Cell Histology
1/61
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
62 Terms
Histology
Study of tissues of the body and how these tissues are arranged to constitute organs
Histology levels of organization smallest to most complex
Molecules, Cells, Tissues, Organs, Organ Systems
Light microscope can see how much better than the human eye?
1000X
Preparation of tissue for microscopy steps (9)
Fixation, Dehydration, Embedding, Trimming/Sectioning, Removal of paraffin, Rehydration, Staining, Mounting sections of glass slide, Viewing the tissue (LM)
Fixation
Step 1: Small pieces of tissue are placed in solutions of chemicals that preserve by cross-linking proteins and inactivating degradative enzymes
Dehydration
Step 2: In alcohols, removes all water
Embedding
Step 3: Paraffin-infiltrated tissue is placed in a small mold with melted paraffin and allowed to harden
Trimming/Sectioning
Step 4: Paraffin block is trimmed to expose the tissue for sectioning (slicing) on a microtome
Staining
Step 6
Tissues
Cells that carry out the same general function
Four basic tissues
Epithelia, Connective tissue, Muscle, Nerve
Epethelia
Cover and/or line internal and external body surfaces
Form secretory glands and ducts
Connective tissue
Packing, support, connecting
Muscle
Contractility
Nerve
Irritability, conduction
Plane of section
Cutting the specimen, crucial to how specimen is viewed. Improper plane of section can lead to misdiagnosis
Different plane of sections
Longitudinal section, Cross section, Oblique section
Basic dyes
Bond with acidic molecules. Toluidine blue, methylene blue, Hematoxylin. BLUE
Acidic dyes
Bond with basic molecules. Eosin, fuchsia. RED
Hematoxylin will react with what molecules to stain them blue?
Acidic molecules in the nucleus
Basophilic
Anionic components (acidic components) stain more readily with basic dyes
Acidophilic
Cationic components (basic molecules) stain more readily with acidic dyes
Cytoplasm will stain what color?
Red/pink due to cationic proteins, acidophilic
Nucleus will stain what color?
Blue due to anionic nuclei acids, basophilic
Luxol fast blue stain (LFBS) / Luxol blue and hematoxylin
Shows myelin
Hematoxylin and Eosin stain (H&E)
Red=basic molecules (Eosin)
Blue=acidic molecules (Hematoxylin)
Dissolves fat (adipose tissue) and mucous during preparation process. Appears as white circle (empty )
Trichrome stain
3 colors, shows connective tissue (collagen, fibrin) in green/blue color
Golgi stain
Potassium dichromate and silver nitrate
Gold & Luxol blue & Cresyl violet
Used to demonstrate the Nissl substance in the neurons and cell nuclei
Periodic Acid Scheff Reaction stain (PAS)
Shows mucous, microvilli, basement membrane and glycogen molecules
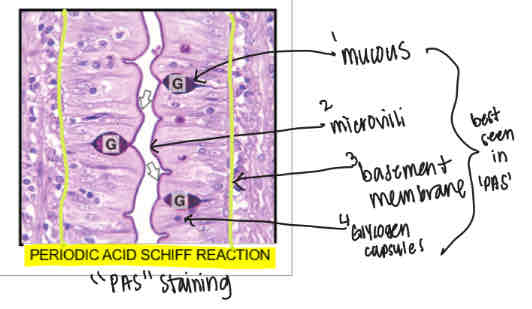
Distortions and artifacts
Result due to problems in interpretation of tissue sections due to either
A) Shrinkage because of fixation, dehydration and embedding
B) Loss of molecules that were not retained after fixation or removed during dehydration (glycogen and lipids)
Two types of electron microscopy
1) Transmission Election Microscopy (TEM)
2) Scanning Electron Microscopy (SEM)
Electron Microscopy
Electron beam passed through a very thin section of tissue
Transmission Electron Microscopy (TEM)
Permits high resolution
Allows magnification of up to 400,000 times
Looks at small portion of the cell
Can view organelles
Utilizes osmium tetroxide
Osmium tetroxide
Reacts with phospholipids and imparts electron density to cell and tissue structures because it is a heavy metal to enhance image formation in TEM
Electron dense
Dark areas in TEM
Electron lucent
Light areas in TEM
Scanning Electron Microscope (SEM)
Shows only surface views
10 nm resolution
Can view large depth of field
Inside of organs are viewed by freezing and fracturing tissues to expose internal surfaces
Membranous organelles
Contain plasma membranes that separate internal environment of organelle from the cytoplasm (cytosol + organelles)
RER, SER, Golgi, Mitochondria, Lysosomes, Peroxisomes
Non-membranous organelles
Lack a plasma membrane
Ribosomes, cytoskeleton and inclusions
Nucleus functions
1) Cellular regulation - houses genetic material which directs all cellular activities and regulates cell structure
2) Produces ribosomal subunits in nucleolus and exports them into cytoplasm for assembly into ribosomes
Outer nuclear membrane
Faces the cytoplasm, contains ribosomes and is continuous as certain sites with RER
Inner nuclear membrane
Faces the nuclear material and is supported on its inner surface by the nuclear laminate
Functions: stability to the nucleus, chromosomes attach
Nuclear envelope
Perinuclear space between outer and inner membranes. Function: Allows molecules in/out of the nucleus.
Molecules >9nm transported by active process mediated by receptors and utilize energy.
Molecules <9 nm (ions and smaller water-soluble molecules) cross via simple diffusion
Nucleosome
DNA sequence wrapped around a core of histone proteins. The basic repeating subunit of chromatin packed inside the cell’s nucleus.
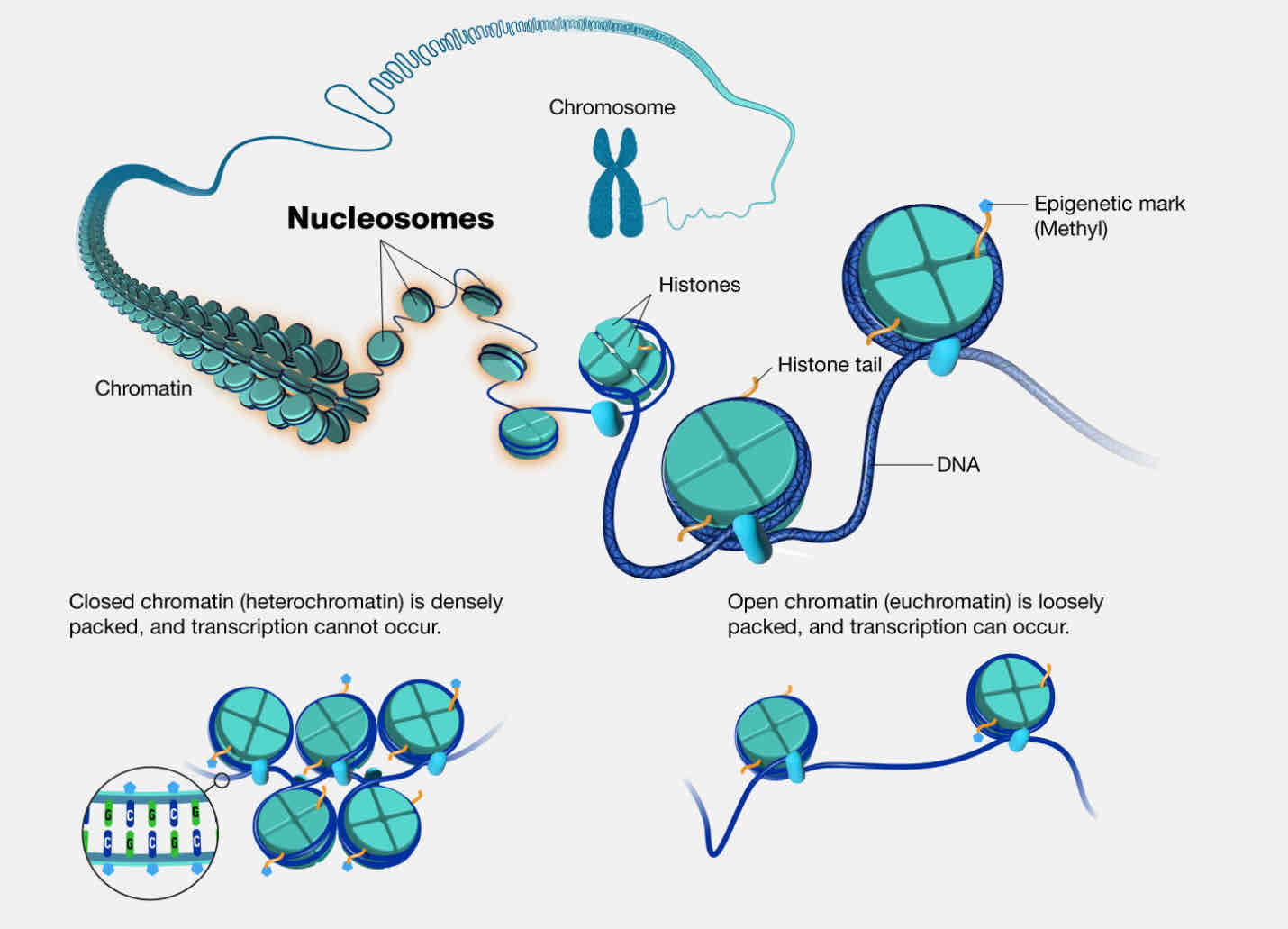
Heterochromatin
Dense staining (dark), highly condensed chromatin (closed). No active transcription
Eurochromatin
Light staining, open conformation chromatin. Active gene transcription occurs
Nucleolus function
1) Site of ribosomal RNA (rRNA) synthesis
2) Initial assembly, once assembled transported to cell cytoplasm via nuclear pores to serve as sites for protein synthesis
Ribosomes functions
1) Site of translation’s
2) Protein synthesis
AKA reads messenger RNA (mRNA) and translates into amino acids to form proteins
Found in cytoplasm or bound to RER
RER functions
Protein synthesis
Post translational modifications of proteins (N-linked glycolysation)
Glycolysation
Modification process in which a carbohydrate covalently attaches to a target macromolecule, typically lipids (glycolipids) and proteins (glycoproteins)
N-linked glycolysation
Begins in RER
Co-translational mechanism
O-linked glycolysation
Occurs in Golgi after protein is translated, occurs post-translationally
SER functions
1) Synthesis and break down of glycogen
2) Detoxification of drugs, metabolic wastes, etc.
3) Synthesis of lipoproteins, cholesterol, bile salts
4) Synthesis of steroid hormones
5) Uptake and release of calcium in muscle cells
Golgi functions
1) Post-translational modification of proteins (from RER via vesicles), specifically O-linked glycosylation
2) Synthesis of lipoproteins
Mitochondria functions
1) Energy production (oxidative phosphorylation)
2) Beta oxidation of long chain fatty acids
3) Steroid hormone synthesis
4) Regulate apoptosis
5) Storage and release of calcium
Lysosomes functions
1) Heterophagy (phagocytosis and degradation of bacteria by neutrophils)
2) Bone remodeling via osteoclasts
3) Autophagy (destructions of worn-out organelles)
Peroxisomes functions
1) Beta oxidation of long chain fatty acids
2) Degrade hydrogen peroxide, a product of oxidative reactions
Lysosome storage dieseases
Caused by mutation in the gene that encodes for proteins in lysosomes (hydrolases, membrane proteins, co-factors). Results in accumulation of the substrates for lysosomal digestion
Zellweger’s/Cerebrohepatorenal Syndrome
Fatal disease due to absence of peroxisomal enzymes. Reduced degradation of cytotoxic hydrogen peroxide and abnormal accumulation of very long chain fatty acids. Skeletal muscle weakness, hypotonia, inability to nurse properly, neurological disorders, enlarged kidney, etc.
Plasma membrane functions
1) Physical barrier: establishes a flexible boundary
2) Selective permeability: regulates entry and exit of ions, molecules, nutrients
3) Electrochemical gradient: establishes and maintains an electrical charge
4) Communication: contains receptors
Carbohydrates functions (in plasma membrane)
1) Attach to proteins, forming glycoproteins
2) Forms cell coat (glycocalyx)
3) Establishes extra cellular microenvironment
4) Aids in metabolism, cell recognition, cell association, receptor sites for hormones, etc.