moduel 4: Proteins
1/17
Earn XP
Description and Tags
amino acid = AA
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
18 Terms
What does is structure influenced by and what does it determine?
thermodynamics and chemical interactions drive folding
folding determines function
levels of protein structure
primary structure
secondary structure
tertiary structure
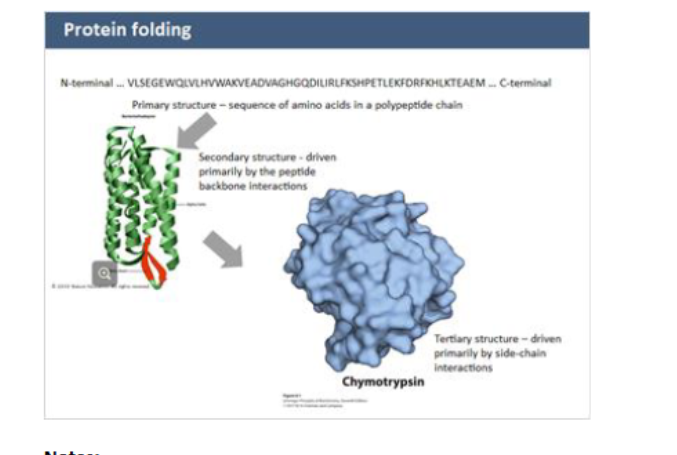
Primary structures
linear AA sequence
dictated by DNA
contains “instructions” for folding
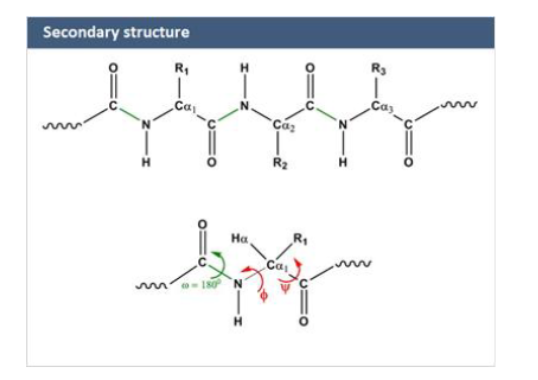
secondary structures
folding of backbone only (not side chains)
governed by:
peptide bond rigidity
H bonding
structural constraints
peptide bond = partial double bond making structure rigid (planar)
only rotation allowed around
phi: N- C(alpha)
psi: C(alpha)-C
angles determine allowed conformations (Ramachandran plot)
Types:
Beta sheets
Alpha Helix
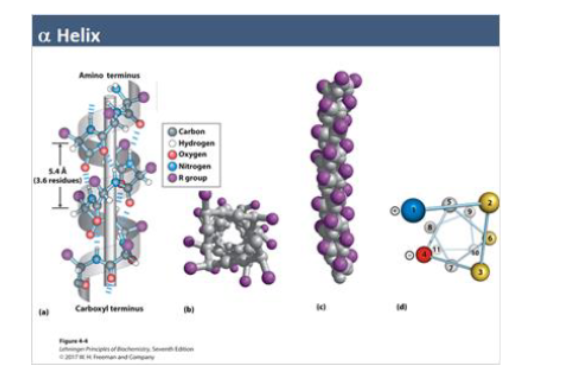
Alpha Helix
helical coil (3.6 residues/turn)
stabilized by H bonds (i → i + 4)
key features:
R-groups pt outward
has a dipole:
N-terminus → +
C-terminus → -
AA effects:
alanine → stabilizes helix
proline → breaks helix (rigid kink)
glycine → too flexible
structure depends on BOTH sequence + residue positioning
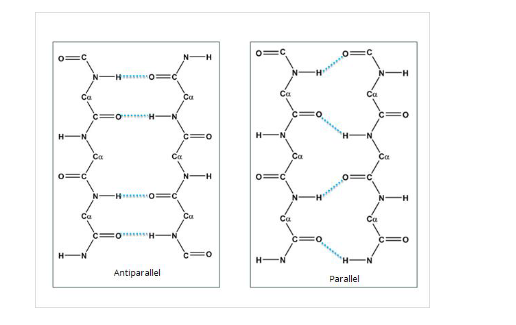
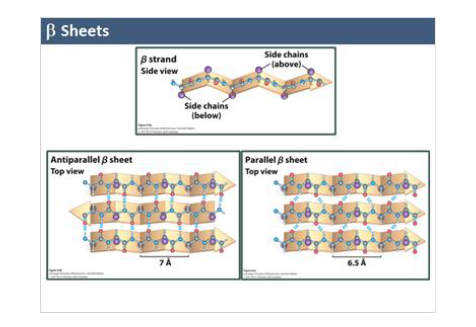
beta sheets
extended zig-zag strands
stabilized by inter-strand H-bonds
still backbone drive (like a-helix), but diff geometrey
types:
antiparallel (more stable)
parallel
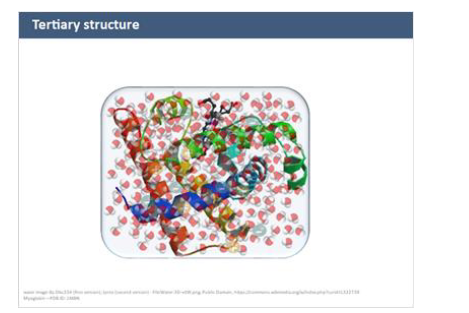
tertiary structures
full 3D folding of entire pp
driven mainly by side chain interactions

Hydrophobic effects and protein folding
folding driven by entropy of water, not protein
water forms cages around phobic residues → entropy decreases
phobic residues limit number of H-bonds free H2O moelcules can make → thermodynamically unfavourable
causes water to surround residues to conserve H - bonds and reduce entropy
folding releases water → increases entropy (favourable)
phobic residues → buried inside protein
phillic residues → exposed to water
how do thermodynamics influence protein folding
folding dec. protein entropy, and increases water entropy (net favourable)
forces involved in folding
folding balances all forces
forces include:
H bonds
electrostatic interactions
van der waals
phobic interactions
thermodynamic forces
cooperativity
why is folding a cooperative process?
it is all or nothing (folding produces very little intermediates)
not independant = formation of one interaction → stabilizes structure → makes next easier
ex. like zipper
few first teeth are hard to close
once started, rest zip up quixkly
ensures fast, accurate and complete folding
usually during folding of secondary structures
cooperative folding mechanism
1. Initial interactions (rate-limiting)
First hydrophobic contacts or H-bonds form
This reduces conformational freedom
2. Reduction in conformational space
Protein has fewer possible shapes
Easier to find correct interactions
3. Cascade effect
More interactions form rapidly:
Hydrophobic clustering
Hydrogen bonds (α-helix, β-sheet)
Electrostatic stabilization
4. Folding nucleus
Small stable core forms early
Acts as a template/scaffold for rest of folding
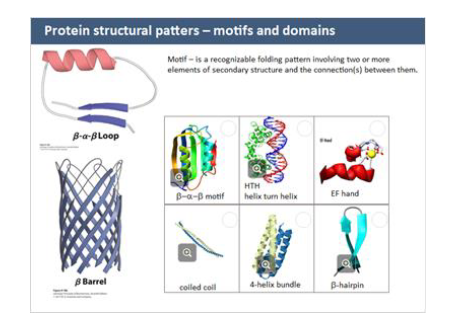
motifs
small, recurring structure (ex. β-α-β loop)
helps predict function
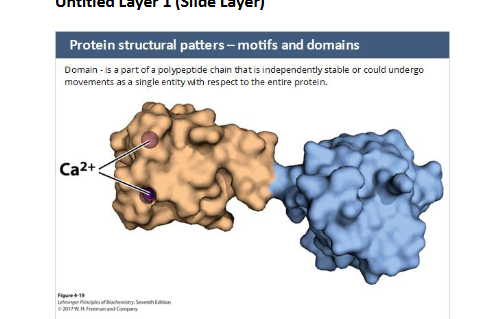
domain
independently stable unit
can function on its own
proteins = modular
protein types
fibrous
globular
membrane
fibrous proteins
structural roles in cells and tissues
structure: elongated/filamentous in shape
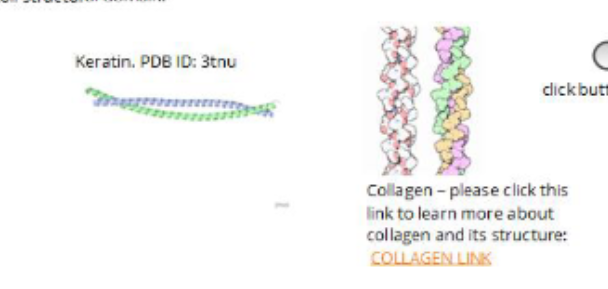
some fibrous proteins are permanent and build to last and not be degraded/modified/regulated (ex. collagen and keratin)
some are regulated (ex. actin and tubulin)
actin and tubulin form regulated fibrous but are not fibrous themselves
EXAMPLES:
colalgen and keratin which both share coiled-coiled structural domain
globular proteins
carry out chxm work in cell (ex. synthesis, transport and metabolism)
structure: compact structures
tertiary structure of flobular portein determines function
phobic AA’s are at core of protein, phillic/polar AA’s on surface
ex. myoglobin and hemoglobin
membrane proteins
has high proportion of phobic AA
allows protein to interact with phobic acyl chains of the lipid bilayer
they do this by resting on top of lipid bilaer (partially buried in lipid membrane - these are called peripheral membrane [rpteons)
could also be integrated in lipid bilayer (integral membrane proteins)
examples: ATP synthase and insulin receptor