bio chem quiz 7
1/52
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
53 Terms
what is heparin and heparan sulfate?
heparin is a linear polymer
heparan sulfate is heparin-like polysaccharide but attached to proteins
highest negative charge density biomolecules
prevent blood clotting by activating protease inhibitor antithrombin
binding to various cells regulates development and formation of blood vessels
can also bind to viruses and bacteria and decrease their virulence
what is the molecular basis of heparin enhancement of the binding of thrombin to antithrombin?
heparin is negatively charged and acts as a tether to bring proteins together and inhibit thrombin
heparin connects thrombin to its inhibitor, antithrombin, then prevents blood clots from forming
heparin is an example of a GAGs: a polysaccharide that is negatively charged and has 2 repeating disaccharides (2 repeating sugars)
what is a glycoprotein as a glycoconjugate?
a glycoprotein is a carb attached to a protein
a protein with small oligosaccharides (3-10 carbs long) attached
carbohydrates attached via its anomeric carbon to amino acids on the protein
common connections occur at Ser, Thr, and Asn
about half of mammalian proteins are glycoproteins
only some bacteria glycosylate a few of their proteins
carbohydrates play a role in protein-protein recognition
viral proteins are heavily glycosylated; this helps to evade the immune system (tricks immune system into thinking its our protein)
what are the two types of glycosidic bonds?
1) O-linked : attaching a carb to oxygen; glycosidic bond to the hydroxyl group
2) N-linked : attaching a carb to nitrogen; N-glycosyl bond
what do glucotransferases do?
glucotransferase are enzymes that recognize amino acids and attach carbohydrates
what are glycolipids as a glycoconjugate?
glycolipids are lipids with covalently bound oligosaccharide
they are parts of plant and animal cell membranes
in vertebrates, ganglioside carbohydrate composition determines blood group
in gram negative bacteria, lipopolysaccharides cover the peptidoglycan layer
often used for cell-cell recognition by changing oligosaccharides attached to the membrane
what are proteoglycans as glyconjugates? and what are the 2 main types?
proteoglycans are GAGs attached to proteins (that are usually attached to cells or extraceullar matrix)
sulfated glucoseaminoglycans attached to a large rod-shaped protein in cell membrane
syndecans: protein has a single transmembrane protein
glypicans: protein is anchored to a lipid membrane
interact with a variety of receptors from neighboring cells and regulate cell growth
what is the interaction of cells with the ECM?
interaction of the cells with ECM:
some internal membrane proteins are proteoglycans
syndecans
other integral membrane proteins are receptors for extracellular proteoglycans
integrins
integrins are a class of receptors that feel the matrix and cells around them (feeling micro-enviornment)
integrins interact with syndecans
these proteins link cellular cytoskeleton to the ECM and transmit signals into the cell to regulate: cell growth, cell mobility, apoptosis, and wound healing
what is the cell signaling mechanism of integrans?
→ integrins interact with the extracellular matrix primarly through the syndecans/proteoglycans, which information can get relayed internally to rearrange intracellular cytoskeleton (mechanism of how cells feel their microenvironment through integrins)
→ integrins have a specific binding site to allow the proteoglycans to attach to them. the integrin then changes its conformation and becomes more linear. structural change gets induced into the intracellular cytoskeleton, which will rearrange or induce a signal
what is lectin?
lectin are proteins that specifically bind carbohydrates to help with cell recognition
what are the two methods of analysis for glycoconjugates?
1) global profiling: all types of oligosaccharides on a kidney cell
2) targeted structure analysis: oligosaccharides in a specific protein/lipid
how to use MS for analysis of glycoconjugates / how to interpret it?
positioning of the m/z peaks = the type of oligosaccharide
height of the peak = abundance / how much
what are the basics of nucleotides and nucleosides?
nucleotide =
nitrogeneous base (pyrimidine: smaller ring or purine: larger ring)
pentose sugar
phosphate
nucleoside =
removal of phosphate group
nitrogeneous base
pentose sugar
carbons AND nitrogen atoms on the nitrogeneous base are numbered in cyclic format
carbons of the pentose are designated N’ to alleviate confusion
nitrogen 1 on pyrimidine and nitrogen 9 on purine are used to connect to a sugar
what are the functions of nucleotides and nucleic acids?
nucleic acids are polymers of nucleotides used for (polymeric form):
storage of genetic info (DNA)
transmission of genetic info (mRNA)
processing of genetic info (ribozymes)
protein synthesis (tRNA and rRNA)
nucleotides are also used in the monomer form for cellular functions (monomeric form):
energy for metabolism (ATP)
enzyme cofactors (NAD+) (Nicotinamide adenine dinucleotides: used to store electrons)
signal transduction (cAMP)
how do pentose forms differ in some nucleic acids and nucleotides?
how pentose forms differ:
beta-d-riboduranose in RNA (has a 2’ OH group)
beta-2’-deoxy-d-ribofuranose in DNA (does not have a 2’OH group, has a hydrogen on 2’ position)
different puckered coformations of the sugar ring are possible
how to tell endo and exo pucker conformations? which do RNA and DNA typically have?
endo conformation = if carbon 2 or 3 is on the same side as the 5’ carbon
exo conformation = if carbon 2 or 3 is on the opposite side (pointing away) as the 5’ carbon
DNA typically takes on a 2’ endo conformation and RNA typically takes on a 3’ endo conformation
what are nitrogenous bases?
nitrogenous bases are derivatives of pyrimidine (smaller ring) or purine (larger ring)
nitrogen containing heteroaromatic molecules
planar or almost planar structures (sp2 hybridization makes the ring flat)
absorb UV light around 250-270 nm (because of conjugated ring system; if you hit with light ~260 nm the amount of light absorbed can be used to determine the concentration)
cytosine, adenine, and guanine are found both in DNA and RNA
thymine is only found in DNA
uracil is only found in RNA
all are good H-bond donors and acceptors (key structural feature to form different H-bonds)
neutral molecules at pH 7
how many hydrogen bonds can the following form based on structure: cytosine, thymine, and uracil?
cytosine: 3 H bonds
thymine: 2 H bonds
uracil: 2 H bonds
what is the nucleoside naming for the following deoxyribonucleotide structures
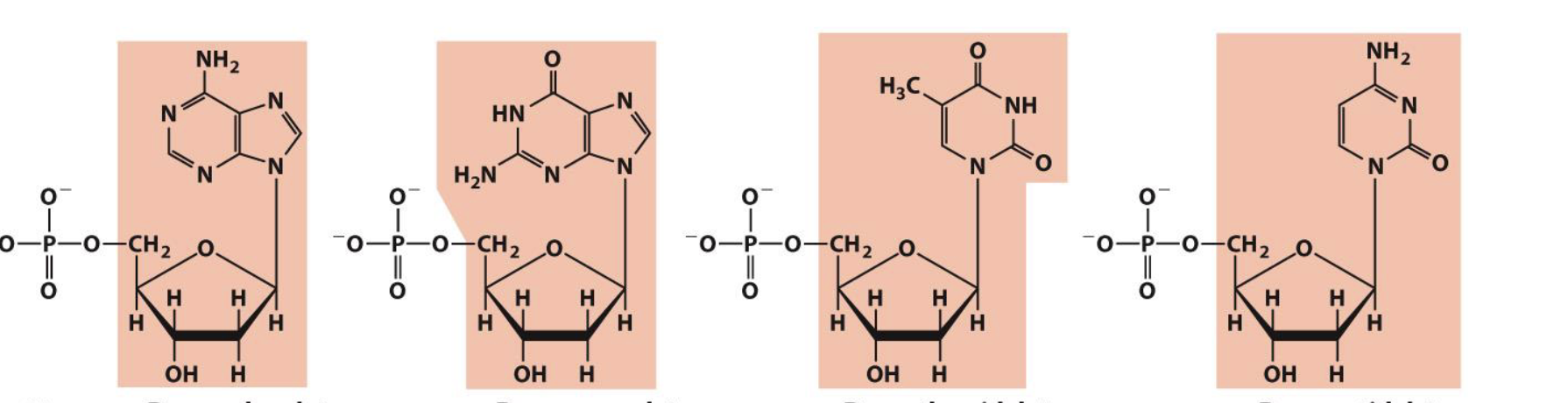
deoxyadenosine
deoxyguanosine
deoxythymidine
deoxycytidine
what is the nucleoside naming for the following ribonucleotide structures
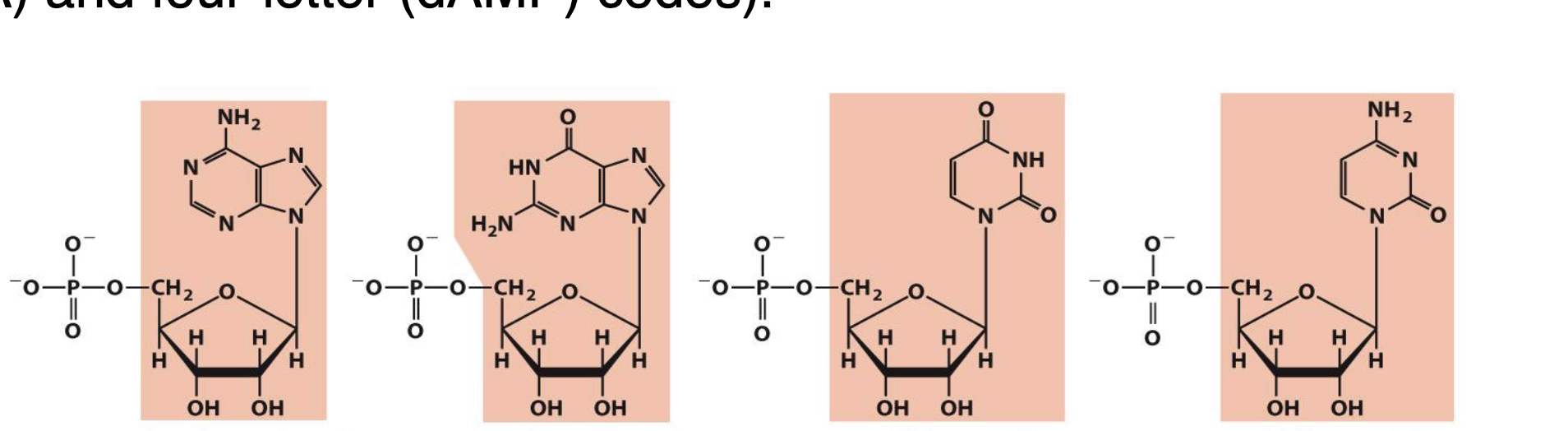
adenosine
guanosine
uridine
cytidine
what is the beta-N-glycosidic bond?
in nucleotides, the pentose ring is attached to the nitrogenous base via a N-glycosidic bond (connects a ribose sugar to nitrogenous base)
the bond is formed to the anomeric carbon of the sugar in beta configuration
the bond is very stable and helps to give DNA its high stability
the bond is formed:
to position N1 in pyrimidines
to position N9 in purines
this bond is quite stable toward hydrolysis, especially in pyrimidines
bond cleavage is catalyzed by acid
in monomeric form, the bond can freely rotate. when connected in strands, it can take on particular angles
syn = on top of each other
anti = opposite
what is the tautomerism of nitrogenous bases?
prototropic tautomers are structural isomers that differ in the locations of protons
keto-enol tautomerism is common in ketones
ketone form is more stable at pH 7 (favors ketone form), by changing the pH you can shift to favor the enol form
Lactam-lactim tautomerism occurs in some heterocycles
both tautomers exist in solution, but the lactam (ketone) forms are predominant at neutral pH
what happens as you change the pH from lactam to lactim?
at lactam, the pH is neutral. as the pH decreases: lactam → lactim → double lactim
changing lactam-lactim form disrupts the hydrogen bonding when holding the nucleotide strands together → this causes weakening of the structure for RNA and DNA
what are minor nucleosides in DNA?
minor nucleosides:
modification is done after DNA synthesis
5-methylcytosine is common in eukaryotes and is also found in bacteria (methylation of DNA creates gene suppression)
N6-methyladenosine is common in bacteria but not found in eukaryotes
epigenetic marker:
way to mark own DNA so that cells can degrade foreign DNA (prokaryotes)
way to mark which genes should be active (eukaryotes)
could the environment turn genes on and off in an inheritable manner? (epigenetic inheritance: modify DNA to determine the degree of expression of genes = epigenetic markers determine by the environment; epigenetic markers = modified nucleosides)
what are polynucleotides?
polynucleotides:
covalent bonds are formed via phosphodiester linkages (to connect nucleotides together)
negatively charged backbone
DNA backbone is fairly stable (because the linkage is very stable)
hydrolysis accelerated by enzymes (DNases: recognize sequence to cut phosphodiester linkage)
RNA backbone is unstable
in water, RNA lasts for a few years
in cells, mRNA is degraded in a few hours (vaccine); has a high turnover in the cell
linear polymers
no branching or cross links
directionality
the 5’ end is different from the 3’ end
we read the sequence from 5’ → 3’
enzymes read / sequence based on this directionality
what is the hydrolysis of RNA?
hydrolysis of RNA:
RNA is unstable under alkaline (high pH) conditions
hydrolysis is also catalyzed by enzymes (RNases)
RNase enzymes are abundant around us
S-RNase in plants prevent inbreeding by preventing pollen tube formation
RNase P is a ribozyme (enzyme made of RNA) that process tRNA precursors
Dicer is an enzyme that cleaves double stranded RNA into oligonucleotides
can be used to target specific sequences (RNA interference tech)
protection from viral genomes
in terms of the mechanism of base catalyzed RNA hydrolysis, what is the role of OH-
the 2’ OH- allows for cleavage of the backbone to create 2 separate nucleotide groups
what are the hydrogen bonding interactions observed?
two bases can hydrogen bond to form a base pair
for monomers, a large number of base pairs is possible
in polynucleotide, only a few possibilites exist (only specific base pairing to give complementary hydrogen bonding)
watson-crick base pairs predominate in double stranded DNA
A pairs with T : 2 H bonds
C pairs with G : 3 H bonds
purine pairs with pyrimidine (Chargaff’s rule: 1:1 ratio of purine to pyrimidine)
basics of AT and GC base pairing?
→ the base pairing of A and T occurs by 2 hydrogen bonds
→ the base pairing of G and C occurs by 3 hydrogen bonds
→ if you increase the number of hydrogen bonds = base pairs get closer together
bacteria growing under hot conditions = more GC base pairing to hold the strands together. colder = more AT base pairing to allow DNA to open up for replication
what is the watson-crick model of B-DNA
B DNA is the most common form
uneven in grooves = major and minor grooves
it is 3.4A as the distance from one base to another on the same strand
base stacking increases the van der waals forces, increasing the stability of the DNA strand
more base stacking = less light absorbed
what are the 2 other forms of DNA besides B DNA?
A form: forms by dehydration of B form, wider diameter and shorter length; shorter, wider, found in dehydrated conditions
Z form: stretching and now a left handed helix; long, skinny, left handed, bases are exposed, found in sites where DNA is being copied
what are some ways to see structure differences in the B, A, and Z form of DNA?
helical sense diameter, base pairs per helical turn, helix rise per base pair, base tilt normal to the helix axis, sugar pucker conformation, and glycosyl bond conformation
what is the replication of genetic code?
for the replication of genetic code:
strand separation occurs first
each strand serves as a template for the synthesis of a new strand
synthesis is catalyzed by enzymes known as DNA polymerases
a newly made DNA molecule has one daughter strand and one parent strand
what is messenger RNA?
messenger RNA is the code carrier for the sequence of proteins
is synthesized using DNA template and generally occurs as a single strand
contains ribose instead of deoxyribose
contains uracil instead of thymine
one mRNA may code for more than one protein
two assumptions: (1) a single RNA strand will make multiple proteins and (2) selection of different exons to create different versions of the protein (alternative splicing)
together with transfer RNA (tRNA), transfers genetic information from DNA → proteins
what is the difference between monocistronic and polycistronic gene strands?
monocistronic: RNA codes for a protein for an individual gene; eukaryotic mRNAs are monocistronic; allows for fine tune regulation of proteins (higher regulation) and tissue differentiation to give them their own phenotypes
polycistronic: single mRNA has multiple genes that can be translated; prokaryotic individuals like bacteria and archaea have this type
what are palindromes?
palindromic sequences can form hairpins and cruciforms (secondary structures for DNA and RNA)
palindromes: words or phrases that are the same when read backward or forward
ex: civic, racecar
what is the difference between a palindrome and a mirror repeat? how does a hairpin form?
palindrome: traditional; think what the complementary strand looks like, complementary strand should read the same
mirror repeat: rotate 180° on y axis, if it reads the same sequence = mirror repeat
hairpin: palindrome sequence that folds in on itself; hairpins come together to form a cruciform
how do RNA molecules have quite complex structures?
RNA molecules can have quite complex structures, such as RNase P. multiple hairpins build on one another, which results in a lot of non-Watson and Crick base pairs in more complex structures
what are complex structures stabilized by?
complex structures are stabilized by non-watson and crick base pair interactions
the rule breaking of Watson-Crick gives ribozymes unique structures and function
ex: tRNA, hammer head ribozyme used to degrade mRNA, and intron (ribozyme) catalyzes its own excision from between exons in an mRNA strand
what is deamination?
deamination is a molecular mechanism of spontaneous mutagenesis
spontaneous: happens due to equilibrium shift of structures
deamination: hydrolysis rxn, amine group is replaced by a carbonyl group
very slow reactions and unfavorable
large number of residues
the net effect is significant: 100 C→U events/day in mammalian cells
repaired through base excision repair pathways (BER): recognizes non-Watson-Crick base pairs and replaces them with the correct one
what is depurination?
depurination is a molecular mechanism of spontaneous mutagenesis
depurination: breakage of N-glycosidic bond
N-glycosidic bond is hydrolyzed
significant for purines: 10,000 purines lost/day in a mammalian cell
what are the two mechanisms for which external factors induce mutations?
molecular mechanisms of oxidative and chemical mutagenesis
oxidative damage
creates ROS (superoxide, hydrogen peroxide, and hydroxyl radical): ROS is way more reactive and may react with biological material
hydroxylation of guanine
mitochondrial DNA is most susceptible
chemical alkylation
add carbon chains/groups to bases
methylation of guanine
cells have mechanisms to correct for most of these modifications but must do so with the correct timing
what are alkylating agents?
alkylating agents donate carbon groups to DNA and RNA
what is radiation-induced mutagenesis?
molecular mechanism of radiation-induced mutagenesis:
UV light induces dimerization of pyrimidines (causes covalent linkage and a change in the phosphodiester backbone); this may be the main mechanism for skin cancers
ionizing radiation (x rays and gamma rays) causes ring opening and strand breaking → these are difficult to fix
cells can repair some of these modifications but others cause mutations. accumulation of mutations is linked to aging and carcinogens
how can nucleotides function as an energy source?
energy source of nucleotides:
→ changing the base will give different types of energy carriers (swap out bases to give different metabolic pathways)
protein synthesis (ribosome function) and signal transduction (G-proteins)
phospholipid and membrane synthesis
carbohydrate/glycogen synthesis
how can nucleotides function as coeznymes?
coenzymes:
carries acyl groups (to start citric acid cycle)
electron carriers (are modified nucleotides and have a conjugated double bond)
how can nucleotides function as regulatory molecules?
regulatory molecules:
cell signaling and signal transduction pathways
what is recombinant DNA?
recombinant DNA is artificially created DNA that combines sequences that do not occur together in nature '
basis of much of modern molecular biology
molecular cloning of genes: gene of one organism is put into another organism
overexpression of proteins
transgenic food, animals
what is organism cloning? DNA cloning and its steps?
organism cloning
creation of genetically identical copies of an organism
organisms may not appear identical due to regulation of gene expression during development
DNA cloning
creation of identical copies of a piece of DNA (gene)
isolate a specific gene from the source organism and amplify it in the target organism
basic steps
cut the source DNA at the boundaries of the gene
select a suitable carrier DNA (vector)
insert gene into the vector
insert the recombinant vector into host cell
let the host cell produce multiple copies of recombinant DNA
what is step 1 to DNA cloning?
step 1: generate recombinant vector
goal: gene from one organism into a plasmid DNA
cloning vector is cleaved with restriction endonuclease
DNA fragment of interest is obtained by cleaving chromosome with a restriction endonuclease
fragments are ligated to the prepared cloning vector
recombinant vector (plasmid) made
what is step 2 to DNA cloning?
step 2: introduce DNA into organism (transformation)
DNA introduced into host cell
propagation (cloning) of transformed cell produces many copies of recombinant DNA
what are restriction endonucleases?
restriction endonucleases are enzymes that target palindromic sequences of DNA
palindromic sequences in DNA are recognized by restriction enzymes. sequences
common in bacteria
eliminates infectious viral DNA
some make staggered cuts
sticky ends
some make straight cuts
blunt ends
large number are known
commerically available
well documented: REBASE
what are DNA ligases?
DNA ligases are enzymes that covalently joins two DNA fragments
it recognizes the phosphodiester backbone and ligases it together, not sequence specific
normally function in DNA repair
human DNA ligase uses ATP
bacterial DNA ligase uses NAD
energy dependent