Biochem midterm #3
1/84
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
85 Terms
evolution of photosynthesis
Oxygenic photosynthesis generates O the atmosphere
Plants MAKE oxygen which heterotrophs USE for cellular respiration
Rise of aerobic respiration occurred after plants made O2
Autotroph - “Self-feeders”
Take simple molecules and make macromolecules to feed themselves and other organisms
Overview of photosynthesis + split into what?
Convert light energy into chemical energy
Plants take in CO2 and water, produce carbohydrates, and release O2
Split into
Light dependent reactions
NADP+ → NADPH
Carbon-assimilation (carbon fixation) reactions
ATP → ADP + Pi
CO2 → Triose phosphates
Compare/contrast chloroplast structure to the mitochondria. How does compartmentalization drive function?
Double mem
Stroma - like the mitochondrial matrix
Thylakoids - membrane-filled spaces (lumen)
Proton Pumping Direction:
Mitochondria: Protons are pumped OUT of the matrix into the Intermembrane Space.
Chloroplast: Protons are pumped IN from the stroma into the Thylakoid Lumen
explain proton pumping direction for photosyn vs. cell resp
Mitochondria: Protons are pumped OUT of the matrix into the Intermembrane Space.
Chloroplast: Protons are pumped IN from the stroma into the Thylakoid Lumen.
properties of excited electrons
caused when a photon hits a ground state e-
Increased potential energy
Transfer energy (excitons)
Easier to transfer (oxidation)
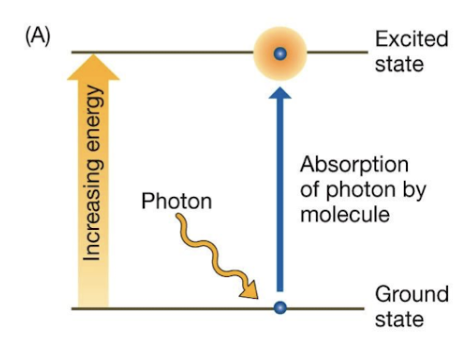
Things that can happen from excited→relaxed state
Heat
Fluorescence
Energy transfer to neighboring P molecule
Redox reactions
Pigments definition
Molecules that absorb specific wavelengths of light
chlorophyll structure
contain a porphyrin ring with conjugated double bonds
Conjugated = alternating single and double bonds
Absorb photons within the visible spectrum
Minimize energy loss through thermal vibration (de-excitation)
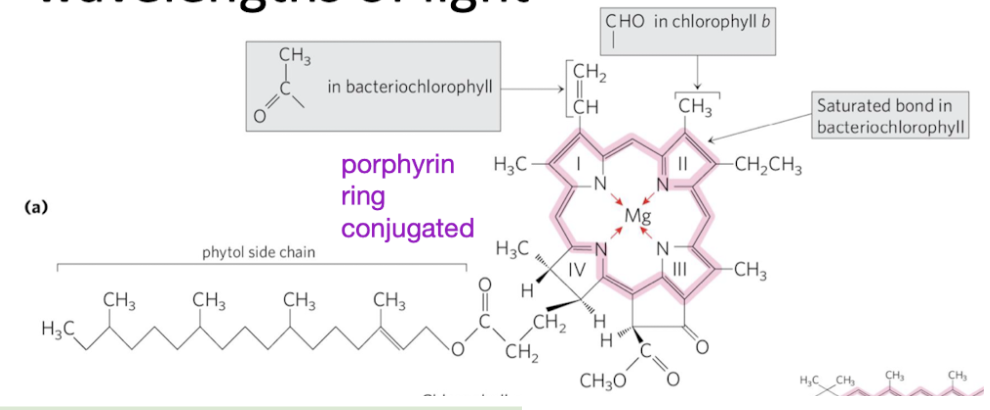
which wavelengths/colors do chlorophylls absorb well
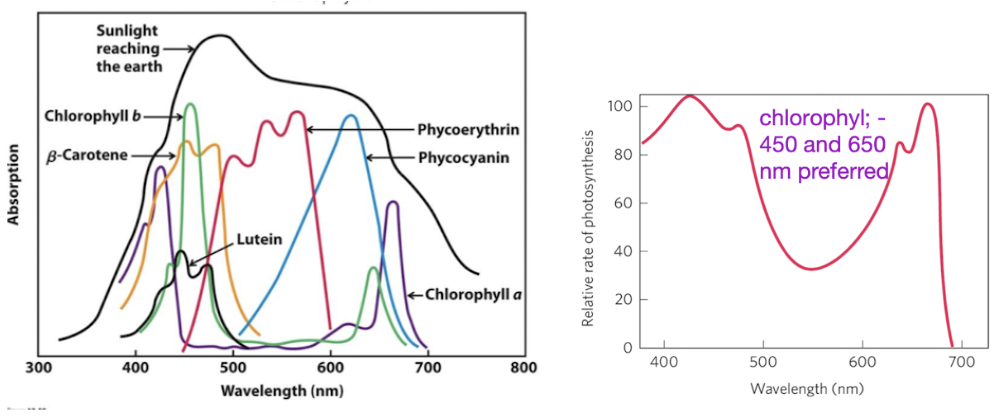
Chlorophyll absorbs well in blue/red regions, reflects green
450 and 650 nm preferred
Accessory pigments help absorb the others
accessory pigments
extend the range of light absorption
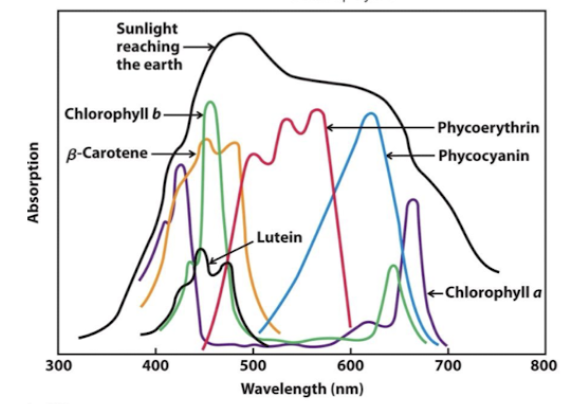
absorption spectrum correlation with photosynthesis
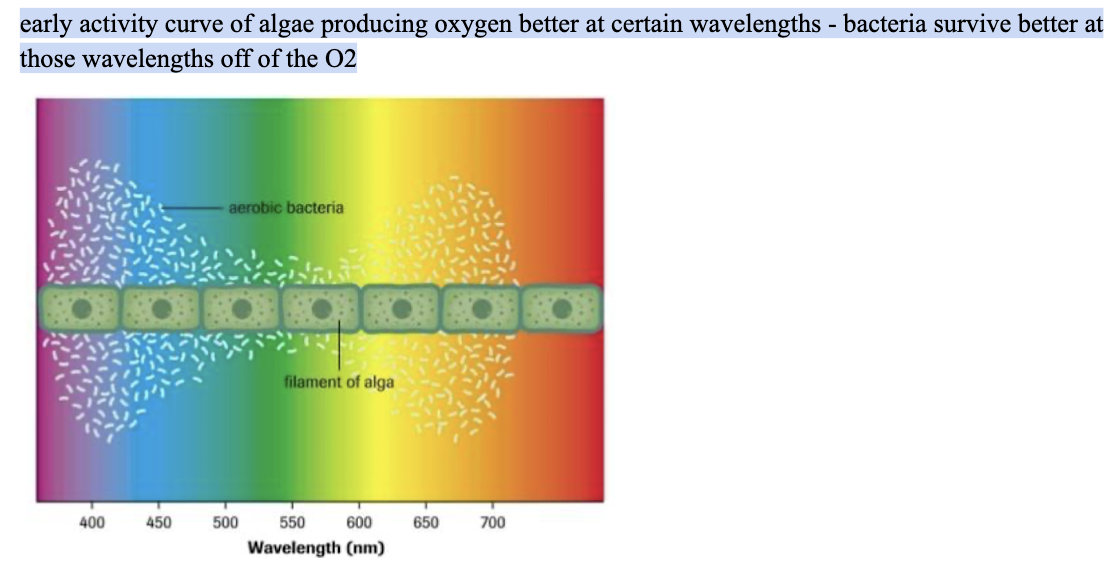
Light Harvesting Complexes location
thylakoid membrane
pigments location
bound to Light Harvesting Complexes within the thylakoid membrane
Antenna complex
provides a network of pigment molecules to capture and transfer energy.
No matter where the light hits, the light can be transferred to neighboring molecules
Will eventually hit the reaction center → like a funnel to the reaction complex
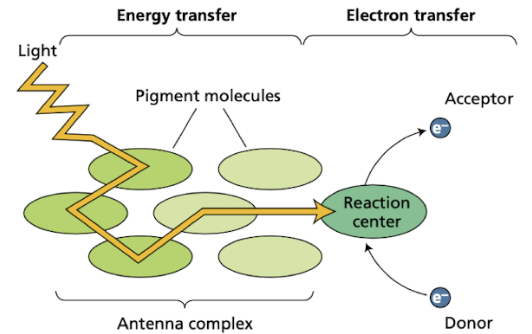
Reaction center
pigments where absorbed light energy drives electron transfer
Antenna complex leads to the reaction center
Reaction center chlorophyll enables efficient redox reaction
Light excites and relaxes e- in antenna molecules until it reaches the reaction center and excites it
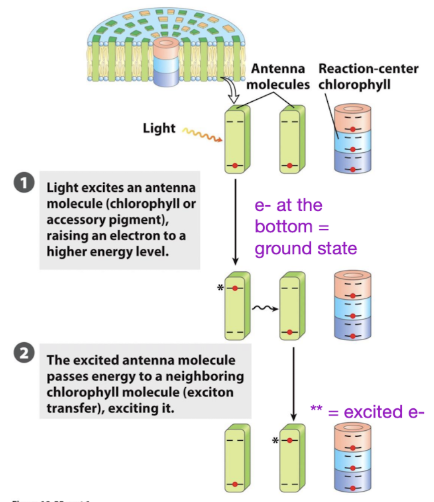
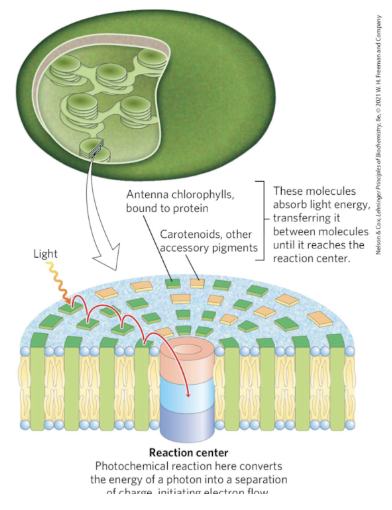
Photosystems
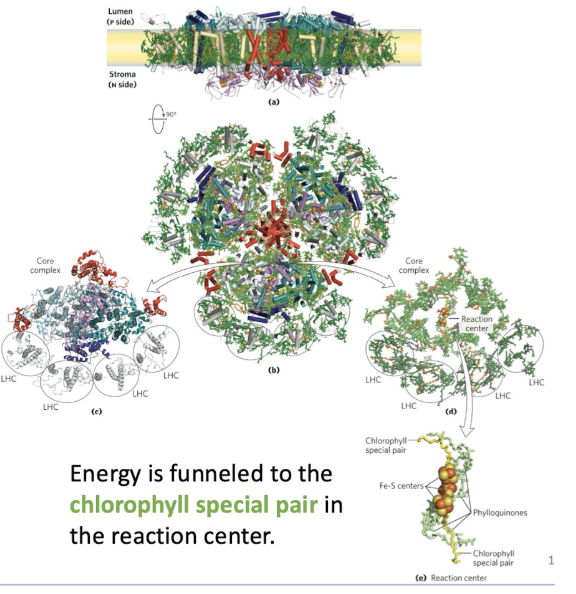
energy harvesting proteins that facilitate transfer of excited electrons
multiple Light Harvesting Complexes (LHC) that direct energy to the core complex.
Separate subunits that contribute to the photosystem
Energy is funneled to the chlorophyll special pair in the reaction center.
Middle of the photosystem has the reaction center

reaction center structure
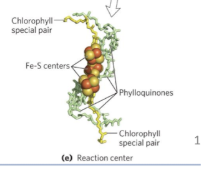
Reaction center chlorophyll enables efficient redox reaction
Excitation causes a separation of charges in the reaction center and initiates a redox chain.
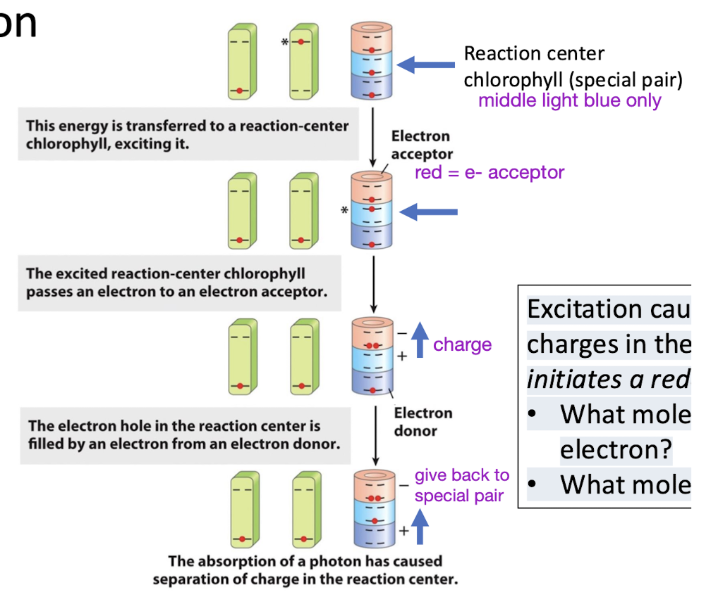
Photosystem II location
within thylakoid membrane
Photosystem II function
first protein complex in light-dependent reactions of photosynthesis
Reduce PQ
Special pair passes e- up to PQ
PQ = plastoquinone
OEC - oxygen evolving complex
Take e- from H2O (oxidize) → produce oxygen
Replace the e- in the reaction center chlorophyll with the e- from H2O
Oxygen-evolving complex in photosystem II (OEC)
Sends the e- from the H2O to the reaction center chlorophyll
Take e- from H2O (oxidize) → produce oxygen
Replace the e- in the reaction center chlorophyll with the e- from H2O
manganese ions (Mn)
Mn complex oxidizes H2O
In lumen (inside of the thylakoid
Every 4 flashes/excitations → produce 1 O2
Water re-reduces Mn atoms in OEC in a single step to produce O2
Mn center donates e-
Getting excited + pulling e- off of water
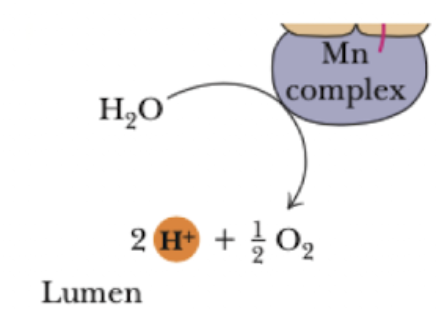
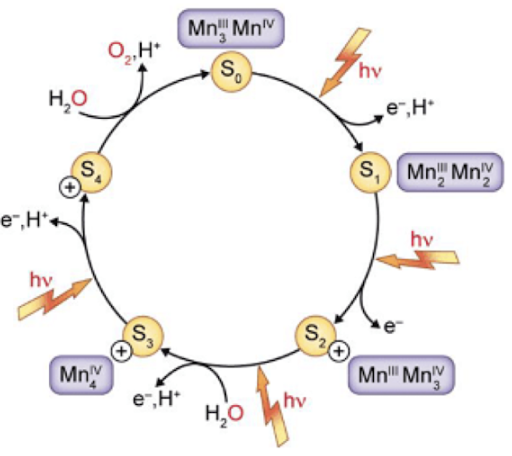
photosystem II importance/facts/structure
first protein complex in light-dependent reactions of photosynthesis
special pair = P680
includes Phe and PQ
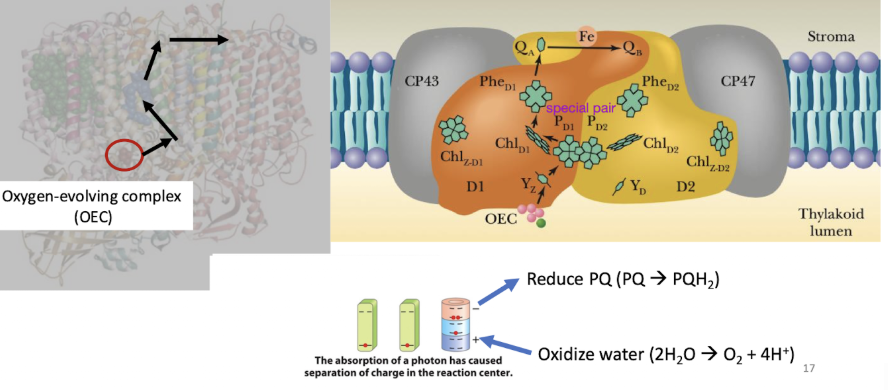
Cytochrome b6f complex
COPY AS COMPLEX III in ETC -
moves H+ from stroma to lumen
Q cycle type thing for PQ instead
Problem:
Plastohydroquinone (PQH 2) transports 2 e-
plastocyanin (PC) can only transport one e- at a time.
flow of e- in thylakoid membrane light reactions
H2O
photosystem II
cytochrome b6f complex
photosystem I
Fd shuttle
NADP reductase - produce NADPH
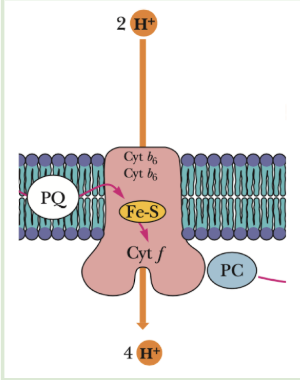
Plastoquinone
Same as ubiquinone :D
Transports 2 e-
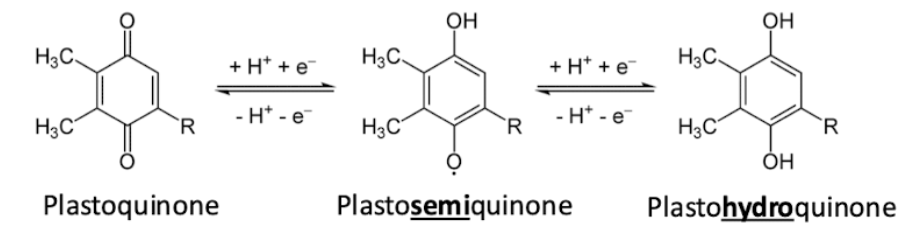
2-step PQ cycle - steps + results
steps
first QH2 oxidized, Q- produced
second QH2 oxidized, regen another QH2 from the Q- radical
Overall
Remove 2 H+ from the stroma
Pump 4 total H+ into the thylakoid lumen
Move both electrons from QH2
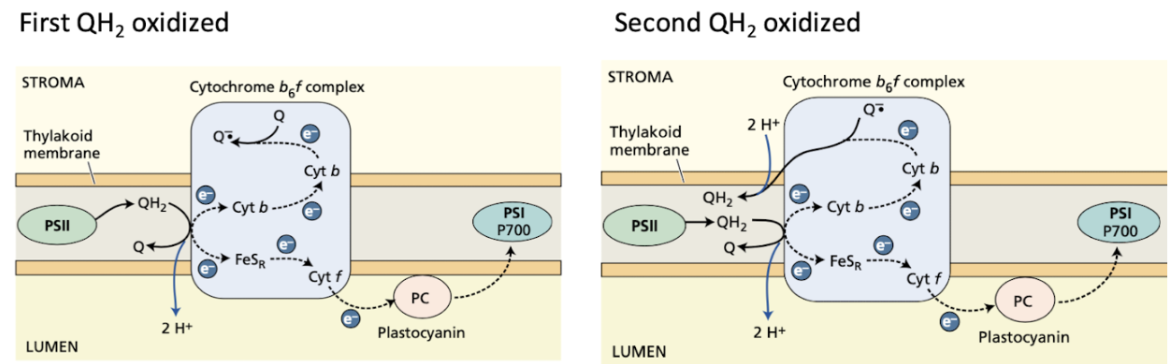
Photosystem I facts
Steps
Accepts e- from PC
Excite e-
Transfer e- to Ferredoxin (Fd)
Ferredoxin (Fd)
Shuttles e- NADP Reductase to produce NADPH.
Shuttle e- to Cyt b6f complex to produce ATP
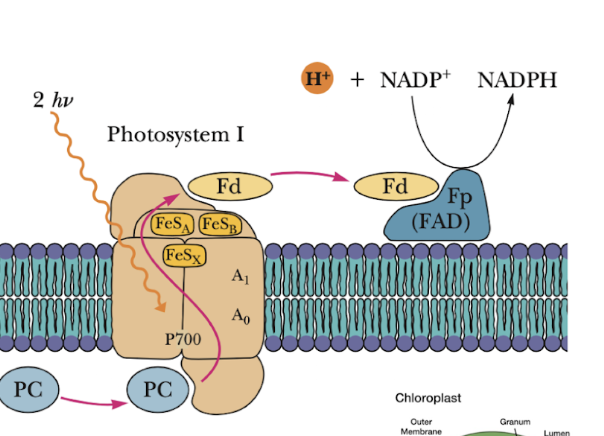
Ferredoxin (Fd)
shuttles between photosystem I to NADP reductase
producs NADPH
ALSO - Shuttle e- to Cyt b6f complex - makes H+ gradient to produce ATP
Z-scheme in light reactions
Tracking e-
Increase in E when e- are excited at the photosystems
Small drops in E when transferring carriers
Photosystems don’t pump any H+, only cytochrome b6f complex
Fd can go to FNR → NADPH
OR back to cytochrome b6f complex
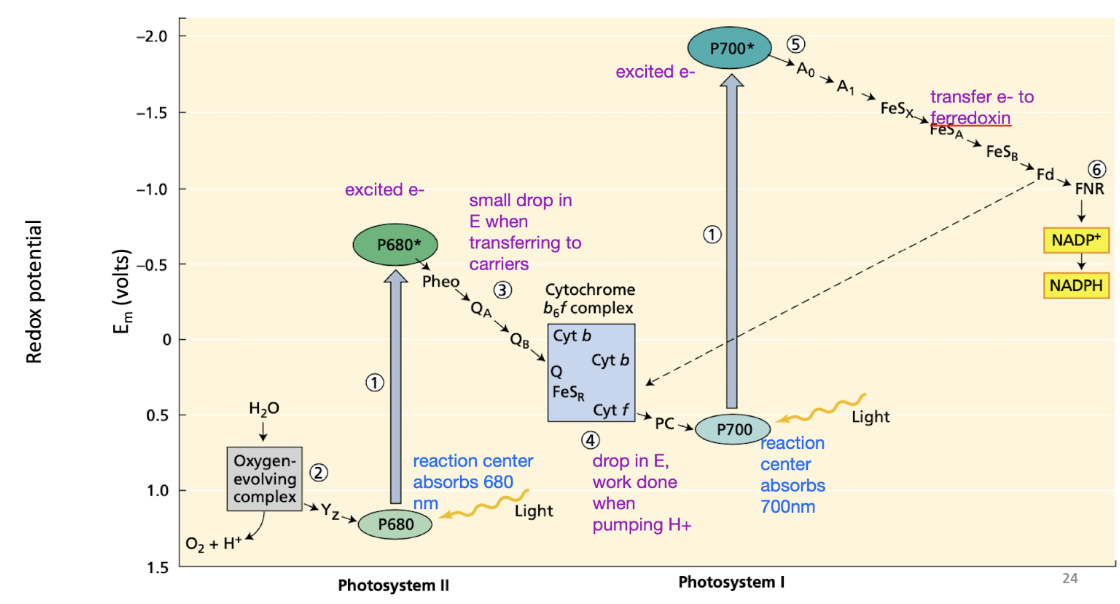
what can pump H+ in thylakoid membrane light reactions
only cytochrome b6f complex
Phosphorylation step of thylakoid membrane light reactions
light generates a proton gradient that drives ATP synthase
via cyt b6f complex
ATP synthase SAME structure
H+ move from LUMEN to STROMA
12 H+ drive full rotation
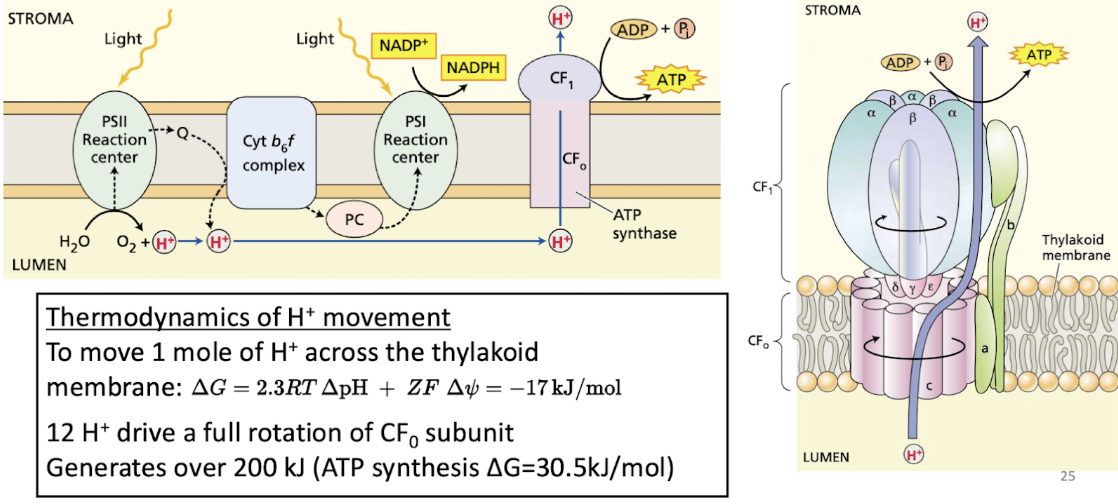
what direction do H+ move in thylakoid membrane light reactions?
cyt b6f - stroma to lumen
ATP synthase - lumen to stroma
linear flow of e- products (light reactions)
1 ATP and 1 NADPH produced
energy requirement of calvin cycle to build sugars?
9 ATP 6 NADPH - 3:2 ratio of ATP to NADPH
Need a proton gradient to make ATP
Need more ATP than NADPH
Solution: Fd → decision making point, it can transport e- to make NADPH or deliver it back to Cyt b6f
2 systems of e- transport for prep for Calvin Cycle
Plants utilize both to maintain 3:2 ATP:NADPH required for carbon fixation.
Noncyclic electron transport
Photosystem II and Photosystem I are required
results in production of NADPH and ATP
Cyclic electron transport
Photosystem I
results in production of ATP but NOT NADPH
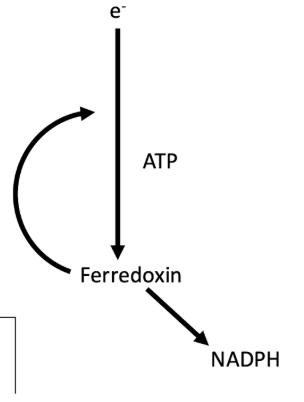
Noncyclic electron transport
Photosystem II and Photosystem I are required
results in production of NADPH and ATP
electrons flow from water —> PSII —> cyt b6f —> PS1 —> NADPH
Cyclic Electron Transport
only Photosystem I required
results in production of ATP but NOT NADPH
Fd shuttles back to cyt b6f complex to pump H+
used by ATP synthase to make ATP
no NADPH made because doesn’t pass by PSII
how are the light reactions regulated?
appressed and nonappressed thylakoids
appressed thylakoids
Stacked tight
Happens when they’re being hit by a LOT of light
Intense production of H+ gradient
Packed with PSII and LHC (light harvesting complexes) to optimize light capture
LHC mediates connection between membranes
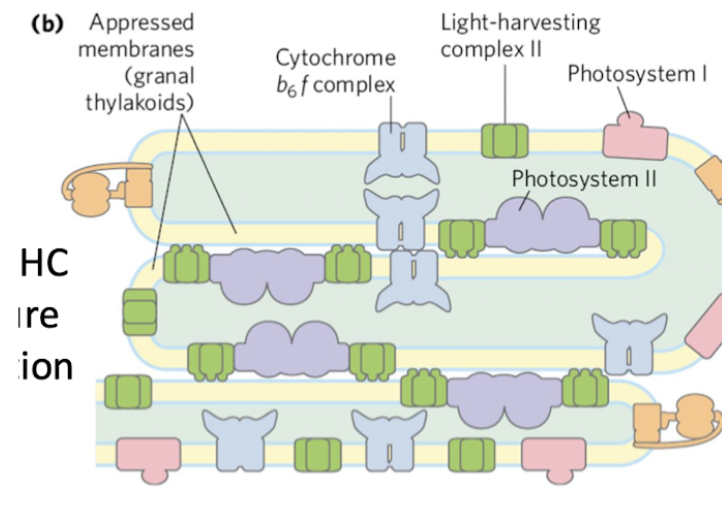
nonappressed thylakoids
Unstacked regions with access to stroma (ferredoxin, NADP+, ADP)
PSI and ATP synthase
More stroma access → more ATP synthase
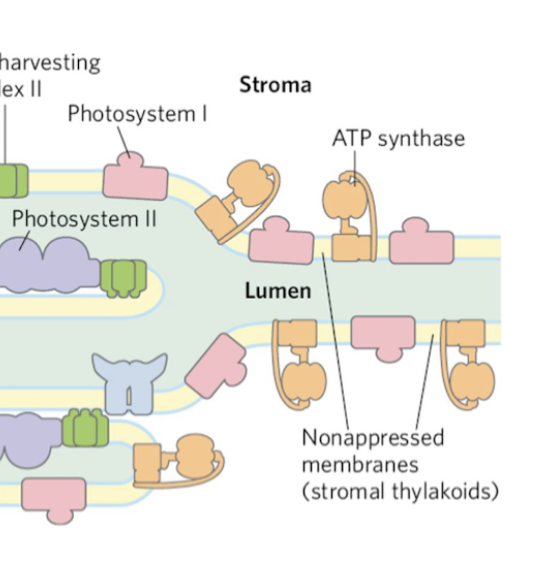
how do the thylakoids stack/appress?
dynamic in response to light
Controlled by covalent modifications - phosphorylation
LHC mediates the connection between membranes
Span thylakoid membrane
Thr-OH residue anchors it
Phosphorylation of the Thr residue releases it to nonapressed state
LHC are mobile and can help funnel energy to PSI when PSII is more active
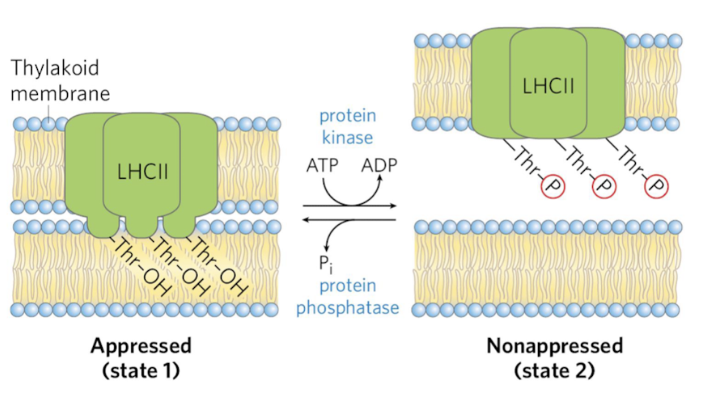
Stoichiometry of CO2 assimilation in Calvin Cycle
For every 3 CO 2 molecules fixed:
9 ATP and 6 NADPH are consumed
1 glyceraldehyde 3-phosphate is ‘produced’.
6 are acutally made, but 5 are re-used for the cycle
Remember: Plants balance cyclic and non-cyclic electron transport to maintain this 3:2 ATP:NADPH ratio.
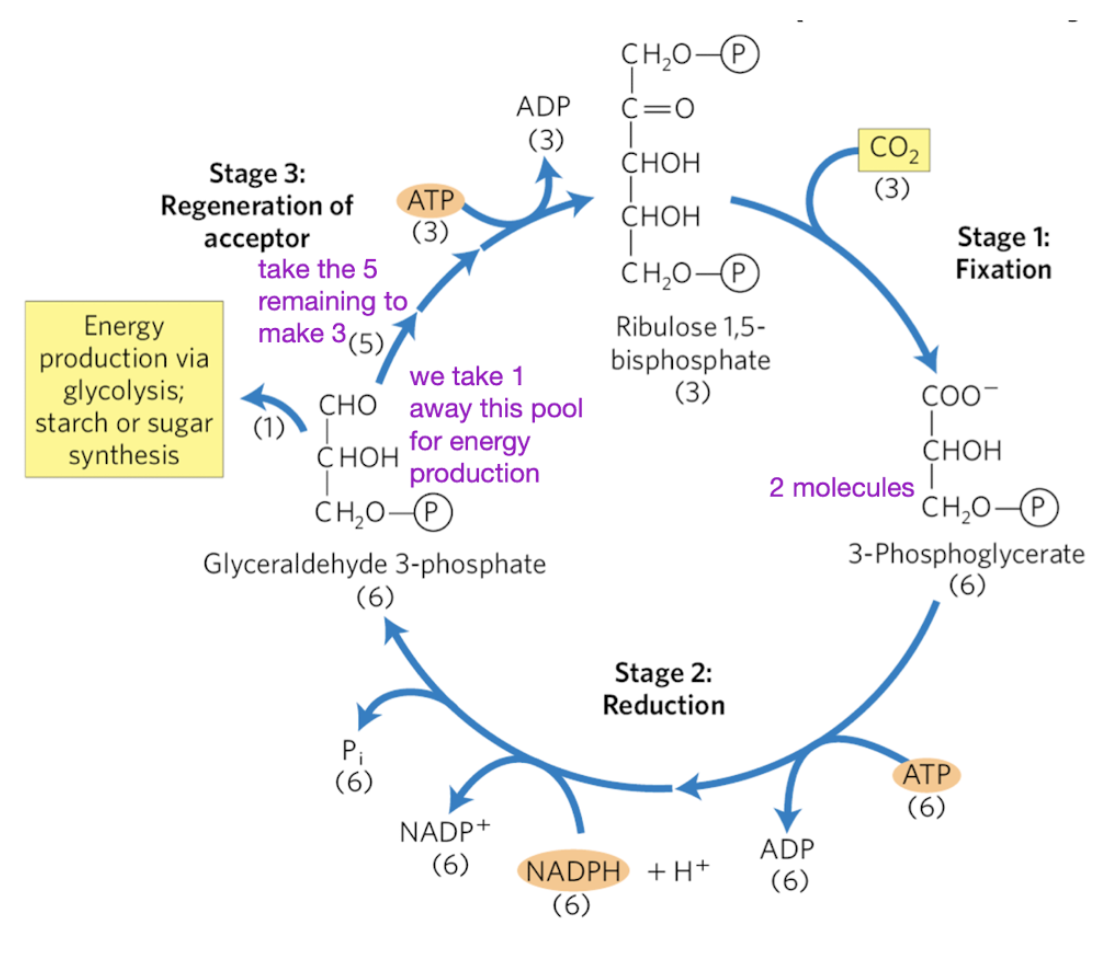
stage 1 calvin cycle
CO2 fixation to 3-phosphoglycerate (3-PG)
3 R1,5BP + 3 CO2 —> 6 3PG
uses RuBisCO - Ribulose 1,5-bisphosphate carboxylase/oxygenase
Very slow: Large amounts are needed to achieve high carbon fixation rates.
~50% of soluble protein in chloroplasts
Forms alternative products in the presence of oxygen
If O2 binds, there will be a reaction still
what is RuBisCO + what’s it used for
Ribulose 1,5-bisphosphate carboxylase/oxygenase
most abundant protein (enzyme) on earth
fixates CO2 onto ribulose 1,5-bisphosphate —> 3PG
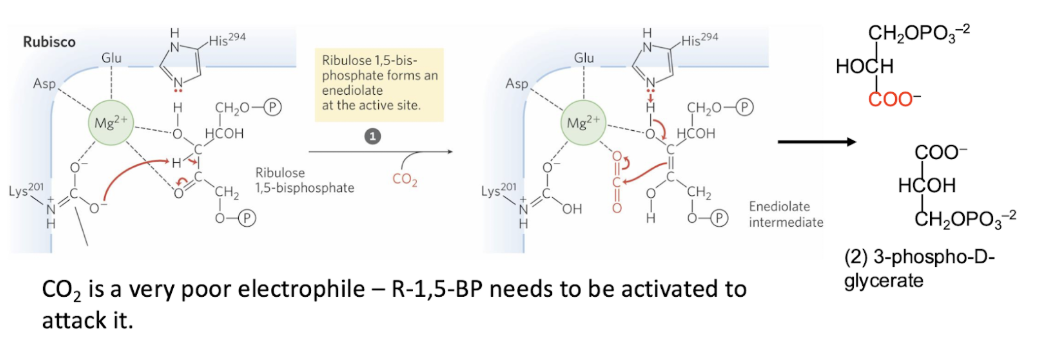
Carboxylase activity of RuBisCO
CO2 is a very poor electrophile – R-1,5-BP needs to be activated to glycerate attack it.
Enediolate intermediate → extremely reactive, used to activate R1,5BP
So that the reaction can more easily proceed
Problem: once it is primed for attack, it cannot discriminate between CO2 and O2
Solution: plants have evolved different mechanisms to increase [CO2] in leaf tissue
stage 2 of calvin cycle
reduce 6 3-PG —> 6 G3P
uses 6 ATP and 6 NADPH
Note: we need 5 G3P to regenerate RuBP
1 G3P is used to make other stuff like glucose, cell wall, etc
Phosphoglycerate kinase
Uses ATP to convert 3PG → 1,3 BPG
Phosphoglycerate kinase
Uses ATP to convert 3PG → 1,3 BPG + NADPH —> G3P
stage 2 of calvin cycle
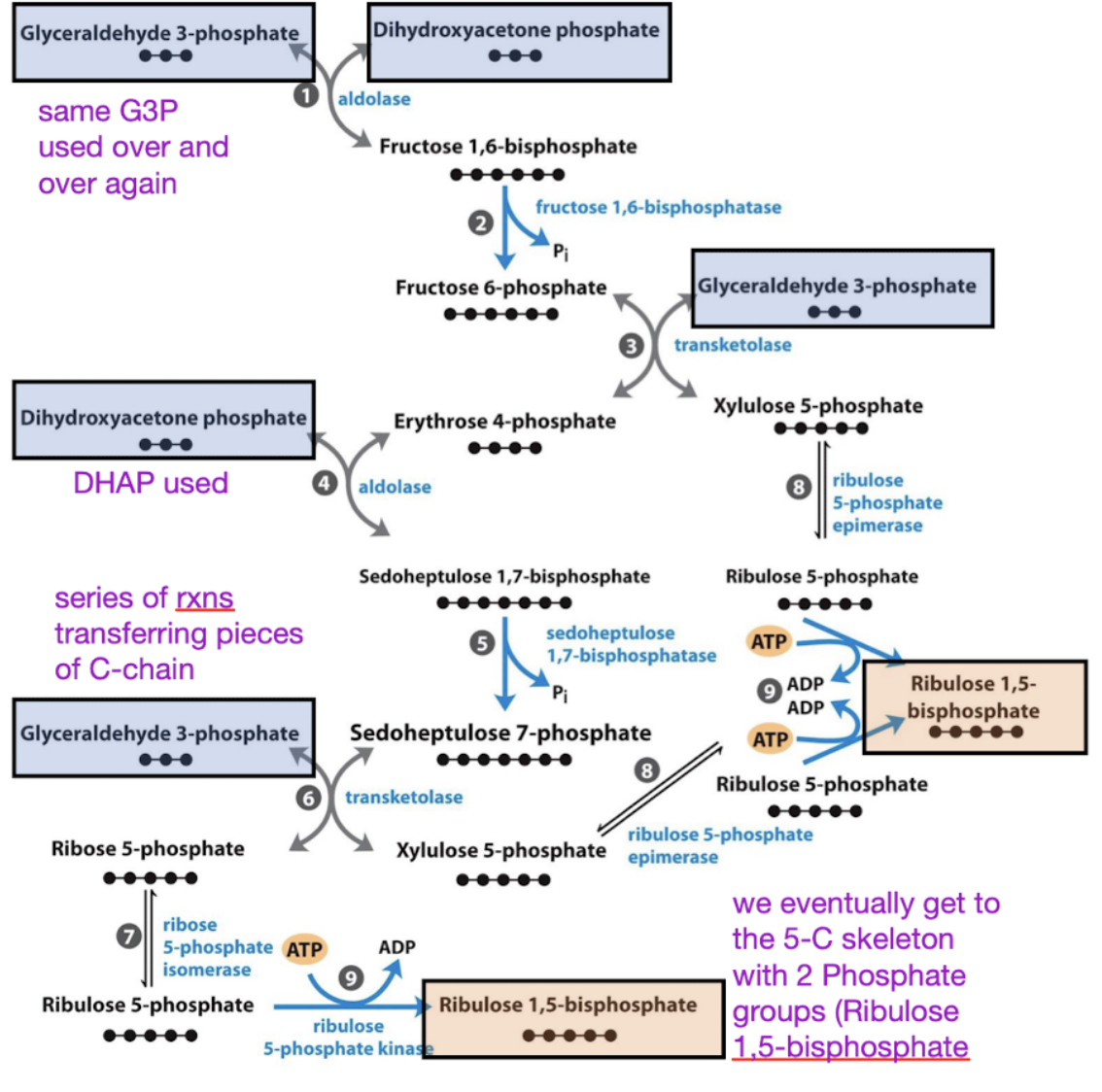
stage 3 calvin cycle
regeneration of 3 ribulose-1,5-bisphosphate using 3 ATP
3 ATP + 5 G3P —> 3 R1,5BP
Problem: RuBP is a pentose (5C), but G-3-P and DHAP are 3C.
Take home: Triose phosphates interconvert to form pentose phosphates.
Note: Blue arrows are exergonic steps that make the entire process irreversible.
G3P and DHAP transfer C’s between each other to make a 5
ribulose -5-phosphate kinase
Uses ATP to convert ribulose-5-phosphate → ribulose-1,5-bisphosphate
ribulose -5-phosphate kinase
used in generation of R1,5BP
Uses ATP to convert ribulose-5-phosphate → ribulose-1,5-bisphosphate
how are Enzymes in Calvin Cycle are indirectly activated by light
Remember: the thylakoids shift between the appressed and nonappressed states
Thylakoid lumen can regulate the amount of Mg
This changes the amount of RuBisCO active sites
Increasing Mg2+ increases RuBisCO activation
Fructose 1,6-bisphosphatase (Stage 3) is pH dependent and requires Mg2+.
In light conditions, its activity increases more than 100- fold.
When chloroplasts are illuminated:
Stromal [Mg2+] increases (2-3mM → 6-8mM)
Stromal pH increases (ph 7 → pH 8)
What happens when chloroplasts are illuminated? impact on calvin cycle?
stromal Mg2+ and pH increase
activity of stage 3 enzyme fructose 1,6 bisphosphatase + RuBisCO INCREASE
what are the 4 fates of glucose?
synthesis of structural polymers - cell wall, matrix
storage - glycogen, starge, sucrose
PPP - form ribose 5-phosphate
glycolysis - pyruvate + reverse=gluconeogenesis
list of tissues diff function for glucose fate + takehome
brain - depends on blood sugar
liver - 90% of GNG
skeletal muscle - glycolysis
kidney - 10% of GNG
brain usage of glucose
High-priority glucose user.
No glycogen stores
cannot do GNG.
Depends on blood sugar.
liver glucose usage
Stores glycogen to share and performs
90% of GNG to maintain blood sugar
Provides the blood glucose for the brain and other parts of the body
skeletal muscle glucose usage
Burns glucose for ATP.
Stores glycogen but won’t export glucose (won’t share with anyone else, cannot give glucose to other organs like brain)
Mostly performs glycolysis using the glycogen
Will export lactate under anaerobic conditions.
kidney glucose usage
Performs about 10% of GNG, mostly during prolonged fasting.
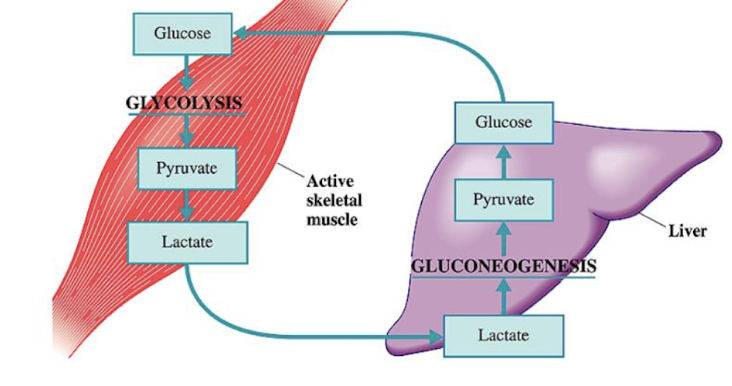
Cori Cycle
liver does GNG to produce blood sugar
blood sugar used by brain and muscles
skeletal muscle uses the glucose in glycolysis
Uses O2 and glucose so quickly → switches to anaerobic respiration
Muscles export lactate into blood → liver uses it to make glucose
how are sugars catabolized?
Glycolysis is central to all of metabolism
Each mono or disaccharide does not have its own catabolic pathway
They feed into glycolysis in the shortest path possible
Digesting carbohydrates → try to get it into a form that is a glycosidic intermediate
If you convert into a glycosidic intermediate → can be digested via glycolysis and harvest E
Polysaccharides are also broken down into monosaccharides that feed into glycolysis
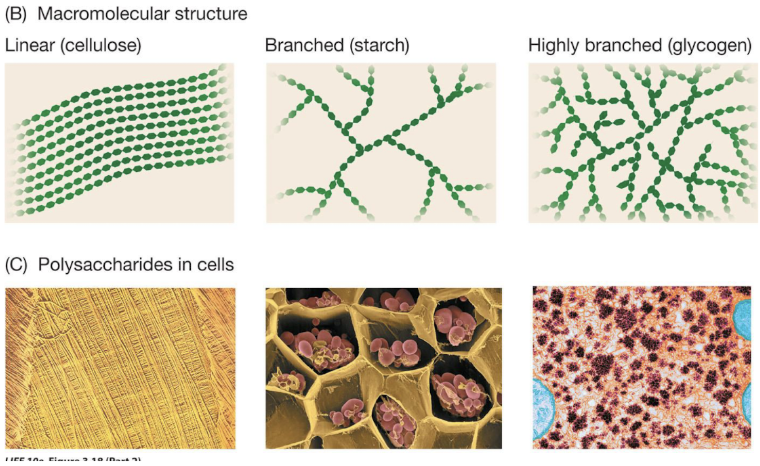
marcomolecular structure of carb polymers
Macromolecular structure
Cellulose - linear
Starch - branched
Glycogen - highly branched
Types of glycosidic linkages differ, as do the branching patterns
Glycogenesis overview
makes glycogen
20% liver, 80% muscle
Solves a key osmotic problem for cells:
If stored as free glucose, the amount of water required would burst the cell!
Enables parallel processing:
thousands of non-reducing ends can be cleaved simultaneously when the cell needs glucose
glycogen
produced in glycogenesis
main storage polysaccharide in animal cells
alpha 1-4 linked glucose subunits
heavily BRANCHED
alpha 1-6 branches
contains glycogenin protein seed to start polymers
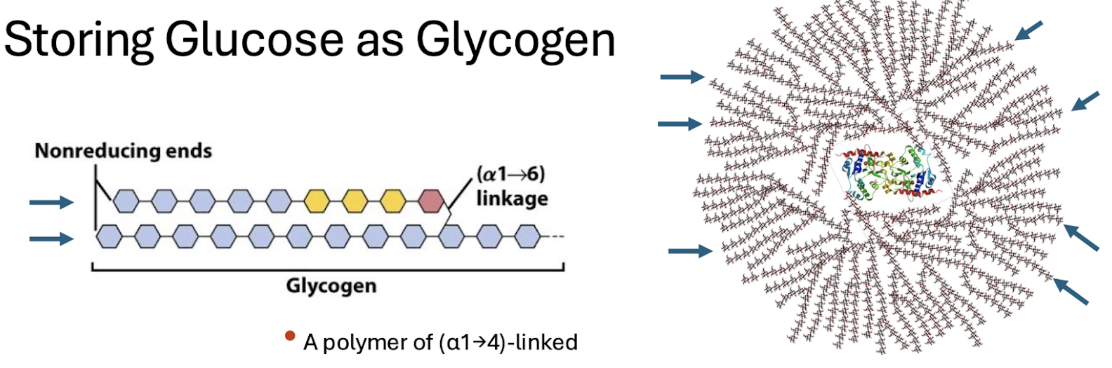
glycogenin
protein core = “seed” in the center of glycogen that can help start polymers
how do we prime glucose for glycogenesis
activate it
Isomerize to glucose-1-phosphate Building a polymer of (α1→4)-linkage
move phosphate from 6C to 1C for the a1—>4 linkages
add uracil phosphate —> UDPG
adds another phosphate group to the C1 phosphate
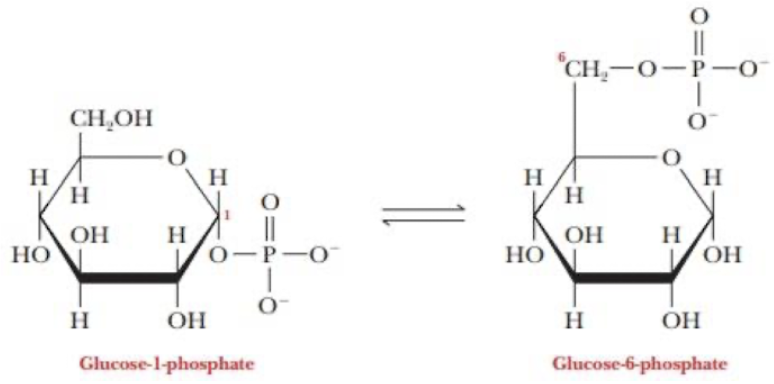
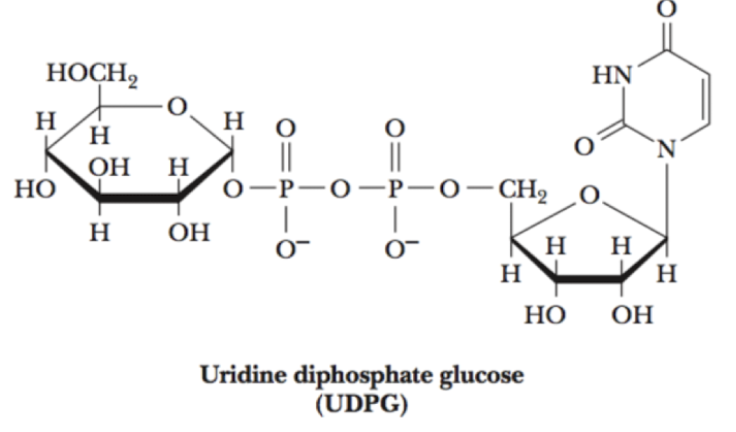
why do we add P and UP to glucose?
we add phosphate and uracil phosphate —> UDPG
Binding Energy:
the nucleotide group forms favorable interactions with enzymes, and the free energy of nucleotide group binding enhances catalytic activity.
Reactivity:
the UDP is a good leaving group. The attached sugar carbon is activated for nucleophilic attack.
Separate Pools:
tagging hexoses with nucleotides sets them aside for biosynthesis separate pools are available for energy production and storage.
what is the role of sugar nucleotides in biosynthesis
Making sugar nucleotides is a common substrate for polymerization
Formation of sugar nucleotides is highly exergonic and essentially irreversible, driven by the large free energy of PPi hydrolysis.
Break off the PPi
Sugar phosphate attacks the NTP → removing the PPi
PPi
Concentration of pyrophosphate very low
Very high energy
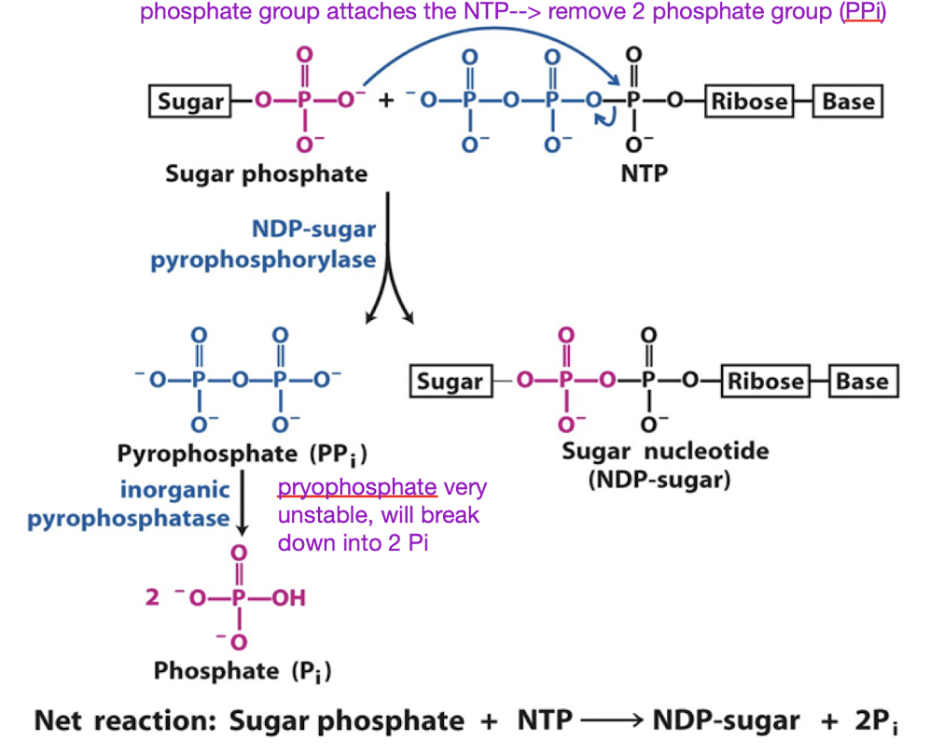
Glycogen synthesis (glycogenesis) steps by glycogenin
Glycogenin adds the first 7 glucose molecules
Remains covalently bound to the reducing end of the completed glycogen molecule
UDP-glucose
Tyr on glycogenin attaches to C1 of UDP-glucose
Glucosyltransferase adds another UDP-glucose
C4 of the 1st UDP-glucose attaches to C1 of another one
repeats 6 times
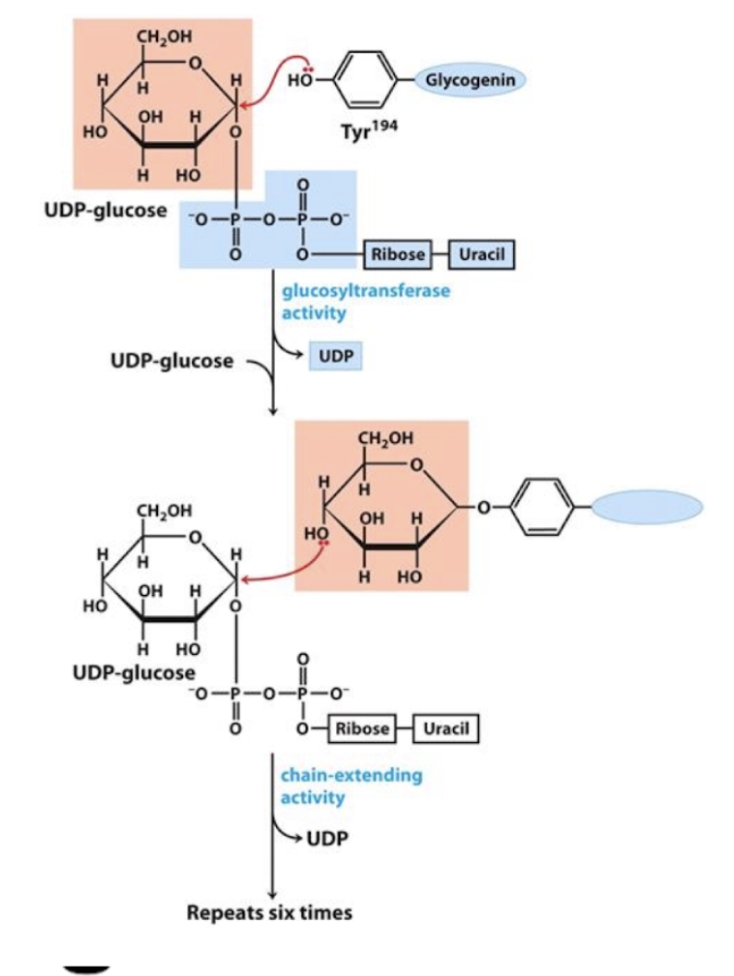
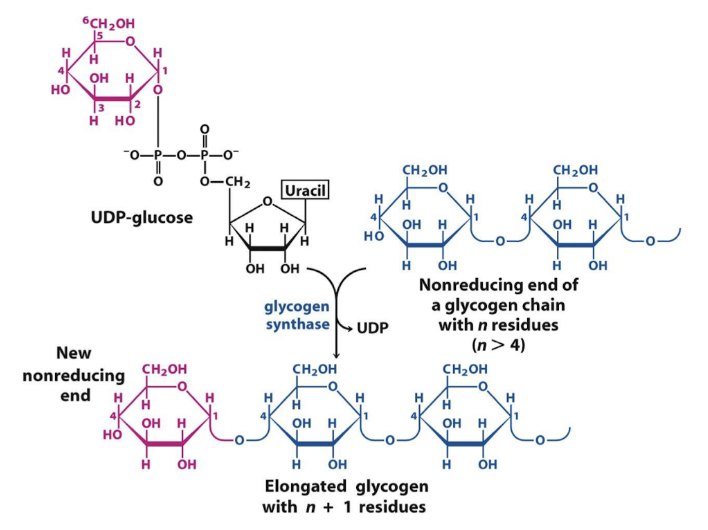
glycogen synthase
catalyzes alpha 1,4 linkages
Glycogen synthase does NOT do BRANCHING, just lengthens the chain
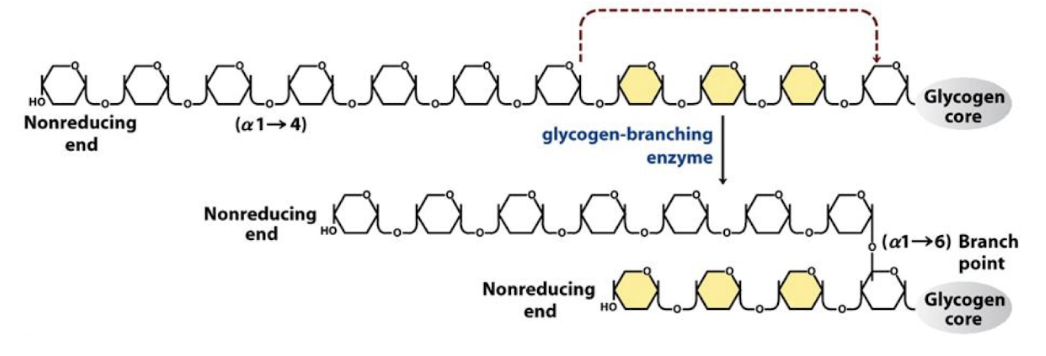
Glycogen-branching enzyme
Synthesizes alpha 1-6 linkages → BRANCHING
Glycogen-branching enzyme catalyzes:
transfer of a terminal fragment (6 or 7 residues long) from the nonreducing end of a branch to the C-6 hydroxyl group of a glucose residue on the same chain or another chain
creating a branch with an (α1→6) linkage
Cut alpha 1-4 link and make it alpha 1-6
Have 2 nonreducing ends → can add more reducing glucose to these ends
overall steps of glycogenesis
take glucose 6-phosphate —> glucose 1-phosphate
glucose 1-phosphate + uracil-phosphate —> UDPG
7 UDPG connect using glycogenin
Glycogen synthase extends linearly a1-4
glycogen branching enzyme adds branches a1-6
Phosphorolysis overview
degrade glycogen
Formation of a phosphate ester preserves some bond energy
uses 3 enzymes
glycogen phosphorylase
transferase
glucosidace
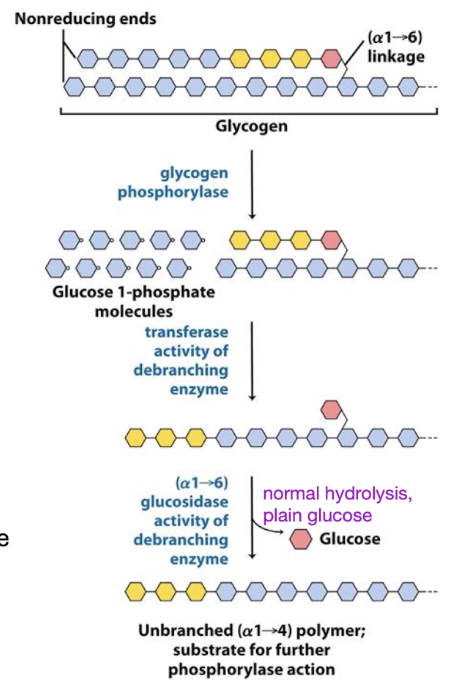
glycogen phosphorylase
degrades glycogen
processive enzyme - catalyzes consecutive reactions without releasing its polymeric substrate
Sequentially removes terminal residues → G1P
Stops 4 residues from branch point
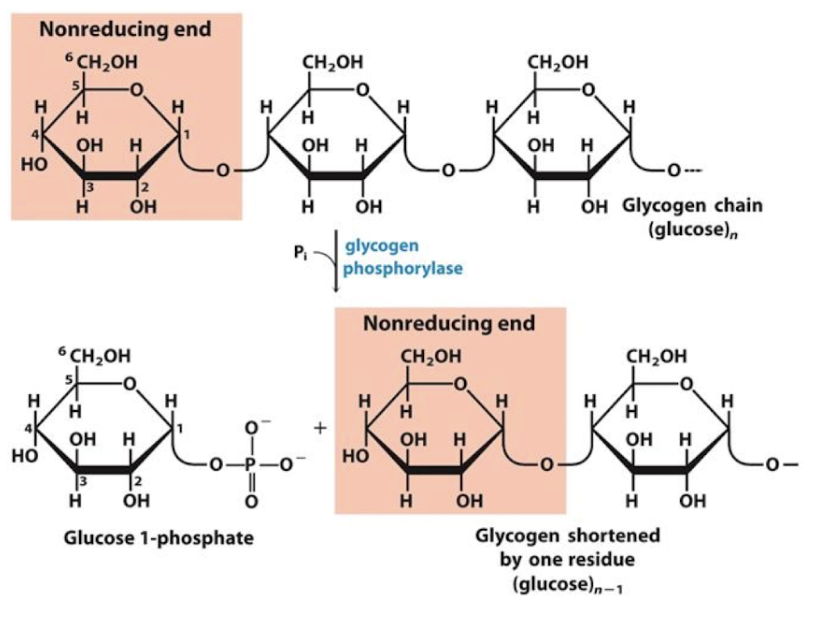
processive enzyme
catalyzes consecutive reactions without releasing its polymeric substrate
Transferase
glycogen breakdown
debranching enzyme
shifts a block of 3 glucose residues to nonreducing end of the same or a different glycogen molecule
leaves 1 glucose behind
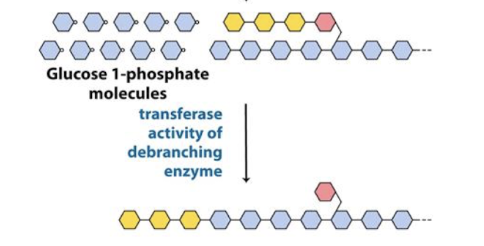
glucosidase
glycogen breakdown
debranching enzyme that resolves
a(α1→6) linkage → glucose (not G-1-P)
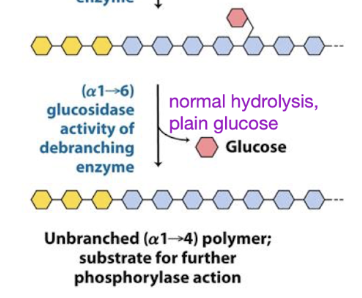
reciprocal regulation of glycogen metabolism
Glycogen phosphorylase (breakdown) and glycogen synthase (production)
Opposing pathways → if one is activated, the other is inhibited
Glucagon = low energy, need to increase blood sugar
Glucose = high energy
The conditions that activate glycogen synthesis deactivate glycogen breakdown
phosphorylase
inhibited by glucose, insulin(high energy, need less glucose)
activated by glucagon (low energy, need more glucose)
synthase
inhibited by glucagon (low energy, need to store less)
activated by glucose, insulin (high energy, store and make glycogen)
why can’t we just reverse glycolysis to make glucose?
In the cell, steps that are close to equilibrium are reversible
Highly exergonic steps are irreversible (1, 3, 10)
bypass reactions
The three irreversible reactions of glycolysis are bypassed to synthesize glucose from pyruvate in gluconeogenesis
Use a different set of enzymes
Are exergonic and effectively irreversible
Are reciprocally regulated
3 bypass reactions:
First bypass: (step 10 glycolysis)
Pyruvate → phosphoenolpyruvate
Second bypass: (step 3 glycolysis)
Fructose 1,6-bis-P → fructose 6-phosphate
Third bypass: (step 1 glycolysis)
Glucose 6-phosphate → glucose
First Bypass overall reaction and 2 pathways
Pyruvate → phosphoenolpyruvate
Pyruvate into matrix → oxaloacetate → malate → export out into cytoplasm using malate-aspartate shuttle → oxaloacetate
Have to move the reducing equivalents from matrix into cytoplasm
NADH
Lactate in cytoplasm → oxidize into pyruvate → export into matrix
Use NAD+ to lactate → pyruvate + NADH (in cytoplasm)
Lactate better because NADH generated DIRECTLY in cytosol
Cori cycle → USED BY LIVER
Both pathways are required to balance redox equivalents and PEP to start gluconeogenesis
Overall: energy expensive
Requires 2 ATP to synthesize 1 PEP
In glycolysis, we gain 1 ATP from 1 PEP
problem about the first bypass
Pyruvate is imported into the mitochondrial matrix, but the rest of our pathway is in the cytoplasm.
Pyruvate carboxylase produces OAA that is available for anabolic reactions.
Produced in anaplerotic reaction
Other OAA is consumed in CAC
Also part of the shuttling system → malate-aspartate shuttle
Malate-aspartate shuttle → moves the malate + NADH out of the mitochondria
requires NADH - which has a low concentration in cytosol
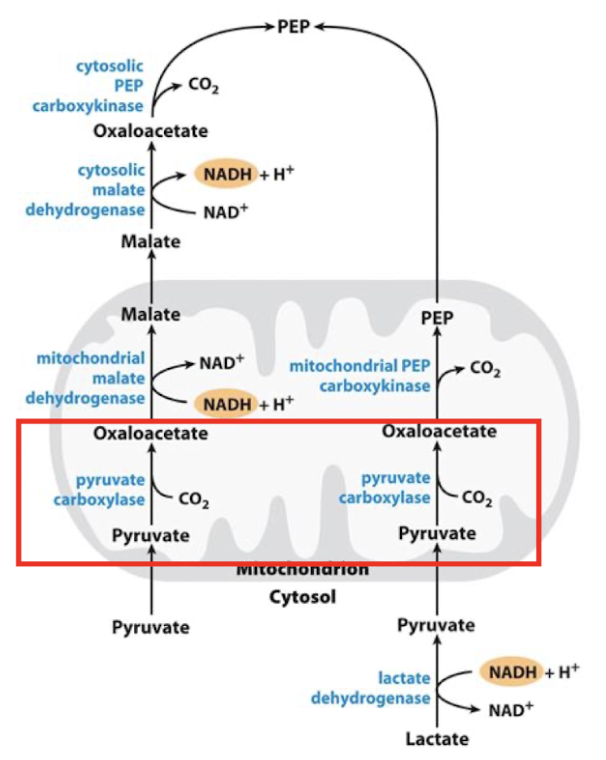
Step 1: Pyruvate → oxaloacetate for 1st bypass in GNG
First, pyruvate is transported into the mitochondria
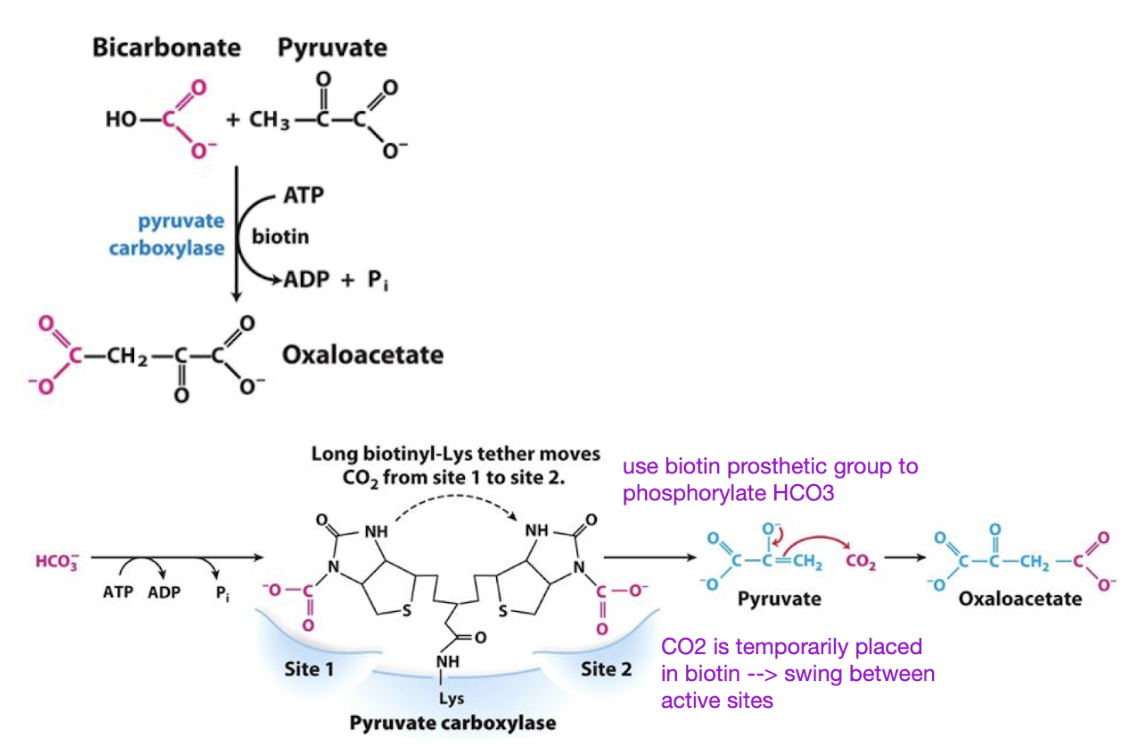
Pyruvate carboxylase is the first regulatory enzyme in these pathways and is stimulated by acetyl-CoA. Why?
High acetyl-CoA = energy rich, CAC not running = ready for anabolism to store energy
Pyruvate oxidation = makes acetyl CoA
Biotin - swings between 2 active sites to place the CO2
Add carboxylic acid to pyruvate = oxaloacetate
Costs 1 ATP
Step 2: oxaloacetate (OAA) → PEP in GNG
Happens either in cytosol or in mitochondria
Longer pathway in cytosol
Shorter pathway in mitochondria
PEP is the highest-energy phosphorylated intermediate in the cell
requires two highly exergonic steps to drive its synthesis.
Use 1 GTP
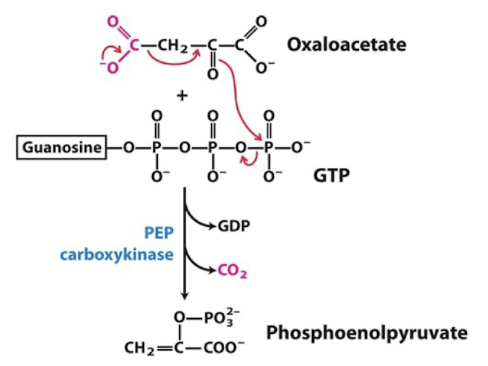
first bypass is highly regulated