MCB 3
1/150
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
151 Terms
____ _____ cells are __ ____ than ______(gram negative or positive for options )
is the periplasmic space smaller /narrow in gram positive or negative by abt 20 nm
after the periplasmic space in gram positive cells, we have the ___ ___
gram positive have a ___ layer of peptidoglycan
gram positive, less complex, gram negative
gram positive
cell wall
thick
teichoic acids are present in ___ ____ cells
they interact ____ with _____
they have a ___ ____ ___ which allows for the embedding in the ___ ____
teichoic acids are found in ____ which are found in a. thin layer in the _____
gram posiitve
covalently peptidoglycan.
fatty acid tail, plasma membrane
peptidoglycan, periplasma
which type of gram do you find fewer enzymes in
in the periplasmic space, there are enzymes involved in ___ ____ _____
overlapping layers of peptidoglycan
lipoteichoic acid is good for anchoring ____ to the ____
lipoteichoic acid carries a ____ charge, can act as a ___ ____ to…..
anything up to what kiladoltan can come in
gram positive
cell walll synthesis
peptidoglycan , membrane
negative, can act as a physical barrier preventing things from entering cells
25,000
gram negative
they vary from ___ to ___ nm
____ takes up 20-40 percent of the cell
after the periplasmic space, wa have the ___ ____, which has an inner leaflet and outer leaflet
the inner leaflet has the same ____ in the plasma membrane
the outer membrane is made up of ___
30-70nm
periplasmic space
outer membrane
phospholipid
lipopolysaccharide
there are more enzymes in the periplasmic space for gram ____ than posiitve
enzymes here are involved in ____ ___, ___ ____, enzymes can also hydrolyze them and neutralize them
Braun’s lipoprotein is the most ____ protein in the outer membrane. It’s embedded in the ___ leaflet of the outer membrane and peptidoglycan anchors the outer
what are porins
they allow gram negative bacteria to have more options than gram positivee
negative
nutrient acquisition, toxin modification,
abundant protein, inner leaflet
channel proteins only in gram negative bacteria that allow substances to pass through via facilitated diffusion and active transport across the outer membrane /concentration gradients
3 components of LPS, what are they
explain what each portion that both of them do
lipid A, core polysaccharide, and O antigen
composed of 3 parts: lipsd A-inner most part of LPS, portion that embeds with the inner leaflet lipids
core polysaccharide, smaller part of LPA, connects lipid A to O antigen
the most external section and biggest=O
what is lipid A made up of
what does this help to maintain?
what is the core polysaccharide made up of , around how many, what’s an example of a sugar that would be found here
what Is O antigen made up of, how many, repeating sets of sugars
lipid A: made up of 2 glucose sugar derivatives, 3 fatty acids, phosphate or pyrophosphate
structure and integrity,
sugars, around 10, slightly unusual sugar, might find heptose,
O antigen-made up of sugars, up to 200 sugars, repeating sets of sugars
LPS importances
which part of LPS is responsible for stabilizing the outer membrane structure
____ ___ is sticky and aids in the attachment to surfaces and biofilm formation
LPS contributes to negative charge on cell surface
helps stabilize outer membrane structure, job of lipid A specifically
helps with attachment to surfaces so org can
polysaccharide portion
creates a permeability barrier
prevent host cell immune response
protection from host defenses,
some can modify sugars that are found in O antigen
when host detects gram negative, it amounts an immune response to the O antigen or sugar that it’s seeing
what happens if you can change the O antigen
what is another big importance of the LPS system, due to which part of the LPS
when you treat infection that is gram negative, you don’t want to treat with smth that will cause enzyme to ___ bc lipids will be _____ and acts s an endotoxin
ONLY IN GRAM NEGATIVE
other orgs can’t use it’s immune response correctly allowing the bacteria to escape the immune response
acts as an endotoxin-toxin that is found inside the cell,
lipid A: when lipid A is carried into peptidoglycan, does not trigger septic response, when cell is lysing, it triggers it bc lipid A carries the dangerous lipids
lyse, released
with porin proteins in E.coli for example
they are ____ permeable than plasma membrane due to the presence of porin and transporter proteins
more
do gram posiitve have an outer membrane, if not what is the most external facing layer of the cell
in GP, the peptidoglycan has ___ ____ throughout it’s matrix
gram negative has systems for facilitated diffusion and ____ ____ to move things against concentration
gram neg also havs ___ and ____ ___ in the outer membrane to transfer molecules into the periplasmic space
no, cell wall
large pores
active transport
porin, TonB
__ ___ secretion systems are ____ and specialized type secretion systems and are associated with gram ___ bacteria
they have two domains, the ____ and ___ domain
domain inserts itself into the _ ___, translocates to and then is released
Type V, autotransporters(transport themselves), negative
passenger, barrel domain
barrel domain inserts inteself into outer membrane and then is released
passenger domain is the actual space where the thing that needs to get out sits
for gram positive gram stains, their ___ layer of ___ traps ____t, when adding alcohols you ____ the cells with shrinks the ____ and ___ them shut so crystal violet can’t get out
for gram negative gram stains, have an ___ _____ layer, some crystal violet sticks here, alcoho ____ the _____ out of outer membrane lipids
gram positive-thick, peptidoglycan, violet, dehydrate, pores, seals
outer membrane, crystal violet, pulls, violet
study question, similarities and ddifferences between gram positive and gram negative cell walls
gram posiitve contains ___ ___ in their cell while gram negative does not
____is only present in gram __
____ interacts with plasma membrane so the cell ….
gram ___ has a smaller ___ space
teichoic acids
LPS, negative
techie acid, doesn’t go floating out into space
positive, periplasmic space
blank and blank contribute to the motility of bacteria
what are the three layers thet you might have around a cell wall
flagella, pili
capsule, surface array layer, and slime mold
blank is the most common appendage
do all bacteria have it
they can differentially express their flagella in response to ___
the blank allows it to spin
it’s blank and blank, also a blank
flagella-locomotion
no
enviro
motor
thin and hollow, spiral
note, but don’t memorize: salmonella enteric can only have petrichou’s flagella
spirillum volitions always have polar flagella
salmonella enteric can only have petrichou’s flagella
spirillum volitions always have polar flagella
define
monotrichous, polar flagellum, amphitrichous, lophotrichous, and peritrichous
monotrichous-one flagellium
polar flagellum-F at end of cell
amphitrichous: one F at each end of cell
lopotrichous-cluster of F at one or both ends
peritrichous-come off everywhere on cell surface, surround surface of the cell
what are the three parts of bacterial flagella
blank is the blank and most blank part of flagella
blank is the most blank part, also a blank to the cell envelope
blank protein is a blank coupler, blank sits inside the blank, allows for connection to the bank
from TEM image-3 distinct components
filament, longest, obvious
basal body, internal part, anchor F
hook protein, flexiblee, filament , hook, basal body
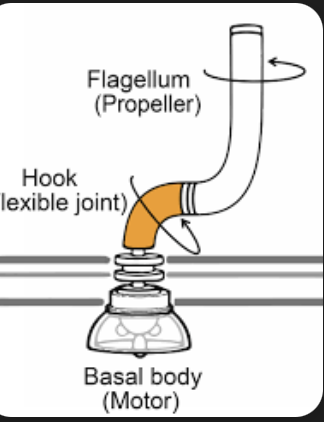
gram negative flagella vs gram positive flagella
WHAT are the 3 rings function
is gram negative more or less complex than gram positive and why
P and N is always more ___ which is important for ___
there are also subunits of…
the basal body is embedded in the ….
how many rings are embedded between diff letter of the cell envelope
where is the P line, MS ring, and C ring found/embedded in
which ones actually spin around to support motion
remember that hook must be wider thane element and remember that the central shaft is hollow too
the rings of the basal body that anchor the flagellum into the cell envelope
negative F-more complex,
bc basal body must span a more complex cell envelope
hollow, construction of filament
protein flagellin
cell envelope
4 rings
P ring is embedded in peptidoglycan, MS ring is embedded in plasma membrane, C ring is found in cytoplasmic side of plasma membrane o.
MS ring and C ring
positive-so
gram positive
basal Is the same function in which two groups
how many rings are in gram positive
where is the inner ring embedded
where is the outer ring associated with
are there more or less layers here than in gram negative
gram positive
gram P and N
only 2 rings here
plasma membrane
peptidoglycan layer
less layers so less rings
gram positive
how many genes does it take here to build a flagellum
operons are used by which two domains, what does it allow us to do
filament extends to outer entire by using ___ ___. Basal body is modified __ ___ ___ ____.\
making flagellum: we shove them up hollow ___ ____ and they pop out at the top
do flagella grow at the tips or the base
once flagella subunits reach the itpe, they do __ ___, meaning they don’t need external organizing force. Interactions between flagellar subunits allow them to make up the configuration
20-30 diff genes to do this
genes are encoded in operons
bacteria and archaea, allows us to transcribe genes as a single RNA molecule/put multiple genes together at the same time to make the same structure
filament extends to outer enviro by using basal body. Basal body is modified type 3 secretion system.\
making flagellum, we shut them up hollow basal body and they pop out at the top
tips
undergo self assembly-don’t need external organizing force, interactions between flaglellan subunits that allow them to make up the configuration
flagella moves in bacteria by …..
how fast depends on bacteria and ….
E. COLI does 1100 revolutions per sec
counter-clockwise and clockwise rotation is what kind of movement(both each
what direction can most bacteria not do
which kind of bacteria can move backwards
spinning like a propellor
which way your’re spinning
counter-clockwise rotation =directed run=forward -
clockwise rotation=tumble
bacteria can’t do backward movement
Archaea can do backwards though
with tumble remember that something happens and disrupts run causing cell to stop and tumble
chemotaxis
what 2 things do you need to do this
what do chemo repellents bind to
what do chemoreceptors do, what can influence them
directed movement towards or away from chemical attractant
NEED to have a flagella to do this, must also have chemoreceptors-inthe plasma membrane,
chemo repellents bind to chemoreceptors
chemoreceptors sense presence of things, diff in concentration that have influence too
define positive and negative chemotaxis
positive chemotaxis-towards a chemo attractant
negative chemotaxis-running away from a chemo repellent
is chemotaxis a flagellar movement
when chemoattractants are high and chemorepellants are llow what do bacteria do
what does a zone of clearing around a disk mean
no, regulation
high chemoattractants=bacteria move towards it
high chemorepellants mean bacteria moves away from it
bacteria has run away from the thing, chemorepellant concentration is high
high concentration of attractant means
decreasing concentration of attractant means
high concentration of repellant means
decresasing conentration of repellant means
high arts of runs
increased tumbles(reorienting)
more tumbles (need to reorient away)
more running (more running moving away from it successfully keep going
bacteria have blank movement when there isn’t a conc of anything
high conc of repellent means more run and fewer blank, and once you get away you can blank more and less blank
random
chemotaxis system is made from __
what specific thing switches you from running and tumble
what determines whether or not the bacteria runs or tumble
when CheY-P is phosphorylated what happens
what happens when CheY is unphosphorylated
explain the steps of the signaling chain
what happens if CheZ removes the
what happens when there is a high concentration of attractant
what happens when conc of attraction drops,
proteins
molecule switch does this
the phosphorylation state of CheY
it interacts with the motor switch, causing a clockwise rotation/tumble
CheY-P has no interaction with switch and a counter clockwise rotation occurs meaning run
chemoreceptor detects a chemical signal in the environment
receptor transmits that signal to CHeW
CheW passes it to CheA
CheA phosphorylates CheY, then it becomes CheY-P
CheY-P travels to the motor switch causing a clockwise rotation/tumble
cheZ removes the phosphate from CheY-P resetting it back to CHeY and the run resumes
CheA activity is suppressed, less CheYP means that the motor stays CCW so bacteria keeps running toward the attractant
cheap becomes active, phosphorylates CheY, CheY-P hits the switch and tumbles to reorient
flagella is 2 part motor, allows us to produce torque-torque allows us to spin smth
roto part-composed of C ring or MS in gram neg, and inner and outer in gram positive
C and MS ring turn and stator part-S part is stationary, made from Mot A and Mot B proteins get energy from potential energy stored in proton gradients
motA and motB creates channels so ions can flow down concentration gradient to allow energy for torque
located n basal body
MOT proteins form channel so protons can flow down concentration gradient
gives energy to spin MS ring and C ring
allow spinning of shaft up until filament
FIM and FIN make part of C ring
FLig-believe motB
MOTS not embedded in ring, kind of in the outside
external flagella can def move
other mechanisms of movement though, some that are not external
spirochete-notoruiosu fo rhaing internal flagella
don’t see them externally bc flagella wraps around periplasmic space or organism
when it does movement-it does corkscrew like motions
non flagellar motility
gilding motility-few diff types
performed by orgs that make a slime layer-layer of polysaccharide. orgs producee smooth polysaccharide and then org glides over it
twitching motility-for those that have pili. just bc you have pili does not mean that you can twitch
good for adherence , secondary function that some bacteria have evolve from there I pili
bacteria can extend their pili and those pili stick to surface and retrace their pili. hold them selves along
polymerization of actin
bacteria that will infect eukaryotic cells, like shigella. it will be endocytosedinto host cell, escaped from the endocystic vesicle, it will use the hosts actin to mvke actin tails, uses actin tails to push between eukaryotic cells, don’t have to exit the cell, you can spread from cell to cell
coopting host cellular machinery, and allowing bacteria to spread inside cells
priamary function of pilis is adherence
made out of protein subunit called pillin
have a protein called ashesi-portion of pili that is sticky
fimbriae-structuraly both are the same thing, it is also made of pilling, have same thing at it’s tips,
bacterial cell surface
stalks-made by certain subset of bacteria, low nutrient enviro bacteria
stalk is an extension of cell membrane
tip of stalk is holdfast. fast is composed of polysaccharides,s sticky, allows for sticking to other cells
find steaks in orgs that liven lownutent enviro bc it gives more expansive surface area fr new nutrients
3 layers
capsule, slime layer, or surface array layer or S layer
they’re glycocafyx-layers of polysaccharide that these types of cell secrete, if you secrete capsulee or slime layer
S-layers aren’y glycocalyx bca ren’t yamde up of polysaccharide, made up of proteins and glycoproteins
capsules and slime layers do
both-aid in attachment to solid surfaces, polysaccharide, think sticky
to other cells, help in biofilm formation,
capsules continued
composed of polysaccharides-thick and tightly organized, well organized, adheres very tightly to cell surfaces, aren’t physically removed easily from cells
can see them insight microscope
capsules help protect the cell from essication, bc they have a high water conc
help hide the bacterial cells out in the host cell immune response
ex: smooth strain pathogenic-has a capsule so host immune cell can’t see it, so it won’t have an immune response
, rough strain non pathogenic-don’t contain a capsule and host cells can see them and mount a response on them before they have a chance to do anything
biofilms have glycocalyx later, slime mold or capsule
other cells can stick to other bacteria
form this big 3d community
it’s protected for Theo orgs
single cells can be easily killed, harder to kill the orgs in the inner portion
plaque=a bifilm that hangs out in the tooth
biofilm that contains archaea and bacteria
slime layer
it is much thinner than capsule
it doesn’t adhere as tightly
can’t see a slime layer under a light microscope, yes with electron though
they can protect the cell from desiccation due to high water content, aid in attaching
can protect the cell somewhat form host cell immune response, not as good though
great for gliding motility, capsule cannot do this
capsule-is soft and gel like
slime layers-in gram neg and pos orgs
if you have a capsule you don’t have a slime layer or s layer
if you have slime layer or s layer you don’t have capsule
CHECK THIS
s layers
in gram pos and gram neg
gram neg-S layer sticks to other membrane
gram pro-associated with peptidoglycan surface
S layer functions
acts as a physical layer, makes later highly impenetrable
protect it from ion and pH fluctuations, osmotic stresss, enzymes and predation
-shape and rigidity, highly structured, not softt or flexible
-can protect from host defenses but not as great as capsule or slight-does it by blocking antimicrobial comps from being able to make it’s way into the cell
-potential uses in nanotechnology
come together via self assembly
helps to prevent with damage
bacterial endospore
dormant structure formed by some bacteria
not metabolically active
very resistnat
desiccation, heat, chemicals, radiation, antibiotics
hard to get things through an endospore
some endospores are some of the most nasty toxins,
majority are soil dwelling, bc soil is a dynamic enviro, nutrients washed out and washed into it
spore is made and then released into the enviro by bacteria
spores are made in the mother cell
central=bacillus subtillius
bacillus anthracis=subterminal
terminal=all the way at the end
clostridium tetani do swollen sporangium
where spores sporm are characteristic of the org that forms
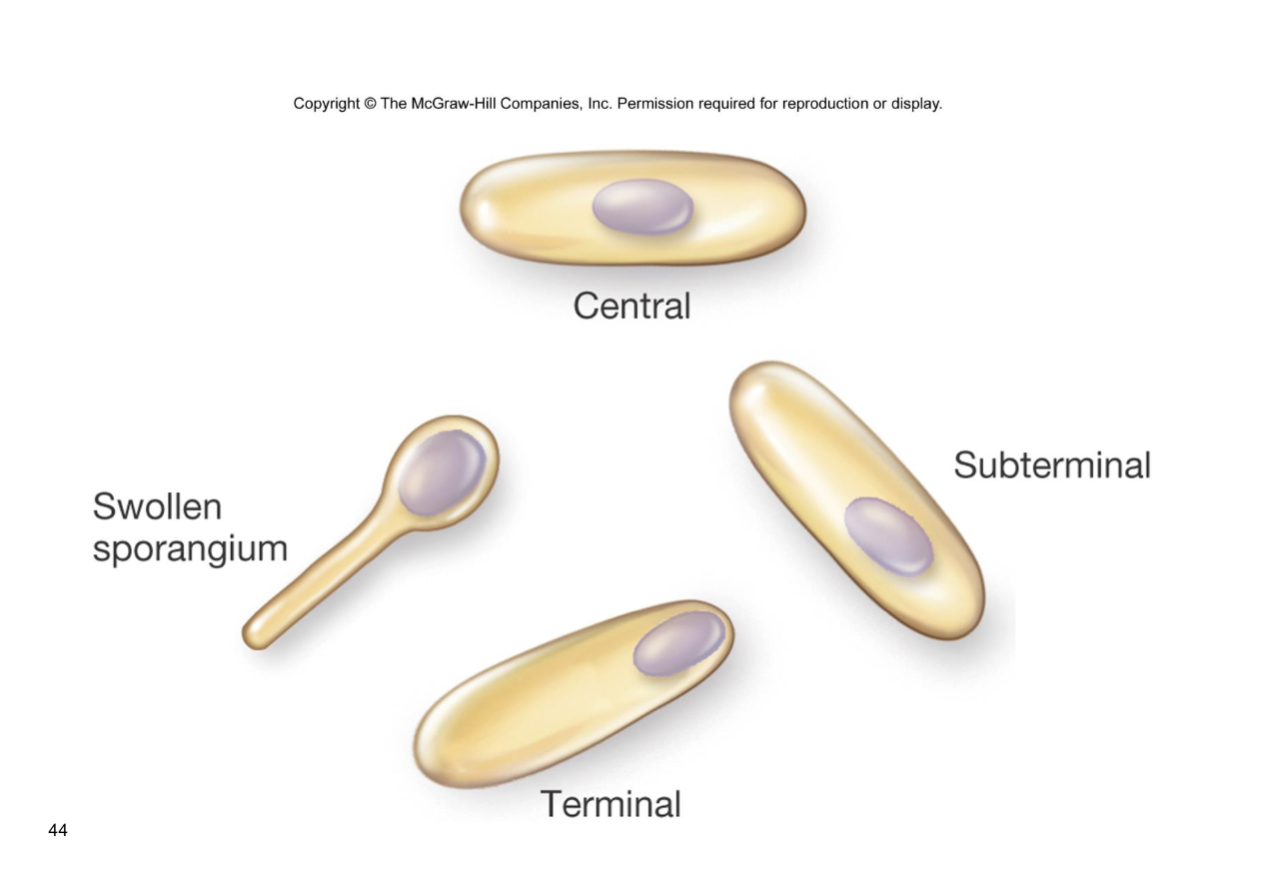
core is where DNA is found, doe snot have water, has calcium dipicenoic acid, CaDPa, double membrane structure inner and outer membrane
inner membrane of spore =highly impermeable, proteins are tightly crossed here, phospholipid bilayer, germinate receptors are located here
core wall/cell wall
then cortex-can occupy half of cell volume, has ppetidoglycan, they are not fully crossliknked though it’s kind of weekend, difference in cross linking contributes to heat resistant properties
then outer membrane-slightly more impermeable than inner membrane
then we have inner coat and outer coat-made out of 70 diff inner proteins, tightly cross linked, some orgs have. a layer made up of glycoproteins-epsosporeum, bacillus and anthracite have this
outer membrane act as physical Barries due to cross linked barriers
why they’re so resistant
have a low metabolic activity
during sporulation calcium DPa moves into core, displaces water so water moves out, causes enzymes not to function,
they are resistant to ___ due to The actions of SASPS, small acid soluble DNA, they physically bind to DNA, proteins block DNA forms being damaged by the UV
dehydrated core
specialized cross inking of cortedxt which contributes to heat reisstnace
spore coats and exosporium=just physically barriers to prevent things form moving across it
how to make a spore-sporulation
triggered by nutrient deprevaition, doesn’t have orgs to. eat can either dye or hang around and make spores, that’s what they do
long process, can take 10 hours, all or nothing, can’t stop in the middle or go back, it’s tightly controlled
controlled by master orgs Spore 0a, once it’s phosphorylated it starts the response and edouens’t stop
lack of nutrients trigger sporulation.
500 proteins that are involved
megatherium is the largest of bacillus species
cell division-cell divides in step 1-also have axial filament formation-DNA stretches out the length of the cell,
step 2: septum formation, forming a wall int he cell, turning it into 2 separate compartment, 1 compartment will be smaller, another will be larger
spore develoeps n smaller comportment
step 3: mother cell membrane comes down and circulates around, engulfing of spore by membrane of mother cell, smth of DNA to spore foremother cell
step 4: core is formed in the middle, membrane is 4, laying down our cortex here
step 5: synthesis of coat synthesis inner and outer
step 6: complete spore coat formation nd exosporium formation
we''ve made out spore
step 7: lysis of the mother spore, spore produces enzymes that lyse the mother cell so that it can be released into the enviro
germination
happens when we hav enough nutrients in enviro
we unwind everything that we did in sporulation
complex process, all or nothing response, multi stage process as well, not as many as sporulation,
last stage is outgrowth
fully developed vegetative org is cracking spore so bacteria can come out
activation -process where you wake yourself up, not germinating here, allows cell to response appropriately to nutrients when nutrients are present in the enviro
ex: temp rises, triggering activation, if nutrients come into our enviro we can do germination:
nutrients bind to germination receptors-inner membrane which triggers germination, opposite steps of sporulation. water rushes into the cell, calcium DPa rush out, metabolic activity increases, activation of diff enzymes, which degrade the coats and exosporium, water rushing in breaks peptidoglycan
germination
outgrowth-acitv enzymes degrade the spore codes allowing bacteria to be released into the enviro
species: groups of strains sharing common features, while differing considerably from other strains
domain is broadest, work our way down to genus species, which is more narrow
gram positive is more related to itself than gram neg
if you’re likely to form nasty groups colonies, more to look like one that forms that too
spore forming orgs are more related then non spore forming
similar metabolic pathways
culture collections to store diff organisms
store it in -80 for sleep
ppl like to store strains so everyone can u
archaea have both eukaryotic and bacterial factors
they liv win some of the most inhospitable places on earth-what they’re characterized as
e
methanogens-first area-can produce methane in a way that is unique to Archeaa
halo bacterium salinarium-it is an archaea, it likes salt-between 3-5 molar
prococcus furiosus-likes intense heat, hydrothermal vents
picrophilus oshimae-bitter loving-loves sulfur ph of .7
methanogenium frigidum-loves chilly enviro
A things similar in bacteria
.5-5 micrometers in diameter
similar shapes, cocci, rod shapes, spirals, also some that are unique to them
singular, circular chromosome, rods, spheres too
lacks a true membrane bound nucleus-DNA is in area but not constrained by membrane
eukaryotic structures
how DNA is organized-DNA is complexed with histones, wrapping DNA around them
transcription and translation machinery is highly conserved, homologues of DNA replication enzymes
their plasma membrane-organization-HAVE monolayers, not in eukaryotes or bacteria
cell wall-composition
archaea has irregular shapes
they have rectangular shapes like in thermoproteus spp
squared too-not present in bacteria
cell envelope-plasma membrane and everything outside of that
has 6 organizations of envelope-start with plasma membrane but diff organizations
majority have a cell wall
temp determines what the bilayer or monolayer will look like
hydrocarbon tails don’t have fatty acids, they use isoprene unit-five carbon molecule with short branched chains-makes the tail portion of lipids in plasma membrane
-iso unit links to glycerol-we’ll see Ether linkages in archaea-less impacted by temp changes, more stable at higher or colder temps in comparison to ester in bacteria and eukarya
bilayer for archaea-use isoprene unit for 20 carbon molecules called phytanyl, linked to. glycerol via Esther linkage, connected via 1 carbon
phytanyl groups interact with each other
monolayer -2 hydrophilic head groups, isoprene units connected from one to the other
more likely to find monolayers for orgs that like high temps-ex those that can survive autoclave
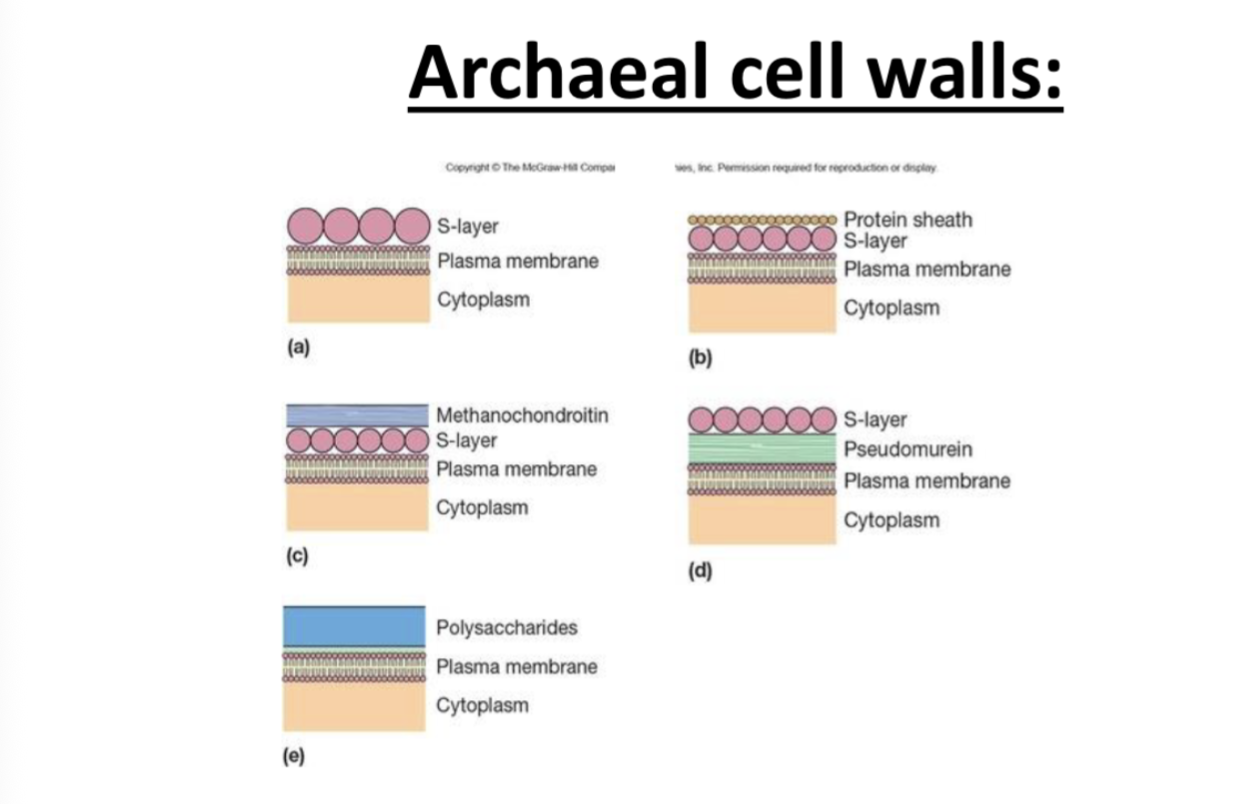
Rachael cell wall
5 distinct configurations
most have s layer, it’s part of the cell wall
configuration e does not have an s layer, but others do
if you have f configuration for Racheal cell envelope you ve two lipid layers so no celll wall
archael cell walls provide the physical barrier, preventing things from making way into archaean cell walls, help us maintain our cellular shape
help in osmotic protection
configuration e-have pseudo marry, look like murine but is not murine. lots of similarity in the structure of both and differences
NAT in pseudomary and NAM in peptidoglycan
archaea have beta 1-3 linkage and pseudomurien have beta 1-3
penta peptide that gets cut off in both archaea and bacteria
L and D form in bacteria no D form in archaea
lysosome won’t work against archaea bc it’s beta 1-3 not beta 1-4
beta lactic work yes or no
archaea have plasmids and inclusions
dna in cytoplasm nuclear region
how DNA is packaged is diff in archaea bc it gets wrapped around
DNA is neg charged and histones are pos charge, wrapping prevents it being exposed to nucleases hanging out in enviro
diff in archaea nucleosome, it’s a 6 length pair not 8 bc genome in archaea is smaller than eukaryotes
cytoskeletal for cell division, cell wall support for shape, organizing internal space of cell
homologs of bacteria and some homologs to eukaryotic , depends where you are in evolutionary timeline, further you are the more closer they are to eukaryotes, earlier on you’ll find cytoskeletal elements somewhere in the middle you have some that have cytoskeletal elements of both bacterial and eukaryotic, play the same role though
slime layers and capsule are rare in bacteria, mostly just s layer in archaea
unique: have cannulae-hollow glycoprotein tubes that link cells together to form a complex network, allows them to communicate maybe, sharing of nutrients, surface structures that we only find in archaea
another surface structure-hami or hamus-filamentous structures coming off surface of the cell they aid in archael attachment, grappling hook configuration that helps aid in attachment
great for biofilms bc you have to have cells that attach to each other
helps preventing archaea from being removed from enviro
flagella in Arachne or archella
appendage of locomotion
rotate like propellar
differences-they are thinner, not hollow like bacterial ones, -that’s what causes differences
they are built at the base here
2 or more diff versions of flagella protein, only 1 in bacteria
movement is diff
archaea have forward and backward not spin or tumble like bacteria
bacteria gets energy from ion gradients to move rotors
archaea hydrolyzes ATP and uses that to spin
eukarya have flagella but they don’t spin around,
metabolism is the sum of all the chemical reactions in an organism (catabolic and anabolic)
catabolism is the energy releasing processes -gives fuel for rxns
anabolism is the energy using processes
enzymes mediate each step in reactions
metabolic p
laws of thermodynamics
1-energy can’t be created or destroyed, only converted from one to another
2-physical and chem processes happen, and randomness occurs-universe tends to go to disorder, higher to lower energy state causing disorder
entropy-measure of randomness or disorder in system
free energy
free energy change =delta G
exothermic=neg delta G, spontaneous -don’t add energy in system for it to happen
pos delta G-endergonic, non spontaneous, need to add energy to system to make it start
transition state-energy peak of reaction, most unstable point of reaction, right before we have bonds break
activation energy-the amt of energy it takes to hit the transition state
ushe absorbs energy from enviro to hit this peak
the more activation energy you need, the slower ithe reaction will run
not every reaction need san enzyme to run
enzymes speed up rate of a rxn,. some actions w/o enzymes are so slow it wouldn’t even be useful,
enzymes lower the Ea to make it quicker
non spontaneous reaction-adding an enzyme won’t make it spontaneous
enzymes don’t change the freee energy that is available in system
free energy is determined by free energy available in reaction and product
converting reactant to product MUST:
enzyme and substrate must encounter each other in prop orientation, enough force, enough energy, when adding enzyme in you take the chance that this will all happen out of the equation
enzymes have an affinity for the substrate , increase the concentration of substrates ou are promoting the likelihood that they will enocounter each other, enzymes grab the substrate, orient them properly, and bring them together with enough force to do bond breaking
take chance out of equation
one enzyme have one specific substrate
confirmation of active site, structure component of substrate confirms this
lock and key model-now we use induced fit model, better and uses things form lock and key
lock and key model-have to have substrate with specific confirmation to fit,
diff-lock and key is rigid model, doesn’t allow for any flexibility in structure of enzyme itself
but there IS some flexibility
induced fit model-there is still specificity, specific confirmation of active site,o only specific substrate will fit, when substrate bind to active site, the enzyme active site binds to it and changes the final confirmation of it a bit
enzymes are catalysts, don’t get modified or used int he process
they have a turnover number that is associated with it
can be anywhere from 1-10,000 mols
hturnover number=how fast can enzyme grab a substrate and convert it to product
active site=where substrate bonds to on enzyme , is very specific to substrate
substrate is unique to active site too, they fit together great
some enzymes are only made up of polypeptides-only protein component
some enzymes need non proteins components, it’s gnats be essential for it’s function, if you do not have it , it won’t work
cofactors=non protein components
coenzyme MUSt bind to enzyme to create active site that has a structure that is complimentary to substrate
coenzyme-organic, more loosely attached to enzyme
prosthetic groups=inorganic, ex zinc or magnesium, ore firmly attached ot enzyme
important coenzymes
NAD+
NADP+
FAD
coenzyme A
coenzymes again
they give us active site
coenzymes acts as carriers
it pops off a functional group, and it shuffles over functional group to substrate to, modifies substrate and modifies it so it can be released into the enviro
two factors effecting enzyme function
temperature
pH
when enzyme denatures ad unfolds, substrate can’t find an active site
all enzymes have a. temp range that they are optimal in, you can get too high or too low
too high or too low will cause enzyme function loss
too high-you’re denaturing,
too low-you’re loosing function for a diff reason, bc enzyme and substrate need to encounter each other and when there’s barely heat it sows down rate of motion decreasing the chance for them toe nocunter each other, also won’t have energy we need to reach transition state
ph-you’r loosing function too high vs too low the same way, it’s not like temp ewer it’s diff you’re loosing structure both ways
concentration of substrate-the more substrate available, the faster you’re products gnats form, increase product formation until hint v max
v max is where you plate bc enzyme is saturated, won’t matter if you grab more enzyme is grabbing as much as they can
affinity-being able to grab and pull the substrates to them
km-substrate concentration required by the enzyme to operate at half of it’s maximum velocity
low km means high affinity for substrate, need ton of substrate around , has low affinity for substrate
enzyme that has high km needs not art of substrate b it has a high affinity for substrate
competitive inhibitors-will compete directly at the active site for binding with he substrate, look a lot like the natural substrate , it binds to active site
noncompetitive inhibitor-don’t compete at active site of enzyme, bind at a diff site somewhere on enzyme that’s not active site, causes a conformational change that changes site of the active site, natural substrate won’t be able to fit into site
affinity for substrate vs how much affinity for competitive einhibitor
concentration-overcome it by dumping more natural substrate into enviro, increases likelihood that enzyme will encounter the one with higher concentration
sulfahydomite-bacteria static-prevent cell form replicating DNA,
folate is used to make DNA -improtant to be able to make this
PABA-binds to dihydroxate synthase,
enzyme promotes change of PABA to product to be used to make folate
Sulfa drug looks like PABA and goes there to bind
enzyme makes a product, but product can’t be used by the cell to make folate, isn’t the correct product
this targets folate synthesis pathway
we get folate from food, we don’t have to worry about disrupting our folate pathway
ways to regulate
metabolic channeling focused on where intermediates and end products are located into cell, moving them from one pallete to the other , ore important tin eukaryotic bc don’t have tons of compartment in prokaryotic, gram neg periplasmic to eukaryotic
regulate synthesis of enzyme-don’t make an enzyme, can’t use it to make product,
direct stim ulation or inhibition enzyme- do you want to turn enzyme on or off,
metabolic channeling
regulating by moving things around to diff places in the cell
compartmentation
used more in eukaryotic cells bc they’re more complex, higher level of compartmentation
bacteria still use it, bu not as frequently, ushe gram negatives
you can have diff distribution of enzymes and metabolites-creates variation of enzyme concentrations and metabolite concentrations within diff compartments
channeling allows us to operate similar pathways simultaneously and independent of each other
allows us to coordinate diff activities via transport of themselves
post translational regulation of enzymes activity
turn the enzyme on or off
2 ways-
allosteric regulation or covalent modification, allows us to turn enzyme on
allosteric regulation
you havve allosteric effector
binds to enzyme on allosteric site -NONcovalently, reversible binding, can come off
when it binds there it changes shape of enzyme
change reaches all the way to the active site
positive efector increases enzyme activity while negative effector inhibits the enzyme
it is VERy specific bindingng to a SPECIFIC site, it’s purposefully, , we can not only inhibit enzyme activity, we can turn on enzyme activity, diff form noncompetitive inhibition-smth form enviro is effecting enzyme, only turning enzyme off non specifically activating enzyme ,