Lecture 2: Overview of Eukaryotic Gene Expression
1/28
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
29 Terms
Purpose of gene control in multi-cellular organisms
Execution of precise developmental and tissue specific program
Proper genes are expressed in proper cells at proper times
True for all cells but RBCs and repro cells
Start of transcription
Takes place on DNA that is wrapped chromatin
Chromatin needs to open for a gene to be activated and for transcription to proceed
Chromatin mediated regulation is a eukaryotic mechanism
One component of epigenetic regulation of gene expression
Organized state of chromatin influences transcription
Euchromatin
Less dense regions of chromatin
Active genes are found in euchromatin
Allows gene regulatory proteins and RNA polymerase complexes to bind to the DNA
Facilitating the active transcription of genes into mRNA
Heterochromatin
Regions of chromosomes that are densely packed
Rich in repetitive DNA
Transposons
Centromeres
Telomeres
Not accessible to transcriptional machinery
Inactive genes are found in heterochromatin
Difference between Prokaryotes vs Eukaryotes
Very elaborate transcriptional control
Open a gene is found within open chromatin
Variety of factors regulate the expression of each individual gene
Pioneer TFs come in and a variety of other factors such as Pol. and GTFs help with transcription
Very common genes will be open
Every eukaryotic gene requires GTFs
Eukaryotic RNA polymerases
Polymerase I, II, III
Pre-rRNA
28S, 18S, 5.85 rRNAs
Ribosome compartments
Protein synthesis
RNA Pol I
mRNA
Encodes protein
RNA Pol II
siRNAs
Chromatin-mediated repression, translation control
RNA Pol II
miRNAs
Translation control
RNA Pol II
tRNAs
Protein synthesis
RNA Pol III
5S rRNA
Ribosome component
Protein synthesis
RNA Pol III
snRNA U6
RNA splicing
RNA Pol III
7S RNA
Signal recognition particles for insertion of polypeptides into the ER
RNA Pol III
Other small stable RNAs
Various functions
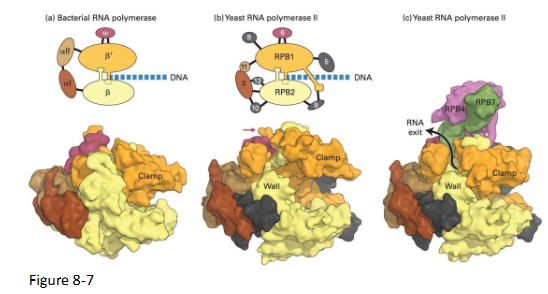
RNA Polymerase Structure → YEAST RNA pol II
RNA pol II consists of 12 polypeptides
RPB1 - RPB12
All other eukaryotic RNA polymerases share very high level of homology with yeast RNA Pol II
RNA Pol Clamp domain
RPB1 accommodates the DNA
After positioning over DNA the clamp is closed by a bridge
Synthesis of RNA takes place at catalytic center with the participation of Mg++
Synthesized RNA exists through a “channel” and is immediately capped by 7m Guanosine
Complexes of multiple polypeptides: Yeast vs Bacteria
Prokaryotes
Beta and beta prime subunits
Eukaryotes
RPB1 and RPB2 subunits
Homologous in eukaryotes and prokaryotes
Structural similarity of the enzyme
Carboxy Terminal Domain (CTD) in RNA Pol II
Specialized domain not found in other polymerases
Prokaryotic and eukaryotic
Involved in multiple regulatory interactions and play a key role in initiation release, elongation and processing of synthesized mRNA
Yeasts: 26 repeats of Tyr-Ser-Pro
Mammals: 52 repeats of Tyr-Ser-Pro
Ser residues in CTD
Phosphorylated upon transition from initiation and elongation
Genes transcribed by RNA Pol II are regulated by
Conserved Basal Promotor Element (Core Promoter sequences)
Promoter proximal binding sites for transcriptional activators (GTPs)
Distal enhancers or repressors
Chromatin Structure
Core Promoter sequences in Eukaryotic DNA
Transcription starts at defined point: Initiation site
Usually an A (adenine) on the coding strand
Four elements direct the positioning of the polymerase at these promoters:
TATA box
Initiator
BRE
DPE
TATA Box
Tight consensus sequence
Prevalent in highly transcribed genes
Initiator
Less conserved element
Some genes contain initiator but no TATA
BRE
Influences activity of promoter
Most upstream
DPE
Downstream promoter element
Influences activity of promoter
Importance of RNA polymerases
Need to recognize promoter to correctly Initiate transcription
Several GTFs assemble preinitiation complex over Core promoter sequence
Other factors that help with preinitiation
DNA helicases help the polymerase initiate transcription
Protein kinases release the polymerase
Elongation factors facilitate movement of polymerase
Additional proteins move nucleosomes out of way
GTFs of RNA polymerase II
TFIIA, B, D, E, F, H