ap bio unit 3 review
Enzymes
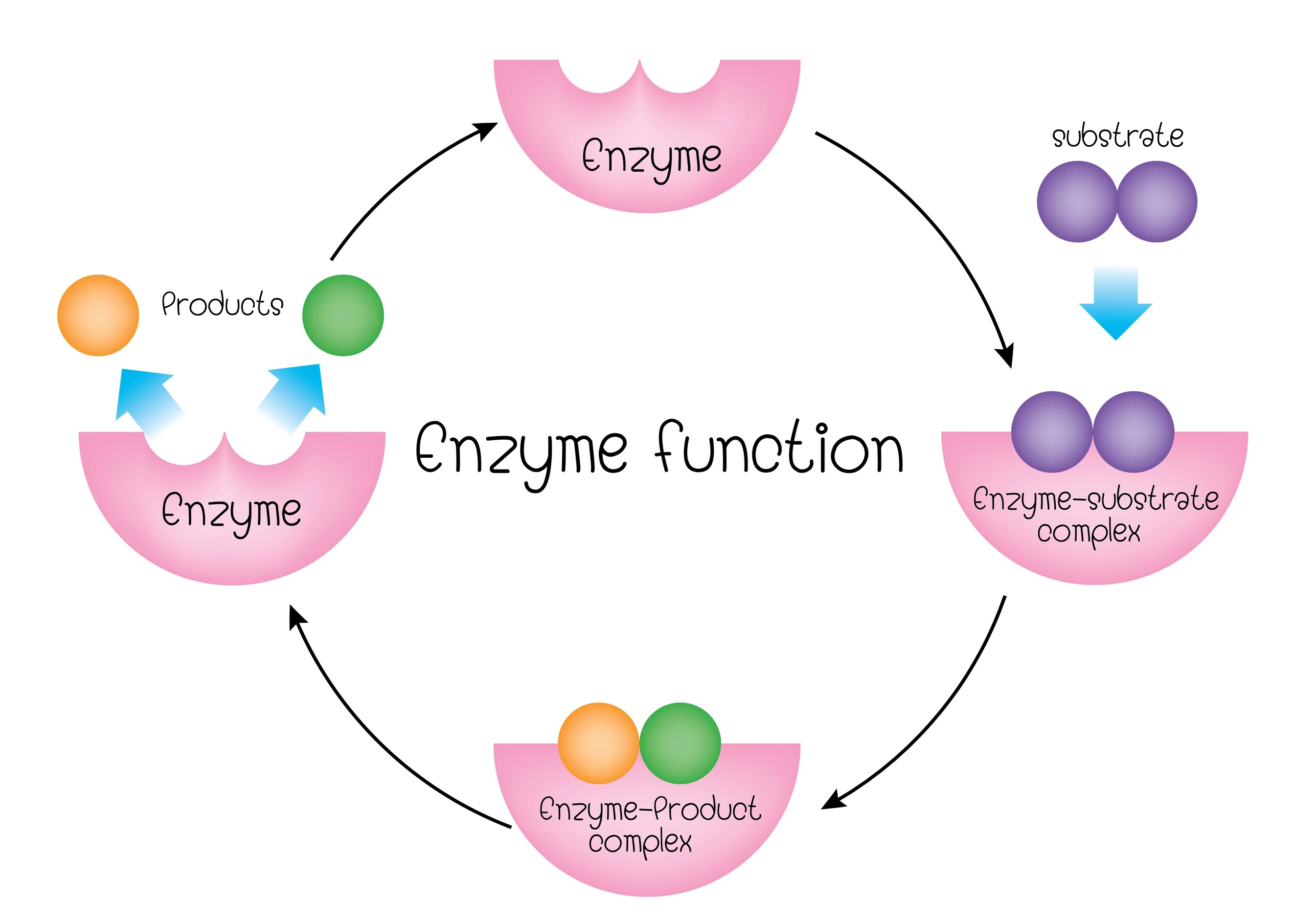
Enzyme Structure and Function
Structure Overview
- enzymes: biological catalysts; used to speed up biological processes
- ribozymes: biological catalysts made of RNA
- active site: this part of an enzyme interacts with the substrate (reactant)
- shape of the active site on an enzyme is specific to the enzyme and its function
- substrate and active site shapes must match in order for enzyme to work
- any charged r-groups on the amino acids within an active site of an enzyme must have compatible charges on the substrate
Environmental Factors that Affect Enzyme Functions
- enzymes catalyze at the most optimal temperature and pH levels
- temp. is too low = less frequent collisions between enzyme and substrate; reaction slows down
- temp. is too high = enzyme can denature; bonds between substrate and enzyme can be affected; enzyme shape can alter
- pH levels are different = bonds in enzyme can be disrupted and tertiary structure of enzyme can change
- denaturation: changes to an enzyme’s structure; can limit an enzyme’s ability to catalyze chemical reactions
- denaturation can sometimes be reversed if the reaction returns to optimal conditions
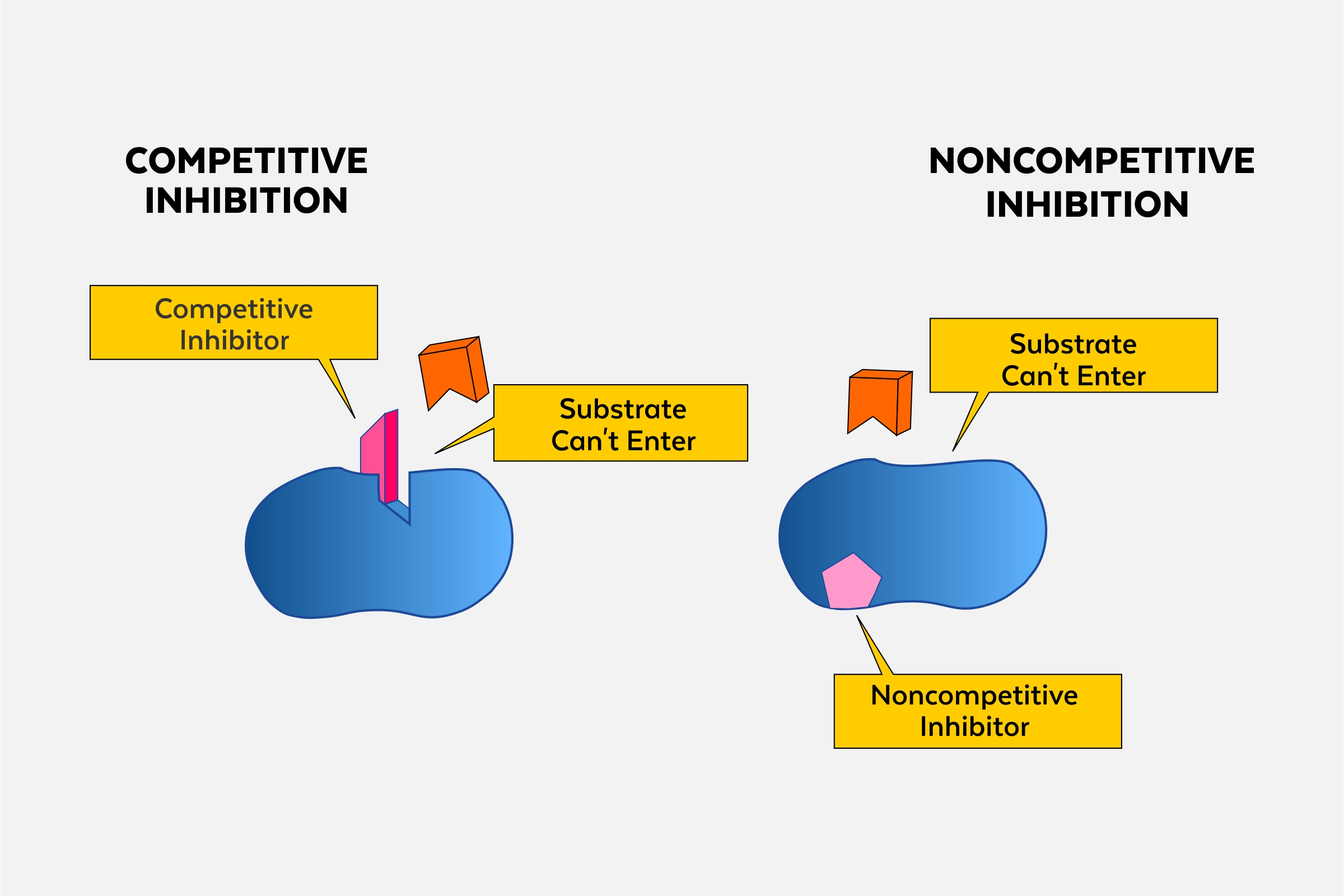
- competitive inhibitors: compete with substrates for the active site of the enzyme
- competition lowers the rate of enzyme-substrate reactions occurring
- effects of competitive inhibitors can be reduced by increasing the concentration of the substrate (reactant)
- non-competitive (allosteric) inhibitors: do not bind to active site and they bind to a different site on the enzyme; the binding changes the shape of the enzyme, which changes the shape of the active site and reduces the amount of enzyme-substrate bonding
- higher concentration of substrate does not affect non-competitive inhibitors
- non-competitive inhibitors function in feedback mechanisms by adjusting the rate of chemical reactions in the cell to suit changing environmental conditions
- cofactors: inorganic molecules and coenzymes (organic molecules): increase the efficiency of enzyme-catalyzed reactions
- they usually increase efficiency by binding to the active site or the substrate to enhance the binding of the substrate to the active site
Activation Energy in Chemical Reactions
- all molecules have a given amount of free energy (G); chemical reactions in biological processes involve changes in molecules
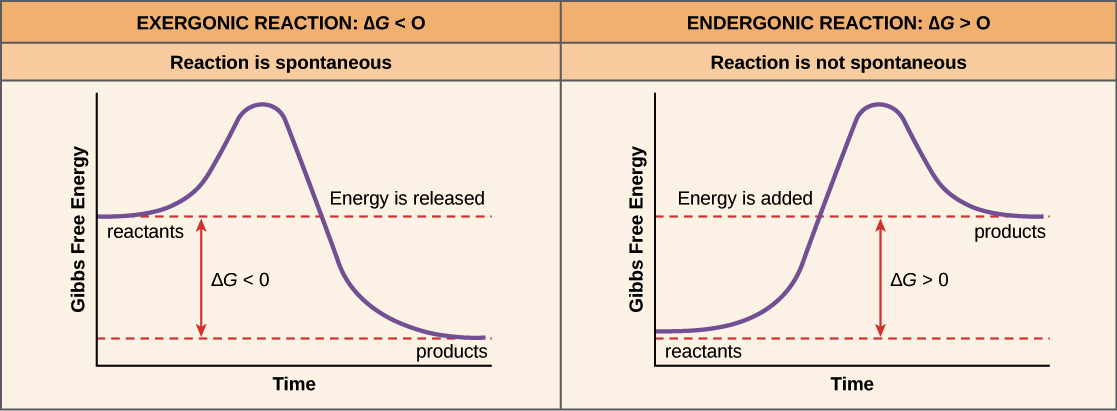
- chemical reactions can be endergonic or exergonic
- endergonic reaction: has products with higher free energy levels than its reactants; considered energetically unfavorable
- exergonic reaction: has products with lower free energy levels than its reactants; considered energetically favorable
- all chemical reactions require an input of energy to reach a transition state/start the reaction
- activation energy (Ea): the difference between the energy level of the reactants and the energy level of the transition state of the reaction
- higher Ea: results in slower rate of chemical reactions
- lower Ea: results in faster rate of chemical reactions
- enzymes can lower activation energy (Ea) in multiple ways:
- bringing the substrates together in proper orientation for a reaction to occur
- destabilizing chemical bonds in the substrate by bending the substrate
- forming temporary ionic or covalent bonds with substrate
- enzymes can lower activation energy but they cannot change an endergonic reaction (energetically unfavorable) to an exergonic reaction (energetically favorable)
Energy and Metabolism/Coupled Reactions
- energy input into cell must be higher than energy requirements for cellular systems
- processes that release energy can be paired (coupled) with processes that require energy
- coupled reactions: occur in multiple steps to allow for the controlled transfer of energy between molecules (leads to more efficiency)
- coupling exergonic reaction with endergonic reaction: allows the energy released by the exergonic reaction to “drive” the endergonic reaction
- example:
- exergonic step: breakdown of ATP into ADP and a phosphate group (Pi (inorganic)); releases approx. 30 kilojoules of energy per mole of ATP
- endergonic reaction: the reaction that combines glucose with fructose to form sucrose requires approx. 27 kilojoules of ATP per mole of sucrose formed
Photosynthesis
Overview
heterotrophs: consume other organisms to obtain energy required for biological processes
autotroph: produce their own organic molecules from inorganic molecules
- photo-autotrophs: autotrophs that use light energy to power the process
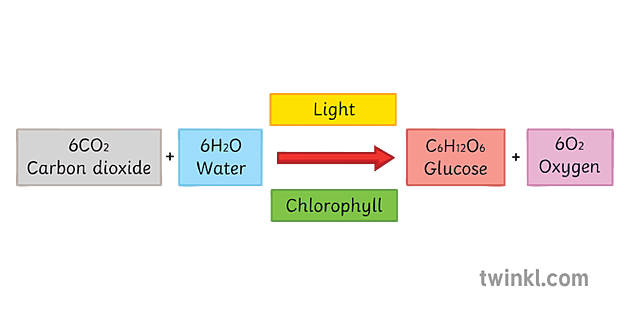
- photo-autotrophs: autotrophs that use light energy to power the process
light-dependent reactions: use energy from sunlight to split water to produce oxygen gas, protons, and high-energy electrons
- oxygen gas: released into atmosphere
- protons and high-energy electrons: used to power the production of ATP and NADPH
- ATP and NADPH are sent to the light independent reactions
light-independent reactions: use ATP, NADPH, and carbon dioxide to produce sugars
- ATP, Pi (inorganic phosphate group), and NADP+: sent back to light-dependent reactions
plants: photosynthesis occurs in the chloroplasts
- chloroplasts:
- outer membrane: filled with liquid called stroma
- stroma: has floating stacks of membranous sacs called grana; location of light-independent reactions
- thylakoid: each individual membranous sac from the grana stack; location of light-dependent reactions
prokaryotes: also perform photosynthesis (ex. cyanobacteria); do not contain chloroplasts
- light-dependent reactions: occur on infoldings of plasma membrane
- light-independent reactions: occur in the cytosol
Light-Dependent Reactions
- photophosphorylation: conversion of ADP to ATP using the energy of sunlight by activation of PSII
- light energy excites the electrons in the chloroplasts to a higher energy level; energy is released as excited electrons move through chloroplasts
- NADP+ accepts the electrons to form NADPH
- NADPH: source of reducing power for light-independent reactions
- chlorophyll: light-absorbing pigment that captures the energy of photons from the sun; found in photosystems 1 and 2
- photosystem: composed of proteins, chlorophyll, and accessory pigments; PSI and PSII contain different types of chlorophyll that absorb most light energy at slightly different wavelengths (PSI = 700 nm; PSII = 680 nm)
- photosystems are located in the thylakoid membrane of the chloroplast; they are connected by an electron transport chain (ETC)
- accessory pigment: other light-absorbing pigments besides chlorophyll
The Process of Light-Dependent Reaction
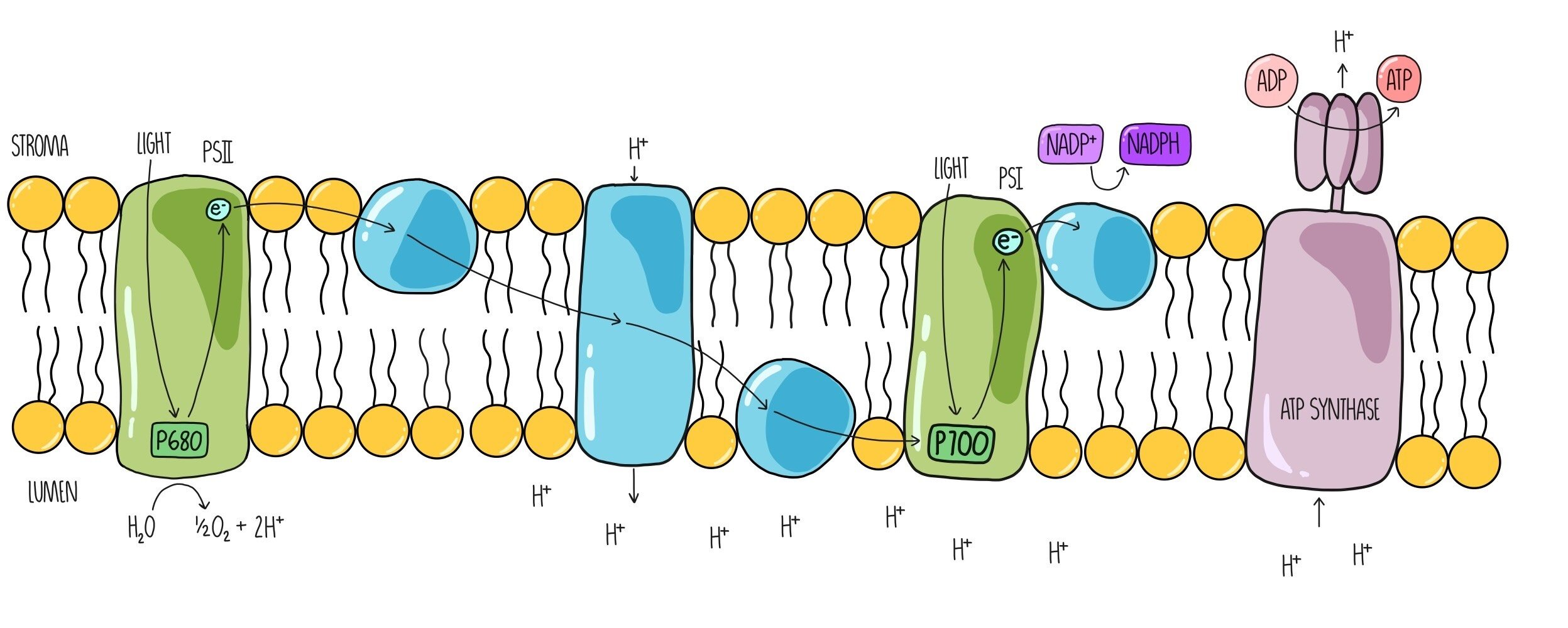
- energy in p+ is used to boost e- in chlorophyll to higher energy level in PSII
- e-s from PSII are passed from one protein carrier to another in a series of redox reactions (like falling down a hill)
- the final e- donor in the ETC passes the e- to the PSI
- as the e-s pass through the ETC, energy is released and used to create a proton (H+) gradient
- H+ ions are actively transported against concentration gradient across the thylakoid membrane
- electrons from PSII that fell down the ETC are now in PSI and need to be replaced in PSII
- photolysis: the splitting or decomposition of a chemical compound by means of light energy or photons
- occurs when splitting of water molecules takes electrons from hydrogen atoms and produces H+ ions, electrons from PSII, and oxygen gas
- proton gradient generated by photolysis of water and the ETC powers the production of ATP by ATP synthase
- chemiosmosis: process of using a proton gradient and ATP synthase to produce ATP; also used in mitochondria to generate ATP during cellular respiration
- the electron from the ETC that is now on PSI is boosted by a photon of light energy from the sun; the electron passes again through a series of carriers (much shorter than ETC) where it is finally transferred (along with a proton) to NADP+ by the enzyme NADP+ reductase
- this produces a molecule of NADPH, which will provide the reducing power for the light-independent reactions
Light-Independent Reactions (Calvin Cycle)
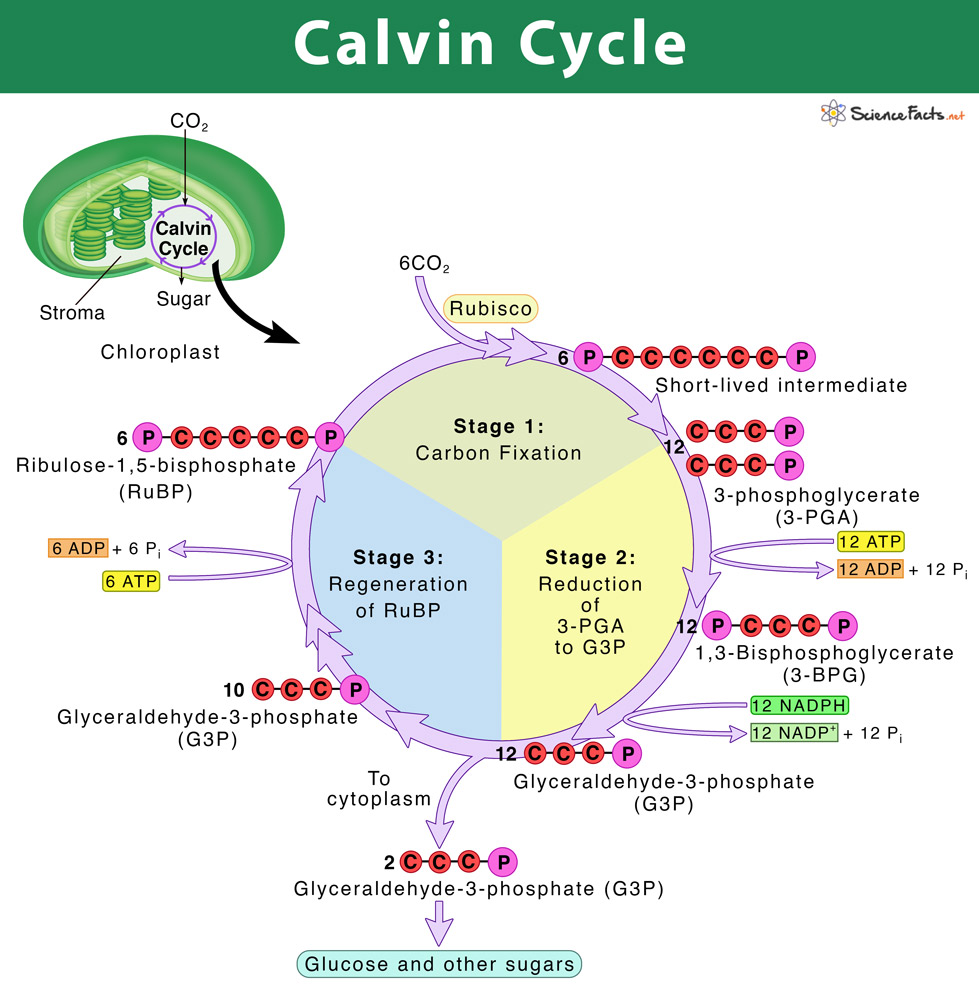
- light-independent reactions occur in stroma of chloroplast
- the process can be broken down into 3 part:
- fixation of carbon
- reduction
- regeneration of RuBP
Fixation of Carbon
- fixation: to turn a biologically unusable form to a usable form
- enzyme ribulose-biphosphate-carboxylase (rubisco) adds one molecule fo carbon dioxide to the 5-carbon molecule ribulose-biphosphate (RuBP)
- this produces a 6-carbon intermediate that is unstable
- the unstable molecule is then broken down further into two 3-carbon molecules
Reduction
- the ATP and NADPH from the light-dependent reactions are used to reduce the 3-carbon molecules into glyceraldehyde-3-phosphate (G3P)
- G3P: can be used to make sugars; some of it is used during regeneration
Regeneration
- 5-carbon RuBP must be regenerated for the calvin cycle to continue
- for every 5 molecules of G3P, there are 15 carbon atoms present
- ATP from light-dependent reaction in used to rearrange the five G3P (3-carbon molecule) molecules to form 3 molecules of RuBP (5-carbon molecule)
- this process requires energy that comes from the light-dependent reactions
Cellular Respiration
Overview
- cellular respiration includes the following processes:
- glycolysis
- oxidation of pyruvate
- krebs cycle (citric acid cycle)
- oxidative phosphorylation
- the presence of oxygen determines whether the process will be anaerobic or aerobic
- anaerobic: without oxygen; anaerobic organisms (don’t have access to oxygen) can perform glycolysis and fermentation
- aerobic: with oxygen; aerobic organisms (have access to oxygen) can perform all processes in the presence of oxygen and only glycolysis and fermentation in the absence of oxygen
- can perform more processes than anaerobic, meaning they can extract more energy from organic compounds
| cellular process | anaerobic organisms (without oxygen) | aerobic organisms (with oxygen) | aerobic organisms (without oxygen) |
|---|---|---|---|
| glycolysis | ✓ | ✓ | ✓ |
| oxidation of pyruvate | ✓ | ||
| krebs cycle | ✓ | ||
| oxidative phosphorylation | ✓ | ||
| fermentation | ✓ | ✓ | ✓ |
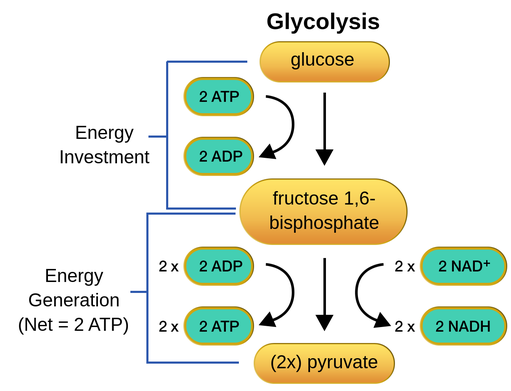
Glycolysis
- occurs in the cytosol of the cell (all organisms have cytosol, so all organisms can perform glycolysis)
- a 6-carbon molecule enters glycolysis alone with two molecules of NAD+ (electron carrier)
- each NAD+ is reduced (loses hydrogen atom and electrons) to NADH
- two molecules of ATP are required for early steps of glycolysis
- four molecules of ATP are produced by glycolysis (net gain of ATP)
- at the end of glycolysis, the 6-carbon glucose molecule is split into two 3-carbon pyruvate molecules
| location | inputs | outputs |
|---|---|---|
| cytosol | glucose (6C) | 2 pyruvate (3C) |
| 2 NAD+ | 2 NADH | |
| 2 ATP | 4 ATP |
Oxidation of Pyruvate
- occurs in the mitochondria
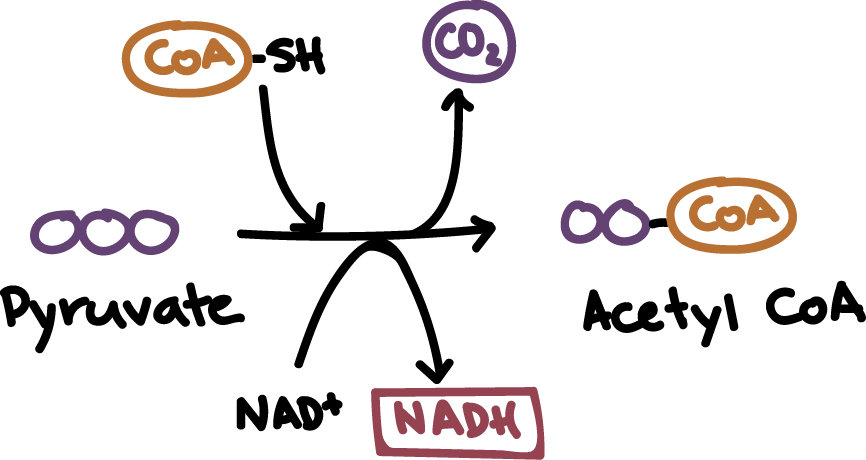
- the 3-carbon pyruvate molecule is oxidized (loses hydrogen atom and electron) and NAD+ is reduced (gains a hydrogen and its electrons) to become NADH
- as this happens one of the carbons of the pyruvate molecule is reduced as CO2 (leaving behind a 2-carbon acetyl group)
- coenzyme A attaches to the 2-carbon acetyl group and delivers the acetyl group to the krebs cycle
- each molecule of glucose that enters glycolysis generates 2 pyruvate so oxidation of pyruvate occurs twice for each molecule of glucose
| location | inputs | outputs |
|---|---|---|
| mitochondria | pyruvate (3C) | acetyl group (2C) |
| NAD+ | carbon dioxide (1C) | |
| NADH |
Krebs Cycle (Citric Acid Cycle)
- occurs in the matrix (liquid center) of the mitochondria
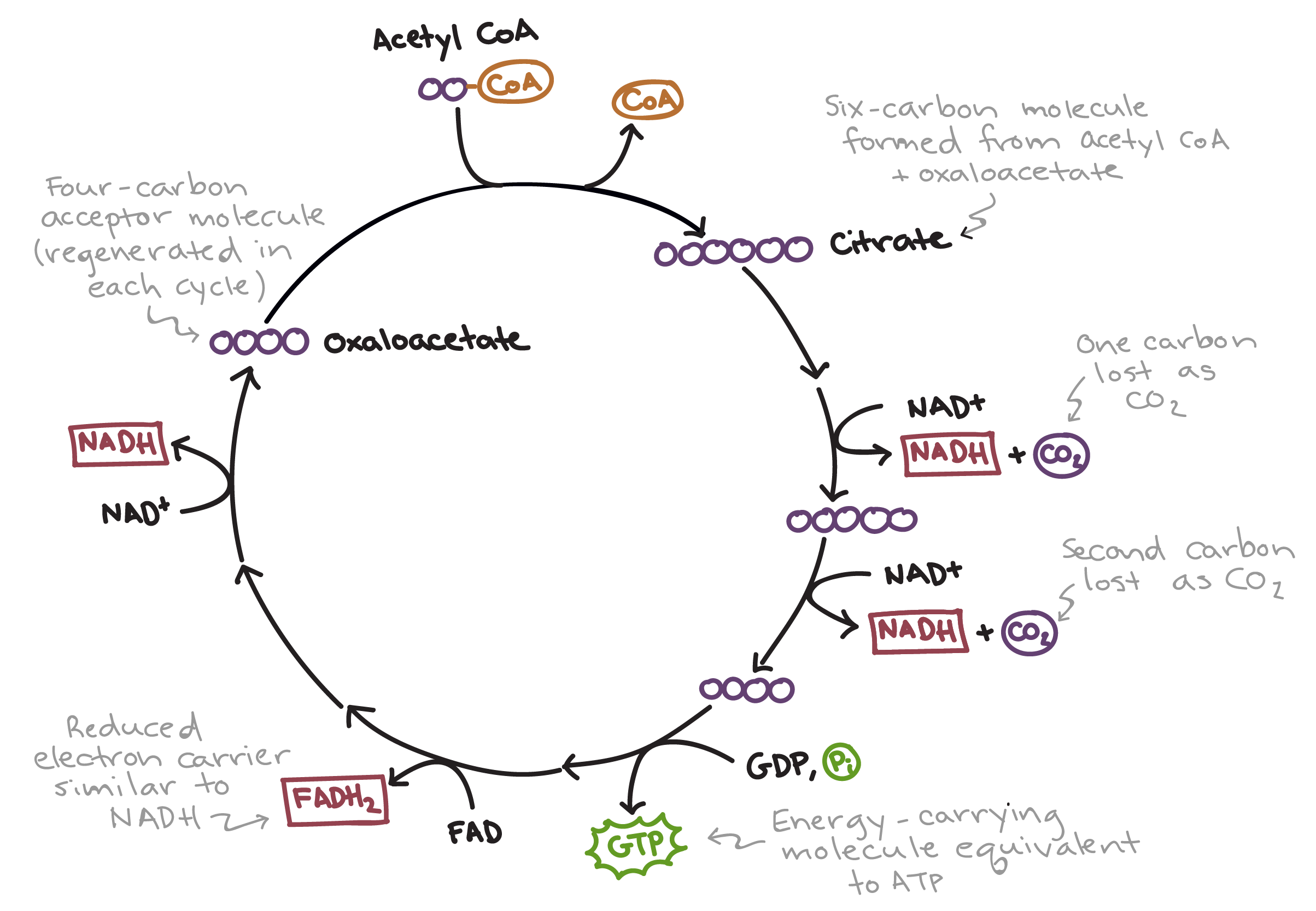
- coenzyme A brings the 2-carbon acetyl group to the cycle (initially attached as 4-carbon intermediate but forms 6-carbon molecule)
- the 6-carbon molecule goes through a series of enzyme-catalyzed reactions to regenerate the 4-carbon intermediate (and produces 2 molecules of CO2)
- at the end, all the carbon that was originally in the glucose molecule (at the start of glycolysis) has been released as CO2
- during the cycle:
- one molecule of ATP is produced from substrate-level phosphorylation
- phosphorylation: direct addition of a phosphate group to ADP without the use of an electron transport chain or chemiosmosis
- three molecules of NAD+ are reduced to NADH
- one molecule of FAD+ is reduced to FADH2
| location | inputs | outputs |
|---|---|---|
| matrix of mitochondria | acetyl group (3C) | 2 carbon dioxides (1 C each) |
| 3 NAD+ | 3 NADH | |
| 1 FAD+ | 1 FADH2 | |
| 1 ADP + Pi | 1 ATP |
Total Products of Glycolysis, Oxidation of Pyruvate and Krebs Cycle for Each Molecule of Glucose
| molecule | glycolysis | oxidation of pyruvate | krebs cycle | total |
|---|---|---|---|---|
| ATP | 2 (net) | 0 | 2 | 4 |
| NADH | 2 | 2 | 6 | 10 |
| FADH+ | 0 | 0 | 2 | 2 |
| CO2 | 0 | 2 | 4 | 6 |
- all 6 carbons in glucose molecule have been released as CO2
- four molecules of ATP have been produced by substrate-level phosphorylation
- a total of 12 high-energy electron carriers (10 NADH and 2 FADH2) have been produced and will enter oxidative phosphorylation
Oxidative Phosphorylation
- involves the ETC and chemiosmosis (both of which occur in the membrane of the mitochondria)
- this process yields the most production of ATP in cellular respiration
- the electron carriers (NADH and FADH2) previously produced carry their electrons to the ETC
- as they deliver their electrons they are oxidized to NAD+ and FAD+, which can be used earlier on in cellular respiration
- as the electrons travel through the ETC their PE decreases and energy is released
- the released energy is used to pump H+ out of the matrix and int the intermembrane space of the mitochondria to create a proton gradient
- at the end of the ETC, molecular oxygen (O2) combines with four protons (H+) and four electrons (e-) to form 2 water molecules (2H2O)
- this means oxygen is the final (terminal) acceptor of electrons in cellular respiration
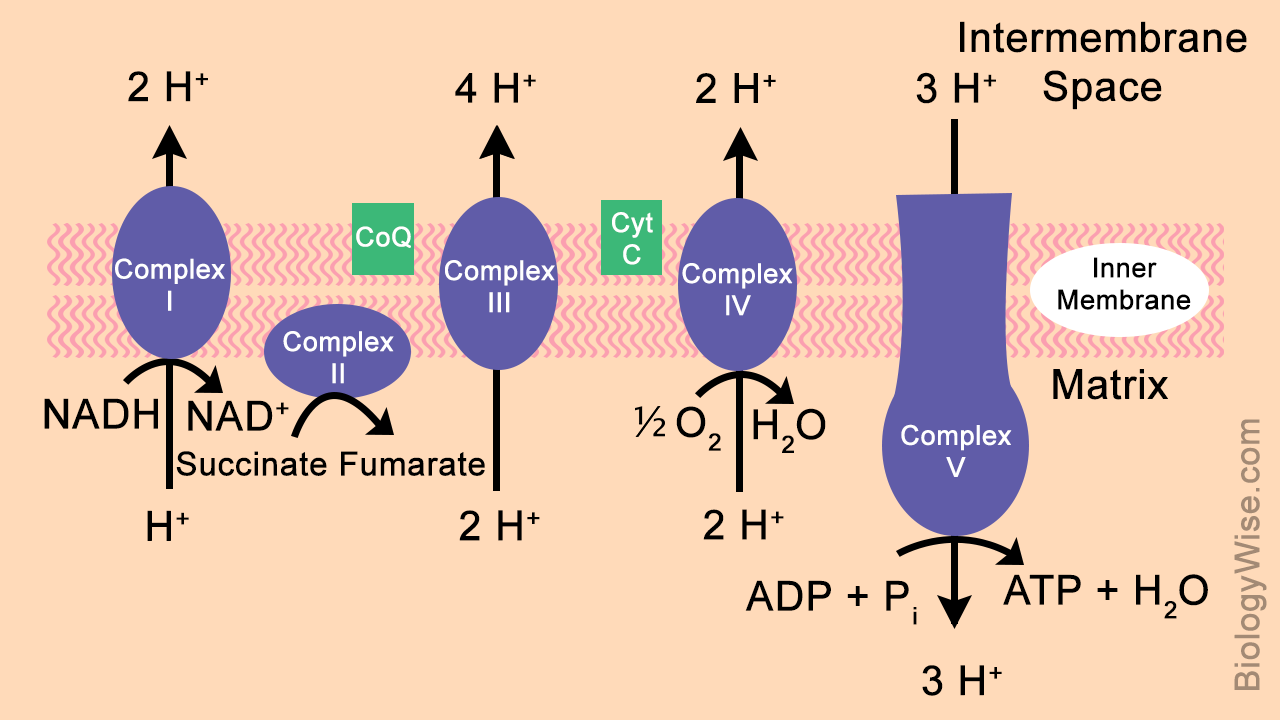
- the proton gradient created by the ETC is used to produce ATP through chemiosmosis
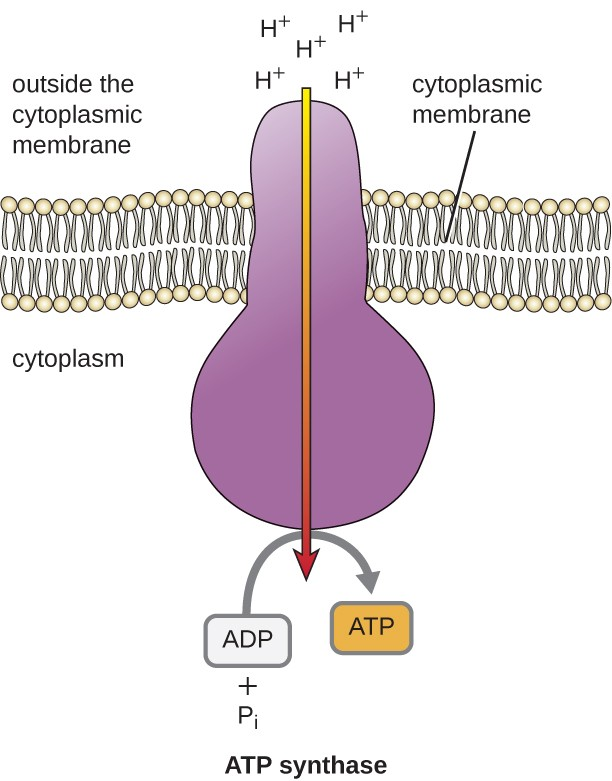
- protons flow from areas of higher concentration in intermembrane space to areas of lower concentration in the matrix through a channel in the ATP synthase enzyme
- this flow of protons leads to a change in the shape of the enzyme; the new shape of the enzyme allows for ATP synthase to catalyze the production of ATP
- (ideally) 34 total ATP molecules can be produced:
- 10 NADH x 3 ATP = 30 ATP
- 2 FADH2 x 2 ATP = 4 ATP
- FADH2 has a lower potential energy than NADH, so it produces less ATP
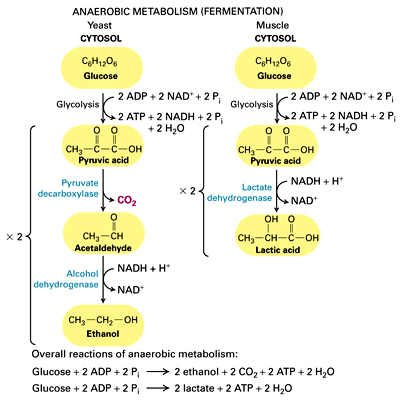
Fermentation
- when oxygen isn’t present, oxidative phosphorylation cannot occur (oxygen is the final electron acceptor)
- anaerobic conditions require the use of fermentation to regenerate NAD+ needed to keep the process of glycolysis going
- fermentation only occurs in the cytosol
- alcohol fermentation: pyruvate is reduced to an alcohol (usually ethanol) and CO2; NADH is oxidized to NAD+
- example: yeast undergoing fermentation for bread to rise
- lactic acid fermentation: pyruvate is reduced to lactic acid (3-carbon molecule); NADH is oxidized to NAD+; no CO2 is produced
- example: can occur in muscle cells if they do not have enough oxygen to carry out oxidative phosphorylation