BIOCH 330 Post Midterm 1 (FINAL EXAM CONTENT)
1/77
Earn XP
Description and Tags
RNA
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
78 Terms
RNA polymerase (prokaryotes)
Protein which 3 Asp residues which coordinate 2 Mg ions to stabilize the 3 phosphates of the NTP. Runs 5’ to 3’ along DNA. Reads the template strand, produces RNA complement to this (the coding strand). Very processive (40 NT/second), can stay on a strand for 16 hours.
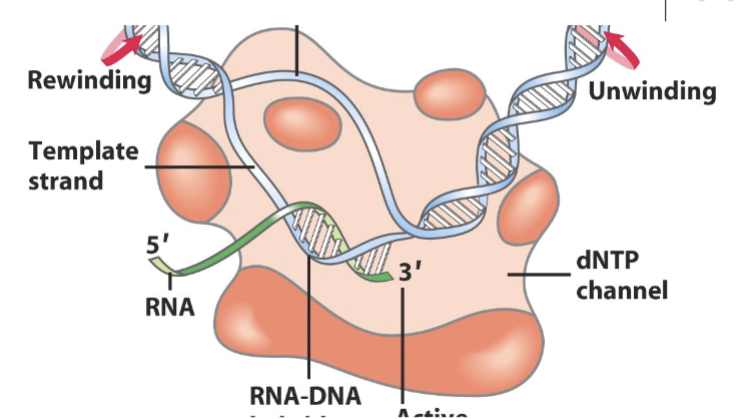
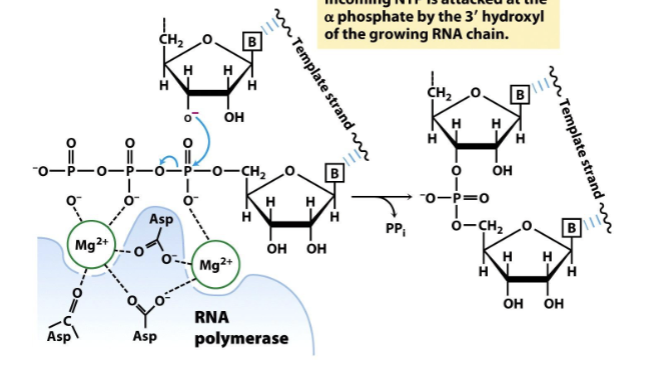
RNA synthesis
3’ OH of growing RNA strand attacks α phopshate of the NTP and attaches it to the RNA strand, producing PPi (ATP hydrolsysis). Always adding to the 5’ end.
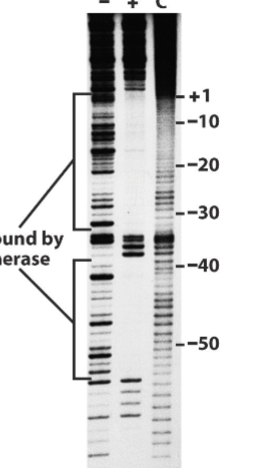
Footprint Experiment
Figured out where RNA poly binded to. Ran a DNAase with two test tubes of labelled DNA (controlled such that it only cut each piece once), a control without poly and the experimental one with it. It then visualized the two experiments using gel electrophoresis. A visible gap appeared in the DNA with the RNA poly, which corresponded to the positions that the RNA poly binds to (-35 to +1 region)
Sigma factors
One of the most important subunits of the RNA polymerase holoenzyme; mainly involved with the specific affinity poly will have to different promoters.
Rho dependent termination
Protein which attaches to DNA and when it reaches RNA polymerase, it bumps it off off of the DNA strand, stopping translation. Uses ATP hydrolysis
Rho independent (intrinsic) termination
70-80% of termination. Long sequences of G-C (palindromic) repeats, which cause RNA poly to pause. While this happens, the strand forms an internal hairpin loop, which interacts with polymerase bound NusA protein. Behind G-C repeats, A rich sequences bind with repeating U sequences in the RNA, which allow for dissociation due to the new stress from the loop and the weaker bonding between U-A.

Eukaryotic RNA polymerase characterstics
Multiple polymerases (I, II and III). Require specific promoter sequences (TATAAA box), binding proteins (TBP), and transcription factors (TFIIA, TFIIB, TFIIH, TFIIF, TFIIE etc). Increased complexity for increased regulation (compared with prokaryotes)
RNA pol I (euk)
For (pre forms) of large rRNA transcription; found in the nucleolus
RNA pol II (euk)
For (pre forms) of mRNA and miRNA transcription
RNA pol III (euk)
For (pre forms) of tRNA transcription and 5s ribosomal RNA (rRNA)
TATA binding protein (TBP)
Eukaryotic protein which recognizes and binds to the TATA box, bending it into a horseshoe shape. Facilitates the assembly of the pre-initiation complex. Attached to the larger TFIID complex
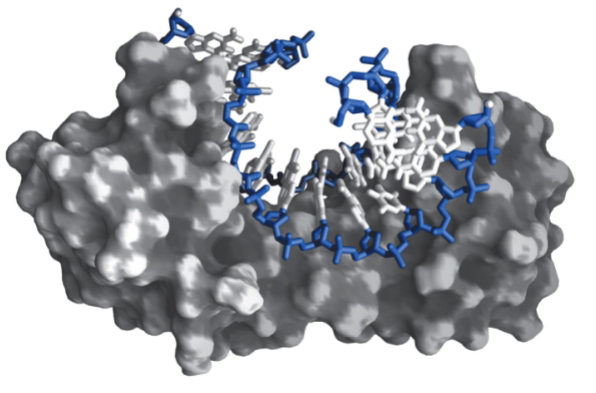
TFIID
An RNA pol II transcription factor. Massive protein complex that contains TBP within it, along with transcription associated factors (TAFS). TAFS includes proteins which act as specific promotors and proteins that interact with basal machinary for stabilization.
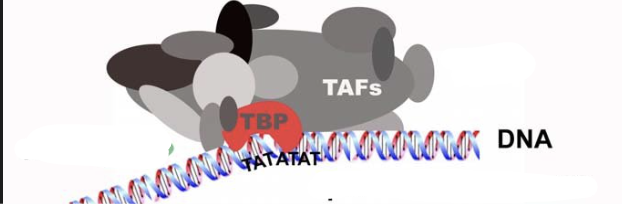
Eukaryotic RNA transcription promoters for initiation
Requires TFIIA + TFIIB and TFIID (Within it TBP) attached to Pol II which is then attached to TFIIH, TFIIF, and TFIIE
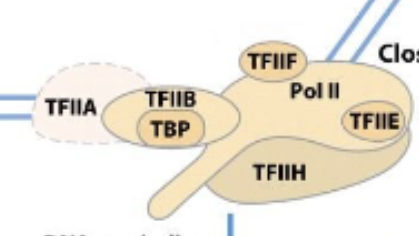
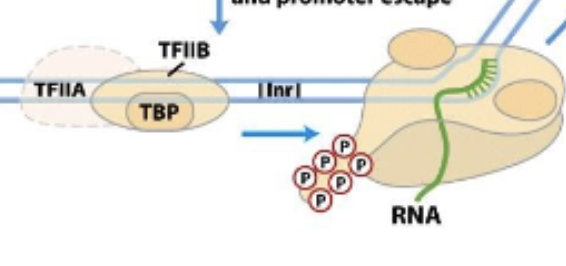
Eukaryotic RNA transcription promoters escape (part 1)
Pol II is phosphoryalated after being successfully attached to DNA strand and starting transcription. TFIIA, TFIIB (with TBP) are then left behind/pushed off by phosphates.
TFIIH
In transcription initiation, acts as a helicase. Also communicates with DNA repair machinary (as it is one of the first things to come into contact with the DNA). Responsible for phosphorylating the C-terminus end of RNA pol II.
Phosphorylation of Pol II
Important for proper elongation and recruitment of other transcription factors
Eukaryotic RNA transcription promoters escape (part 2)
Pol II begins elongation of RNA strand and loses TFIIH and TFIIE. Remains phosphorylated until termination
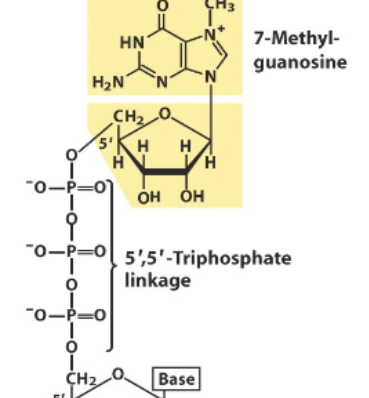
5’ capping
Eukaryotic mRNA feature, where a terminal Guanine at the 5’ end is modified into 7-methylguanosine (done by methyltransferase). Causes there to be a 5’ to 5’ Triphosphate linkage. Protects RNA strands from being degraded by 5’ exonucleases (present roaming the cells, to protect against foreign invaders) and also aids in effecient translation
Influenza key features
Viral envelope (made of 2 glycoproteins); negative strand RNA virus (has to be copied in order to synthesize any proteins); segmented genome and RNA dependent RNA polymerase. Often do not contain a 5’ cap (so susceptible to being degraded by 5’ exonucleases - unless partakes in cap stealing)
Cap stealing
Method by which viral RNA can steal a cap from host mRNA. Causes the host mRNA to be down regulated (transcribed less), protects the invading viral RNA and causes it to be more effeciently translated to express its viral proteins.
Polyadenylation
Done via polyadenylate polymerase = PAP (ATP → PPi + rna-A(n)). Signal sequence AAUAAA at 3’ end recruits CPSF (cleavage poly-A specificity factor) which contains an endonuclease and directs cleavage of the RNA strand downstream. Endonuclease then recruits PAP, to add 100-250 A residues by ATP hydrolysis. A-tail is bound in protective proteins (Poly-A binding proteins = PAB) which protects the RNA from 3’ exonucleases.
RNA vs DNA reaction speeds
RNA is usually faster becuase the 2’ OH can participate in cleavage of the phosphodiester linkage in forming a 2’, 3’ cylcic phosphodiester. This is intermolecular, and thus can lead to faster hydrolysis than possible in DNA
Bacterial DNA methylation significance
Protects the DNA from being cut by endonucleases, whereas phage DNA is commonly unmethylated and therefore is cut up by restriction enzymes within bacteria in order to destroy it.
Restriction mapping
Gives information about where specific restriction enzyme binding sites are on a piece of DNA. Does not actually give info as to what the sequence of DNA is (no NT information)
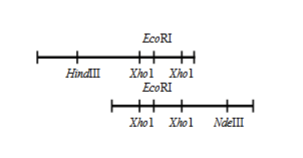
Mapping viral gene structure
Gives information on what proteins a viral piece of RNA will produce (very useful to know, can tell us the function/attack method of the RNA)
Gene structure analysis
Experiment where ssDNA (template) was paired back up with its mRNA strand. In theory, this should have formed a perfect duplex. What actually happened was a bunch of the DNA was “extra” in seemingly random sections all along the helix, which caused it to loop out before reconnecting back to the RNA. Led to the discovery that a large portion of DNA was actually non-coding = genes were not continuous = discovery of introns. (Done with Adenovirus genome)
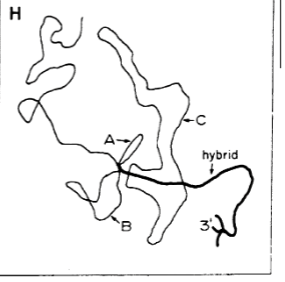
Adenovirus uniqueness
Most viruses/bacteria/simple organisms in general only contain exons. This virus was different due to it containing introns, allowing the discovery to be applied to more complex organisms
Group I and II Introns
Self splicing (autocatalytic). No proteins required.
Spliceosomal introns
Large protein complex required to remove them. Occur in nuclear mRNA. ex. Adenovirus introns
Unnamed introns
Certain tRNA’s which require ATP and an endonuclease for removal. (Not talked about in any detail)
Majority of vertebral genes (including humans)
Is devoted to introns (as much as 80% = 80% of DNA is non-coding and only there for regulation). Can be 50 - 20000 bp
Only vertebral genes without introns
Histone (DNA winding/storage) and interferon (immune system signalling)
Tetrahymena intron removal (GROUP 1)
Two sequential transferesterifications:
G (cofactor) attacks Exon-U-Intron (2’ or 3’ OH of G attacks the 3’ P of the U). Releases an Exon-U and forms new G-Intron-U-Exon RNA strand
Exon-U attacks G-Intron-U-Exon (3’ OH of U attacks the 3’ P of the U-Exon). Forms Exon-UPU-Exon and G-intron-G
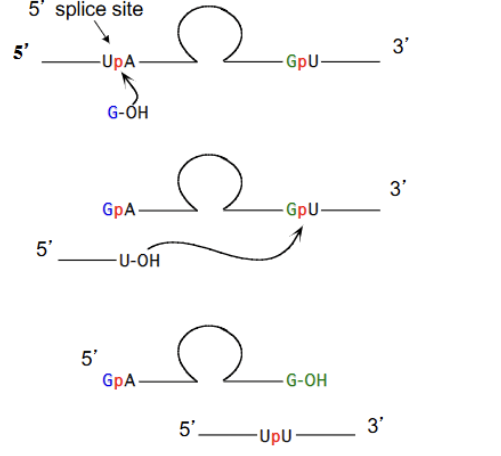
Issues with Group 1 intron mechanism
Thought to be not possible that RNA could autocatalyze, as the rxn occured very quickly (alone, with linear RNA it took thousands of years to occur) and at incredibly specific locations to give the proper introns and exons.
Solution to Group 1 Introns mechanism (how it occurs?)
RNA folds up into specific, enzyme-like strucutres, that enable the reaction to take place both effeciently and at the specific location.
Rolling replication
Involves rna dependent RNA polymerase going around and around circular RNA, generating a ton of linear copies of the RNA. These linear copies form haripin loops via complementary watson crick base pairs. Hammerhead ribozymes can then form and self-cut the RNA at specific places, regenerating circular RNA loops. Used by viroids to replicate (-) RNA into (+).
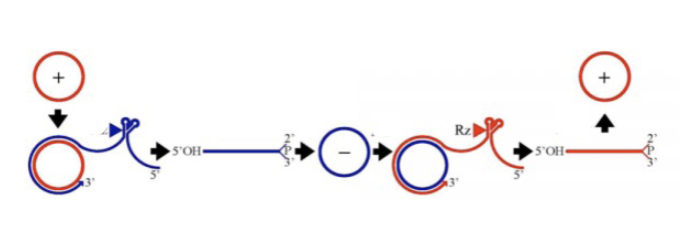
Hammerhead ribozymes
RNA acting as an enzyme (catalytic). Small RNA motif that forms via 3 stems around a core of non-complementary bases. Autocleaves with the help of a Mg ion, forming a 2’,3’ cyclic phosphate (same as alkaline hydrolysis).
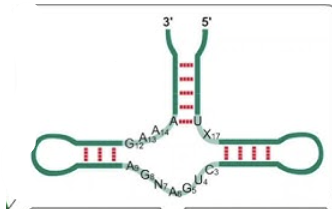
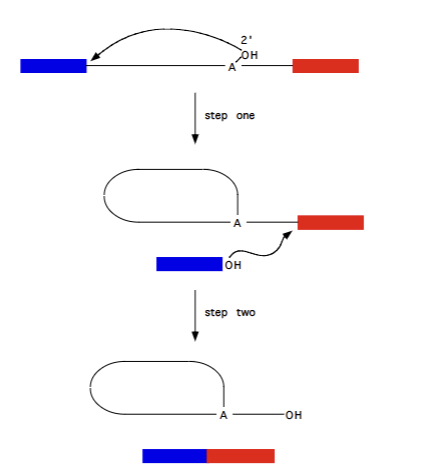
Group II selfsplicing introns
pre-RNA of fungal and plant mitochondria, as well as majority of chloroplast introns. Require Mg2+ , no cofactor. Procedes via formation of lariat structure (lasso = RNA attacking itself to release 5’ exon, and then 5’ exon attacking to connect wtih 3’ exon, releasing intron in lasso shape). Reversible (somewhat)
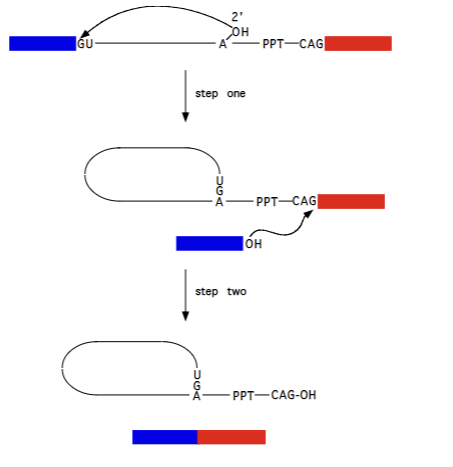
Pre mRNA splicing (splicesomal introns)
Occurs using hte same mechansm as Group II splicing, just requires proteins to facilitate it and uses slightly different molecules (specific sequences, A residue). Lariat structure still formed, 5’ exon attacking 3’ exon to form complete functional exon etc. Irreversible (largely)
snRNP (Small nuclear ribonucleoproteins).
Intermediate-sized RNA strands combined with a protein. 5 that we discuss (U1, U2, U4, U5, U6). Uridine rich.
SM ring (snRNP core)
Protein heptamer of B, D1, D2, D3, E, F and G subunits. Individual units form together following ATP hydrolysis and SMN complex, in stepwise fashion (can self assemble, but not viable due to the potential for 6-ringed and 8-ringed products). snRNA strand fed through the centre hole. A platform around which the SnRNP’s can be organized/fed through.
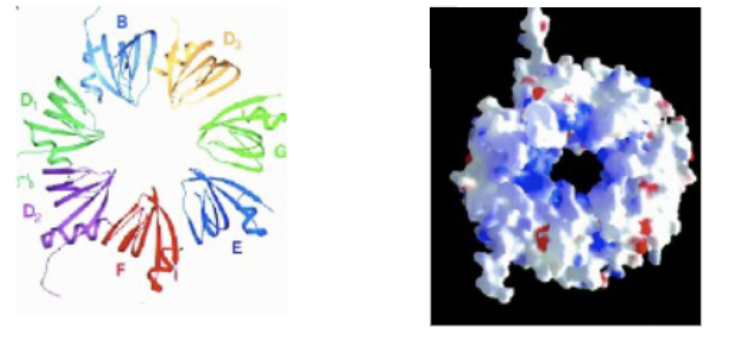
SMN (Survival motor neuron)
Complex which aids in the formation of the SM ring protein. Mutations to this structure cause SMA (Spinal muscular atrophy)
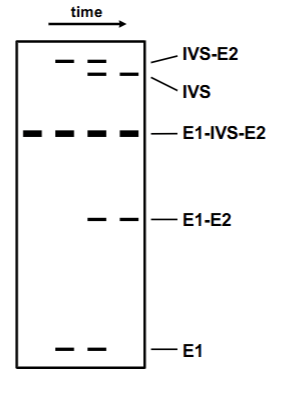
In vitro splicing denaturation experiment
Isolated nuclear RNA (no proteins present) + urea + HeLa cells → splicing reaction able to proceed. Shows the order in which the mechanism occurs. Notes: Due to lariat formation IVS+E2 and IVS by itself is “larger” than E1+IVS+E2 in gel.
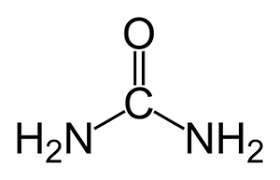
Urea
Denaturing agent of both protein and RNA by outcompeting for H-bonds.
5’ Splicesomal intron site
GU (A attacks here)
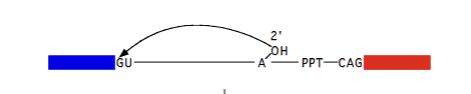
3’ Spliceosomal intron site
AG (Exon attacks here)
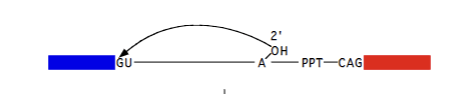
Poly pyrimidine tract (PPT)
C/U containing strands of NT of varying lengths (6-20 NT); characteristic of an intron
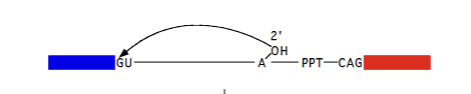
Branch region
UACUAAC (wack-wack). Characteristic of the intron. U2 binds here and causes the A to be pushed out and form lariat structure in Pre-mRNA splicing mechanism.
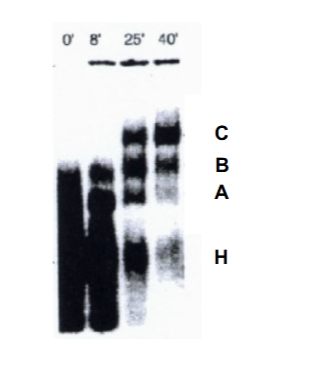
Complex formation RNA experiment
Protein + RNA. Represents the time delay it takes for proper factor association to form snRNP = STEPWISE PROCESS. A is precursor to B, is precursor to C. C is the one which actual splices (causes the splicing to occur late). H=heterogenus mixtures.
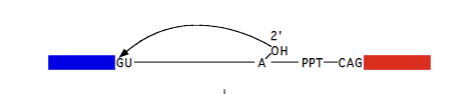
First phase of spliceosomal rearrangement
U1 associates with GU sequence via the internal RNA base pairing. U2 associates with UACUAAC sequence and AG via complementary internal RNA base pairing. UACUAAC imperfect (6 RNA residues for 7 intron residues = A remains unpaired, allows it to attack GU)
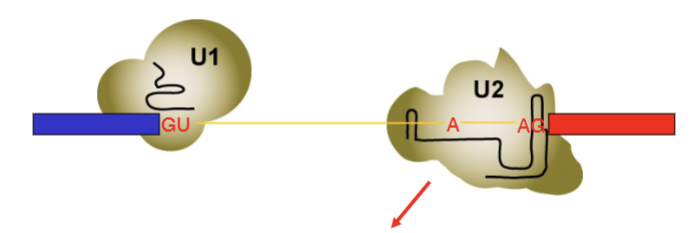
Second phase of spliceosomal rearrangement
U1 is kicked out by U6, U4, U5. U6 now interacts with GU via complementary internal RNA base pairing.
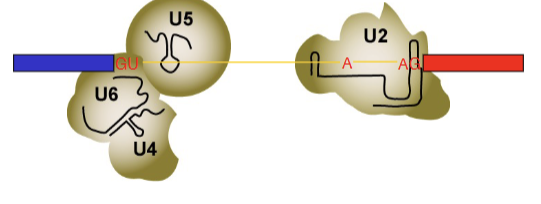
Final phase of spliceosomal rearrangement
U4 gets kicked out by Brr2 enzyme and ATP. U5, U6 and U2 combine together into one dense protein, mRNA structure (Active complex = C complex). Formation of this complex causes the splice to occur (allows the chemistry to happen), after which the lariat structure is able to form.
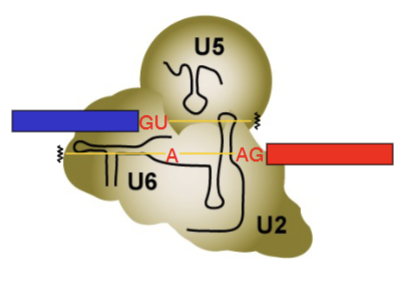
Brr2 (Bad response to refrigeration)
Enzyme involved in spliceosome formation. Helicase which uses ATP to unwind U4 (kicks it off) and also brings together U2 and U6.
Spliceosome proteomics key feature
Across life (between humans and yeast for example) they are extremely well conserved (~90%). This tells us that these proteins are cruicial for any kind of life to occur.
S100 experiment
Taking three test tubes of substancs and seeing if splicing occurs when snRNP’s are added.
Nucelar extract: Splicing
S100 (cytoplasmic extract): No splicing
S100+ (SR proteins from NE + S100): Splicing
Proved that SR proteins are required for any splicing to occur (phosphoserines)
How is S100+ prepared?
Taking Nuclear extract + MgCl2 + spinning. This forms a pellet. Adding this pellet to S100 gives the final product.
U6 stem loop
Highly conserved region of the active site of the C complex (active complex). Critical for function (knocking it out kills yeast cells) because this is where the two Mg ions bind to.
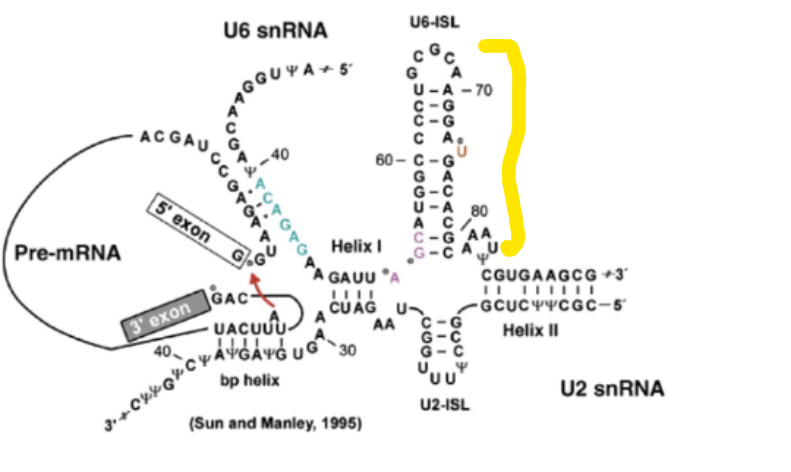
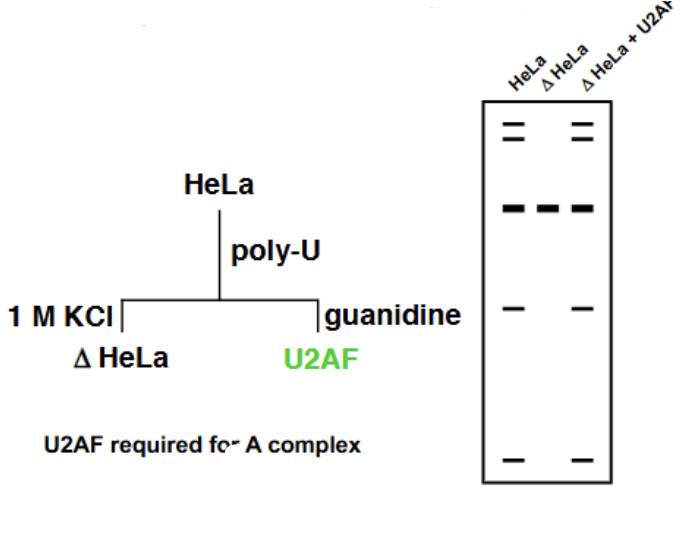
Pyrimidine tract experiment
Took HeLa cells and ran them on a gel containing Poly-U (phosphate), which mimicked the poly-pyrimidine tract and also contained a negative charge. Washed with salt (postivie charge) to remove all the loosely connected substances from the Poly-U gel, giving ∆HeLa cells. No splicing was able to occur with these cells (means that something critical for splicing was still bp with the Poly-U). When the Poly-U was washed again with guanidine, a new factor U2AF was discovered. U2AF + ∆HeLa = splicing. Proved that U2AF was essential for splicing (and likewise, so was the polypyrimidine tract)
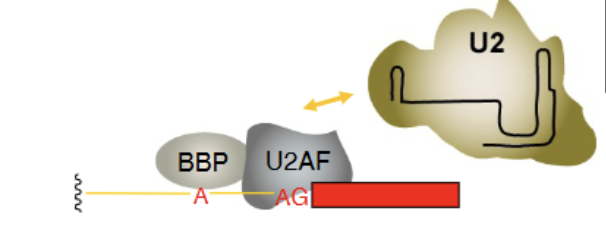
Precursor proteins for U2 snRNP recruitment
BBP (branch binding protein) and U2AF (U2 auxillary factor). Essential
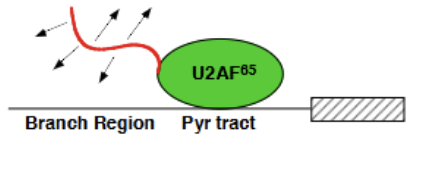
U2AF
U2 Auxillary factor. 2 subunits = heterodimer = U2AF65 and U2AF35. Associates with both the poly-pyrimidine tract (more critical = U2AF65) and the 3’ exon splice site (U2AF35 = less critical).
U2AF65
Contains an effector end (positively charged string portion at N-terminus which recruits the negatively charged snRNA within the U2 complex) and the actual bound portion, which associates with the polypyrimidine tract. Essential for spliceosome formation.
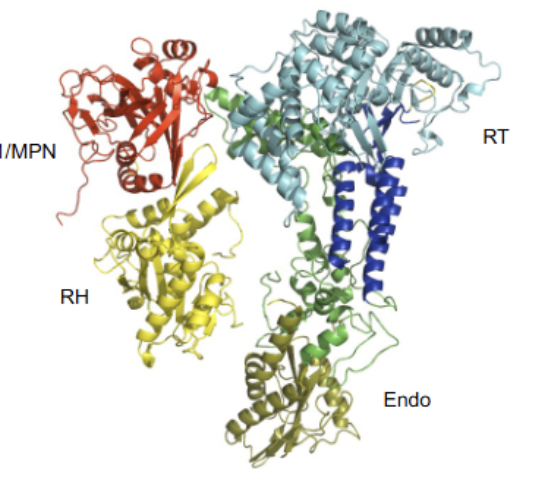
U5 structure
Notably does not perform any catalysis (all active sites have been replaced with non-functional units). Extremly well (most) conserved from humans to yeast = important. Largely performs structural role. Contains many domains, featuring an endonuclease, reverse transcriptase and RNase-H
How was the spliceosome structure solved
Too dynamic to use crystalization methods. Had to use cryo electron microscopy, which was able to solve the strucutre via class averaging and extreme sensitivity (1 electron hitting the protein).
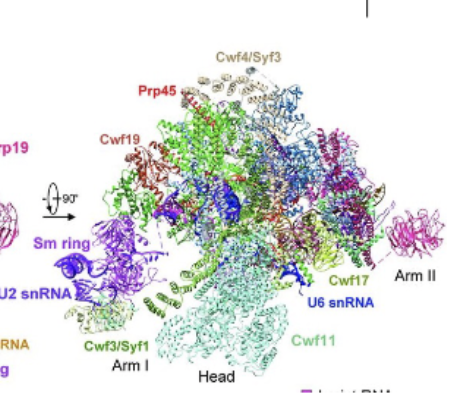
Spliceosome active site mechanism
Using 2 metal ions (attached to U6 stem loop) to activate the incoming group, stabilize the intermediate and then stabilize the leaving group. Exact same mechanism as Group 2 introns, which means that it is an RNA catalyzed reaction = RNA machines at its heart (proteins just there for support).
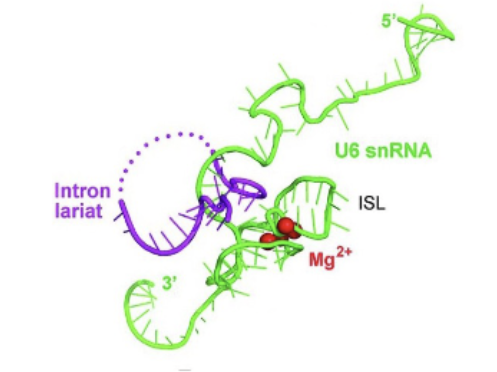
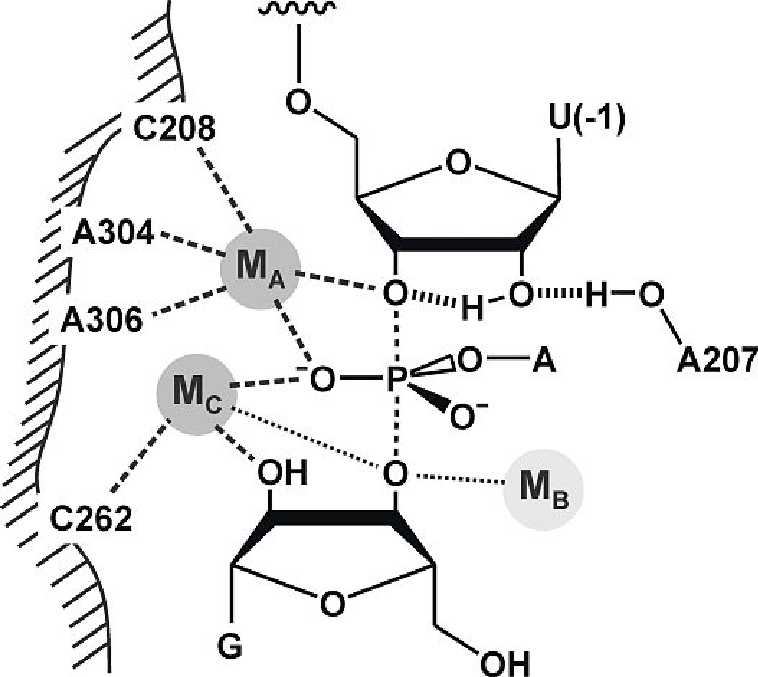
2 metal (Mg) ions required for reaction
RNA polymerase, Group 1 introns, Group 2 introns and the spliceosome all share ______
“Homing” Endonucleases
Certain Group II introns contain ORF (open reading frames) for DNA endonculease and reverse transcriptase activity, allowing them to insert themselves into target DNA
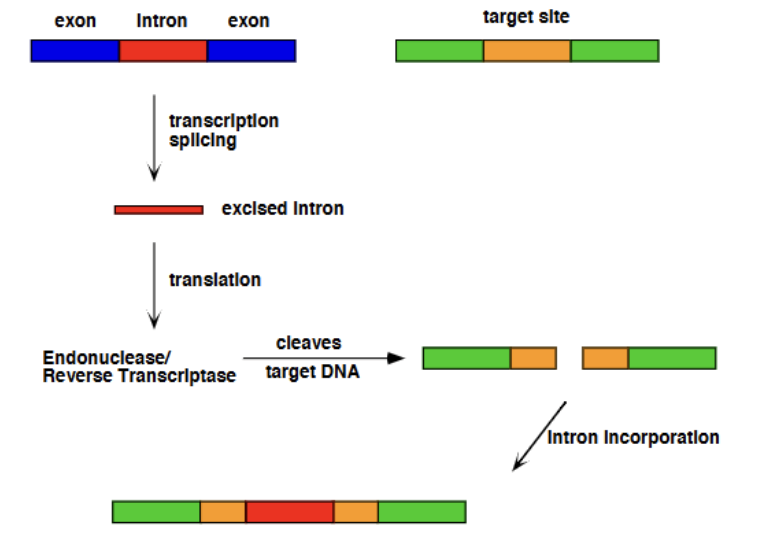
Group II Homing Mechanism (Introns as Mobile Genetic Elements)
Due to Group II reaction being reversible (slightly), the lariat intron product can attack a DNA nucleotide, reversing the lariat formation and looping out as a section of RNA within a DNA duplex. The intron then codes for endonuclease activity, to split up the duplex inthe position where it has inserted itself and cleaves, allowing it to linearize. The intron then codes for reverse transcriptase, coding its DNA complement strand into the duplex. Endogenous RNAse-H cuts out the intron from the duplex, and endogenous Pol II comes to fill in the gap, resulting in the intron being incorportated into the DNA strand.
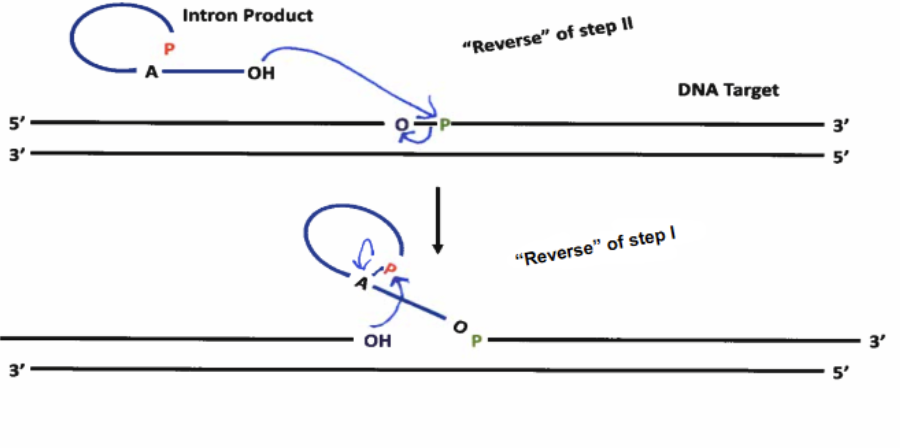
What do mobile RNA genetic elements prove?
That there are certain cases where RNA introns can code for protein activity, such as RT or endo; therefore not accurate to refer to exons as “coding genes” - real life is more complicated than that.
Why is intron insertion into DNA duplexes not a problem?
Because when they are eventually transcribed to RNA, those introns will just self-splice, leaving only the desired exons behind.
RNAse-H
Endogenous protein which cleaves any RNA strand found in a DNA/RNA duplex (heterogenous mixture of DNA/RNA present), releasing the RNA, following the activity of reverse transcriptase
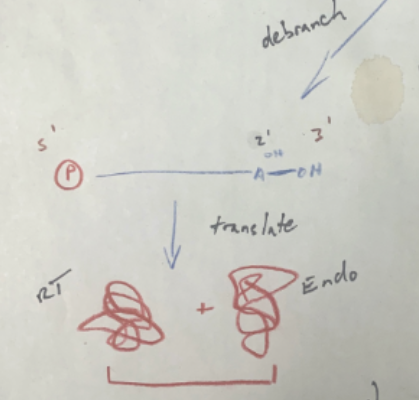
How can RNA genetic elements insert themselves, if they are the ones coding for the proteins required to insert themselves?
Seems to be a catch 22; however, before the RNA intron begins inserting itself into a DNA duplex, it can unbranch itself from the lariat form and be translated by a ribosome, coding for RT and Endonuc. It can then re-loop, and go and attack a DNA duplex, with the help of the proteins it just coded for, and proceed with insertion like normal.
How do a limited number of genes create incredibly complex organisms?
Through differential RNA processing (different cleavage sites, different polyadenylation sites, and alternative splicing), one primary transcript (gene) can many different (but related) mRNA’s, all of which are able to code for different polypeptides. Not 1:1 with gene: protein; many proteins from 1 gene.
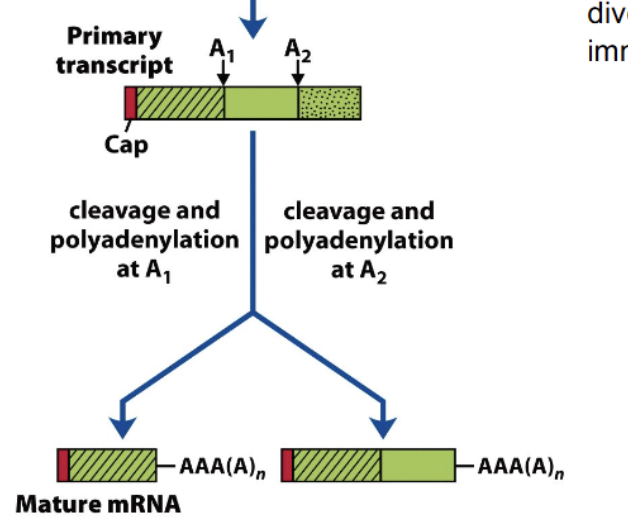
Multiple polyadenylation sites
Choosing where to cleave and polyadenylate a primary transcript has the chance to create a lot of diversity; can form 2 or more distinct mature mRNA’s, from just 2 different poly-A sites.
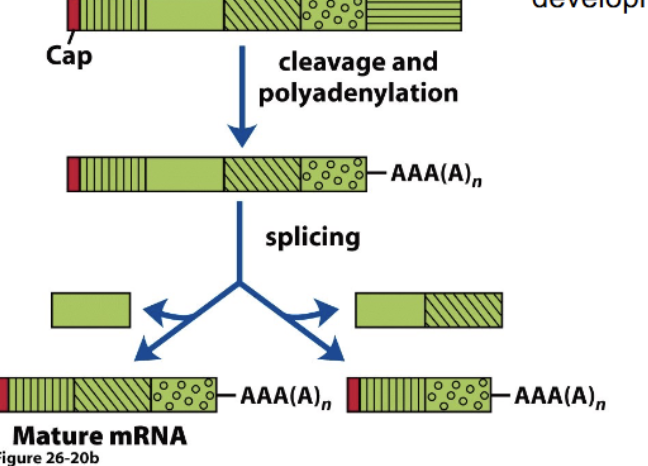
Alternative splicing
Mixing and matching which exons from a primary transcript occur in a mature mRNA. Huge room for variation, from a single transcript with 115 exons we can create 40,000 possible combinations within the mature mRNA
Drosophila gene alternative splicing example
Repressor system in females where sex lethal protein competes with U2AF, forcing it to bind further downstream at a different 3’ splice site (Repressing). (determines fly gender: in males, the sex lethal protein is not coded for and therefore U2AF binds at an earlier 3’ splice site)
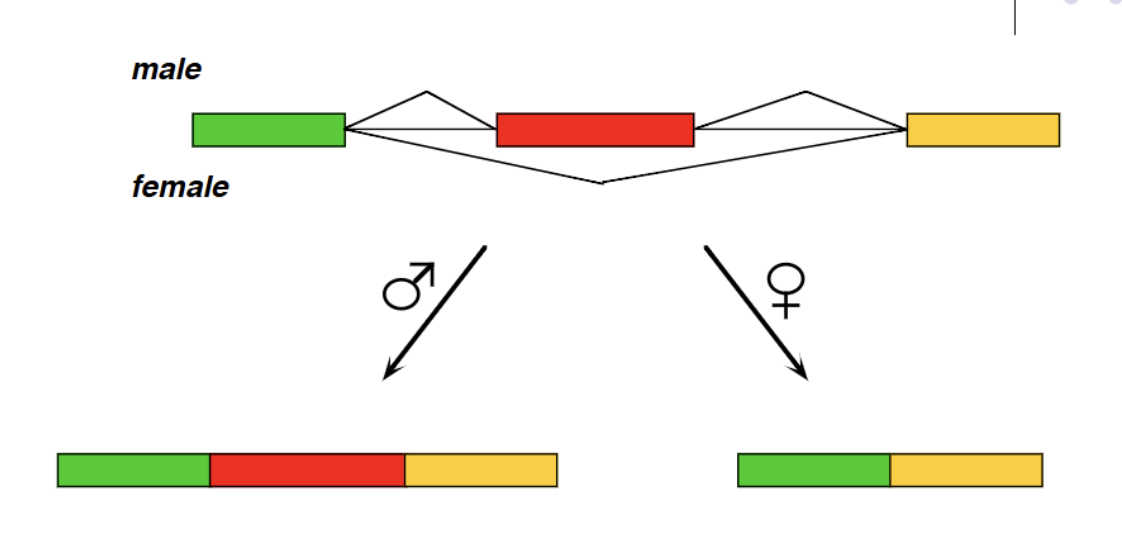
PTB in humans
Polypyrimidine tract binding which competes with U2AF in humans (analagous to Sxl protein in flies). Repressing
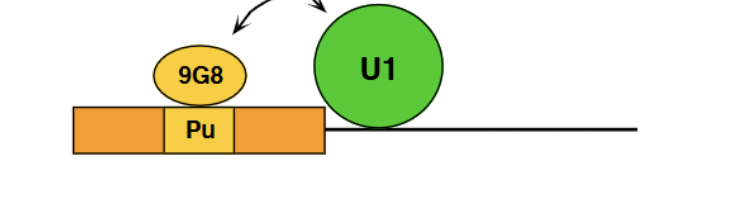
Activation/Enhancement alternative splicing example
9GB = SR protein which causes binding of U1 to the 5’ splice site.
Repression at RNA secondary structure level
Has the ability to fold into strucutres which will block the necessary splice sitesfrom being associated with the necessary snRNP’s. This can be relieved by ATP dependent helicase (another regulation step)