The Genetic Code & tRNA
1/15
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
16 Terms
Translation Overview
translation is the mRNA guided synthesis of proteins and performed by ribosomes
translation occurs after the codons (triplet of nucleotides coding for an amino acid) are read in a successive non-overlapping manner by a tRNA
among the 3 possible reading frames, typically only one will be used in order to translate a given mRNA to synthesize a polypeptide
there’s a signal that indicates where to stop and start translation
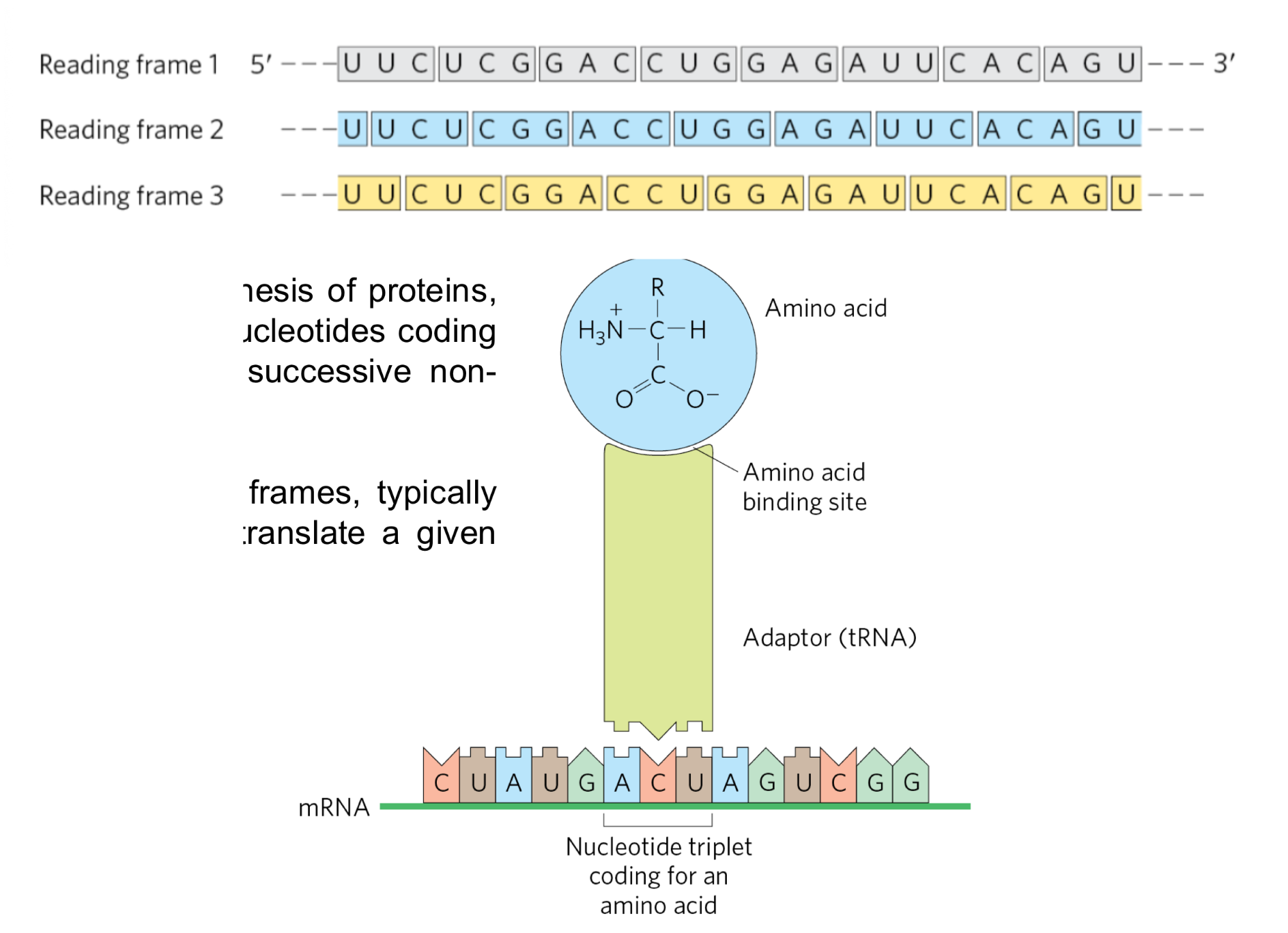
Codon
a triplet of nucleotides on the mRNA coding for an amino acid
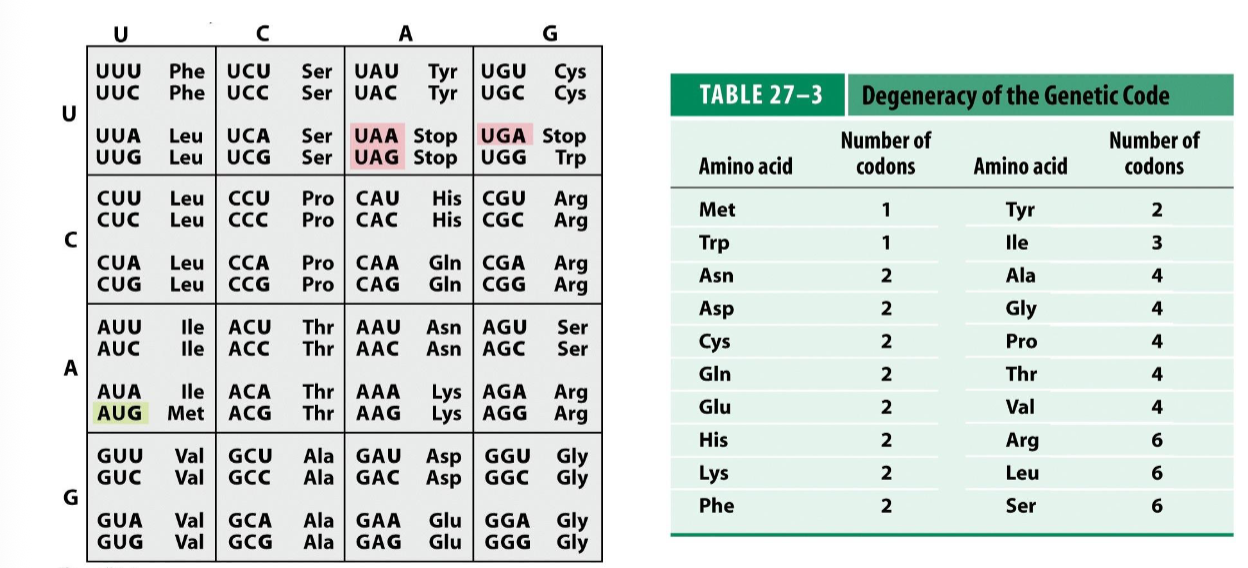
there are 64 codons coding for 20 amino acids and 1 stop signal (proteins are typically composed of 20 a.a’s)
b/c there are more combinations (64) than coded elements (21), the genetic code is considered to be redundant (degenerate)
therefore, several amino acids are encoded by multiple codons
some tRNAs can recognize more than 1 codon
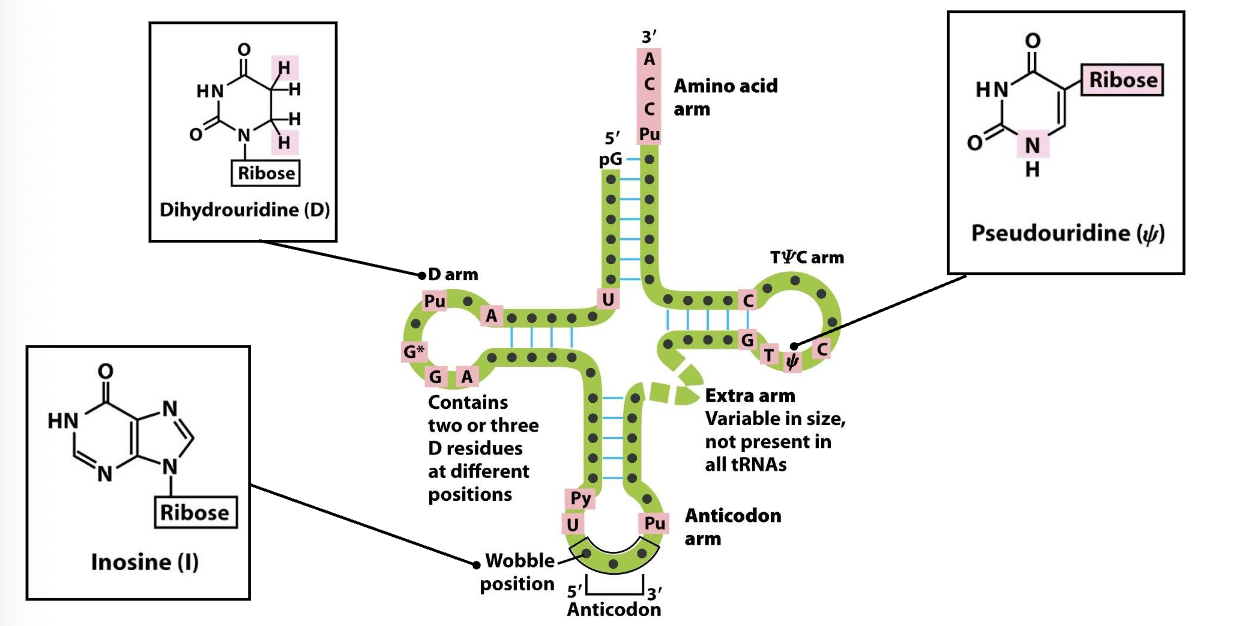
tRNA Structure: 2D
tRNA are small molecules (70-90 nts)
their secondary structure is represented by a cloverleaf
the anticodon arm (bottom) is located opposite to the amino acid arm (top)
there are several modified bases (e.g. inosine) often present in the Wobble position, dihydrouridine in the D arm and pseudouridine
tRNA Structure: 3D
the 3D tertiary structure resembles a twisted L w/ the anti-codon and amino acid positions locate in both extremities
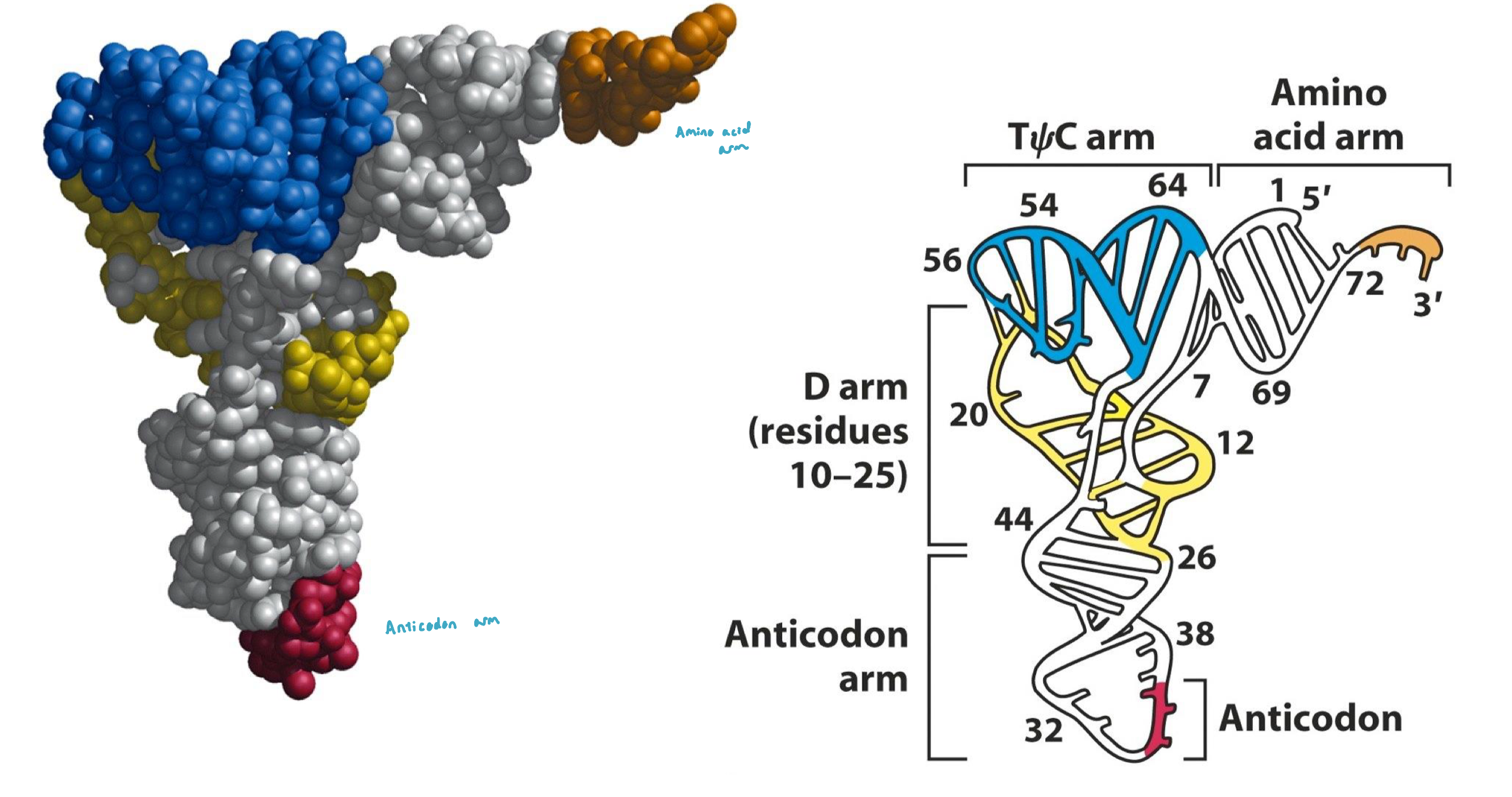
tRNA Structure & Processing
tRNA are transcribed by RNA pol III and are then processed (base modification, cleavage, splicing, 3’ end editing)
all tRNAs have the same CCA sequence at the 3’ end that is added post-transcriptionally
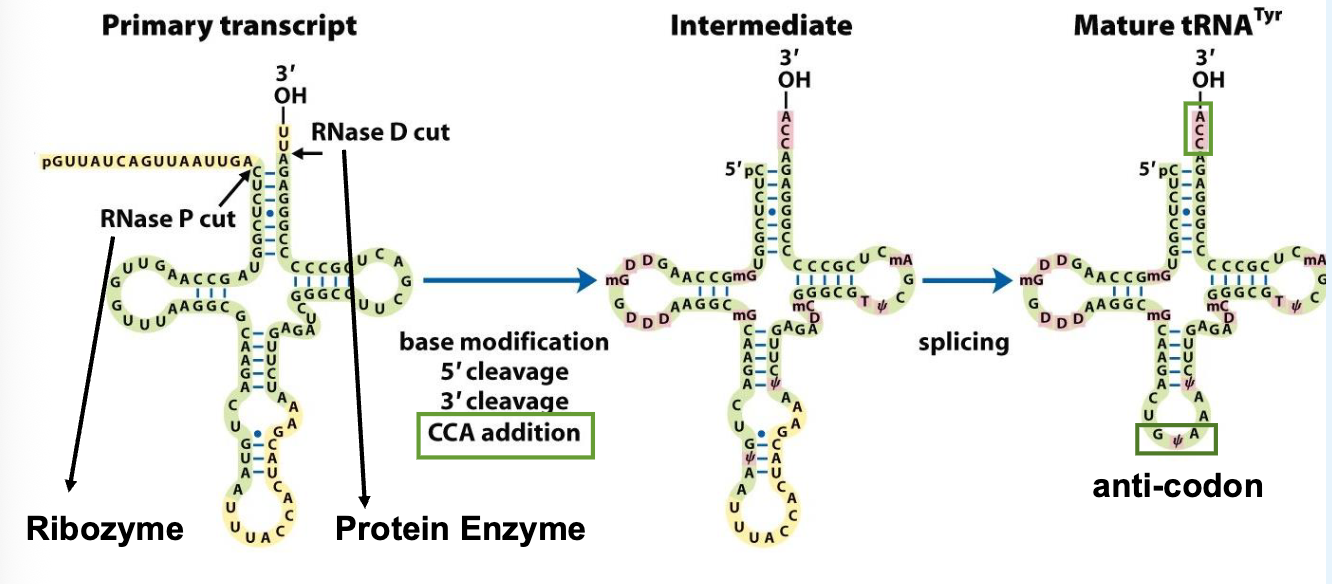
tRNA Processing: Adenosine Deaminase (ADAT2/3)
the tRNA adenosine deaminase (ADAT2/3) modifies adenosine bases on some tRNA anti-codon sequences to create inosine (I)
Inosine can form atypical base pairing with A, C, and U
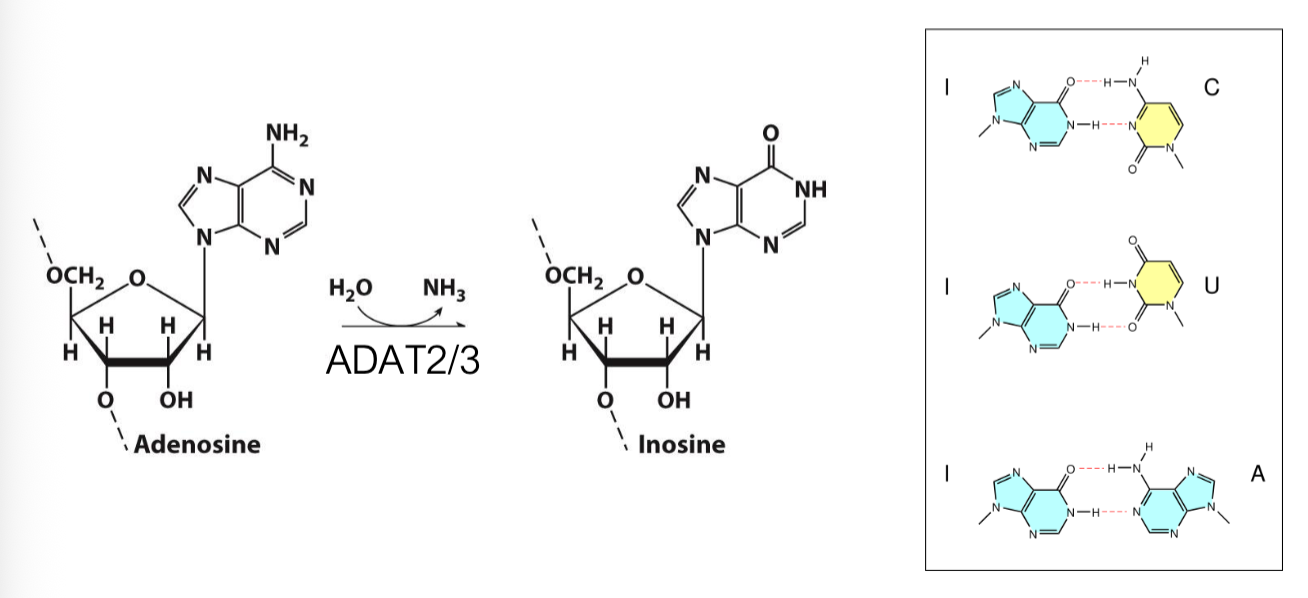
Anti-codon
the anti-codon binds to the codon in an anti-parallel manner
the first nucleotide of the codon binds to the 3rd nucleotide of the anti-codon
the first 2 nucleotides of the codon will form strong W-C base pairing w/ the anti-codon
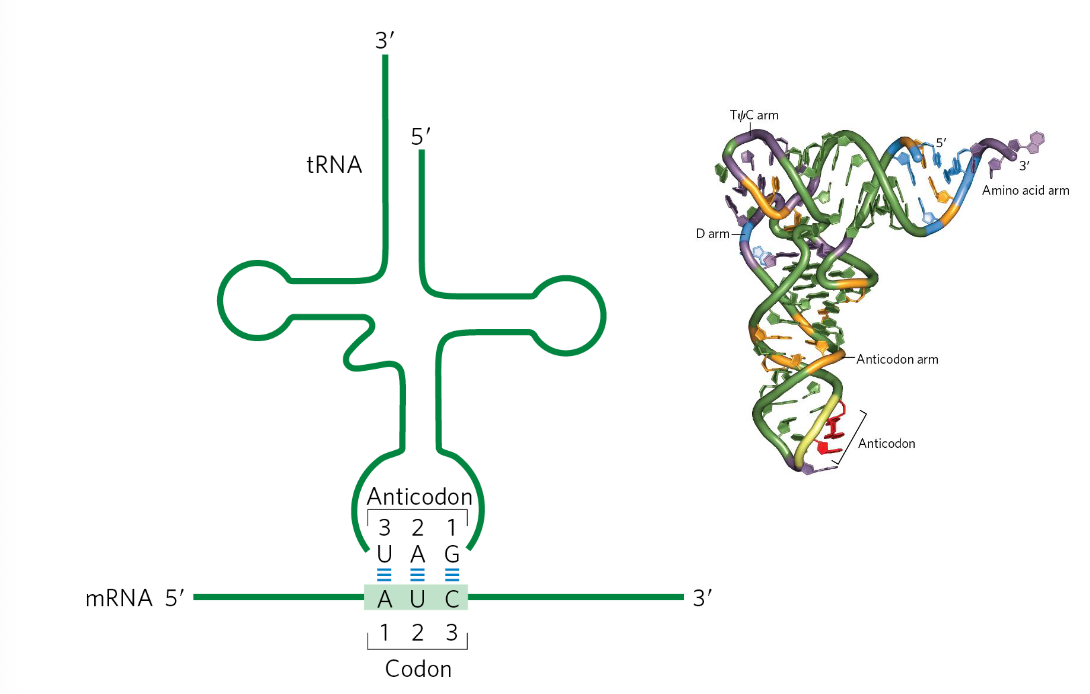
Wobble Base
the first nucleotide of the anti-codon forms a weaker interaction in the Wobble position w/ the third nucleotide of the codon
non-conventional base pairing can occur
e.g. the inosine minor base in the anti-codon can interact w/ A, U, C in the 3rd position of the codon sequence
e.g. U can interact w/ A and G
this allows a more rapid dissociation of tRNA from the mRNA (a higher affinity would significantly reduce the translation rate)
it also reduces the number of required tRNA
while a minimum of 32 tRNAs are required, cells typically express between 40-50 different tRNAs to recognize the 61 diff codons
Wobble Base Figure
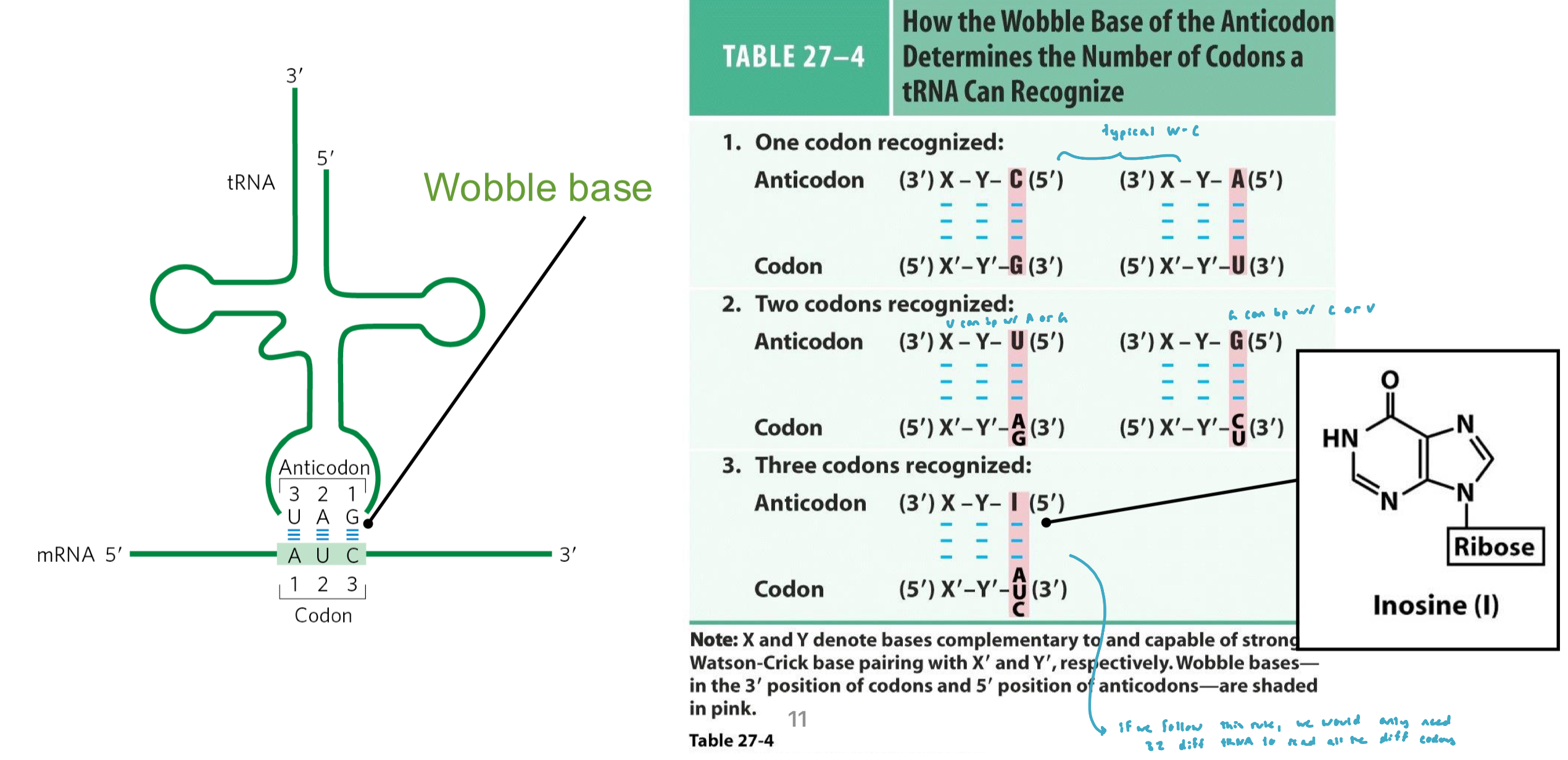
The Codon Flexibility: Genetic Code Variations
whereas the genetic code is considered “universal”, there are exceptions
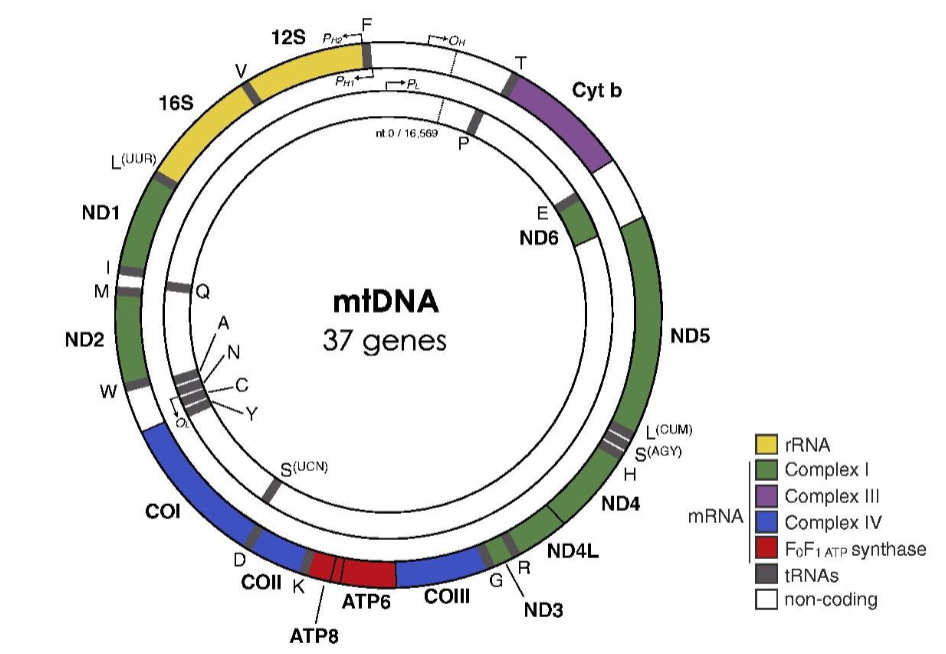
the genetic code is slightly different in mitochondria
mitochondria contains its own genome that encodes a small number of proteins
e.g. the UGA stop codon encodes for a tryptophan (Trp) when translated from the mitochondria
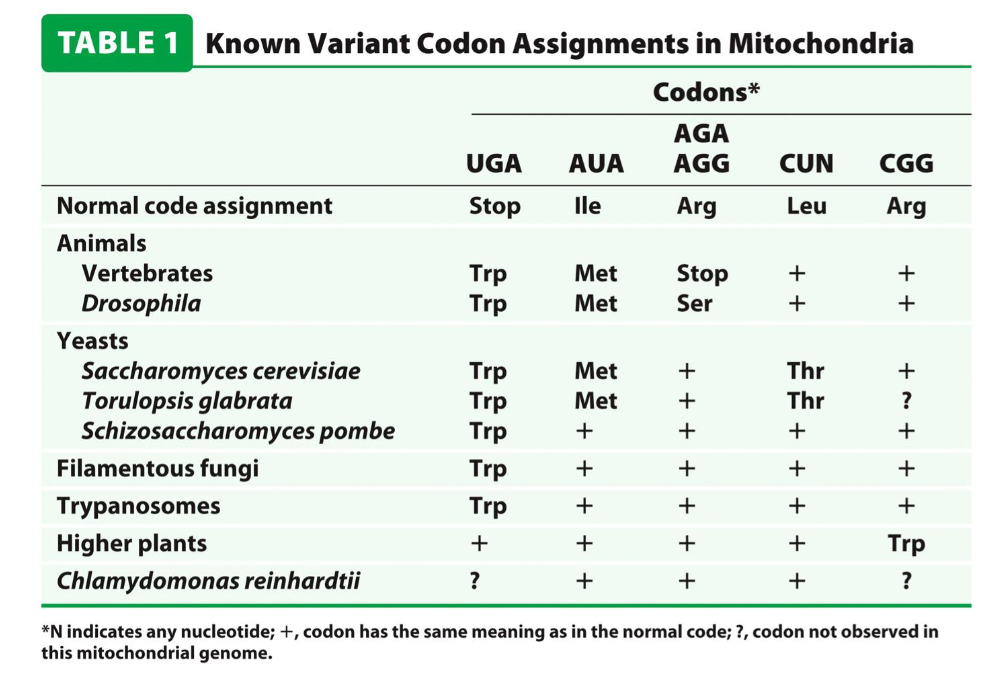
Codon Flexibility: Translational Frameshifts
in most cases, only one reading frame encodes for a protein
there are exceptions where a second reading frame can be used (e.g. viruses)
in this case, the secondary structure of the mRNA can induce a slipping or skipping of the ribosome (ribosome shifts by 1 nucleotide while translating to read a completely diff set of codons → diff protein)
advantage: using one mRNA to encode multiple proteins
Codon Flexibility: Translational Frameshifts Example
in 95% of the cases, the Rous sarcoma Gag protein is normally stopped after the Leu residue
in 5% of cases, the ribosome will slip -1 nucleotide after the UUA codon to place the AUA codon in aminoacyl site and to translate the Gal-Pol fusion protein, that is later cleaved into two polypeptide

Codon Flexibility: mRNA Editing
mRN editing can alter the protein sequence → more diversity in the # of proteins that can be generated by a limited number of genes
editing is mediated by adenosine deaminase that can generate Inosine and cytidine deaminase (C → U), such as APOBEC (apoB mRNA editing catalytic peptide)
APOBEC1 edits a specific codon at position 2153 to convert a Gln (CAA) to a stop codon (UAA)
generates a shorter isoform of the apoB protein
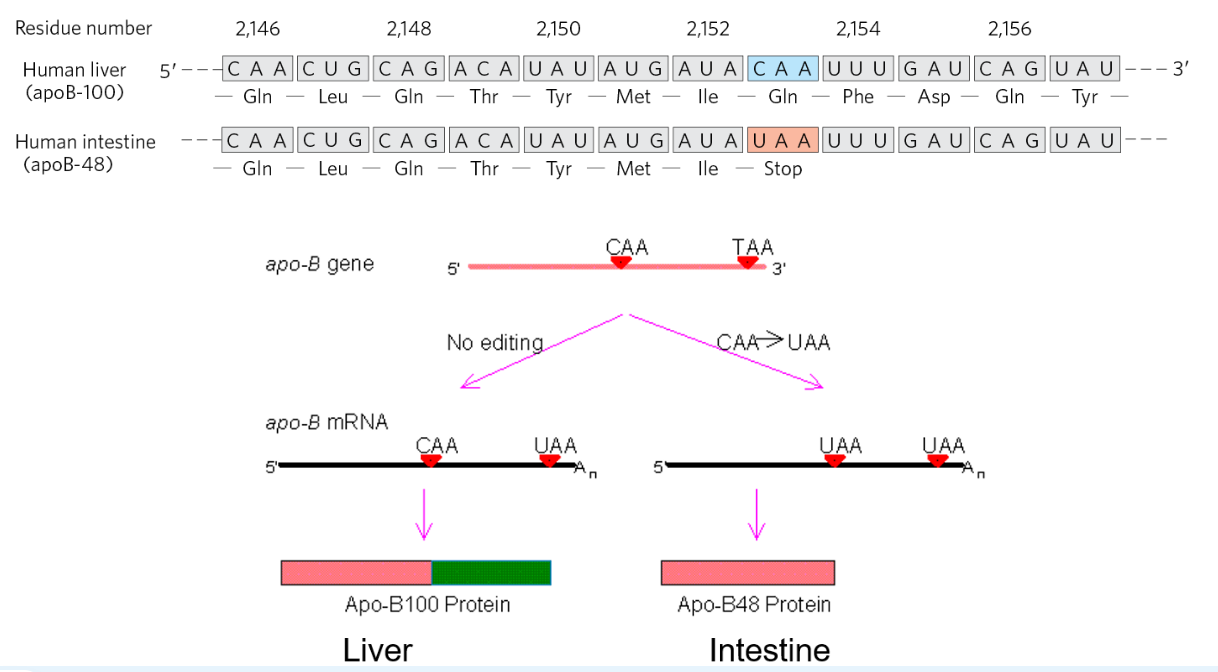
The Codon Usage Bias
while the same codons encode for the same a.a’s, the codon usage is different from species to spcecies
e.g. 22% of Arg codons are AGA in the human genome, while 1% of the Arg codons are AGA in E. coli. Accordingly, there are more tRNA w/ the GCG anti-codon compared to UCU in E. coli
in many fast-dividing organisms, there is a strong correlation b/w codon usage and protein abundance, as translation of an abundant protein will be favored by the higher concentrations of corresponding tRNAs in the cell
there’s a correlation between frequency of a given codon and tRNA abundance.
Genes for which numerous low frequency codons are expressed at lower levels
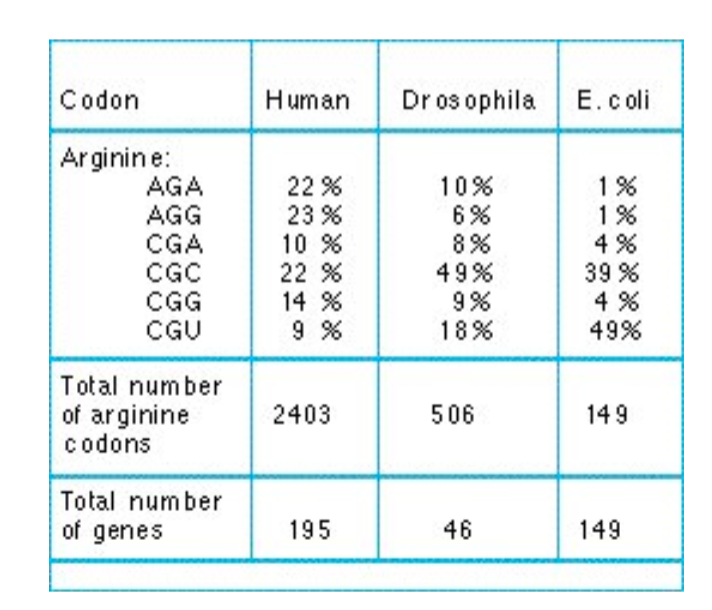
The Codon Adaptation Index (CAI)
CAI is a method for analyzing codon usage bias
CAI measures the deviation of a given protein coding gene sequence w/ respect to the codon usage bias in the genome
CAI can be used to predict expression level of a protein based on its DNA sequence (especially in fast growing organism like the yeast S. cerivisaie)
the codons used in highly abundant proteins are more represented in the genome (~high CAI)
between 0 and 1
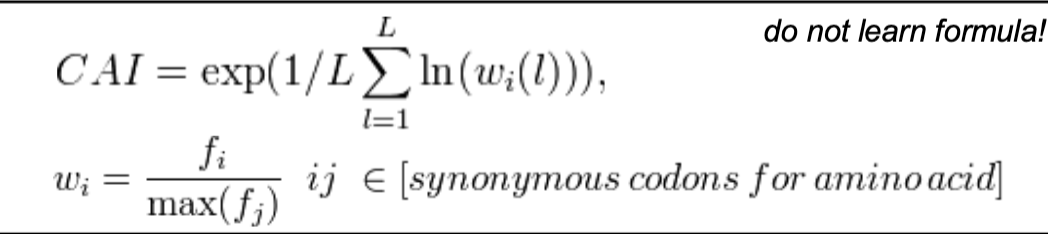
Mitochondria
mitochondrial genome has 22 tRNAs