight (LMW) bands near the bottom (closer to the anode/dye front). Larger fragments move slower through the gel matrix, while smaller fragments travel faster and further.
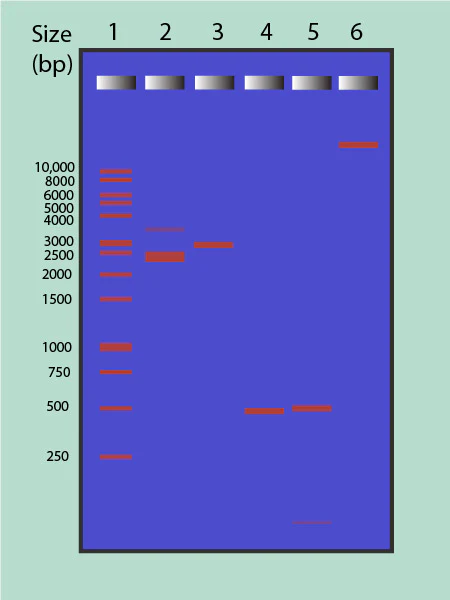
END OF JOURNAL CULB # 1 IF GOING IN ORDER
END OF JOURNAL CULB # 1 IF GOING IN ORDER
Growth factor
• In multicellular organisms, cell division is regulated by growth factors
• Chemical stimuli released by cells that stimulate growth in other cells
• All cancer cells must develop mutations that bypass the need for external growth factors
• Self-sufficiency in growth signals = Hallmark of cancer
(growth factors are signaling proteins, peptides, and other molecules that act as chemical messengers, telling cells to grow, divide, differentiate)
Growth factor receptor
Growth factor receptors (GFRs) are specialized cell surface proteins that bind to specific growth factor ligands, triggering signaling cascades
• Although there are many types of membrane receptors, most growth factor signaling uses receptor tyrosine kinases (RTKs)
• Examples = Epidermal growth factor receptor (EGFR), platelet-derived growth factor receptor (PDGFR), Vascular endothelial growth factor receptor (VEGFR)
Signal transduction
• Signal transduction pathways are the series of events that link an external stimulus to a change in cell behavior
• There are very many different types of signal transduction pathways!
• A “signaling cell” releases some form of stimuli
• Small chemicals
• Peptides
• Ions
• The “receiving cell” responds to the stimuli by changing its function
-Signal transduction pathways link external stimuli to cell behavior
• The endpoint of many signal transduction pathways is the regulation of gene expression by transcription factors in the targeted cell
Ligands
Ligand: A molecule that binds to a site on a larger molecule
>Steroids are ligands/agonists for cytosolic regulatory proteins
Agonists/Antagonists
• Agonist: A ligand that activates the function of the protein to which it binds
• Antagonist: A ligand that inactivates the function of the protein to which it bands
Membrane receptors
Membrane receptor are integral membrane proteins
• Extracellular ligand binding domain
• Intracellular effector domain
• Membrane receptors use conformational changes to transduce a signal across the cell membrane
> Binding on extracellular side changes intracellular side → how the cell “knows” the ligand bound the receptor
(Most chem stimuli are NOT hydrophobic so they cant cross cell membrane meaning they need membrane receptors to initiate transudation pathway)
Second messengers
Receptors activation can produce second messengers, small molecules that diffuse through the cell to affect cell pathways
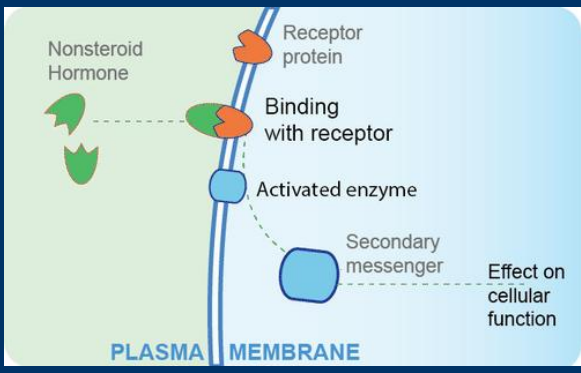
Enzyme cascades
Receptors activation can initiate an enzyme cascade in which a series of enzymes activate one another in sequence leading to changes to cell function
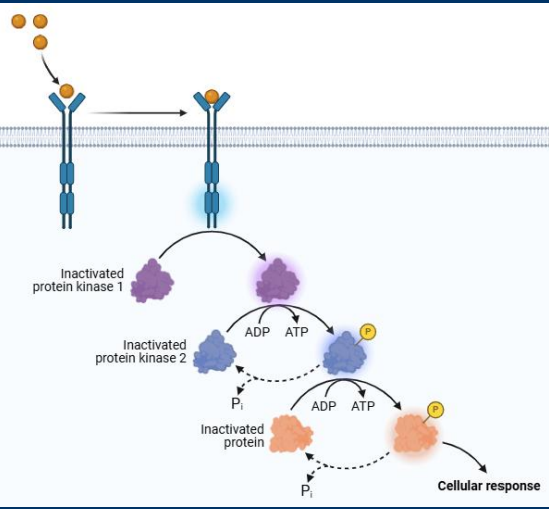
Signal amplification
• Most signaling pathways include both second messengers and enzyme cascades
• Both mechanisms allow for signal amplification
• Each step activates 10s or 100s of events in the next step
• Very small quantities of ligand are often sufficient to elicit a large response from a target cell
(Signal amplification in cell signaling converts weak extracellular signals into robust cellular responses by increasing the number of activated molecules at each step)
Receptor tyrosine kinases
• Although there are many types of membrane receptors, most growth factor signaling uses receptor tyrosine kinases (RTKs)
(Receptor Tyrosine Kinases (RTKs) are specialized membrane receptors that turn external signals (like growth factors) into internal cellular responses such as growth, division, and metabolism)
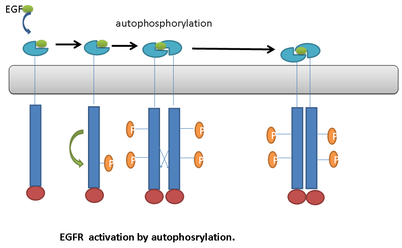
Autophosphorylation
the process where a signaling molecule binds to two neighboring receptor units (monomers), causing them to pair up (dimerize). This proximity enables their intracellular kinase tails to activate each other
-Ligand-bound RTKs undergo intracellular autophosphorylation
-Only when a ligand brings multiple receptors together can autophosphorylation occur
Kinases
• Kinases are enzymes that catalyze the addition of a phosphate group (PO4 ) to other molecules • ~500 kinase genes in genome, and ~20% are associated with cancer!
• When a kinase has activity towards a protein, it generally adds to amino acids with a hydroxyl group
• Tyrosine, Serine, and Threonine
• Kinases signal. By adding a highly negatively charged phosphate to substrates, they affect function, localization, etc.