Chud ass class
1/80
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
81 Terms
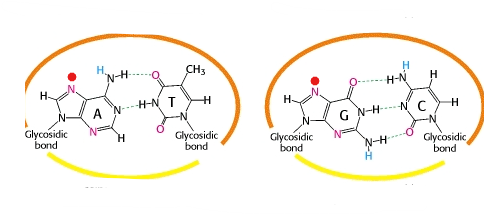
Loop E motif
Reverse Hoogsteen AU pair, bulge important for recognition
K-turn
both types of BPing, Unpaired residues stick out(important for recognition)
Strong acid(pkA)
large Ka small pKA
pKA
-log(Ka)
Buffer region
around were pH=pKa
peptide bond
Condensation reaction forms a H bond donor(NH) and acceptor( O=C)
Peptide bond is
planar due to double bond character(trans favored)
Φ
N-A, Phi
Ψ
C-C=O, psi
at psi 165
Phi can be between -70 to -170
alpha-helix
3.6 per turn, right handed, R groups out
Helix location
often on the outside, but can be anywhere
Amphipathic helix
hydrophobic/philic sides
Helix destabilizing residues
Proline(ring), glycine(spacing)
310 helix
smaller, least favored
pi-helix
largest, 15% of proteins
b-sheets
slight right hand twist. side chain alternate above and below. long range interactions
Loops
usually polar, substrate binding and catalysis. Disordered until they bind to substrate
Helix turn helix
structural motif common in DNA binding proteins
calmodulin
Helix loop helix protein which bind Ca2+(loops binds Ca2+)
Helix loop helix
Loop is longer than a turn(helix turn helix)

Greek-Key
4 anti-parallels beta-strands connected by short turns(loop connects 3 and 4)
Beta-alpha-beta
two parralel beta-strands connected by an alpha-helix. More rigid
Coiled-Coils
2 amphipathic a-helices. homo and heterodimers(a-helices for 2 proteins coil). Left handed superstructure.(7.2)
Heptad-repeats
if i and i+7 are hydrophobic, likely coiled coils.
Coiled-coil “ ‘ “
prime indicates anti-parallels(a-d’)
Alanine
A, Ala, 2, 10
Coiled-coil specificity
Additional interactions(H-bond, charge-charge) gives coiled-coils specificity.
Four helix bundle
Almost parallels a-helices(pairs). Hydrophobic core
a-b barrels
beta-alpha-beta repeats forming a beta strand tube(helices outside)
b-barrels
barrels formed by anti-parallel beta-strands
Glycine
Gly, G. 2, 10
Valine
Val, v. 2, 10.
Leucine
Leu, L. 2, 10
Isoleucine
ile, I. 2, 10
Methionine
Met, M. 2, 9
Proline
Pro/P, P. 2, 10
Phenylalanine
Phe, F. 2, 9.
Tryptophan
Trp, W. 2, 9.
Serine
Ser, S. 2, 9.
Threonine
Thr, T. 2, 9.
Asparagine
Asn, N. 2, 9.
Glutamine
Gln, Q. 2, 9.
Tyrosine
Tyr, Y. 2, 9, 10.
Cysteine
Cys, C. 2, 10, 8.
Lysine
Lys, K. 2, 9, 10.
Arginine
Arg, R. 2, 9, 12.
Histidine
His, H. 2, 9, 6.
Aspartic acid
Asp, D. 2, 10, 4.
Glutamic acid
Glu, E. 2, 9, 4,
membrane helcies
around 20 AA long and mostly hydrophobic residues, held together by Vander walls
Exo vs Endo conformation
Closer(endo) or farther(exo) from C5
C2 vs C3
Which is sticking up(same as base)
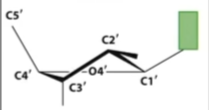
B form
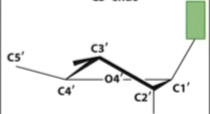
A form
DNA can pucker
at both C2 and C3
A from RNA can only pucker at
C3(due to hydroxyl)
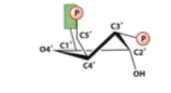
RNA C3 endo(less stretch)
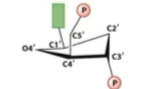
DNA C2 endo(more stretch)
B-Form DNA
Typical DNA form
Major groove
intercalation
Flat aromatic molecules are insert between bases and disrupt transcription
GU wobble base pair
Most common non-conical BPing. Stability is close to canonical AU base pair. Twists and forms a bulge(room for ion)
pseudoknot
Family A
Family B
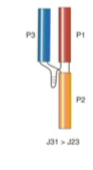
Family c
Coaxial stacking
terminal loops allow helices to stack allowing for 3d structure
Kissing loops
Long range WC interactions between hairpins
Self-cleaving RNA
Hammerhead/Hairpin, 2’ hydroxyl cleaves backbone.
Group 1 introns
2 tetra loop receptors(L2, L9). Pseudoknot belt pulls everything together
Programmed frameshift
Pseudoknot/hairpin and slippery sequence(repeats). Ribosome stopping can cause frameshifting
internal ribosomal entry sites(IRES)
Viral RNA element which mimics eukaryotic 5’ caps
Group 1 IRES
3 pseudoknots and 2 stem loops. Domain 3 mimics AUG start codon in the p-site
Aptamer
Riboswitch binding sites(highly conserved)
Expression platform
active site, intrinsic termination site, or hides shine-Dalgarno ribosome binding site(translation).(poorly conserved)
Guanine/adenine riboswitch
Three way junction. Conserved pyrimidine can BP or base stack with G/A.(regulates G/A concentrations)
TPP riboswitch
Highly conserved and widespread. TPP bridges parallel elements.
SAM 1
4 way junction. cation pi interaction. Intrinsic termination.
SAM 2
H-type, pseudoknot. Sam stretched in the pseudoknot region
SAM 3
Same as SAM, but positive charge is stabilized by sugar and base