Genetics Unit 3
1/39
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
40 Terms
Dominant
Shows when only one copy is present
Recessive
Shows when two copies are present
Haplosufficient
Wild-type dominant if one copy is enough for normal function in heterozygotes
Haploinsufficient
Wild-type recessive if one copy is not enough for normal function
WT baseline
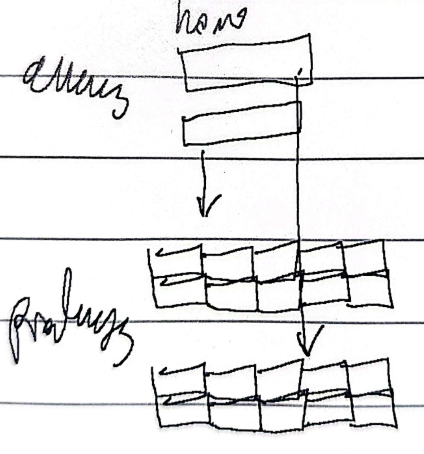
Loss of function
Decrease or loss of functional gene product, can be a null/apomorphic mutation or a leaky/hypomorphic mutation
Null/apomorphic mutation
Loss of function; produces no functional gene product, lethal when homozygous
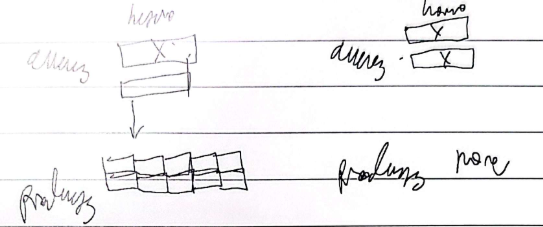

Leaky/hypomorphic mutation
Loss of function; partial loss of function, usually but not always recessive, the same product is produced but there's less of it
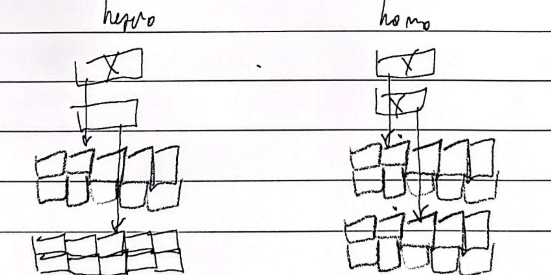
Dominant negative mutations
When homozygous, creates a mutated product that has an abnormal effect with the gene product of another, multiple gene products are impacted and not just their own
Gain of function
Increase or change of gene products
Hypermorphic
Gain of function; more gene products produced
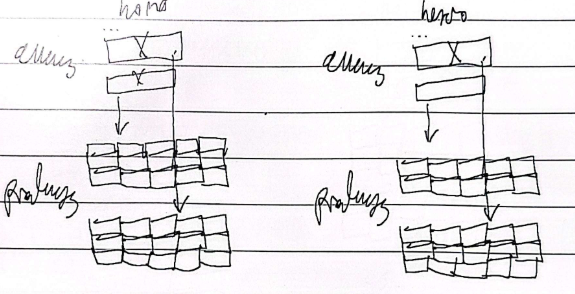
Neomorphic
Gain of function; Mutation causes new functions
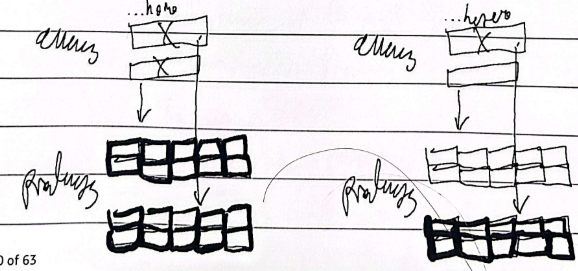
Incomplete/partial dominance
Intermediate phenotype for a heterozygous blend, ex. CrCr is red and CwCw is white, CrCw makes pink, then when CrCw self-fertilizes, 1
Codominance
Both genes are expressed at the same time, ex. white cow and brown cow makes a cow with brown and white markings
Allelic series
Multiallelic genes effect the order of dominance, ex. C is dominant over all alleles, a few alleles are partially dominant then all alleles can be dominant over one allele
Lethal alleles
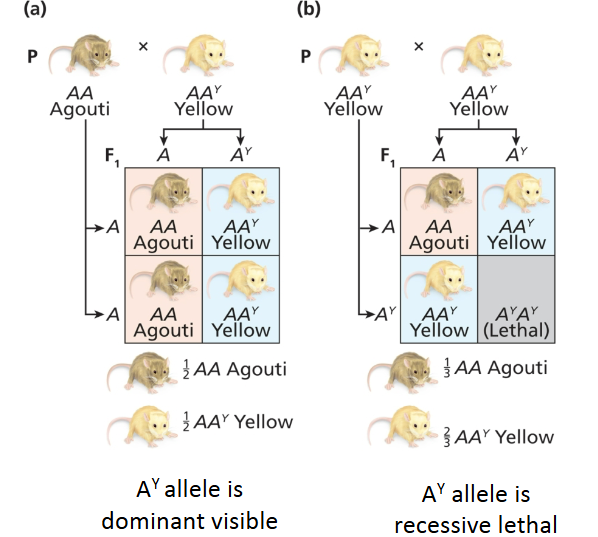
Mostly recessively inherited, only homozygous dies, distortions in segregation ratios, embryonically lethal when recessive, shows later in life when dominant; ex. Ay is a visible dominant mutant allele that gives mice yellow coats instead of brown, Ay is recessive embryonic lethal, AyAy is lethal but AAy is not lethal; recessive lethal and dominant visible
Delayed age of onset
Dominant lethal alleles, ex. Huntington's disease doesn't show until late 30s early 40s, you need only one copy for lethality, you can reproduce before it kills you so it passes on
Sex-limited traits
Sex-specific gene expression, both sexes carry the genes for a trait but only one sex expresses the trait, ex. lactation is female-specific
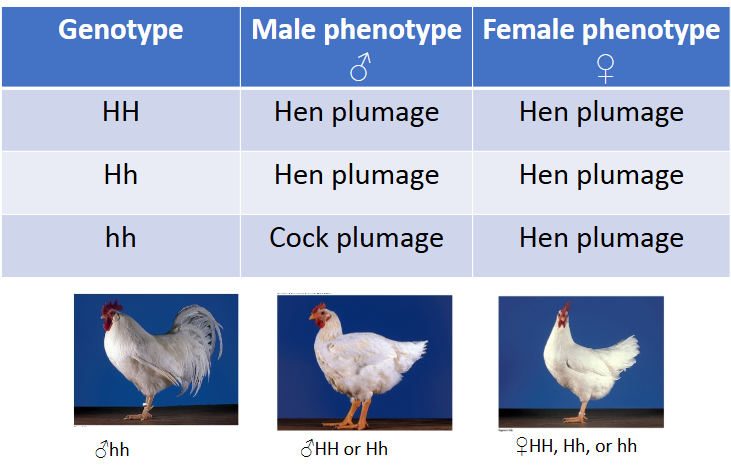
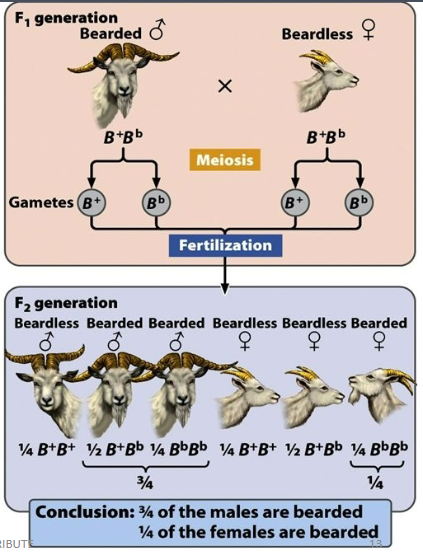
Sex-influenced traits
Sex influences expression of trait, both sexes express the trait to varying degrees, dominant in one sex but recessive in the other sex
Penetrance
When the phenotype is consistent with the genotype, a trait is penetrant, if the genotype is always expressed in phenotype -> fully penetrant, sometimes genotype and phenotype match -> incompletely penetrant; ex. polydactyly, you can have the allele to have extra fingers but you don't have extra fingers
Variable expressivity
Varying degree of severity in phenotypic expression, same genotype with different combination of symptoms
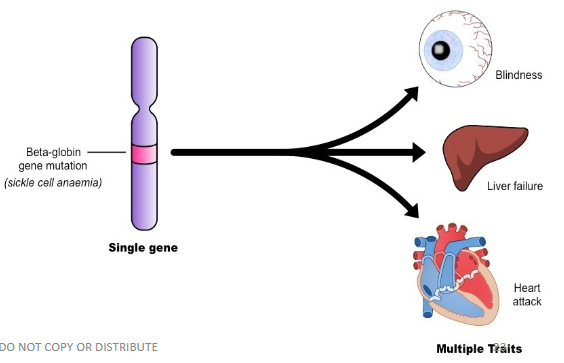
Pleiotropy
One gene affects multiple traits, ex. sickle cell also affects the eyes, liver, and heart
Epistasis
One gene masks or modifies the phenotype of a second gene, dihybrid traits, all phenotypic ratios stem from 9;3;3;1
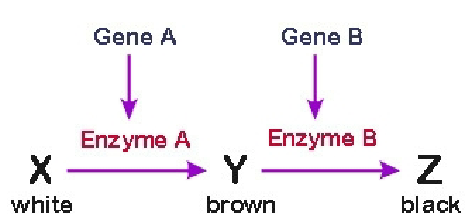
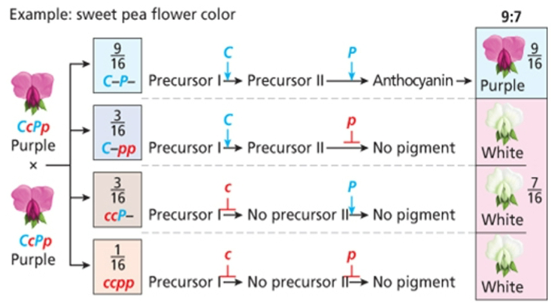
Complementary
9;7, both precursors are needed to get the colored phenotype, two gene interactions needed to get the color
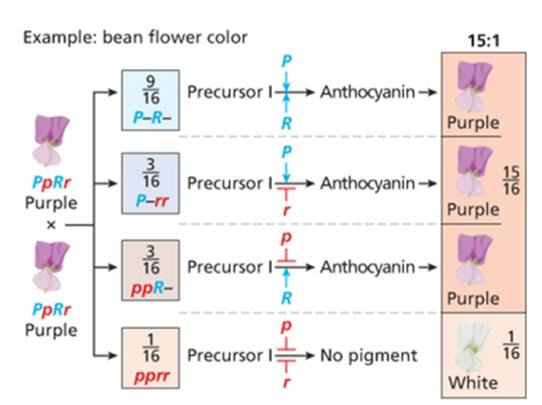
Duplicate gene action
15;1, at least one dominant is needed to get the colored phenotype, double recessive is colorless
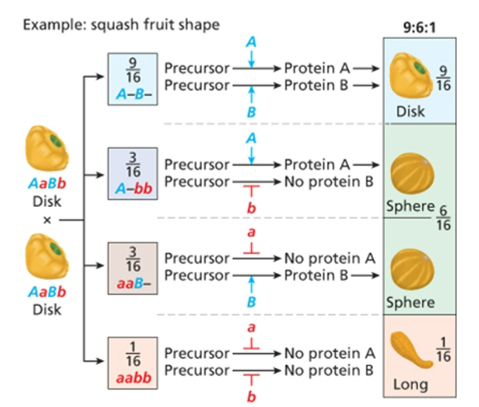
Dominant
9;6;1, two dominant alleles make one phenotype, recessive allele for one gene and dominant allele for the other gene makes one phenotype, double recessive makes a phenotype
Recessive epistasis
9;3;4, homozygous recessive alleles mask the phenotypic expression of alleles at a second locus
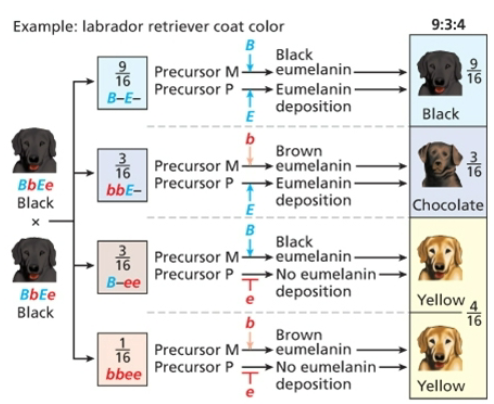
Dominant epistasis
12;3;1, dominant allele will mask phenotypic expression of alleles at a second locus
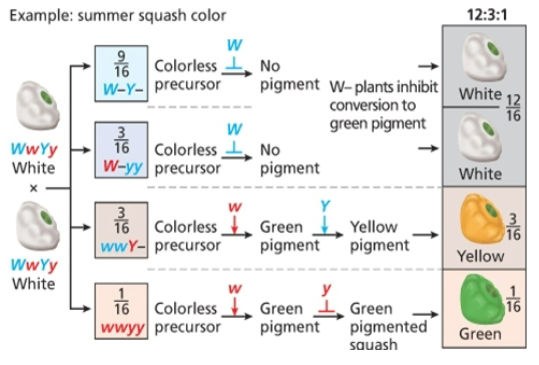
Dominant suppression
13;3, dominant allele prevents expression of the second locus
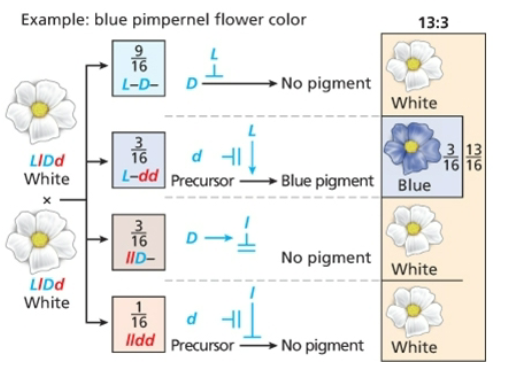
Syntenic genes
Genes on the same chromosome, alleles cannot sort independently and are called linked genes, if the genes are far apart enough independent assortment can happen
Recombinant chromosomes
Homologs cross over and undergo recombination, making them this
Parental chromosomes
Does not undergo recombination, unchanged
Genetic linkage
Frequency of gamete genotype to progeny phenotypes, if linked parental alleles are higher frequency
Complete genetic linkage
No crossover, only parentals
Incomplete genetic linkage
Crossover, parentals are still most common
Recombination frequency
Represented by r, calculated b the number of recombinants over the total number of progeny, higher r means further apart, if r is less than 0.5 then the traits are linked, parental alleles will always be 50% of the progeny at least since crossover only happens between two chromatids not all four
Linkage map
Using r values, sees how far apart genes are from each other, measured in m.u (map units) or cM (centimorgans)
Recombination hotspot
Places where crossover is more common, makes genes seem further apart than they are
Recombination coldspot
Places where recombination rarely happens if ever, usually around the centromere
Haplotype
Two loci close together, two alleles that are often if not always inherited together