Chapter 8: DNA/Genetics
1/45
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
46 Terms
How do we know that DNA is the hereditary molecule?
The process by which cells use hereditary information stored in DNA is very elegant.
DNA must be retained intact, yet copied to make new cells (DNA replication).
It must be turned into multiple “working copies” in the form of RNA to provide instructions to produce enzymes/structural proteins (transcription).
The RNA molecules must be read and decoded to form the enzymes/structural proteins of the cell (translation).
The systems must have the ability to deal with damage to the core DNA molecules of the cell (DNA repair).
What was the discovery of DNA and its role (The Griffith Experiment - 1920s)?
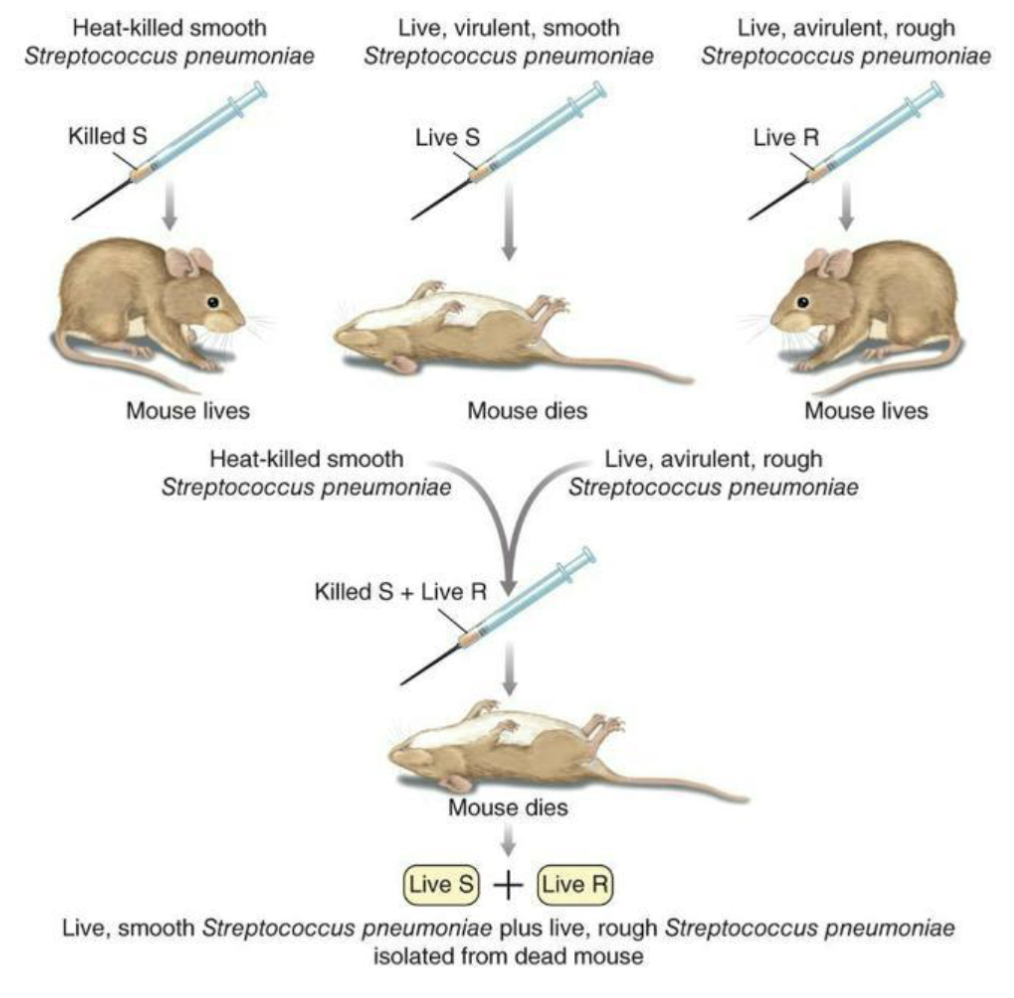
The Griffith Experiment (1920s): One of the earliest evidence that DNA may be the hereditary information in the cell
Image
Bacterial strains
S strain (smooth): Has polysaccharide capsule, Virulent (causes disease)
R strain (rough): No capsule, Avirulent (non-disease causing)
Experimental results
Heat-killed S strain → Mouse lives
Bacteria are dead → no infection.
Live S strain → Mouse dies
Capsule protects bacteria from immune system → infection.
Live R strain → Mouse lives
No capsule → immune system destroys bacteria.
Heat-killed S + Live R → Mouse dies
Unexpected result.
Live S bacteria recovered from mouse.
Key conclusion
DNA from dead S bacteria → taken up by R bacteria → R becomes virulent S.
Biological significance
Demonstrated bacterial transformation (genetic information transfer).
Suggested heritable traits can be transferred between cells.
Transformation: uptake of foreign DNA by a bacterial cell that changes its phenotype.
What was the discovery of DNA and its role (The Avery, MacLeod, and McCarty Experiment - 1940s)?
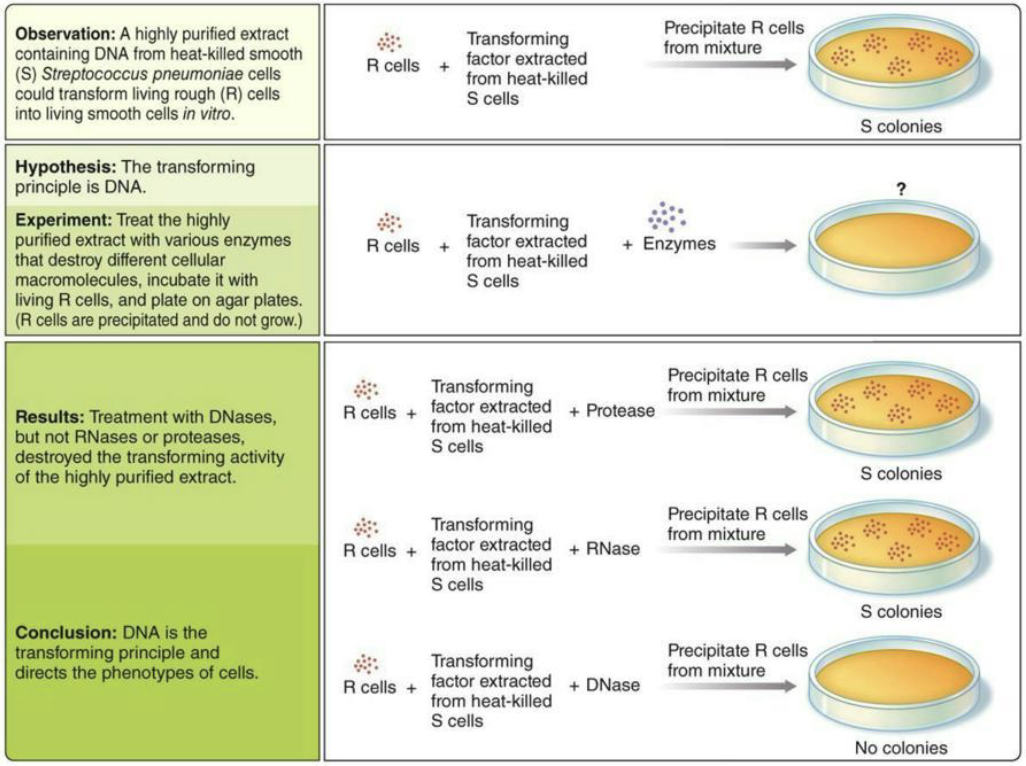
The Avery, MacLeod and McCarty Experiment (1940s): Is DNA, RNA or protein the causative agent of the “transformation” effect observed by Griffith
Image
Tested what molecule caused transformation in Streptococcus pneumoniae.
Extract from heat-killed S cells could transform R cells → S cells.
Added enzymes that destroy molecules: protease (protein), RNase (RNA), DNase (DNA).
Transformation still occurred with protease and RNase, but stopped with DNase.
Conclusion: DNA is the transforming principle (genetic material).
What was the discovery of DNA and its role (The Hershey-Chase experiment)?
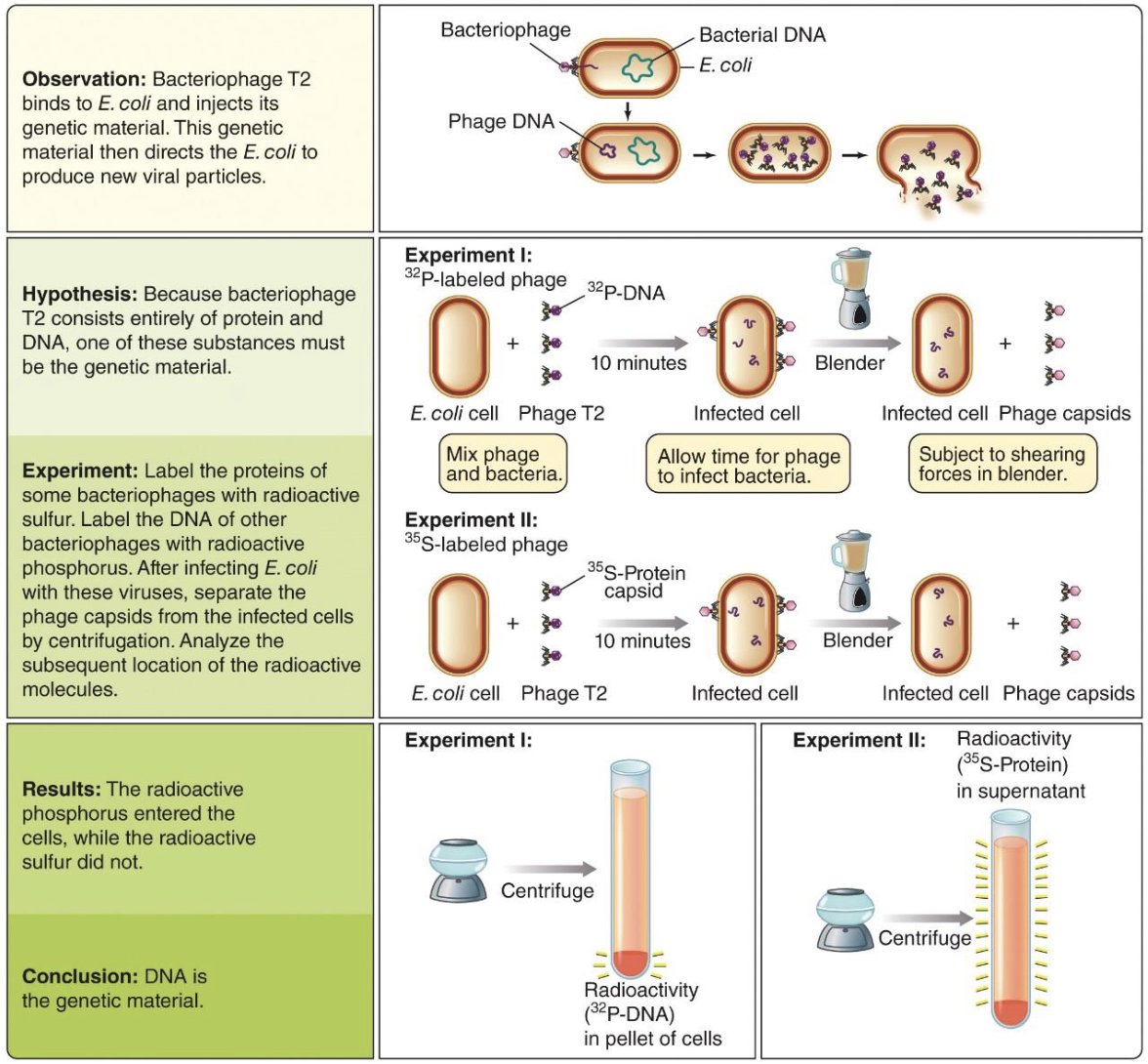
The Hershey‒Chase experiment: Radioactive labelling
Image
Tested whether DNA or protein is genetic material using Bacteriophage T2 infecting Escherichia coli.
Labeled DNA with radioactive phosphorus (³²P) and protein with radioactive sulfur (³⁵S).
After infection, used a blender and centrifuge to separate phage coats from bacteria.
³²P (DNA) entered the cells, while ³⁵S (protein) stayed outside.
Conclusion: DNA is the genetic material.
Who discovered the structure of DNA?
Double helix structure of DNA discovered in 1950s by James Watson, Francis Crick and Rosalind Franklin
What is the structure of DNA?
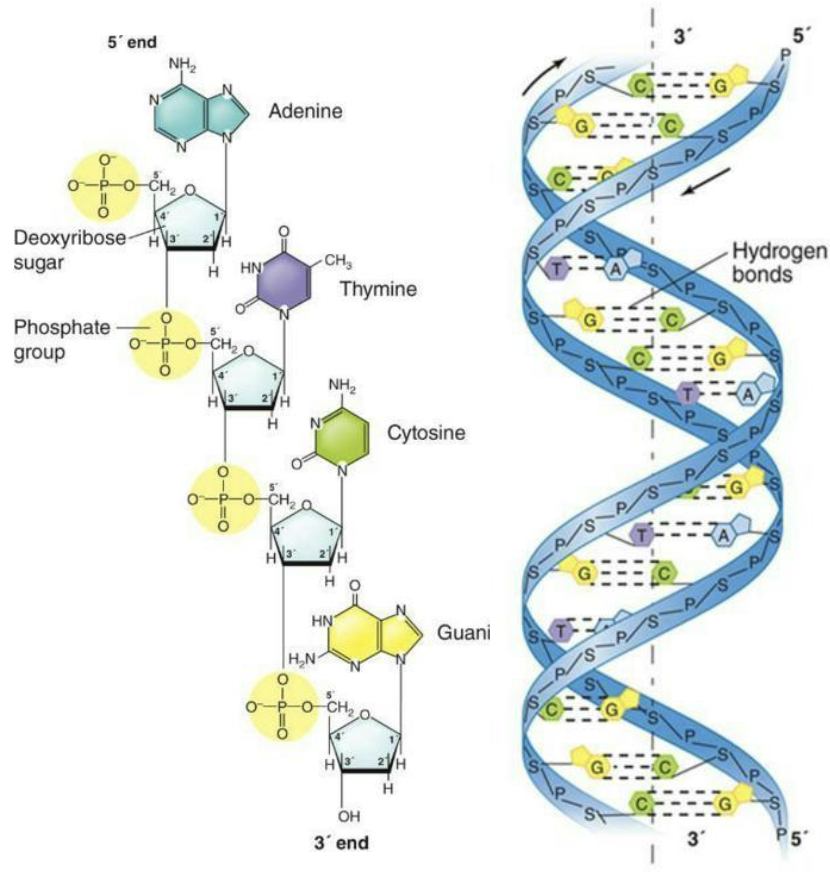
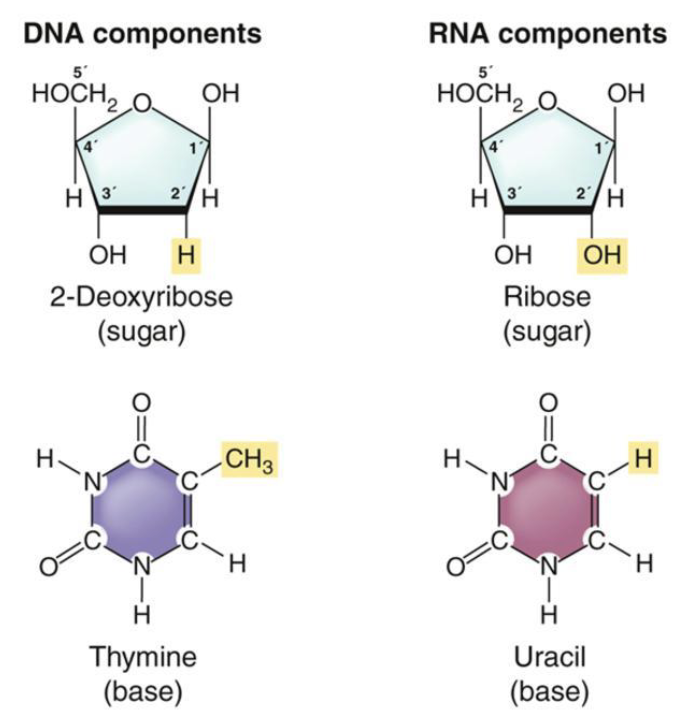
Each nucleotide building block of DNA consists of
A five-carbon sugar, 2-deoxyribose
A phosphate group attached to the 5' carbon of the sugar
A nitrogenous base attached to the 1' carbon of the sugar
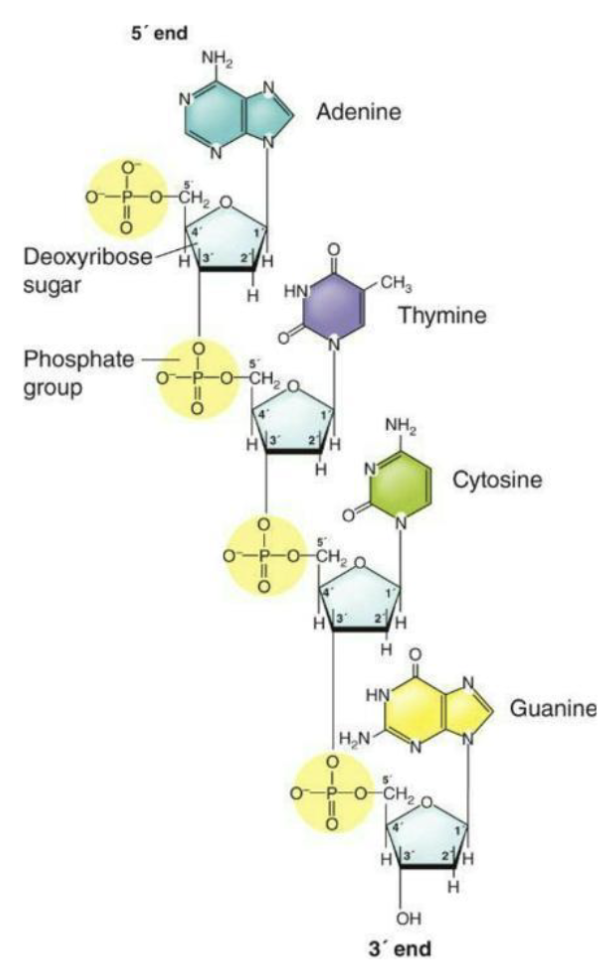
How are the two strands linked?
One strand of DNA is complementary to the other strand.
A pairs with T; G pairs with C by hydrogen bonding.
Phosphodiester covalent bonds form the sugar/phosphate backbone of each strand.
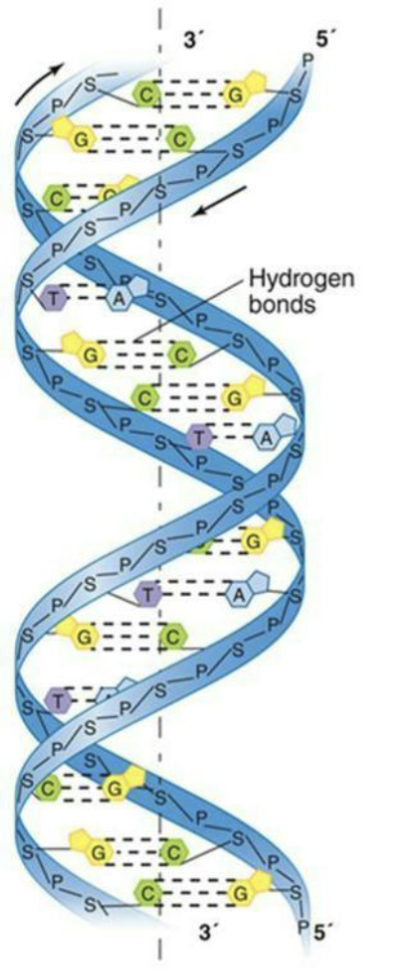
How is DNA packaged?
Structure of DNA is conserved across the three domains but the way its packaged is not.
Bacteria: Single circular chromosome, supercoiled
Archaea: Single circular chromosome packaged around histones
Eukarya: Multiple linear chromosomes packaged around histones. The wrapping of dsDNA around histones helps compact the very large chromosome structures of eukaryotic cells.
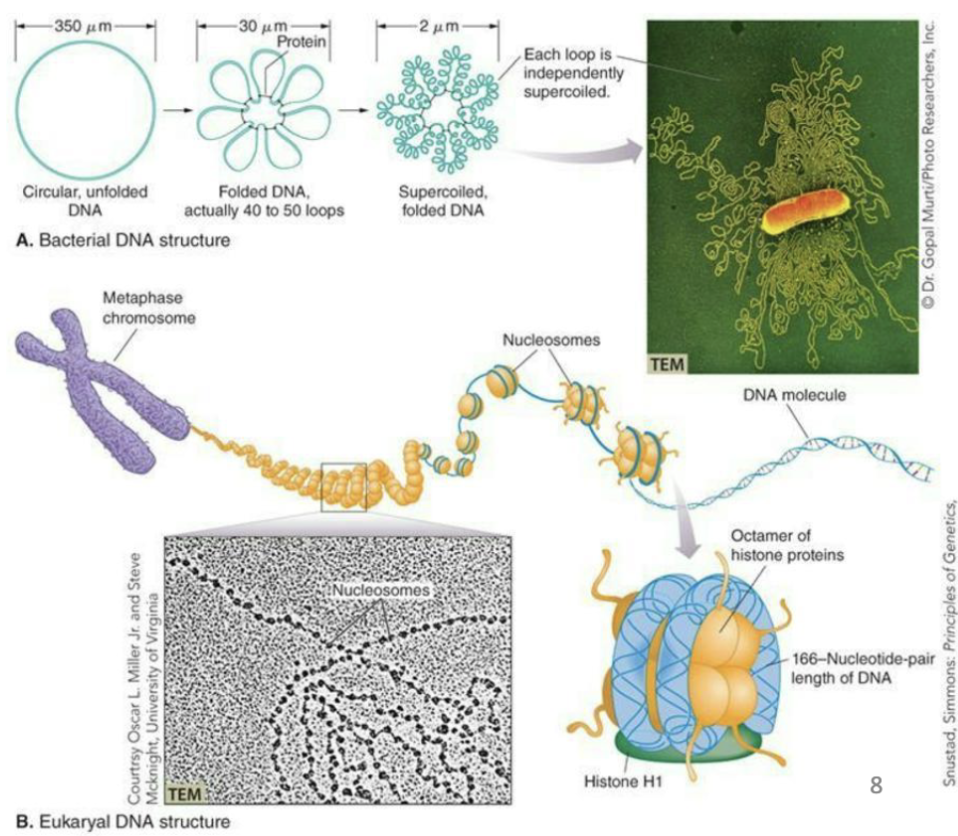
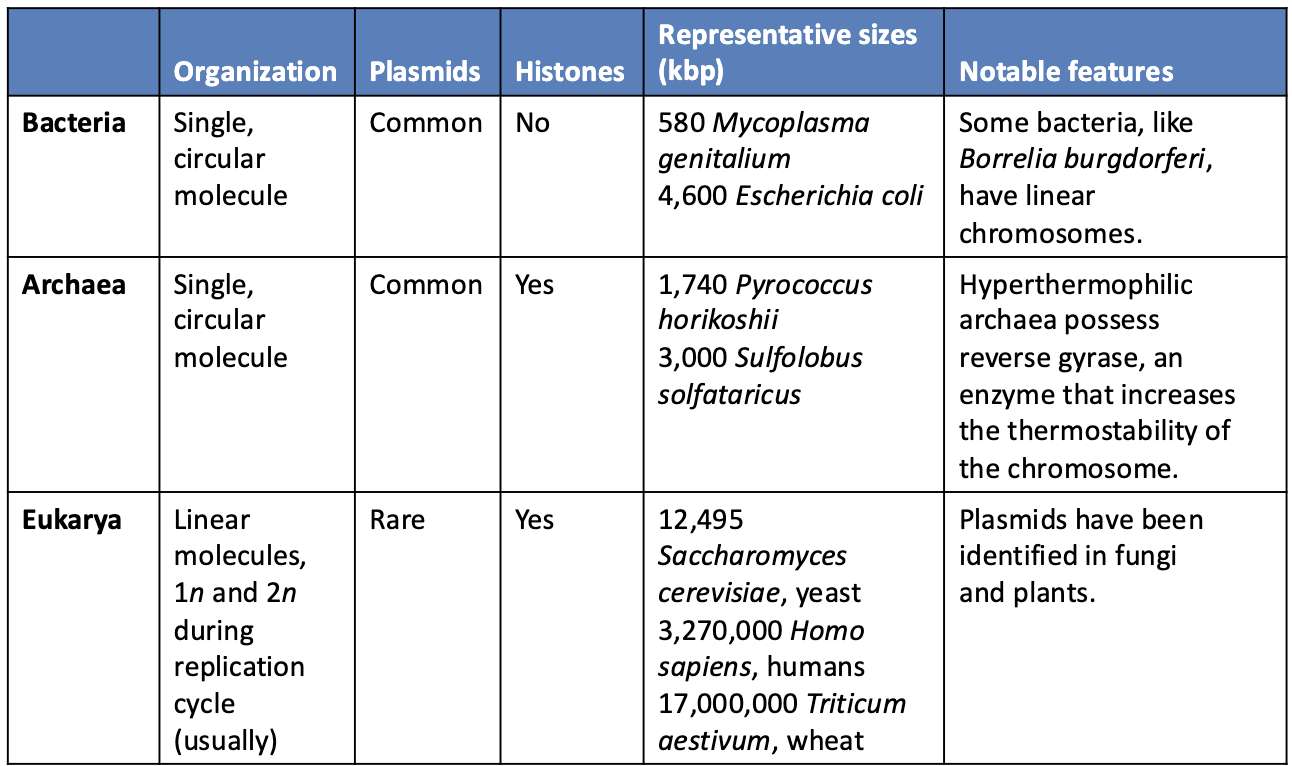
Compare DNA packaging in Bacteria, Archaea, and Eukarya?
Bacteria
Single circular chromosome
Plasmids common
No histones
Genome size ~ 580–4,600 kbp
Some species (e.g., Borrelia burgdorferi) have linear chromosomes
Archaea
Single circular chromosome
Plasmids common
Histones present
Genome size ~ 1,740–3,000 kbp
Some hyperthermophiles have reverse gyrase for DNA stability
Eukarya
Multiple linear chromosomes
Histones present
Plasmids rare
Usually diploid (2n), haploid (1n) at some stages
Genome sizes much larger
Saccharomyces cerevisiae ~12,495 kbp
Homo sapiens ~3,270,000 kbp
Triticum aestivum ~17,000,000 kbp
What are exceptions to DNA packaging?
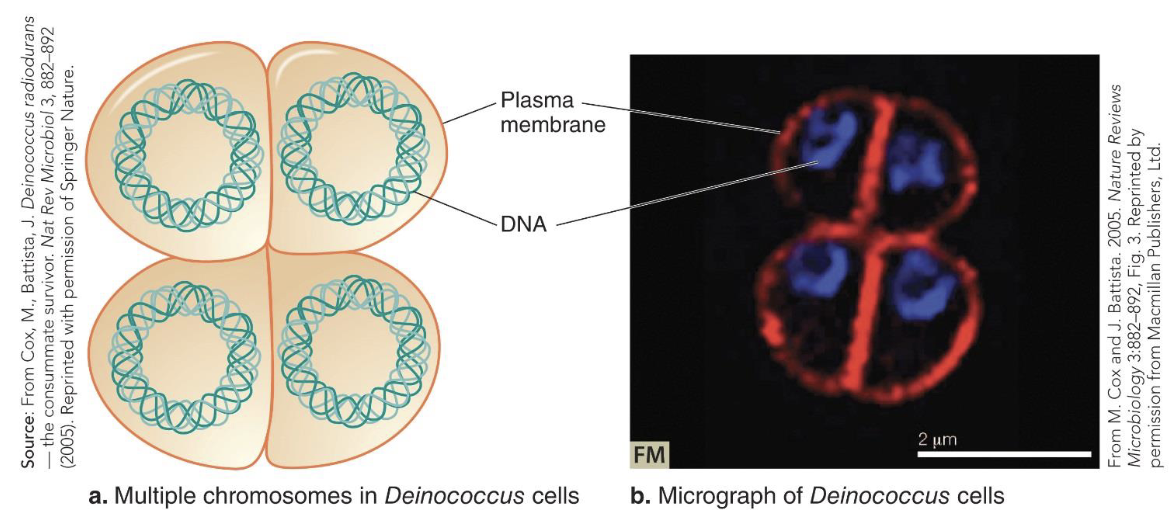
D. radiodurans may possess 4 to 10 copies of its genome.
The stacked genome copies may help give this bacterium its strong radiation-damage-resistance trait
What does it mean by DNA replication is semi-conservative?
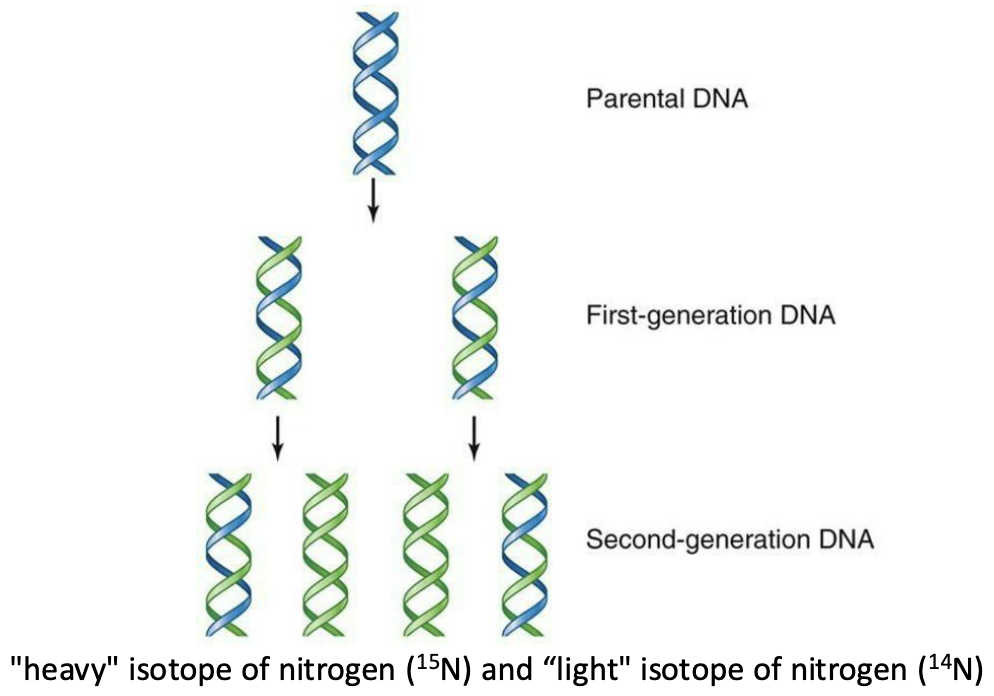
DNA replication is semi-conservative: each strand of a newly copied DNA molecule contains one strand form the original molecule and one newly synthesized
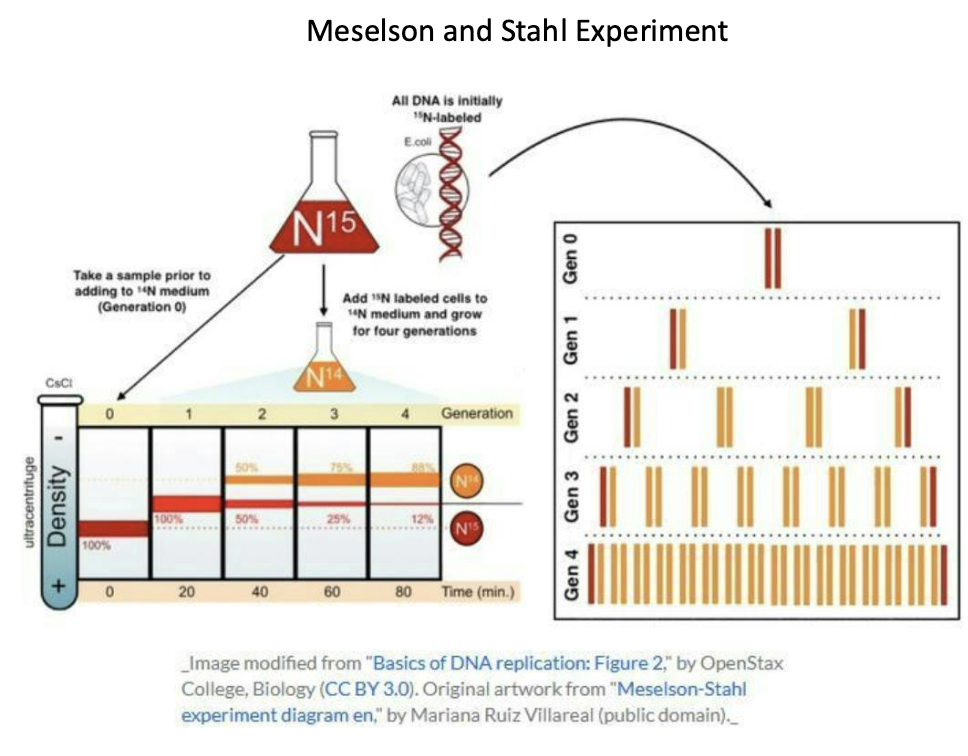
Meselson and Stahl Experiment: Showed that DNA replicates semi-conservatively by growing bacteria first in heavy nitrogen (¹⁵N) and then in light nitrogen (¹⁴N) and observing that new DNA molecules contain one old strand and one new strand
What is the initiation of DNA replication?
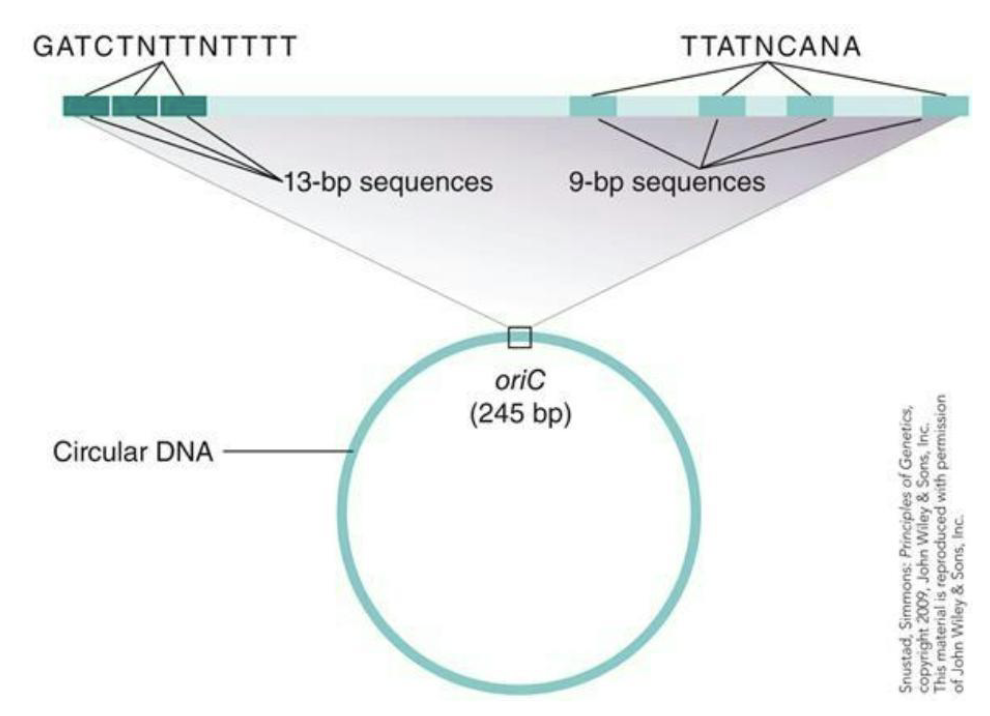
In bacteria DNA replication begins at a specific sequence in the molecule called the origin of replication (oriC)
Jacob Brenner and Cousin discovered how DNA was replicated in E.coli and oriC
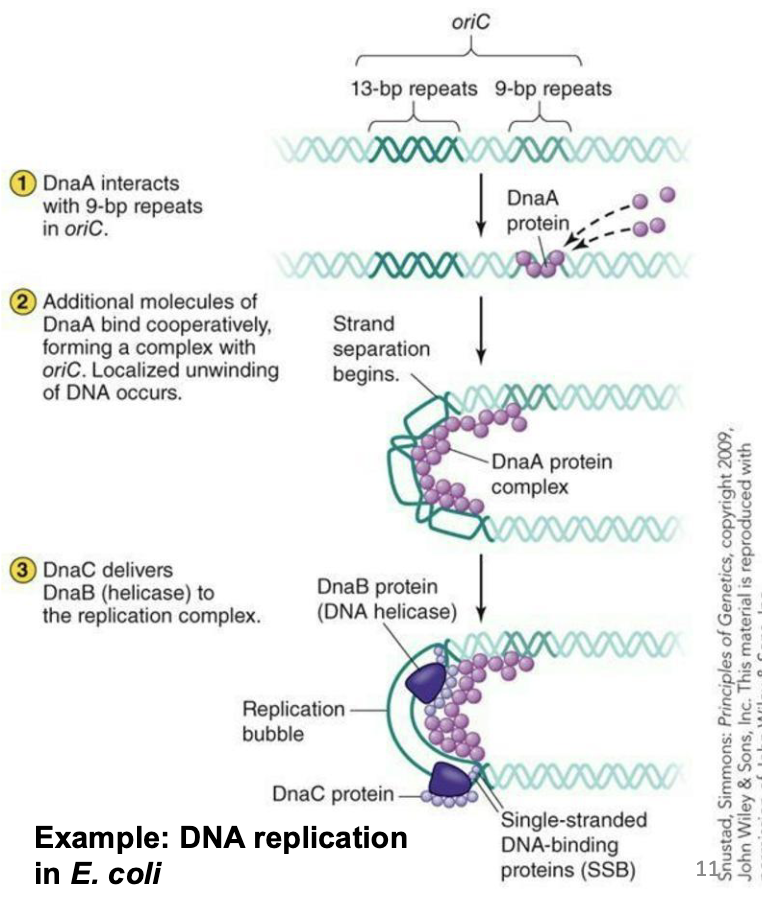
What are the origins of replication?
DnaA protein binds to oriC.
DnaB (a helicase) is recruited with DnaC (a helicase loader).
DnaG (a primase) is recruited to lay down initial RNA primers needed for DNA polymerases to work.
Single-stranded DNA binding proteins are recruited to help keep the DNA unwound.
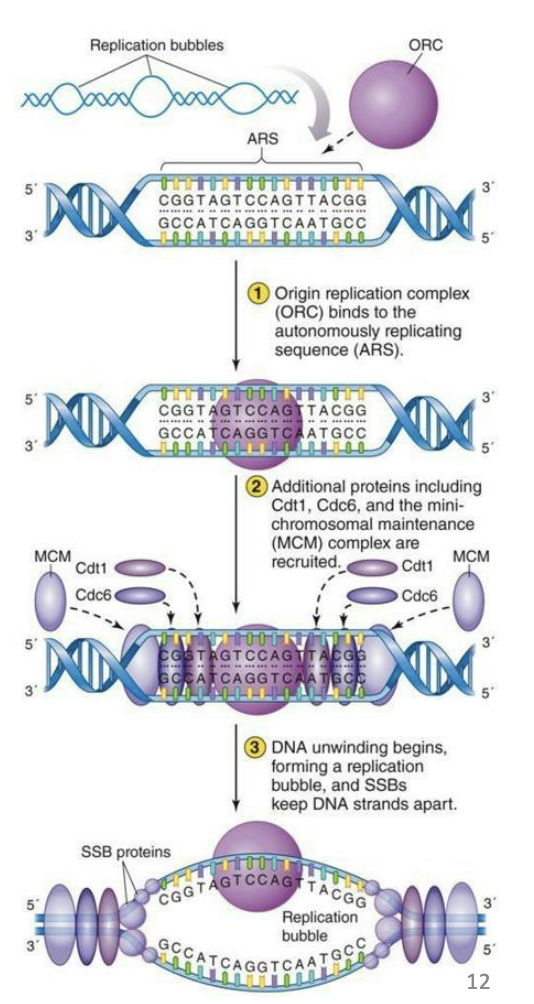
What is an example of Eukaryotic replication initiation?
Multiple origins of replication on each chromosome.
ARS – 10 -100 kb apart
Chromosomes are much larger, so they need multiple starting points for replication.
Studied in yeast.
Very similar to process in bacteria, just using different proteins.
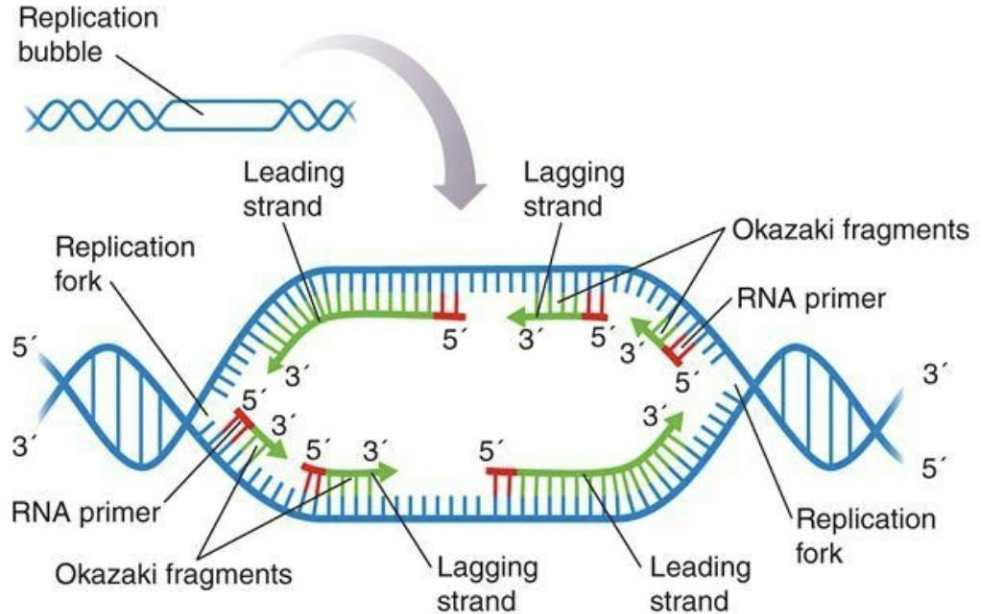
What is elongation in DNA replication?
Initiation and elongation: DNA polymerase III adds nucleotides to RNA primers (synthesized by primase)
Continuous leading strand and discontinuous lagging strand (forming Okazaki fragments)
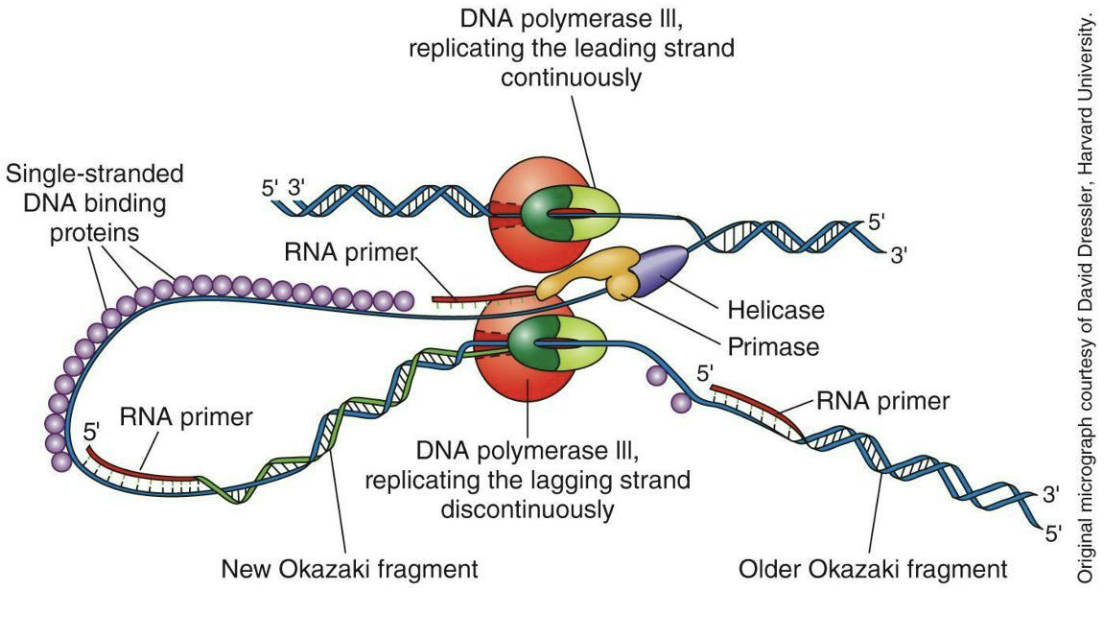
What is at each replication fork?
Initiation and elongation: DNA polymerase III adds nucleotides to RNA primers (synthesized by primase)
Continuous leading strand and discontinuous lagging strand at each replicating fork - a single fork is shown below
What is the role of DNA ligase in elongation?
RNA primers removed and replaced by DNA (DNA polymerase I)
Sealed by DNA ligase joining phosphate to sugar backbone
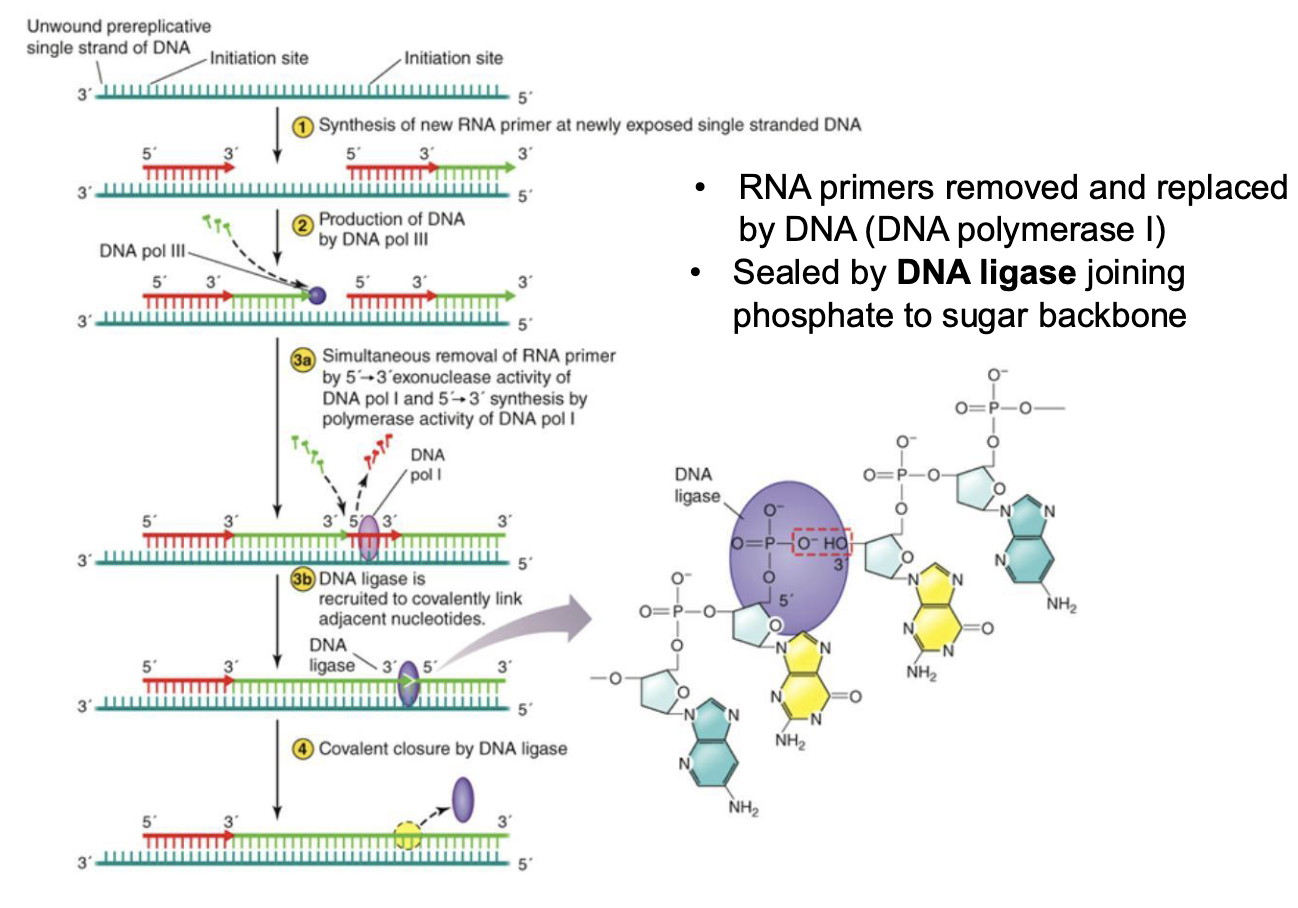
What are the interesting features regarding DNA replication?
Processivity ..... the ability of DNA polymerase to carry out continuous DNA synthesis on a template DNA without dissociation. A sliding DNA clamp protein or clamp processivity factor, is found in prokaryotes, eukaryotes and archaea. In E. coli β clamp –
E. coli DNA polymerase can synthesize as fast as ~ 1000 bp per second
Without proof reading – 1 error in 107 bp
With proofreading 1/109 nucleotides
What is termination in DNA replication?
Termination of circular chromosome replication
1) Bidirectional replication begins at oriC
2) Tus proteins bind to ter sites on the chromosome, stopping elongation
3) Topoisomerase is recruited
4) Newly replicated circular molecules are disentangled
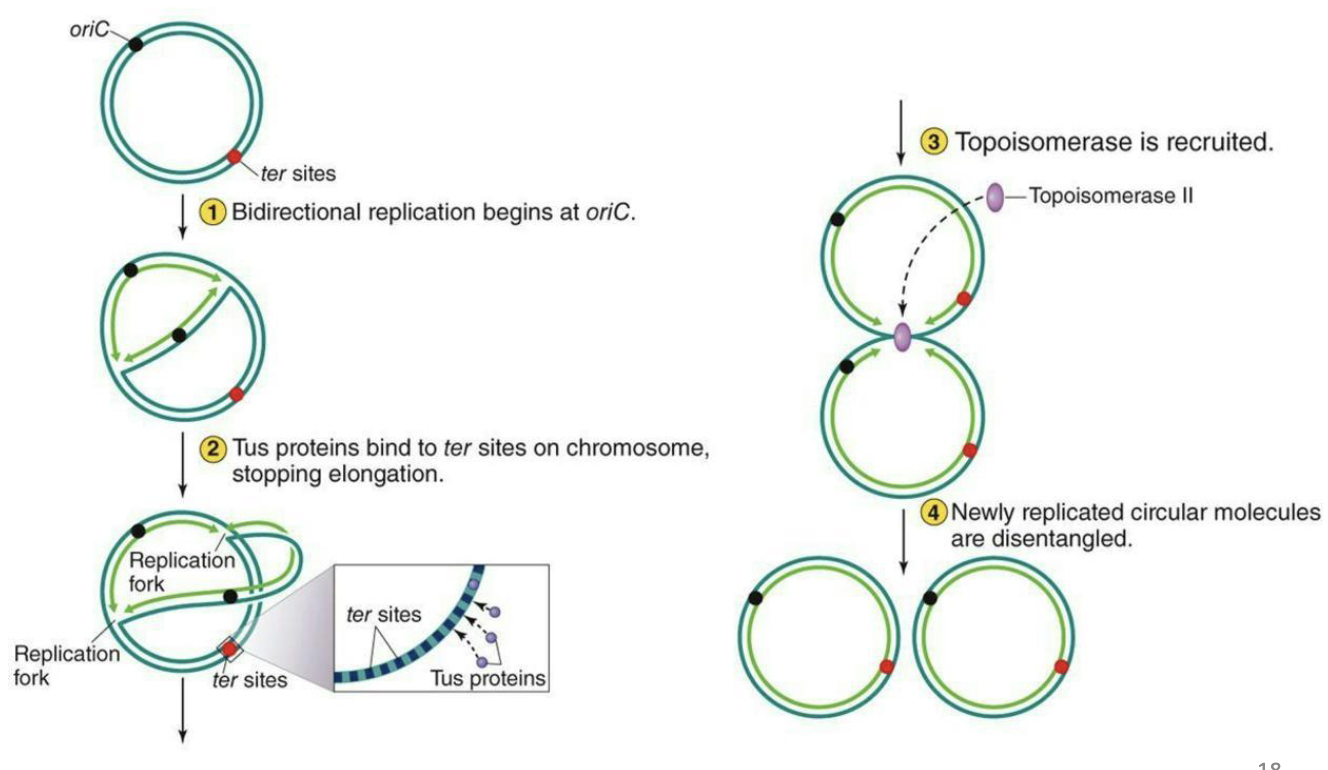
What are the steps of linear termination?
RNa primer removed
RNA primer removal produces 5' end that cannot be extended by DNA polymerase.
Telomerase binds.
Telomerase extends 3' end (RNA-templated DNA synthesis).
Translocation
Repeat extension and translocation several times; release telomerase.
Completion of the complementary strand by DNA polymerase (DNA-templated DNA synthesis)
DNA ligase is recruited to link adjacent nucleotides.
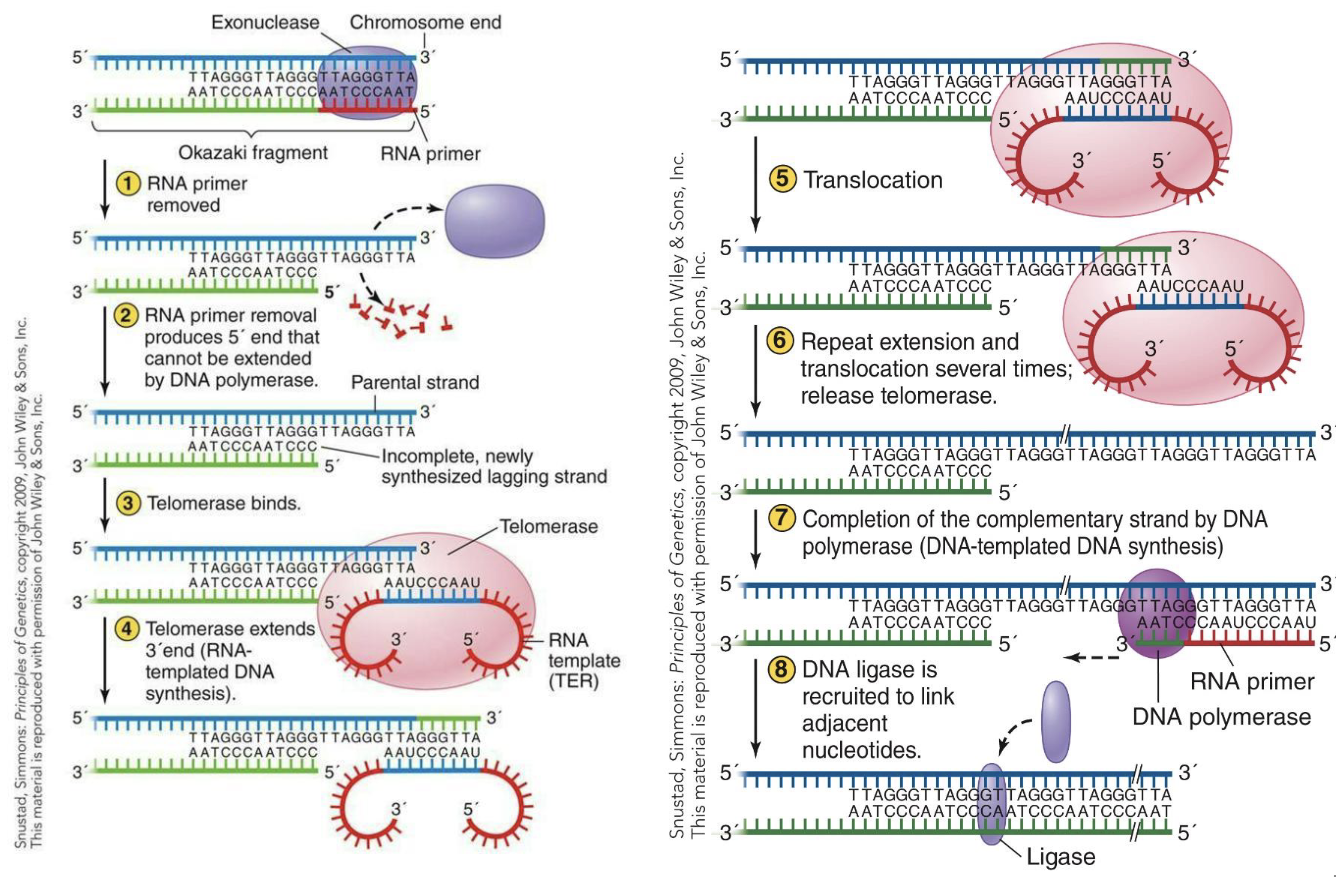
Summary
Experiments by Griffith (~1920) and Avery, McLeod and McCarty (~ 1940) showed DNA from one organism could alter (transform) the characteristics of another
DNA (A,T,C,G) is organized as antiparallel, complementary strands that form a double helix
DNA replication is semi-conserative
Begins at origin of replication, with proteins forming a replication bubble – one oriC in bacteria multiple in eukaryotic chromosomes
DNA polymerase extends an RNA primer by adding nucleotides to the 3’ end i.e. synthesis is 5’ → 3’
Extends continuously on leading strand and discontinuously on lagging strand
Ends by disentanglement of circular chromosomes by topoisomerase or extension of linear chromosomes in eukarya by telomerase.
What is Ribonucleic acid?
Transcription is the copying of a segment of DNA into single stranded RNA molecule
Can be transcribed to messenger, transfer, ribosomal or regulatory RNA molecules
What is messenger RNA?
Messanger RNA (mRNA) are approx. 500-10,000 nucleotides
Contains information to translate into proteins (genes)
5’ and 3’ untranslated regions carry features involved in initiating and terminating translation
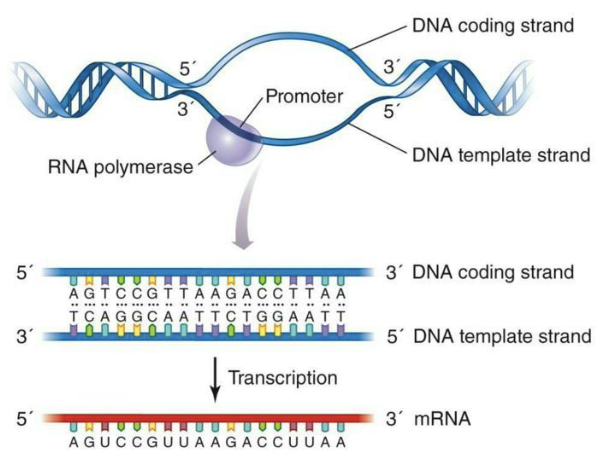
Template strand: the DNA strand RNA polymerase reads to build mRNA.
Coding strand: the DNA strand with the same sequence as mRNA (except T → U), not used as the template.
Bacterial genes are about 1 Kb in size, but mRNA may be much longer because it is polycistronic (can encode several genes on same RNA)
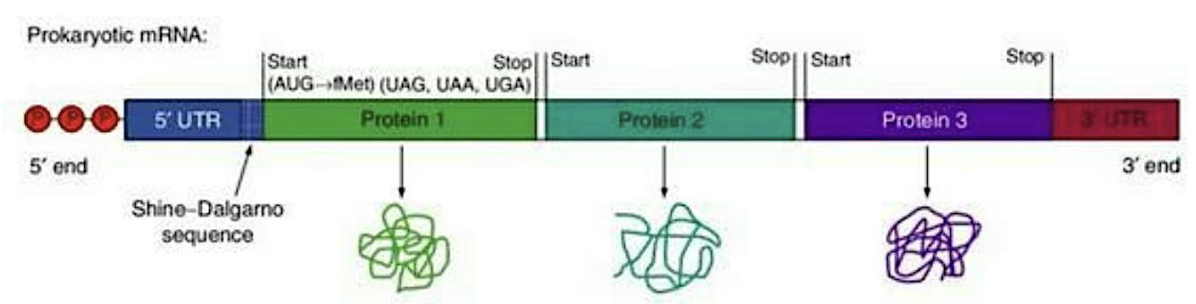
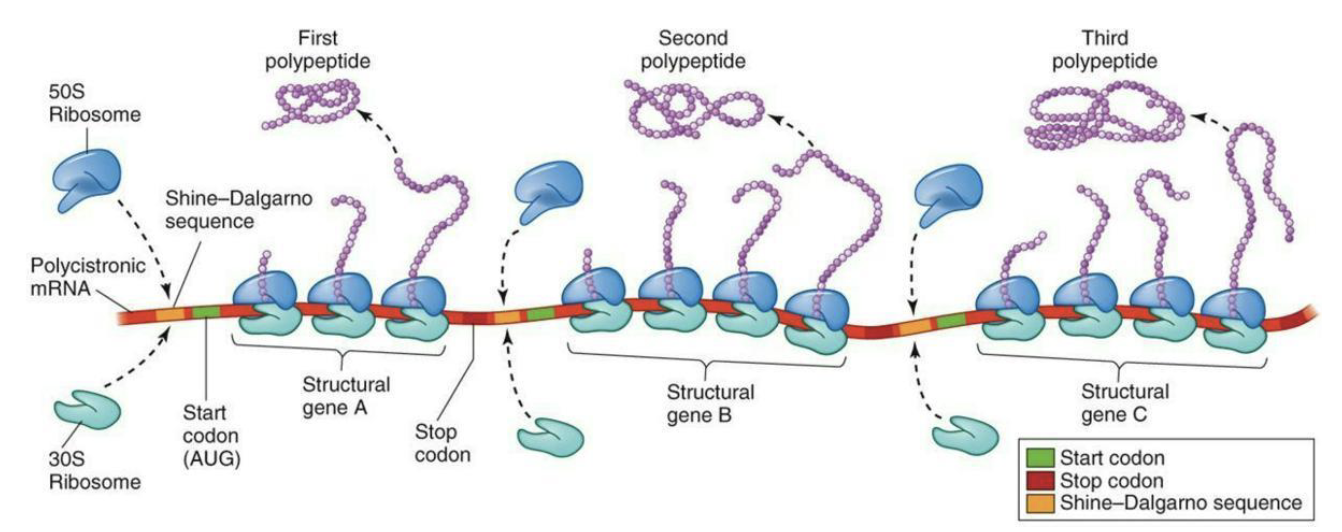
What does the diagram of prokaryotic polycistronic mRNA show?
Polycistronic mRNA → one mRNA codes for multiple proteins.
5′ UTR at start; 3′ UTR at end.
Shine-Dalgarno sequence = ribosome binding site before each gene.
Each protein has its own start codon (AUG) and stop codon (UAG, UAA, UGA).
Ribosome can start translation multiple times on the same mRNA → produces Protein 1, 2, 3.
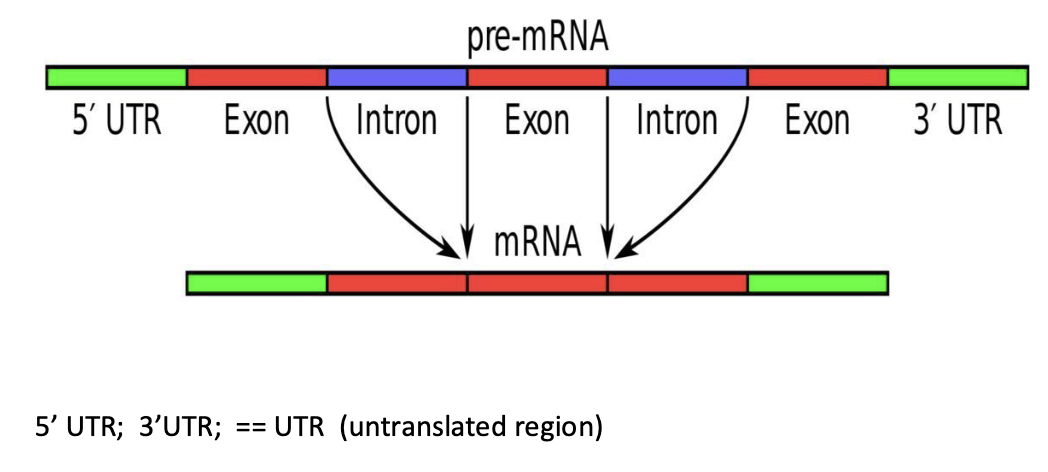
What is eukaryotic messenger RNA?
Eukaryotic genes often contain introns (non-coding regions) (and hence can be extremely large >10 Kb)
Introns must be processed and removed before mRNA can be translated
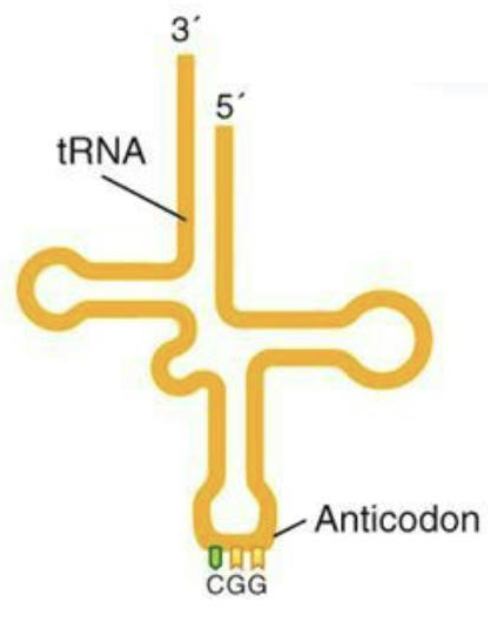
What is transfer RNA?
The transfer RNA (tRNA) are 75 - 100 nucleotides in size
Form secondary structures that deliver amino acids to the Ribosomes
Note the 3’ to 5’ orientation!!
mRNA is read 5′ → 3′ by the ribosome
The anticodon on tRNA binds antiparallel, meaning it pairs 3′ → 5′ to the codon
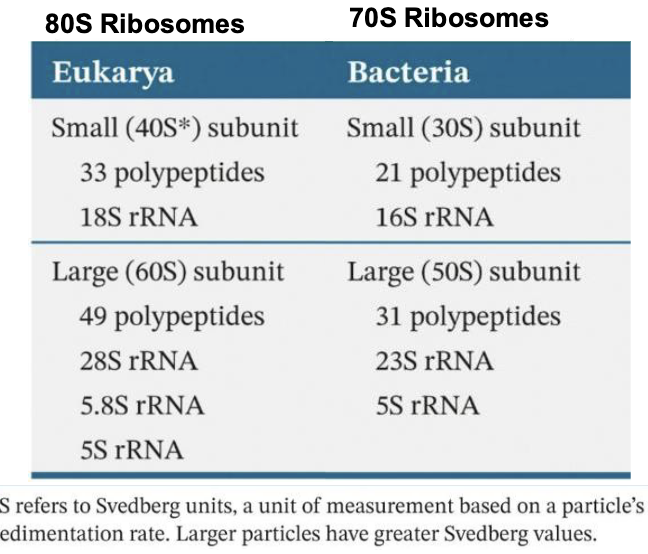
What is ribosomal RNA?
The ribosomal RNA (rRNA) are structural components of ribosomes (about 1500- 1900 nucleotides for small subunits and 2900-4700 nucleotides for large subunits
Vary between eukarya and bacteria, but overall similar organization
Eukarya 80S Ribosomes: Small 40S + Large 60S
Bacteria 70S Rubosomes: Small 30S + Large 50S

What is the promoter?
Promoters direct RNA polymerase to initiate transcription at specific positions on the DNA
Bacterial promoters have elements that are bound by sigma factors (-10/-35, for eg. for the main sigma factor RpoD in E. coli)
Alternative sigma factors can direct the RNA polymerase to the promoters of different sets of genes important under particular conditions (ex. Heat shock, starvation, nutrient depletion)
**Memorize image
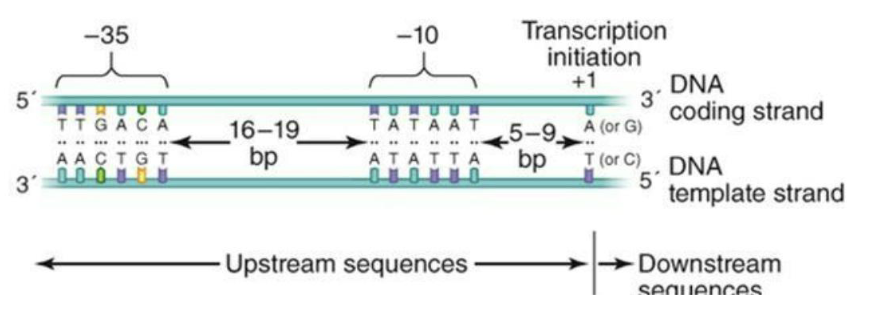
What are eukaryal promoters?
Eukaryal promoters have several different elements that are bound be several different transcription factors (TATA binding protein for eg.) to direct transcription initiation.
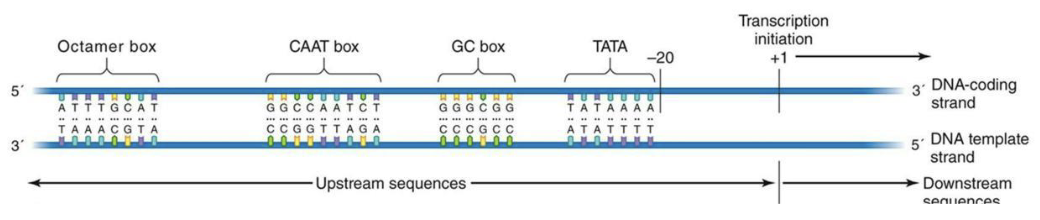
What is initiation and elongation like for bacteria?
In bacteria sigma factors direct the core RNA polymerase to the promoter
Only one RNA pol in bacteria
Different sigma factors can direct RNA pol to a different set of genes that are co-regulated (heat stress genes for example)
**Memorize image and steps
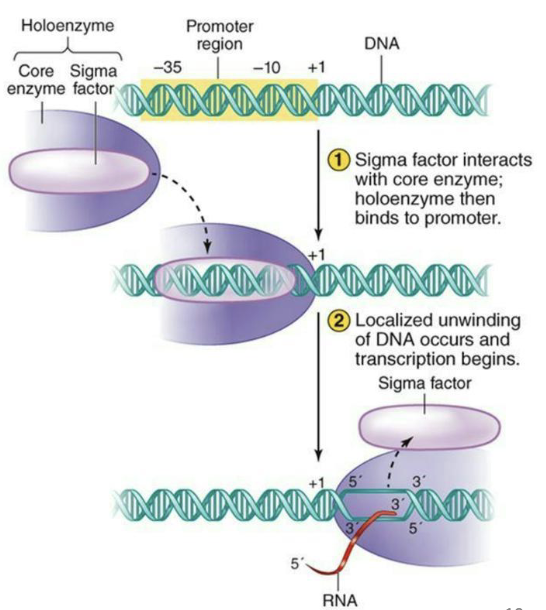
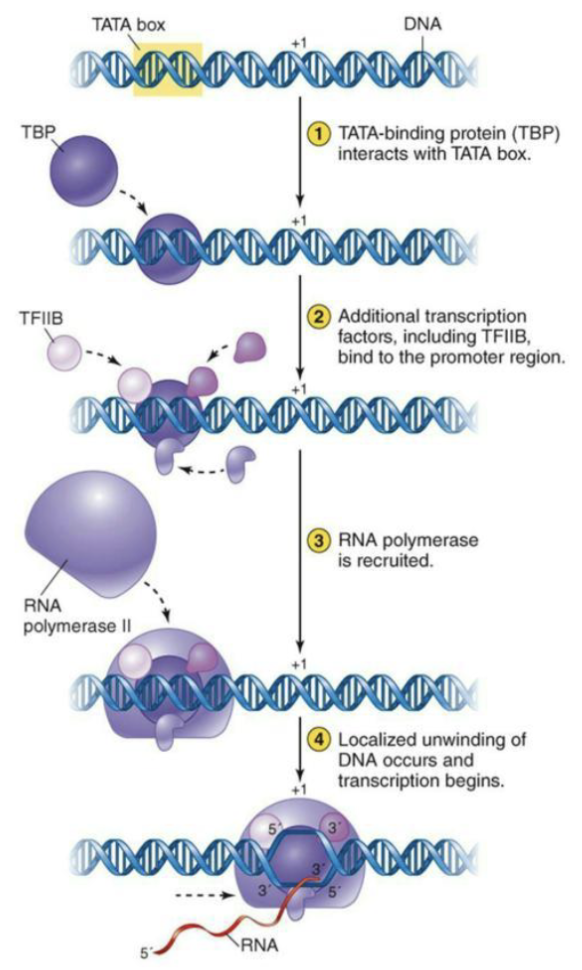
What is initiation and elongation like for eukaryotes?
In eukaryotes, transcription factors bind the various upstream elements
One of three RNA polymerases is directed to the site of transcription initiation
RNA pol does not bind directly to DNA
Binding is facilitated by various transcription factors
This binding initiates the unwinding of the DNA and the start of the actual transcription process.
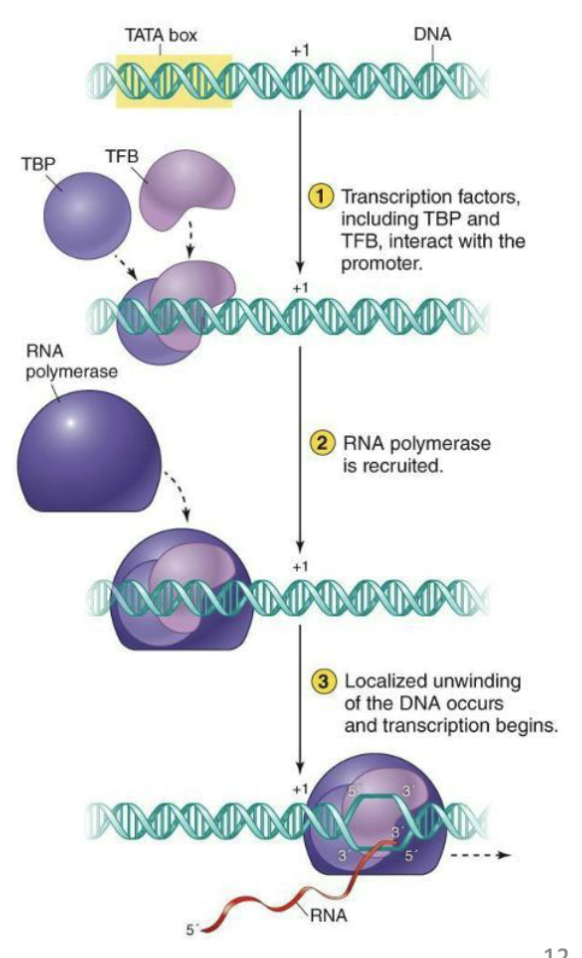
What is initiation and elongation like for archaea?
Single RNA polymerase like in bacteria, but looks more like eukaryal RNA pol II
Not well understood.
Resembles eukaryal transcription
Transcription factors direct RNA polymerase to promoter regions
TFB – transcription factor B – resemble eukaryal transcription factor TFIIB
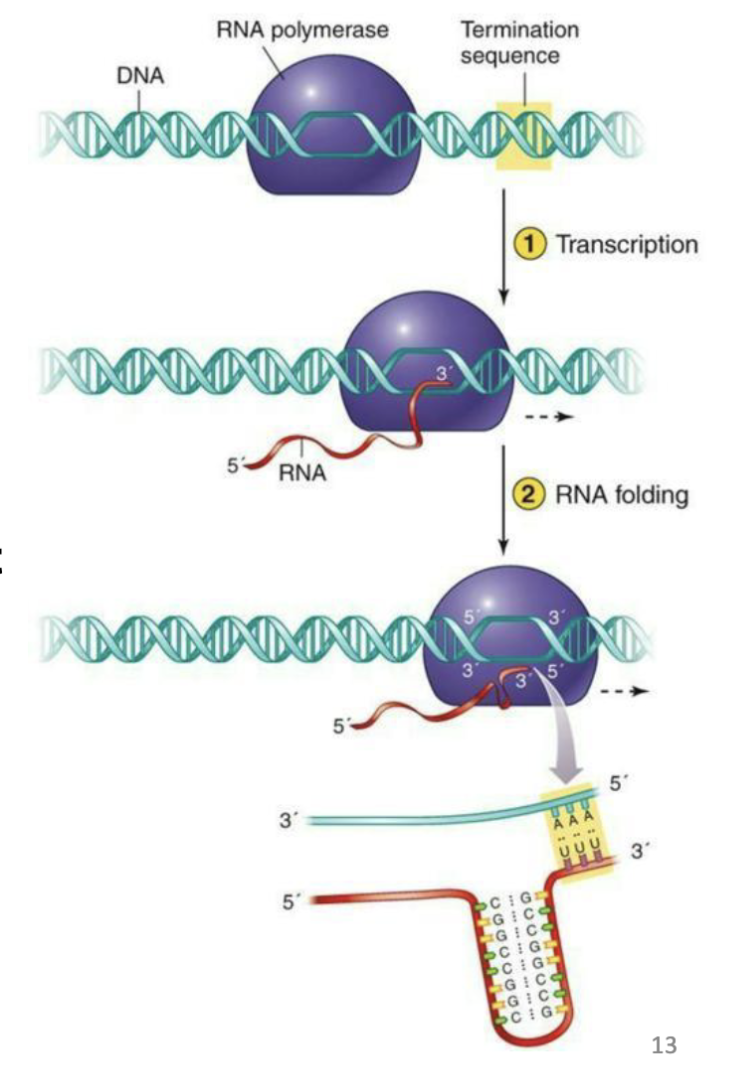
What is termination for bacteria?
In bacteria transcription can be terminated by a rho protein that causes RNA polymerase to pop off DNA at specific sequences
Alternatively, Rho- independent termination can be catalyzed by hairpin formed by complementary stretches of RNA, rich in guanine and cytosine followed by a segment rich in uracil
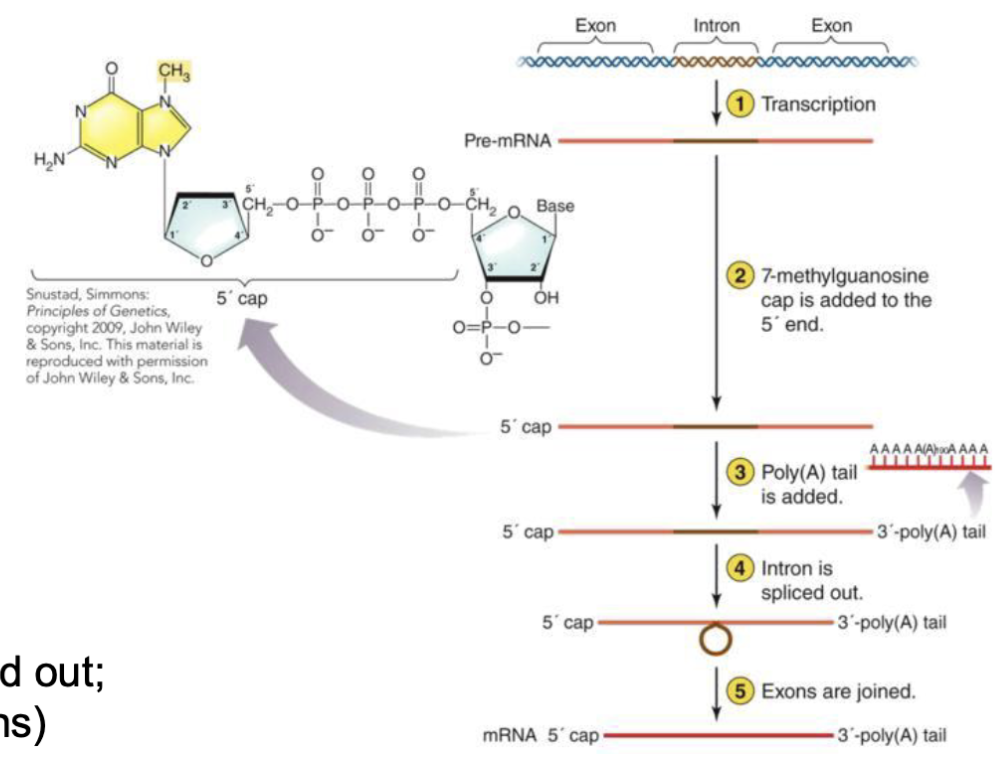
What is termination for eukarya?
In eukarya different mechanisms are used for termination by different RNA polymerases
RNA is modified after termination in eukarya
5' cap added (7-methylguanosine)
Poly(A) tail added
Introns (non-protein coding region) spliced out; exons (coding regions) joined together
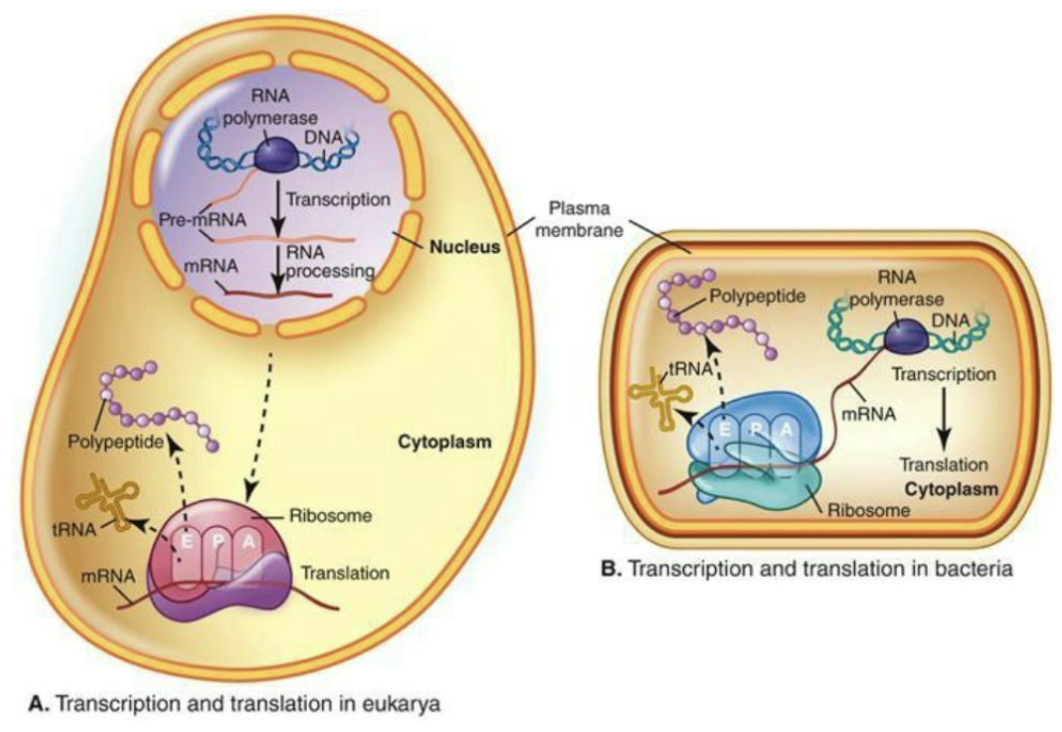
How are transcription and translation linked in bacteria?
Transcription and translation coupled in bacteria thanks to lack of nuclear membrane (independent in eukarya)
The lack of introns and post transcriptional modification of mRNA also make coupling easier.
In bacteria, mRNA has a short half-life so cells can rapidly change protein production in response to environmental changes.
What is the role of ribosomes in translation?
Ribosomes interact with mRNA and “charged” tRNA to assemble amino acids in a particular order
Anticodon of tRNA corresponds to codon in mRNA sequence
Amino acids delivered by tRNA are assembled with peptide bonds
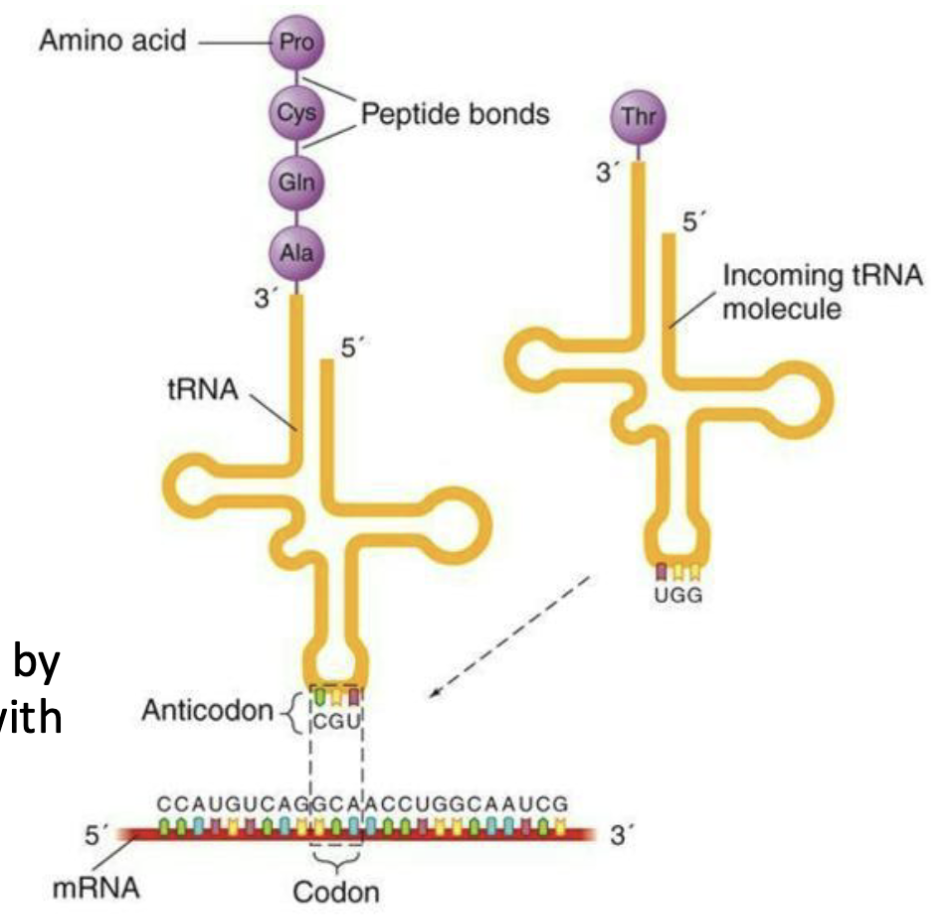
What are the special features of the amino acid code?
64 codons – 20 amino acids - code is degenerate
Several codons for each amino acid - third nucleotide in the codon is often variable = “wobble codon”
Specialized N-formyl methionine initiates translation at first AUG in bacteria
Unique initiator tRNA initiates in eukarya
Stop codons terminate translation of the protein
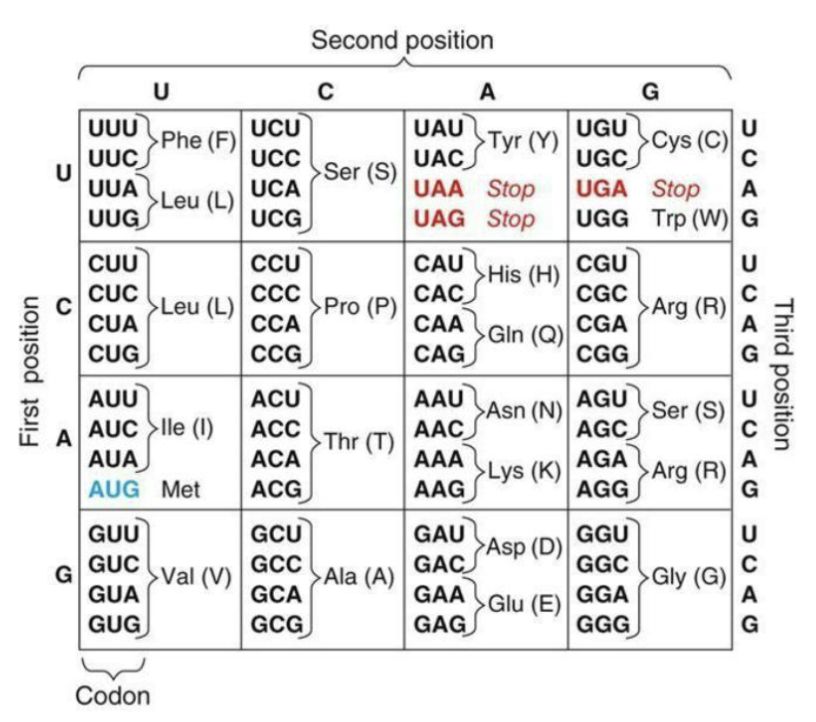
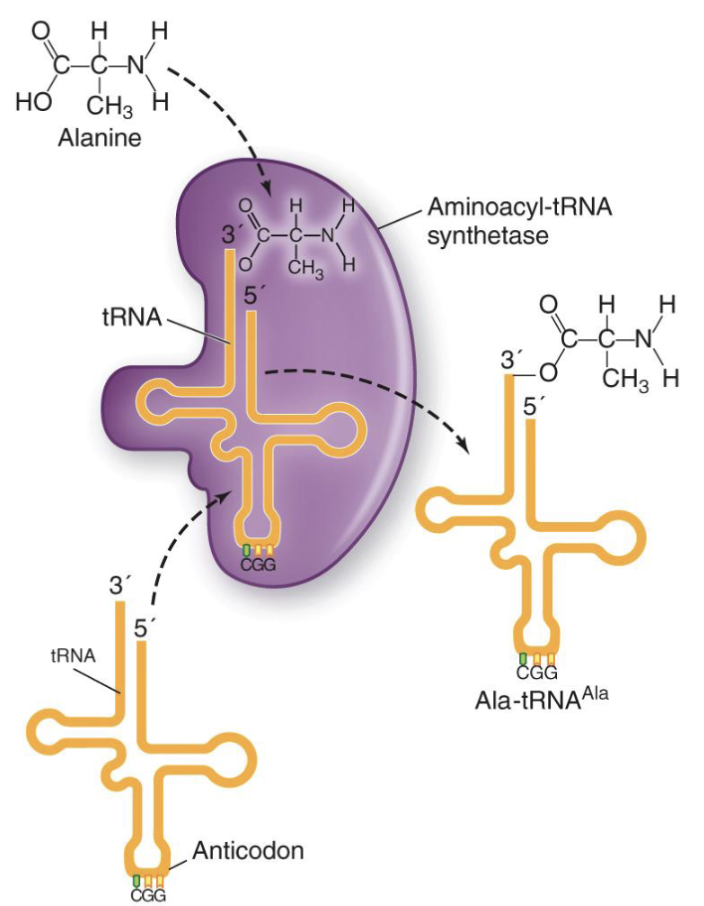
How is tRNA charged?
Prior to initiation of translation, each tRNA that will bring an amino acid to the ribosome needs to be “charged” with that amino acid
Highly specific aminoacyl-tRNA-synthetase enzymes achieve this charging
Create ester linkages
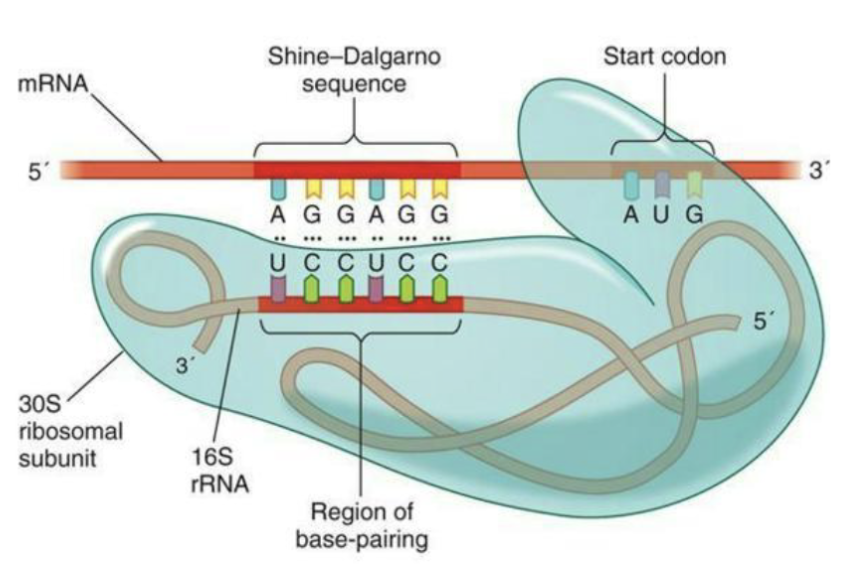
What is the Shine-Delgarno sequence?
In bacteria, interaction between the Shine- Delgarno sequence (ribosome binding site) and specific site in 16S rRNA in the small subunit of the ribosome
In eukarya mRNA is bound by several polypeptides including one that binds the 5’ cap before being bound by the ribosome
What are the different types of mRNA?
Bacterial mRNA can be polycistronic using multiple Shine-Delgarno sequences on the same mRNA molecule to encode multiple proteins
Eukaryal mRNA is monocistronic and TL begins at first start codon
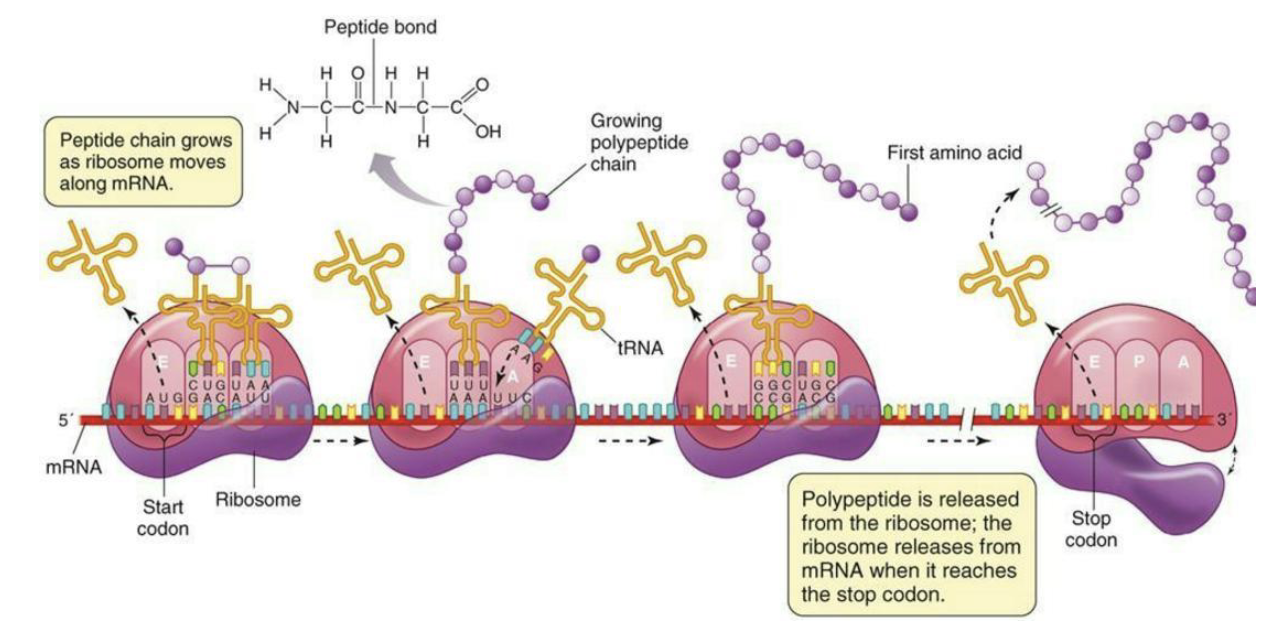
How does the ribosome move along codons in mRNA for elongation?
Ribosome moves along codons in mRNA. Charged tRNA’s enter A site, peptide bond is formed, and uncharged tRNA exits the E site.
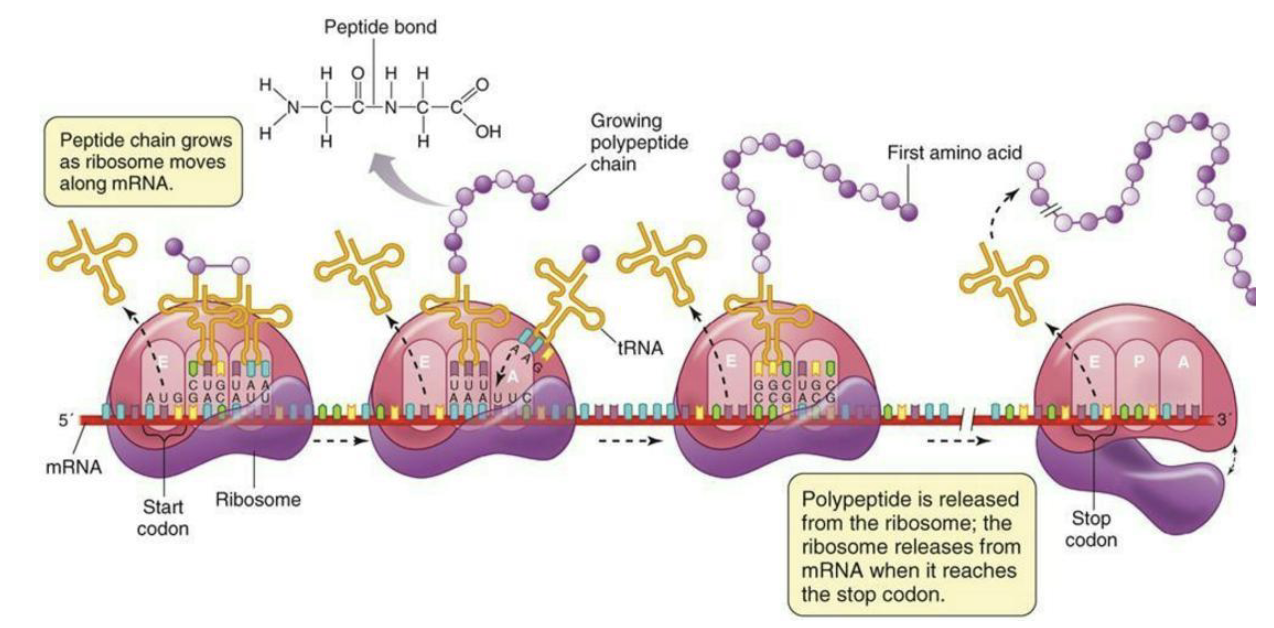
What is the function of release factors in termintion?
Release factors in both eukarya and bacteria cause the complex to fall apart when a stop codon is reached
How are proteins folded?
Proteins are folded upon leaving the ribosome, sometimes with the help of chaperones.
Chaperones (in all domains of life) assist in correct folding and refolding of polypeptide sequence
Eukaryal proteins are often modified by the addition of chemical groups
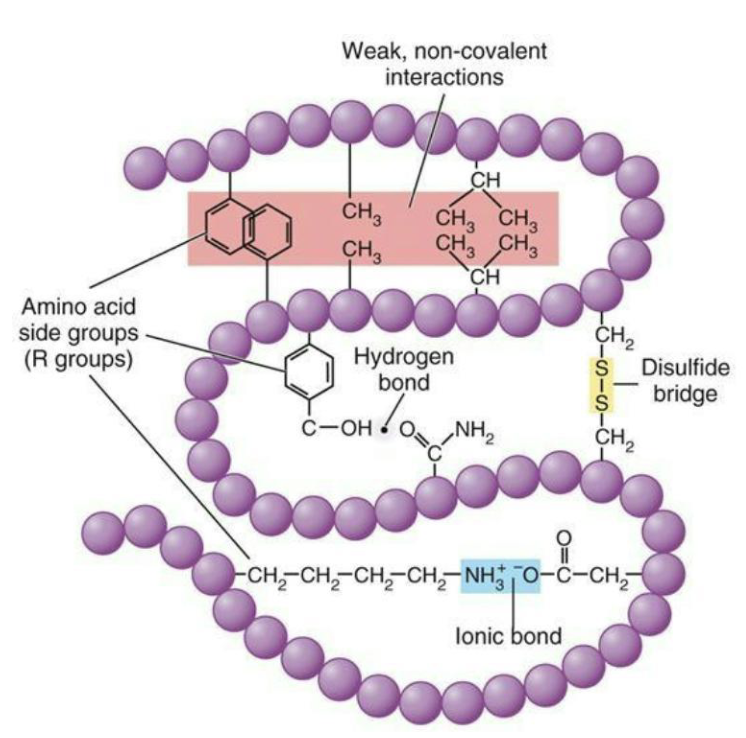
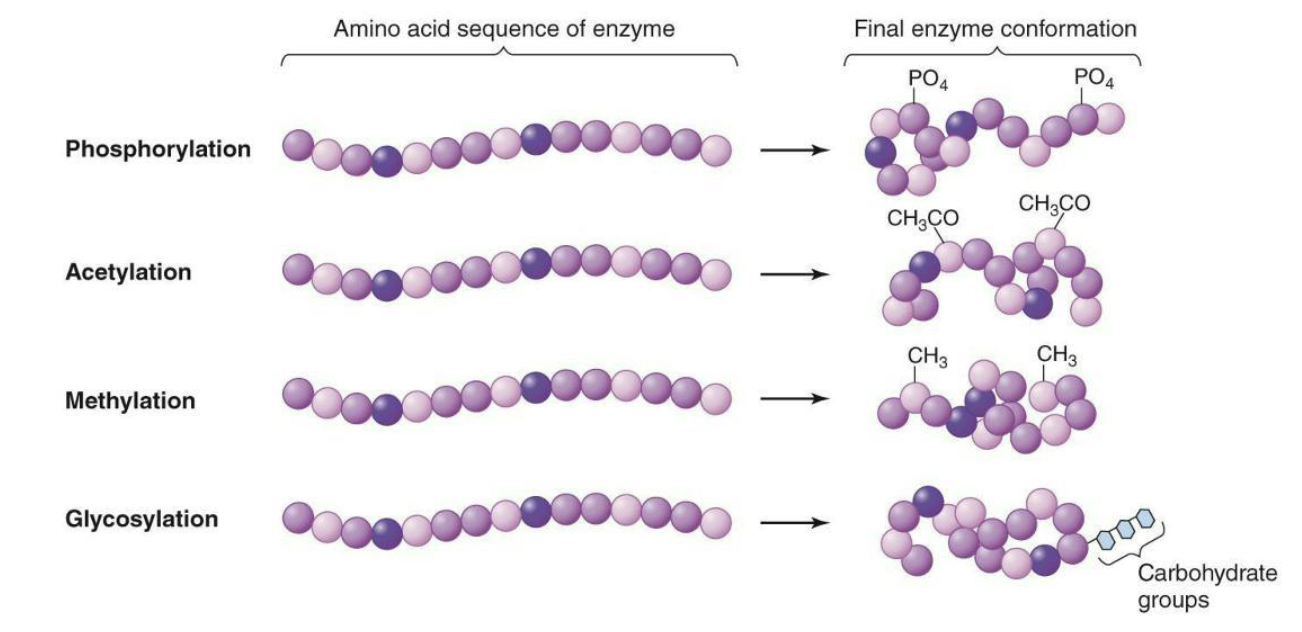
How are proteins modified?
Modification of proteins with different chemical groups or carbohydrates can affect their folding, and serve to activate or inhibit protein function
Phosphorylation
Acetylation
Methylation
Glycosylation
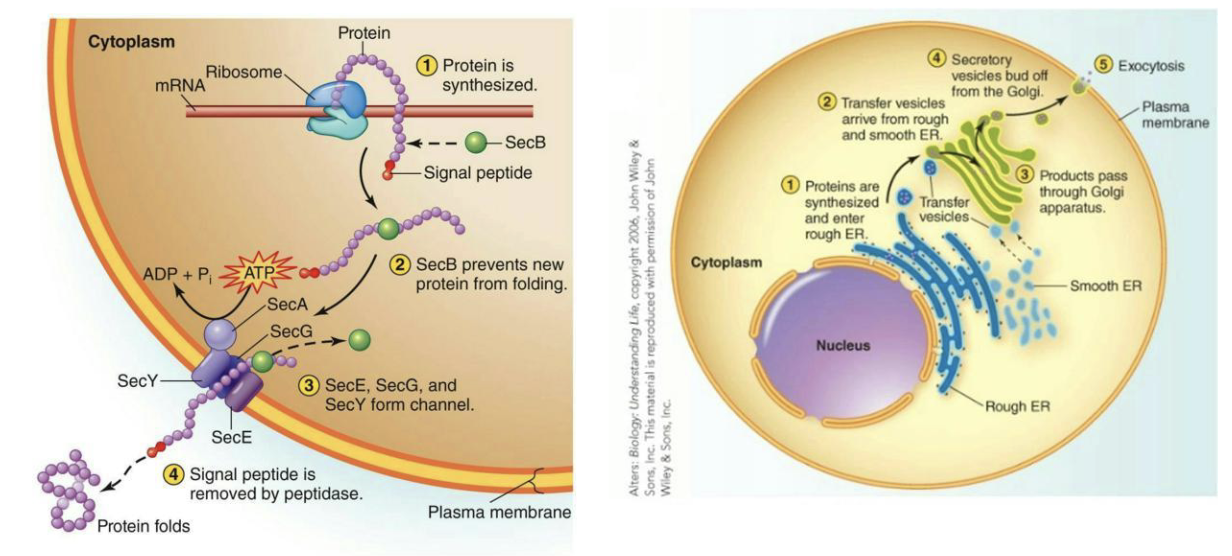
How are proteins transported?
Signal peptides (short AA sequences at N-terminus) direct proteins to appropriate cellular location
Transcription and Translation Summary
RNA (A,U,C,G) made by transcription into mRNA, tRNA, rRNA and regulatory RNA
Sigma factors direct RNA polymerase to promoters in bacteria (other transcription factors used by eukaryotes)
Alternate sigma factors direct transcription of alternate subsets of genes
Eukaryotic mRNA modified by 5’ cap, 3’ polyA tail and introns spliced out
Transcription halted in bacteria by Rho-dependent or Rho-independent termination
Shine-Dalgarno sequence directs ribosomes to start of bacterial genes
Translation carried out by ribosomes that “read” mRNA codons
tRNA with “anticodons” add specific amino acids
Stop codons signal translation to end