Alternative Splicing and Protein Isoforms
1/23
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
24 Terms
Alternative Splicing
form of post-transcriptional, pre-mRNA processing in eukaryotes
a gene can be spliced in different ways to produce variants of the same protein
generates more diversity of proteins
mutations occur at splice donor and acceptor sites, leading to exon skipping/intron retention → alternative splicing
alternative splicing differs between species
Splice Sites
special recognition sequences on the pre-mRNA are located at the intron-exon junctions and within the intron
required for intron removal
recognized by SnRNPs
Splice donor site (5’ end of intron) → usually starts with GU
Splice acceptor site (3’ end of intron) → usually ends with AG
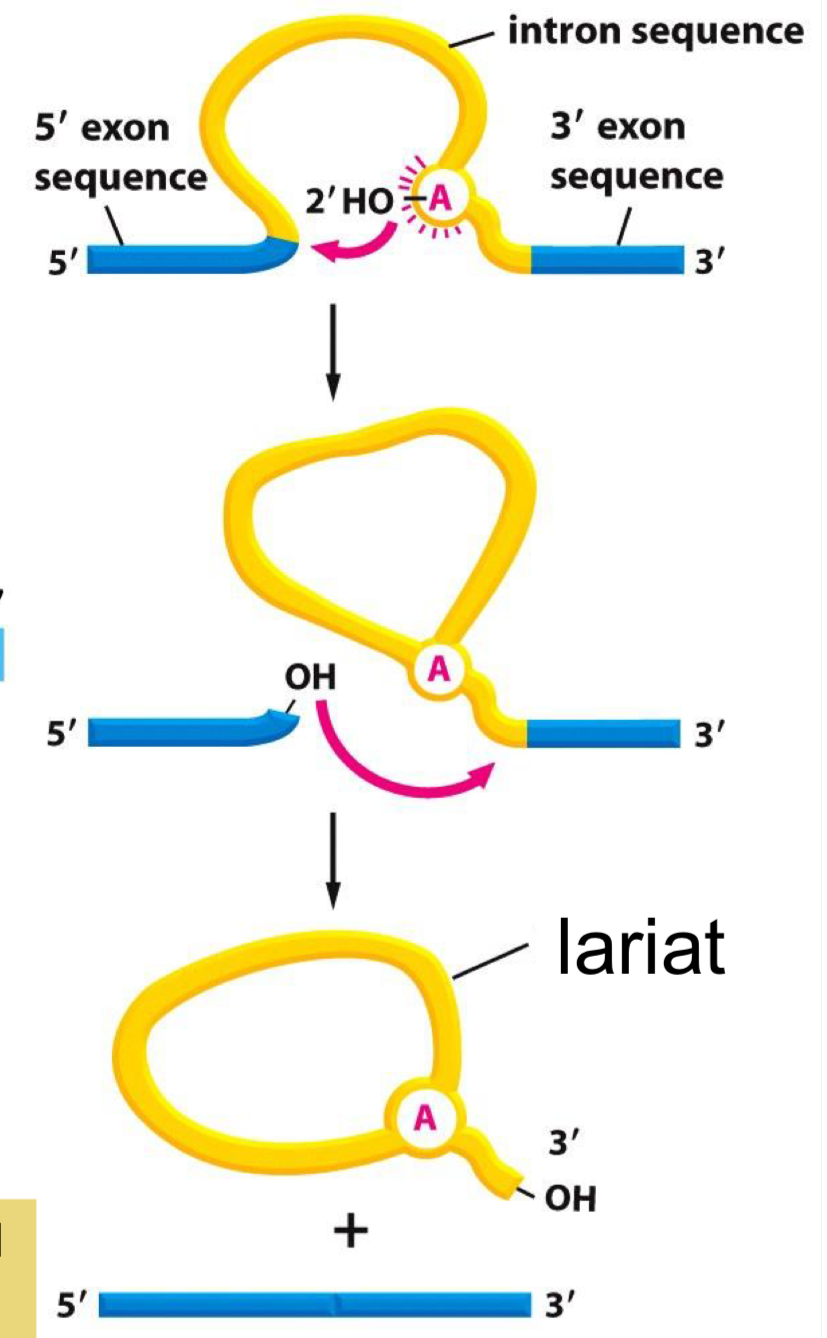
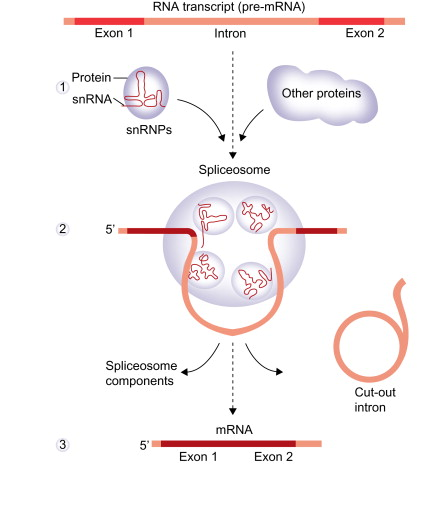
Spliceosome
ribonucleoprotein (RNP) complex found in eukaryotic nuclei that removes introns (non-coding regions) from pre-mRNA and joins exons (coding regions) to form mature mRNA
built from five major small nuclear ribonucleoproteins (snRNPs): U1, U2, U4, U5, and U6
Alternative Splicing Products
alternative splicing can lead to many alternative products and alternative expression patterns
mutually exclusive alternative exons: only include one or the other
alternative 5’/3’ splice site: within an exon, the sequence looks like a splice set and can get recognized as one
alternative promoter and first exon: can have diff “first” exons with a gene that has diff promoters to drive expression
still result in transcription of the same gene, but w/ diff exons
alternative poly A site and terminal exon: can have diff exons; exons have poly A sites
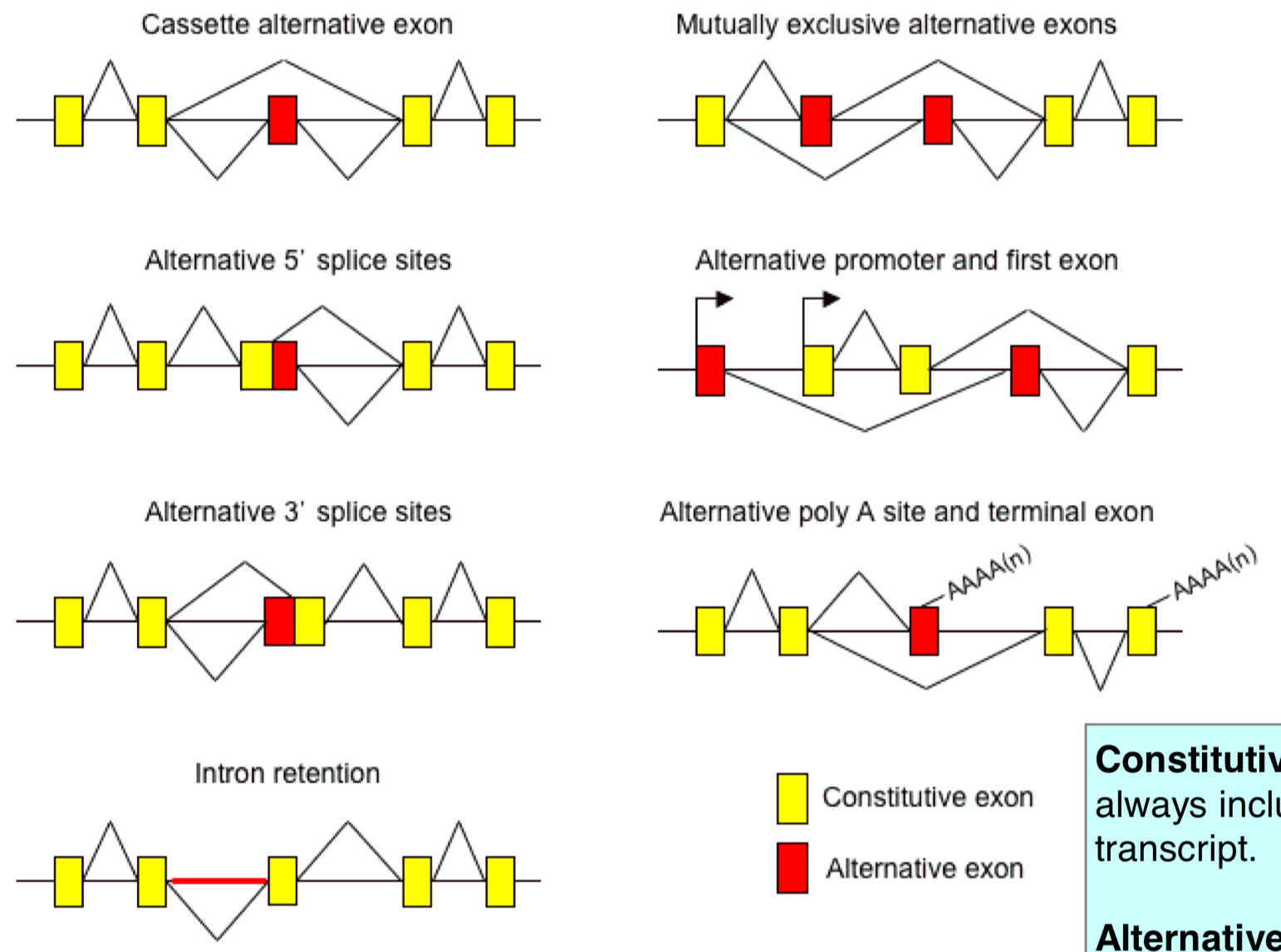
Constitutive Exon
exon is always included in the transcript
Alternative Exon
exon is not always included in the transcript
Weak consensus splice sites
weak junctions: sequences don’t really look like splice sites
might result in the exon getting skipped over along with the introns
Splicing Control Elements
splicing control elements provide another level of regulation
sequences within introns and exons that recruit factors can promote or suppress recruitment of the spliceosome to the splice sites (help or block spliceosome w/ binding or cleaving)
analagous to TFs and enhancers that can promote or suppress recruitment of RNA pol to the promoter
two components that regulate alternative splicing: cis-elements and trans-acting splicing factors
Cis-acting splicing regulatory elements
defining an exon involves the specific stabilization or destabilization of splice site recognition
Enhancers
stabilization: the splice site is recognized more often and the exon is included
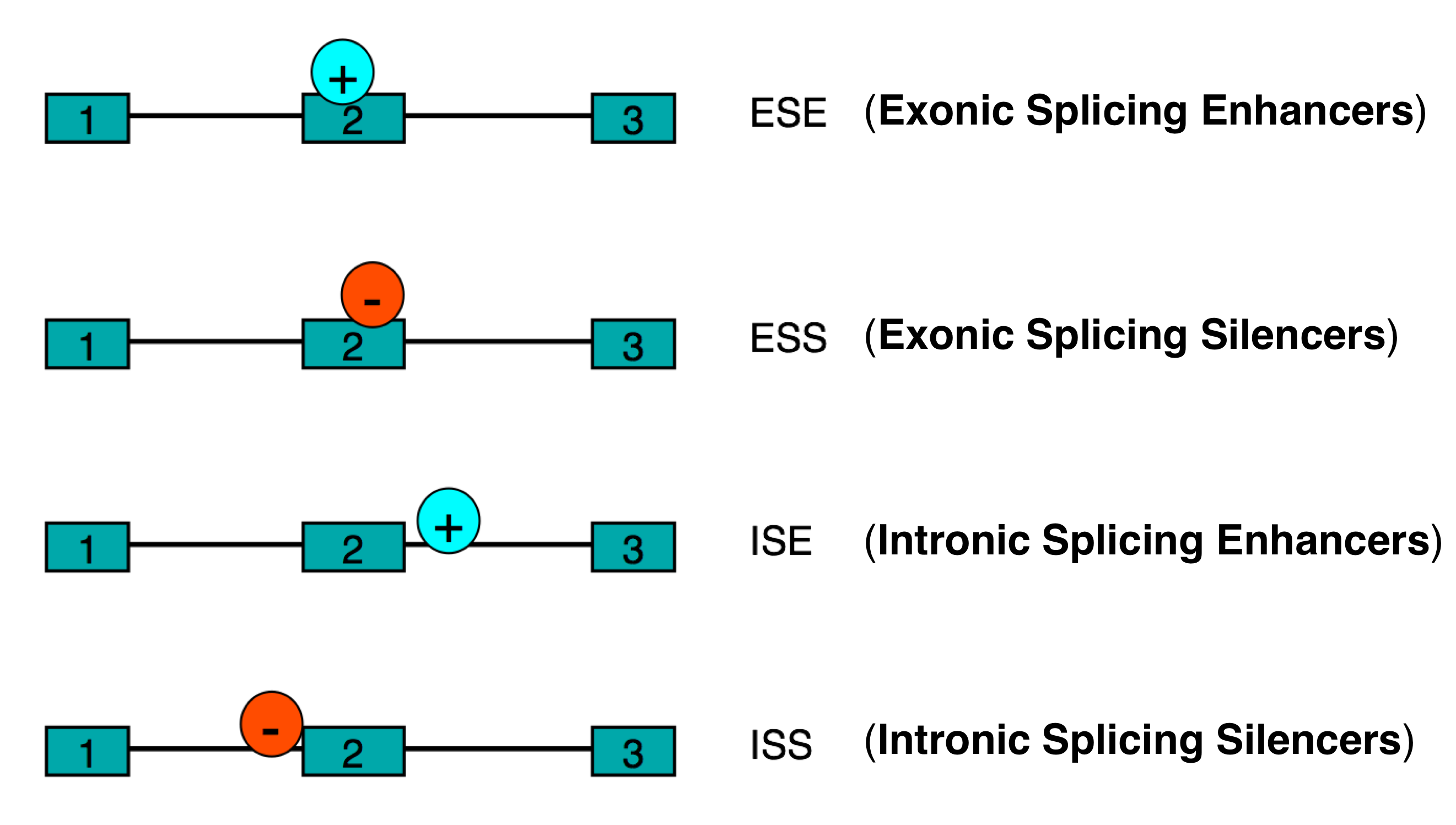
exonic splicing enhancers (ESE), intronic splicing enhancers (ISE)
Silencers
destabilization: the splice site is suppressed and the exon is skipped
exonic splicing silencers (ESS), intronic splicing silencers (ISS)
Classes of splicing regulatory elements figure
Alternative Splicing Regulation in Drosophila Sex: Sex-Lethal
development of sex characteristics depends on the expression (or not) of a master regulator gene called sex-lethal (Sxl)
functional Sxl protein is only produced if 2 X chromosomes are present
XX:AA typically develop female phenotypes (Sxl is produced)
XO:AA or XY:AA typically develop male phenotypes
Alternative Splicing Regulation in Drosophila Sex: mRNA splicing cascade
in Drosophila, female and male biological phenotypes determined first by X dosage and then an mRNA splicing cascade
functional Sxl protein initially expressed by an alternate promoter in XX embryos
slightly later, Sxl transcribed in all embryos (XX, XO, XY) from a maintenance promoter
However, splicing outcome differs depending on whether Sxl protein is already present
females: Functional Sxl protein is present → Sxl protein binds pre-mRNA and blocks inclusion of exon 3 → functional Sxl protein continues to be produced (positive feedback)
males: No functional Sxl protein initially → exon 3 contains a premature stop codon → produces truncated, nonfunctional Sxl protein
therefore Sxl must be a splice factor that recognizes an intronic splice silencer
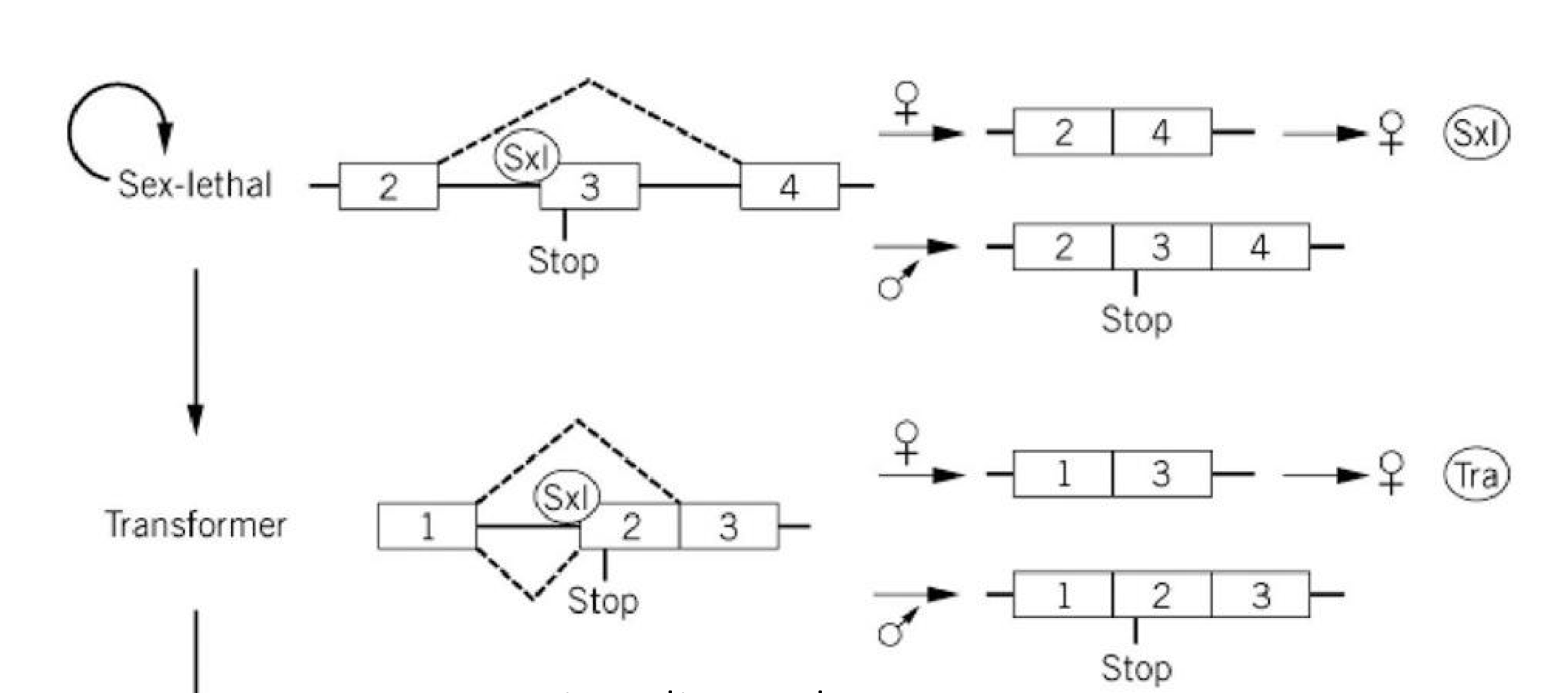
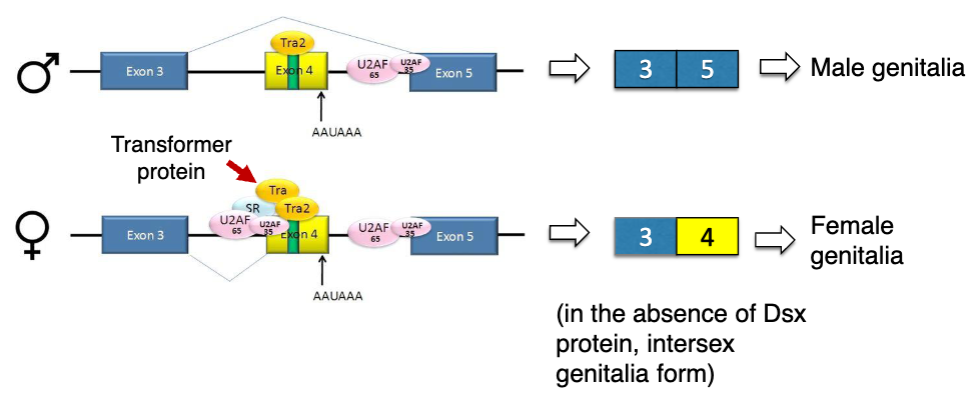
Alternative Splicing Regulation in Drosophila Sex: Transformer
functional Sxl protein only produced in XX (female) Drosophila
functional Sxl regulates its own splicing as well as splicing of Transformer (Tra) transcripts
functional Tra regulates splicing of Double-sex transcripts
if Sxl and Tra are present: version of Dxs that regulates development of female genitalia
if Sxl and Tra are absent, version of Dsx that regulates development of male genitalia
Tra must be a splice factor that recognizes exonic splice enhancers (binds to exon 4 and promotes recognition of splice site
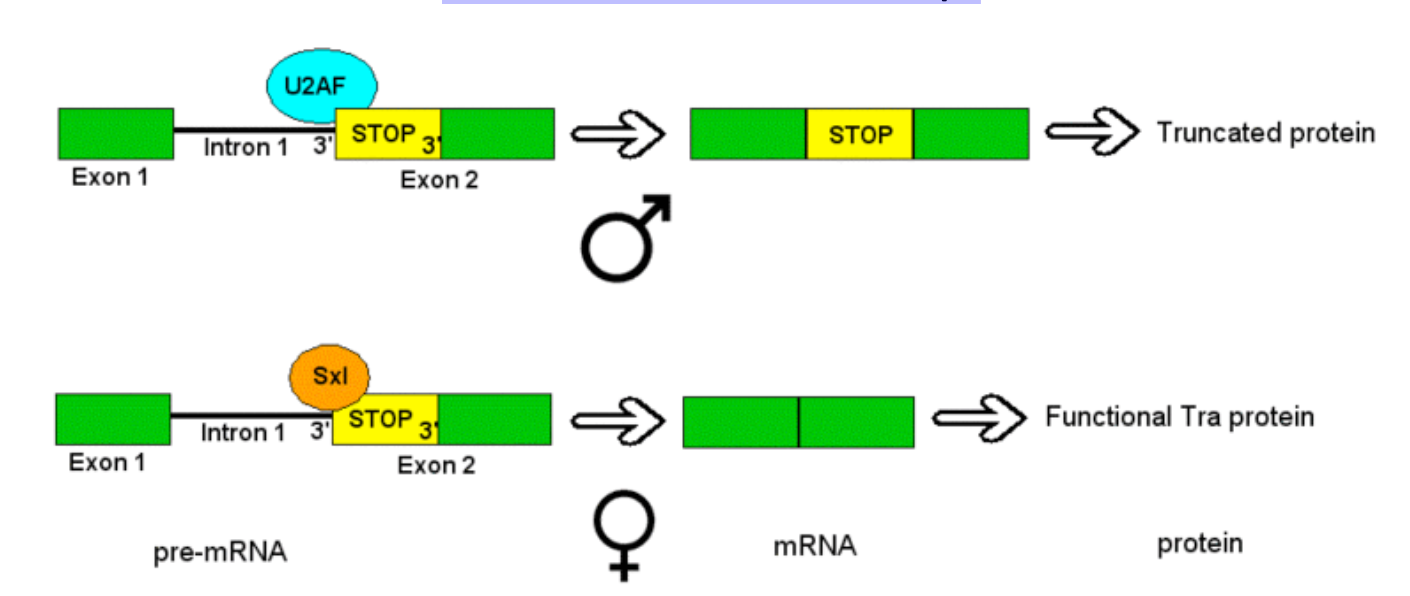
ISS in Transformer Transcript
males: Sxl is absent; splicing factors recognize the splice acceptor site within intron 1, so exon 2 is included in the mRNA
exon 2 contains a premature stop codon, resulting in non functional Tra protein
females: Sxl recognizes and binds to the ISS within intron 1, causing exon 2 and its premature stop codon to be spliced out of the mRNA
result: transcript for a functional Tra protein
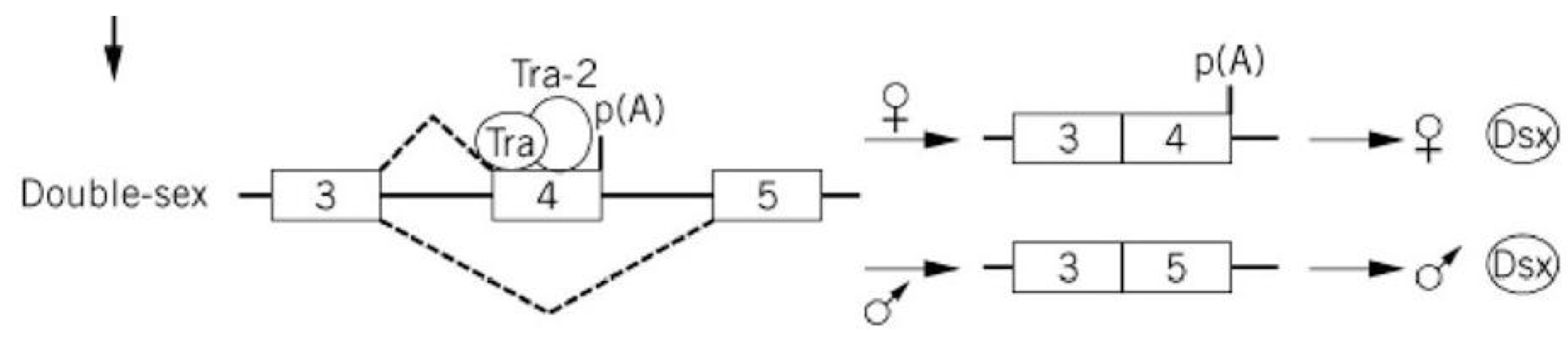
ESE in Double-Sex Transcript
the intron between exon 3 and 4 has only a weak consensus for binding the spliceosome complex, so exon 4 is skipped in males
females: binding of Tra enhances spliceosome binding and promotes splicing between exon 3 and 4
exon 4 has its own polyadenylation signal, so exon 5 is excluded
How can you detect alternative splicing regulatory sequences and events? Large-scale genome-wide approaches
RNA-seq (next gen-sequencing) to identify transcript variants and their relative amounts
comparison of cDNA sequences to reference genome to ID exon-exon junctions (and introns)
ie. comparison with an existing exon-exon junction library
DNA microarrays using exons instead of entire gene sequences
cross-linking and immunoprecipitation: crosslink splicing factors and RNA transcripts, then use antibodies to immunoprecipitate the complexed RNA for sequencing
How can you detect alternative splicing regulatory sequences and events? Smaller-scale approaches
RT-PCR and gel electrophoresis to detect alternate splice variants thru differences in size
Northern blots also possible
EMSA: establish binding of spliceosome factors to RNA transcripts
use of splicing reporter genes
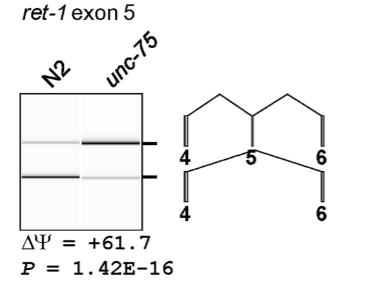
Example of RT-PCR to detect presence or absence of splicing events: Ret-1
C. elegans RNA binding protein UNC-75 is responsible for splicing of neuronal transcripts
N2: Wt worms
unc-75: worms w/ lof mutation
ret-1 is a transcript found in neurons. PCR primers span the indicated exons and introns
from the data, we can say that UNC-75 protein is binding to an exonic/intronic splicing silencer
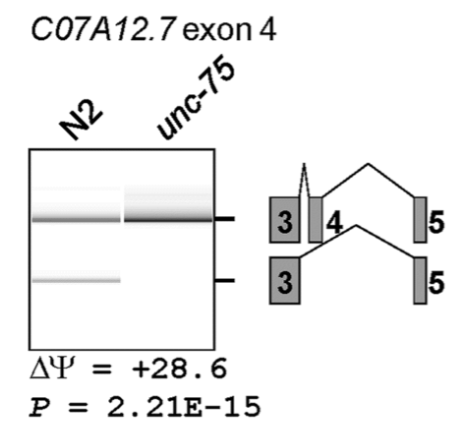
Example of RT-PCR to detect presence or absence of splicing events: C07A12.7 exon 4
C. elegans RNA binding protein UNC-75 is responsible for splicing of neuronal transcripts
N2: Wt worms
unc-75: worms w/ lof mutation
C07A12.7 is a transcript found in neurons. PCR primers span the indicated exons and introns
from the data, we can say the UNC-75 protein binds to an exonic/intronic splicing silencer
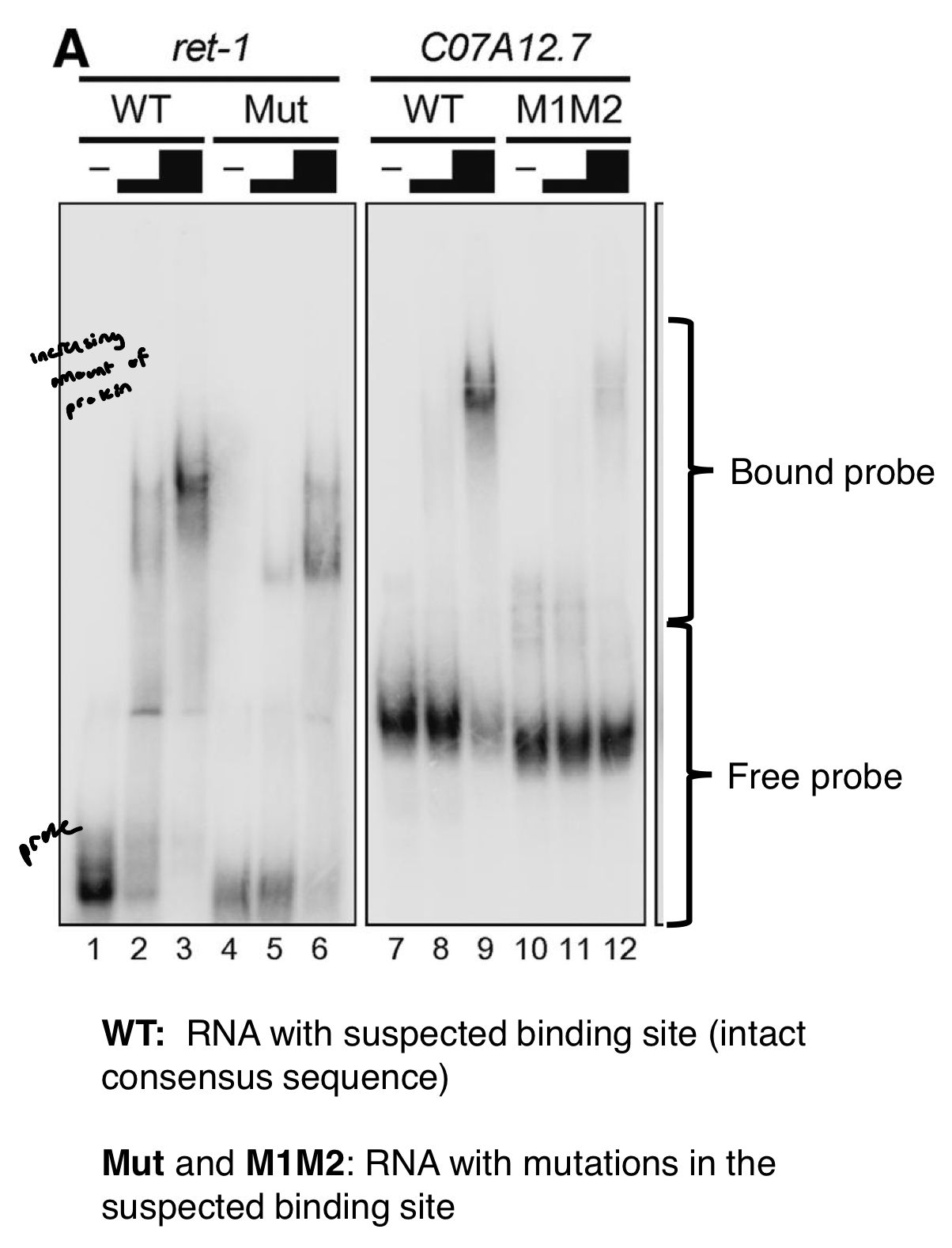
Strategies for pinpointing the regulatory elements bound by UNC-75 protein
comparison of orthologous exons in other species of Caenorhabditis to find common sequence motifs. Consensus sequence: (G/U)UGUUGUG
gel shift assay (EMSA) to esablish if UNC-75 can bind to this motif within these transcripts. Increasing the amounts of UNC-75 are added to the RNA
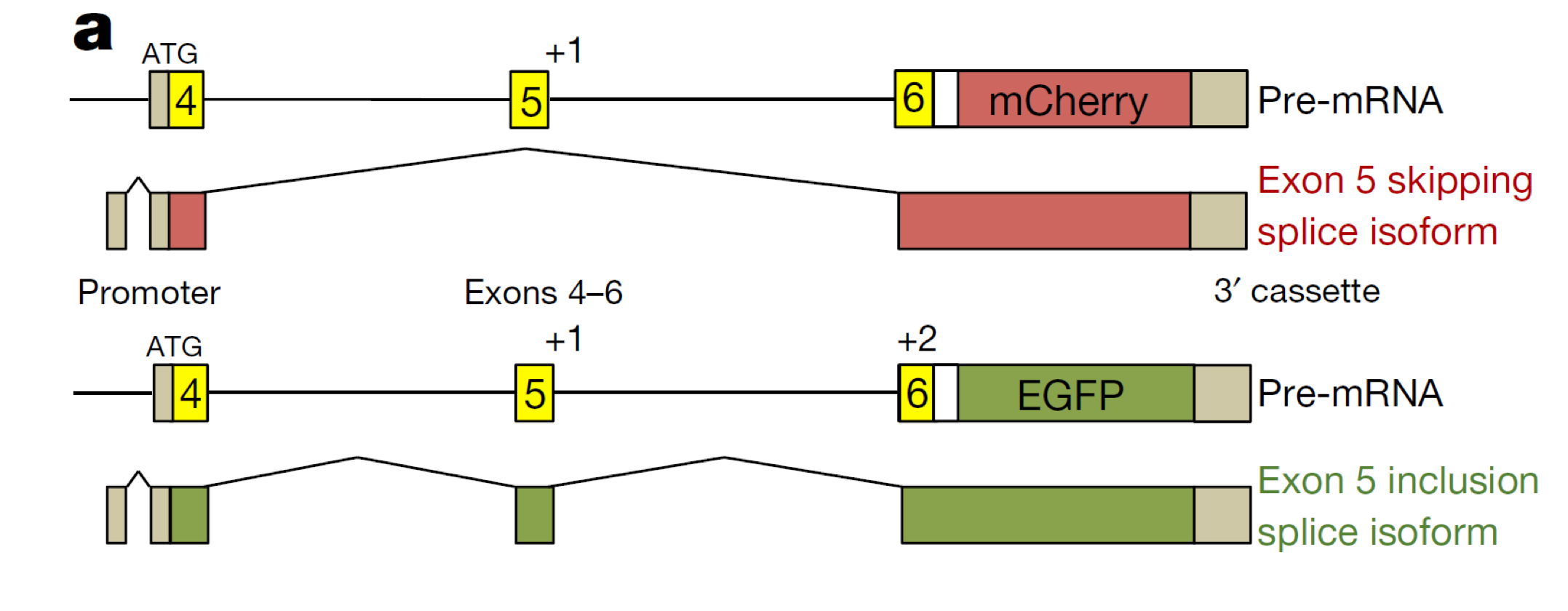
Bi-chromatic (two color) fluorescent splicing reporters
tissue and temporal specificity of alternate isoforms can be investigated by generating reporter constructs expressing distinct fluorescent markers dependent on the expressed isoform
this allows for studying alternate splicing in living cells
the color you get in the final protein depends on what exons are kept
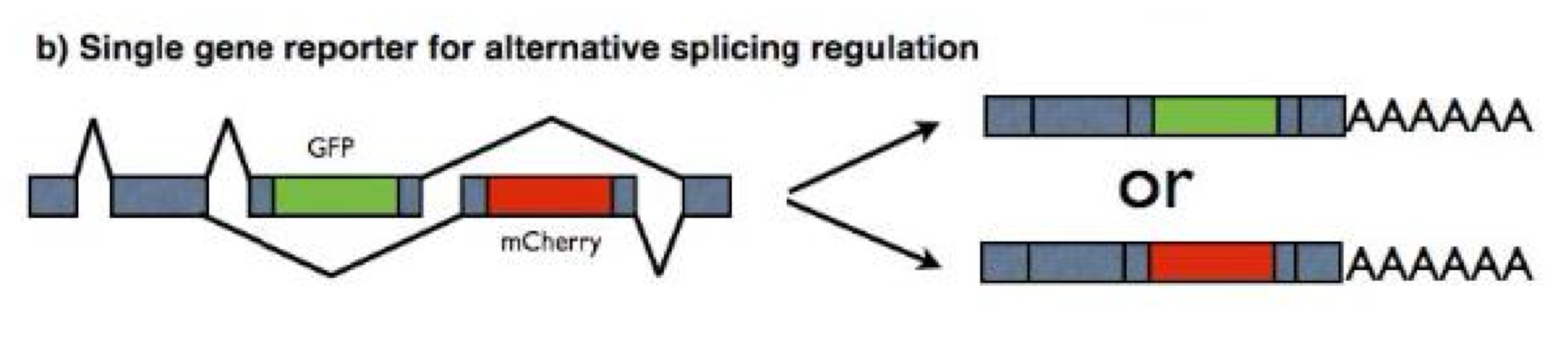
Bi-chromatic (two color) fluorescent splicing reporters: Example
The ret-5 exon splicing reporter construct
reporter construct contains reporter sequences for mCherry (red) and EGFP (green) in different frames
neither reporter is expressed in the absence of splicing
if exon 5 is skipped, mCherry sequences are in frame and cells turn red
if exon 5 is retained, EGFP sequences are in frame and cells turn green