Microbial Processes and Metabolism
1/15
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
16 Terms

Bacterial GROWTH and METABOLISM
bacterial growth
Bacterial Growth
Bacteria grow as colonies on solid media
A colony is a clone of bacteria (clone: every individual cell growing in the colony is identical)
A population of bacterial cells is a culture (when a single colony is transferred to a lipid medium, it is called a culture)
A culture is pure when the population is genetically homogeneous
Generation time: doubling time
E. coli 20 min
Logarithmic growth
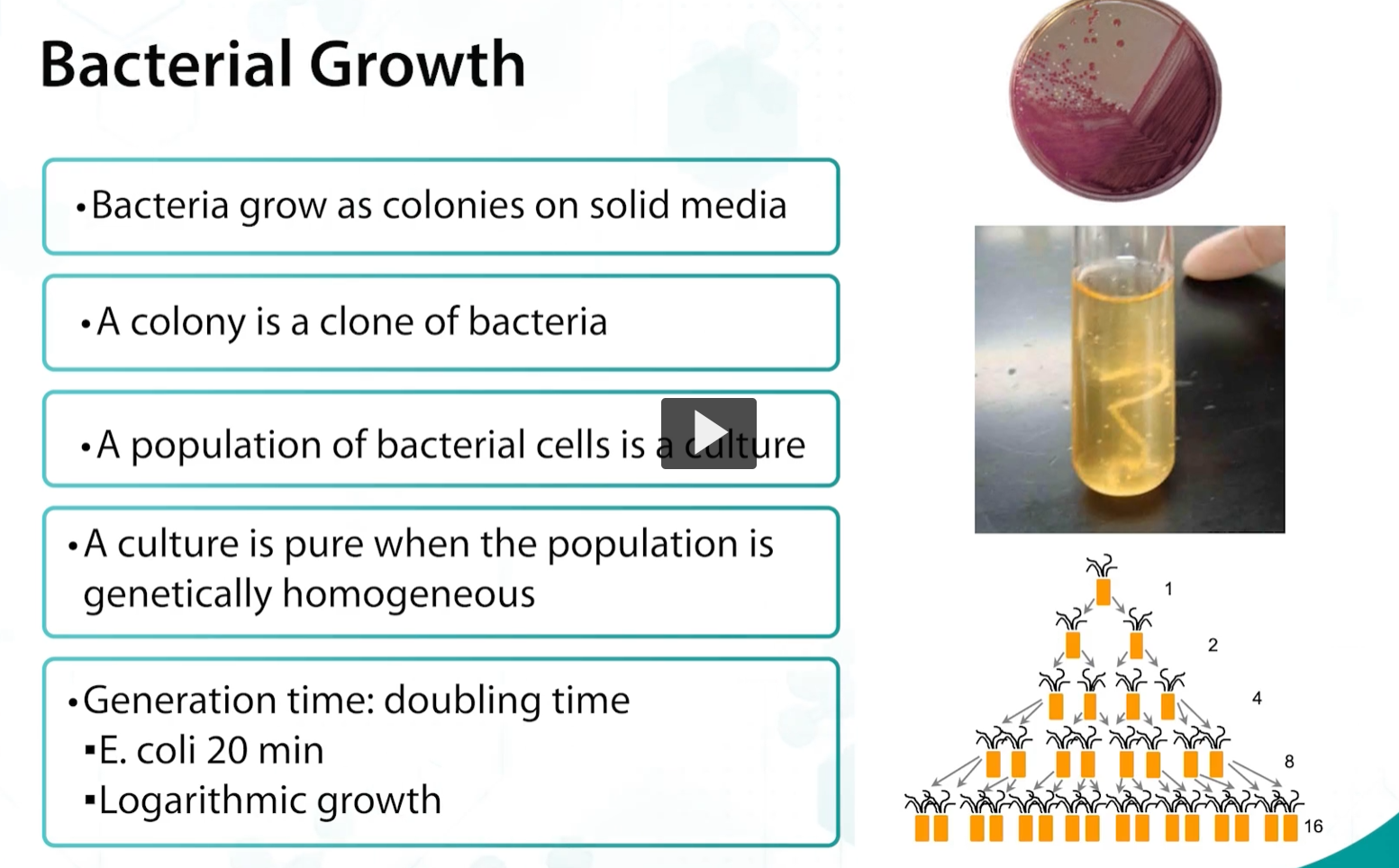


bacterial growth (transverse binary fission)
chromosome replication (copying the chromosomes)
polar separation of daughter chromosomes
cell division (septum)
formation of wall and envelope
separation of the new cells
this rapid process of cell division occurs continuously and eventually leads to exponential growth.
Multiplication by transverse binary fission
binary fission is a tightly regulated process involving more than 30 genes.
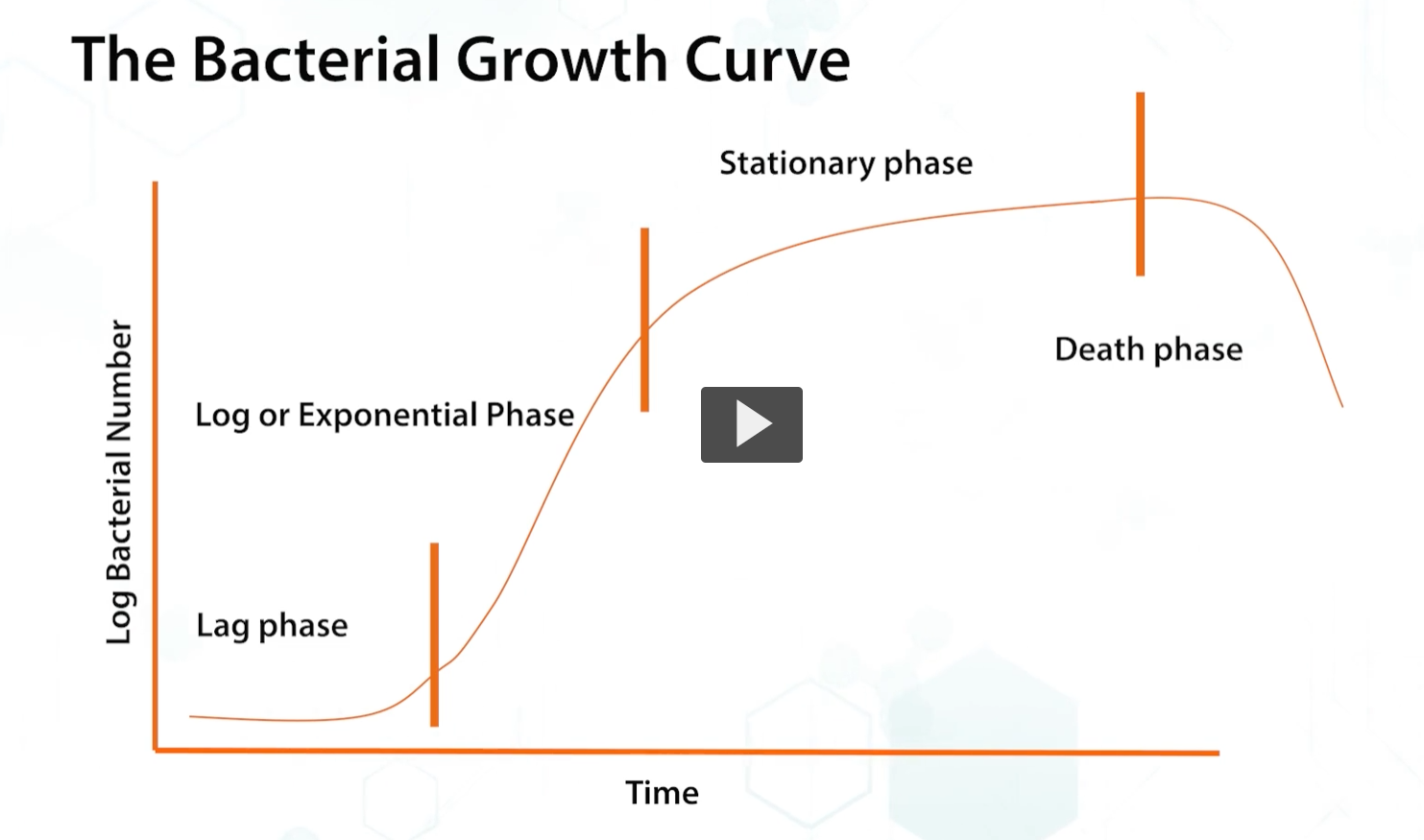
This diagram shows the bacterial growth curve
lag phase: bacteria senses the environment and determines the available nutrients for metabolism, adjusting the gene expression accordingly.
the lag phase is followed by the logarithmic or exponential phase where bacteria grow rapidly until they reach the…
stationary phase: this phase is determined by the remaining nutrients and pH and metabolites in the medium.
Death phase: finally, bacteria enter the death phase, where the bacteria begin to die.
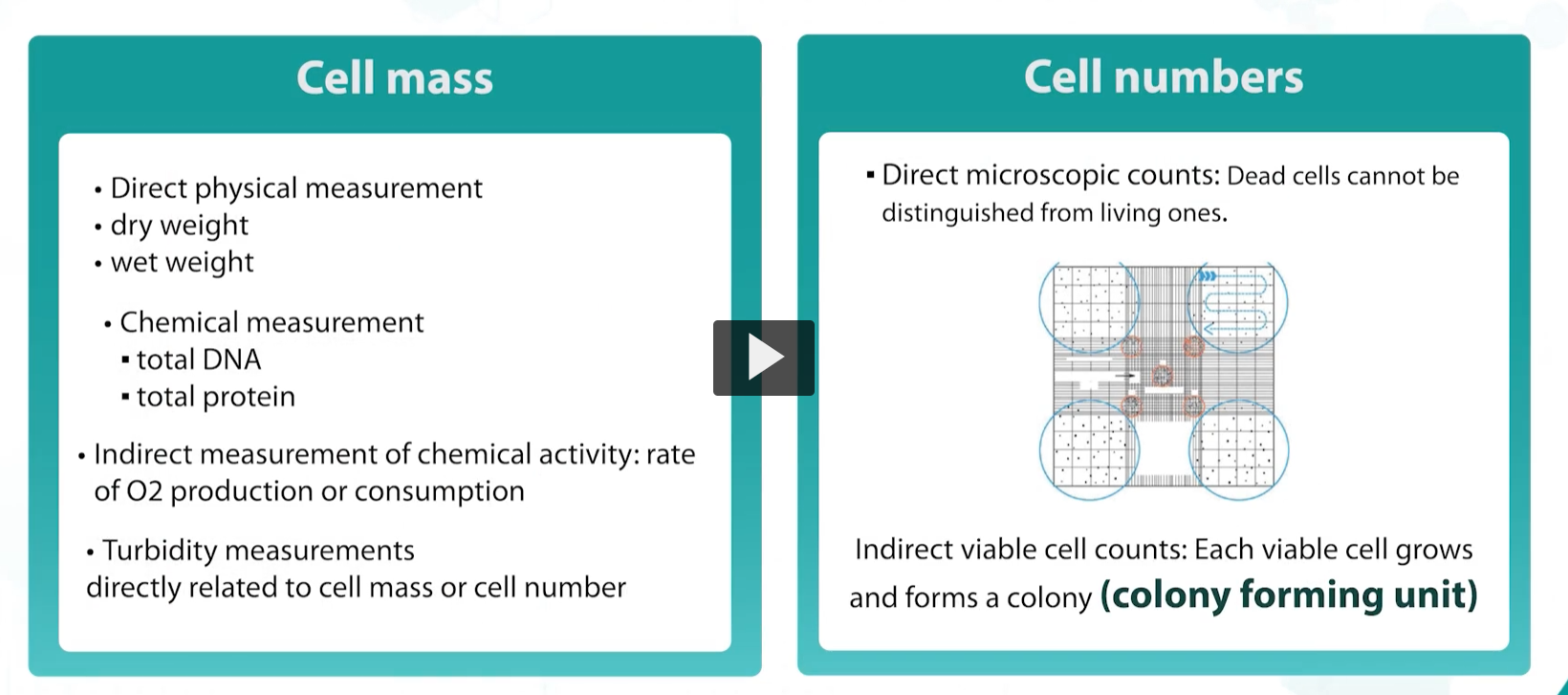
bacterial growth is measured in various ways in the laboratory, physical and chemical measurements can be taken, including cell mass, DNA content, or turbidity OR through direct counting over a microscope using counting chamber or automated subcounters.
Counting live cells is achievable through cell plating (viable plate count). Only living cells are capable of forming a colony, and each visible colony is called a colony-forming unit (CFU).
Cell mass
Direct physical measurement
dry weight
wet weight
Chemical measurement
total DNA
total protein
Indirect measurement of chemical activity: rate of O2 production or consumption
Turbidity measurements directly related to cell mass or cell number
Cell numbers
Direct microscopic counts: Dead cells cannot be distinguished from living ones.
Indirect viable cell counts: Each viable cell grows and forms a colony (colony forming unit)
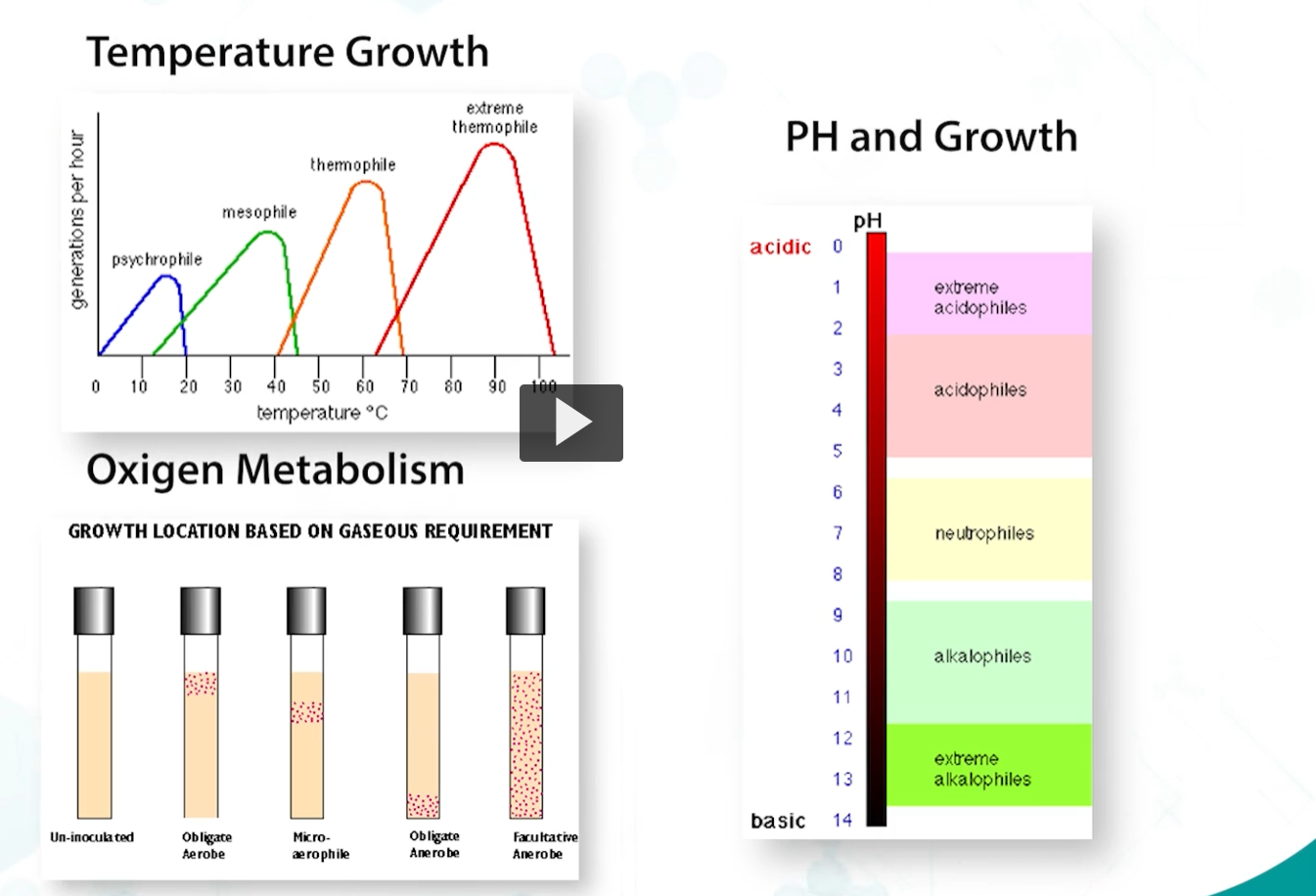
bacteria are capable of growing in a wide range of temperatures, from the artic to hot springs, in various and very wide pH ranges, and in different concentrations of oxygen.
These extreme environments have required the evolution of bacterial and protein structures that can function at such hard conditions.
oxygen metabolism
bacterial growth in oxygen is dependent on the expression of detoxifying enzymes. Because oxygen radicals are produced by bacteria, when they respire, detoxification of these reactive metabolites is crucial for survival.
This requires enzymes such as superoxide dismutase and catalase, the main enzymes responsible for oxygen radical detoxification.
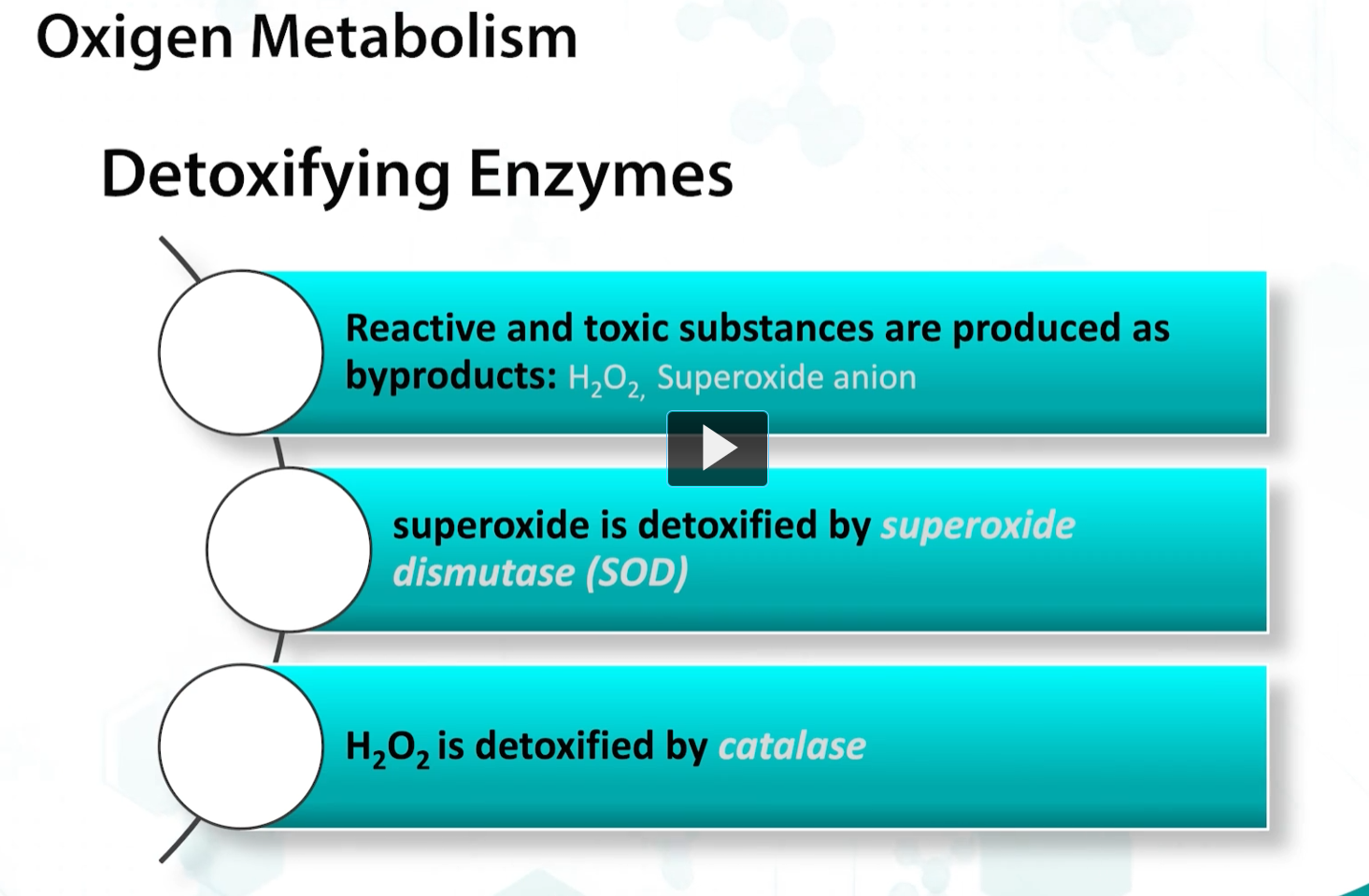

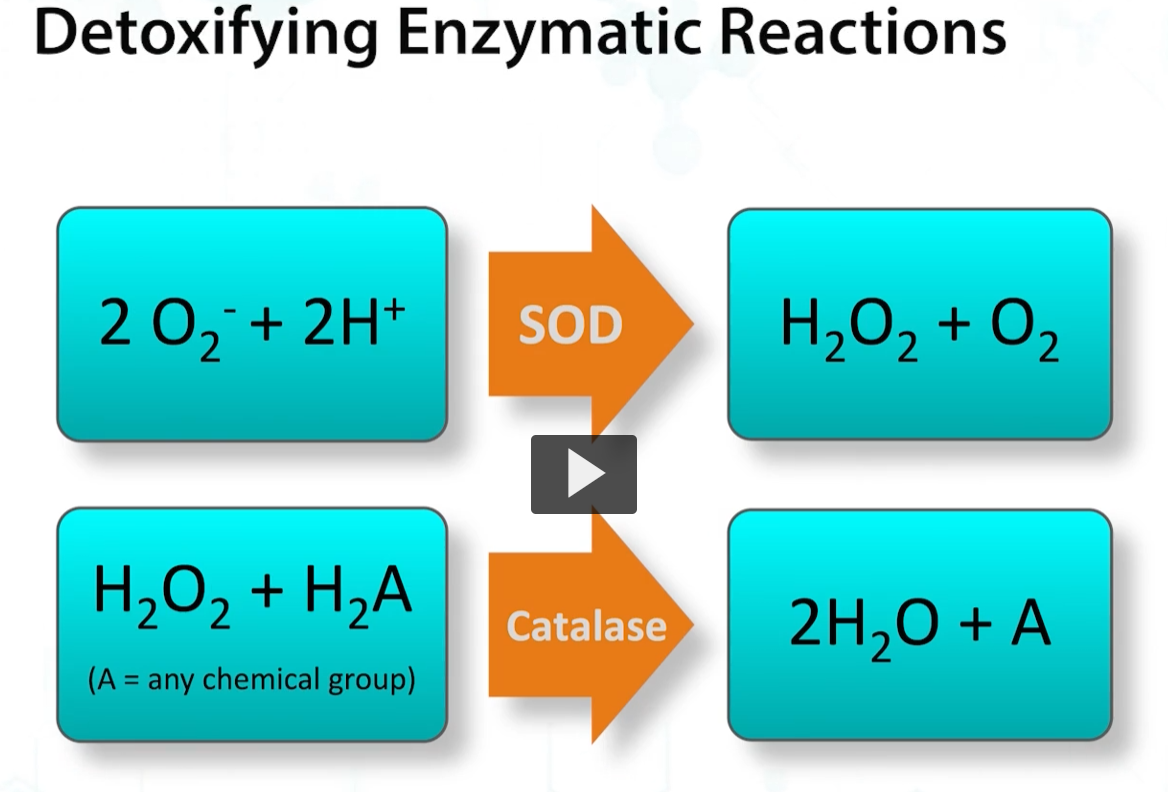
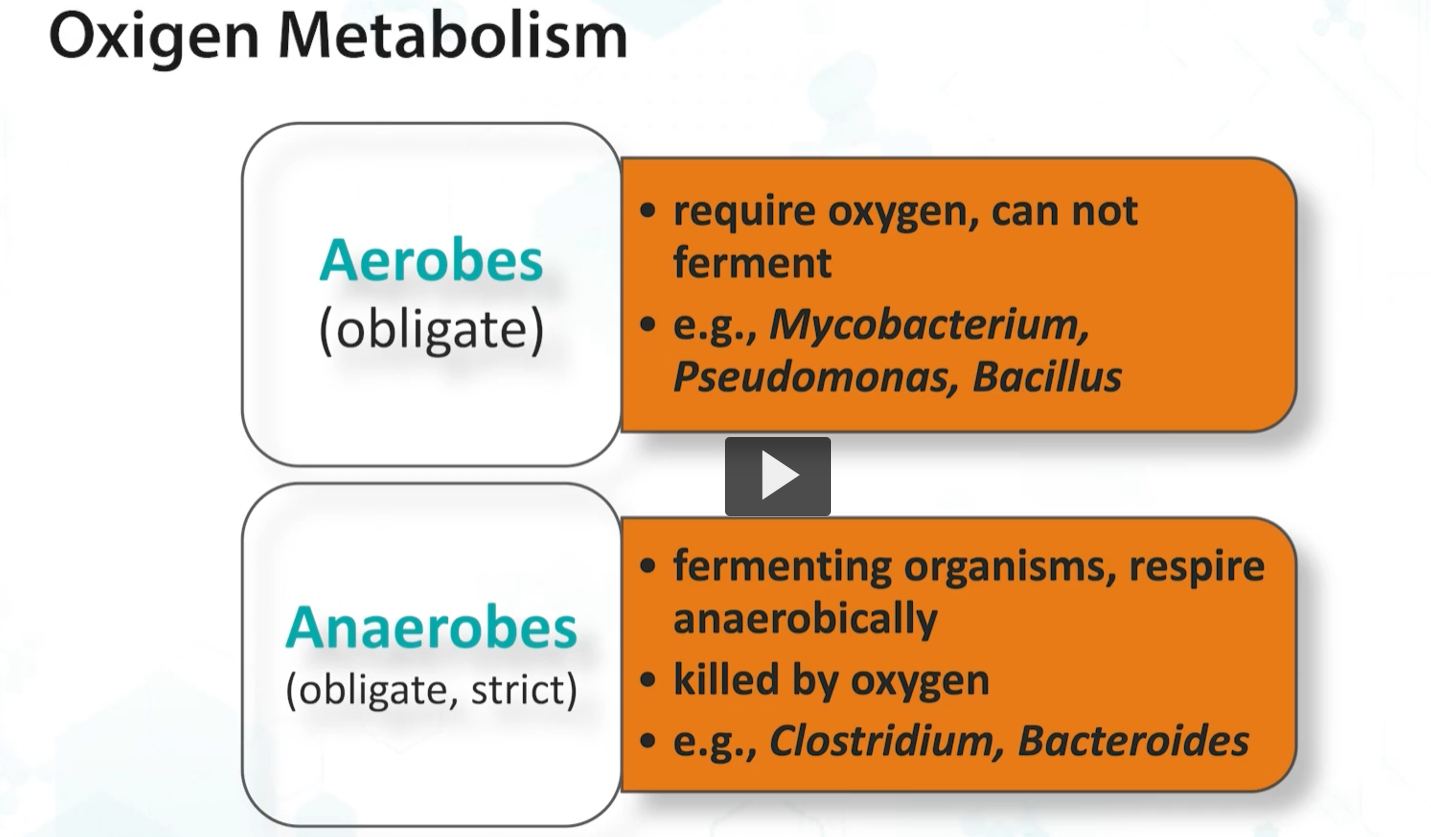
Aerobic organisms such as microbacteria and pseudomonas require oxygen and cannot ferment. Therefore, they MUST express CATALASE and SOD.
Obligate and strict anaerobes are fermenting organisms respire only anaerobically and are killed by oxygen.
oxygen metabolism
faculative anaerobes can respire when there is oxygen AND FERMENT in the ABSENCE of oxygen.
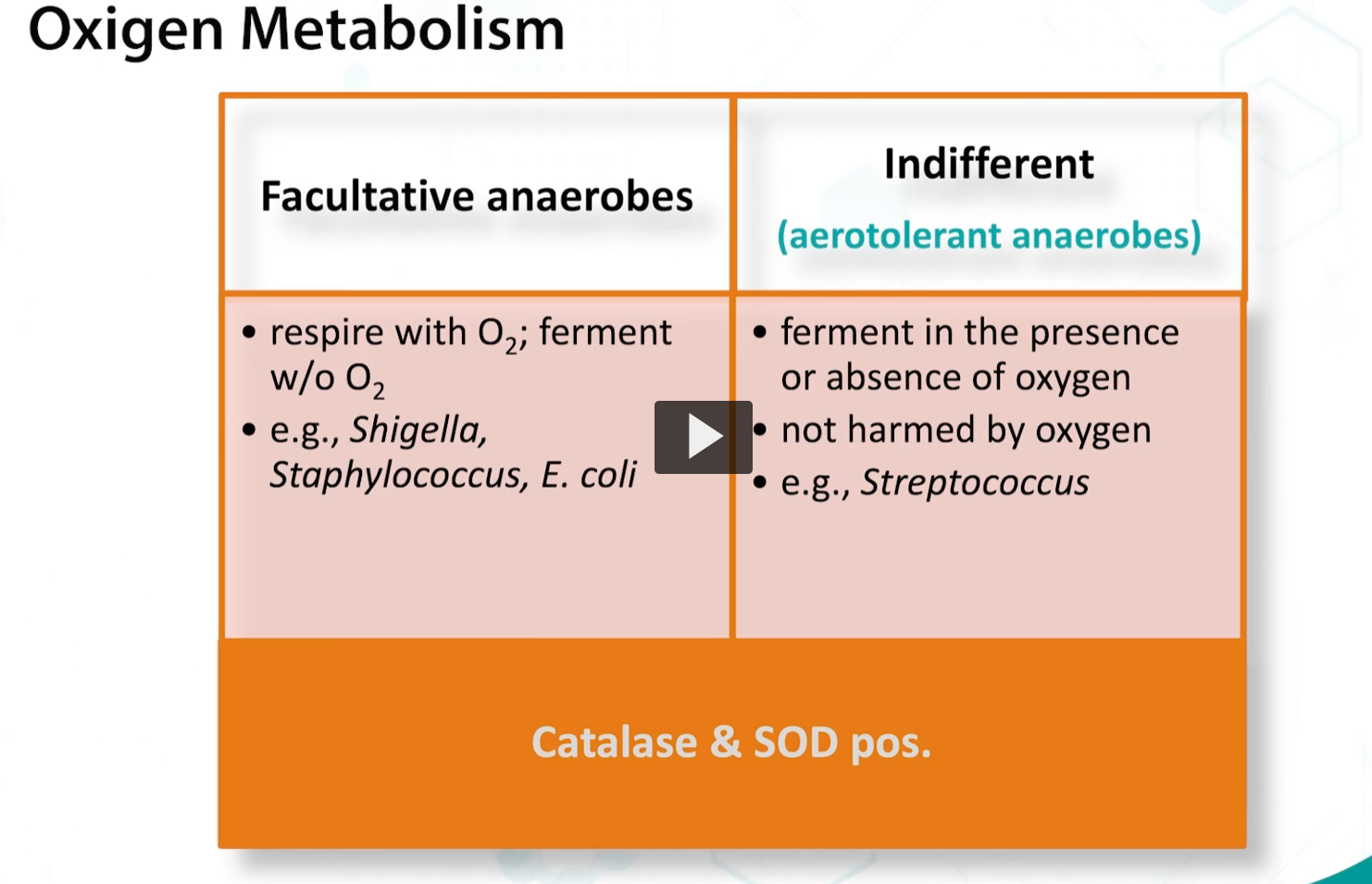


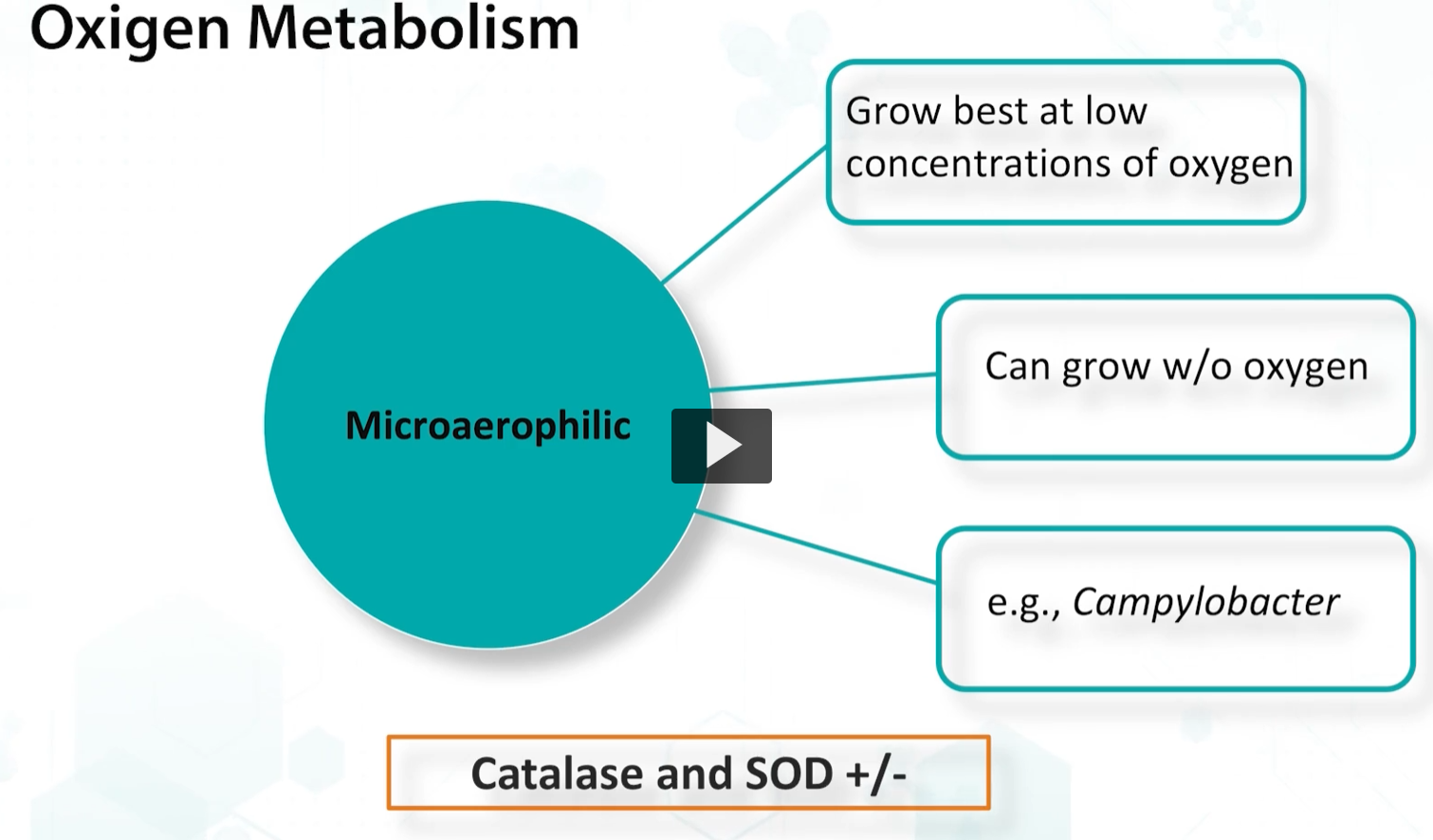
Finally, microaerophilic organisms grow best at low oxygen concentrations, and they express low quantities of catalase and superoxide dismutase,
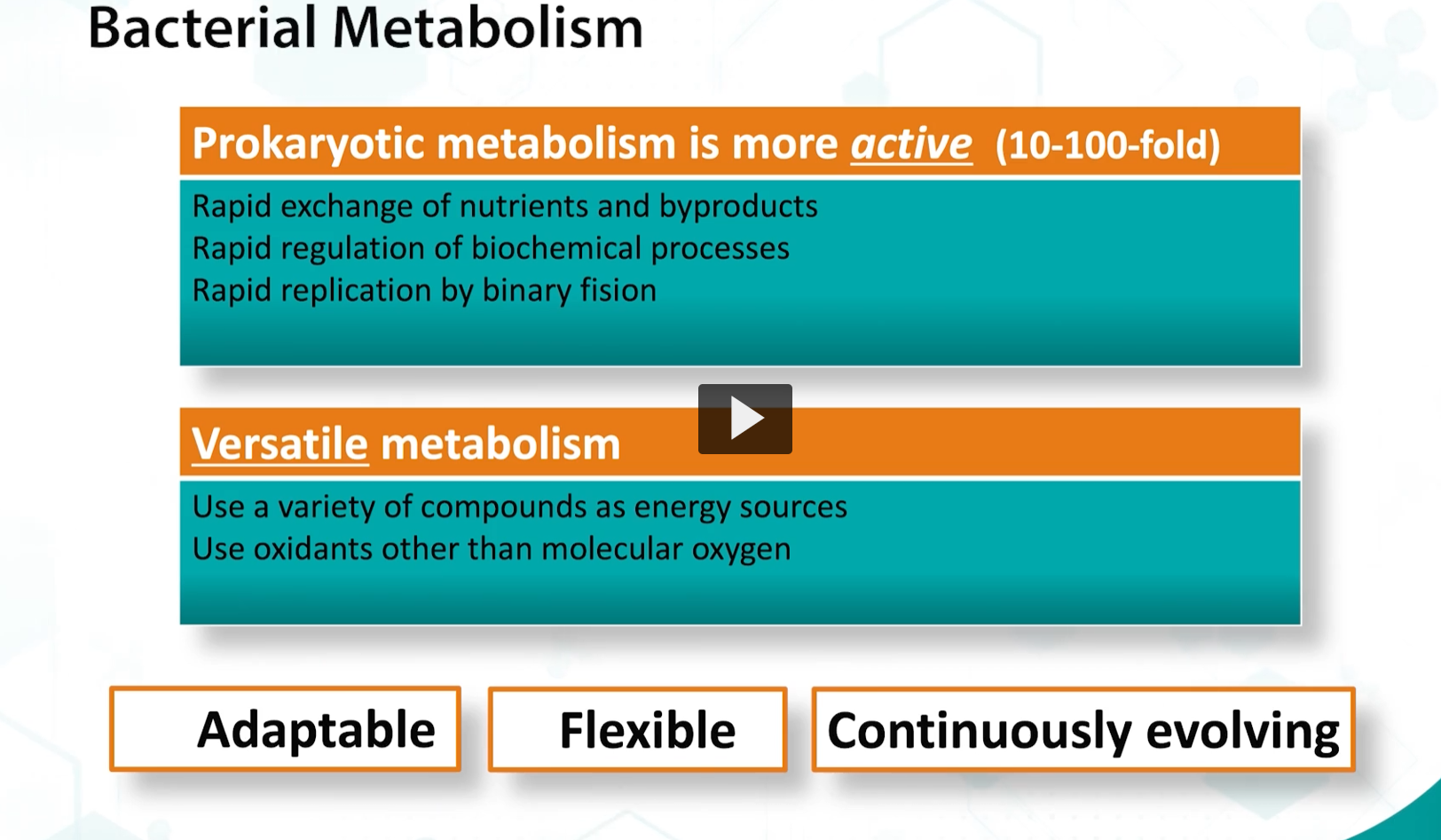
compared to eukaryotic metabolism, prokaryotic metabolism is much more ACTIVE and relies on rapid exchange of nutrients and metabolites.
There needs to be quick relation of these processes. These bacteria are (unicellular?) organisms that need to be adaptable, flexible, and continuously in order to survive in a changing environment.
Therefore, bacteria are must be versatile, and bacteria is always sensing their environment and adapting their metabolism accordingly.
The bacterial cell is considered a specialized energy transformer that uses the nutrients found around it.

energy is generated by substrate oxidation.
The substrates for oxidation include sugars, lipids, and proteins.
Energy generated is stored in high energy compounds, such as ADP, ATP, and acetyl-CoA.
It is important to know that enzymatic reactions that support life processes are VERY SIMILAR in prokaryotic and eukaryotic cells.
Bacterial metabolism consists of catabolism and anabolism.
Catabolism involves breaking down substrate molecules, such as carbohydrates, lipids, and proteins, to generate energy rich compounds, such as ATP, and reducing equivalents such as NADH, which will be later converted into energy.
This process also produces NEW MACROMOLECULES are simply organic compounds that are building blocks for the processes that bacteria need for growth and metabolism.
Catabolism in bacteria can be aerobic or anaerobic, and they can choose from different catabolic pathways to convert substrates into their needed energy.

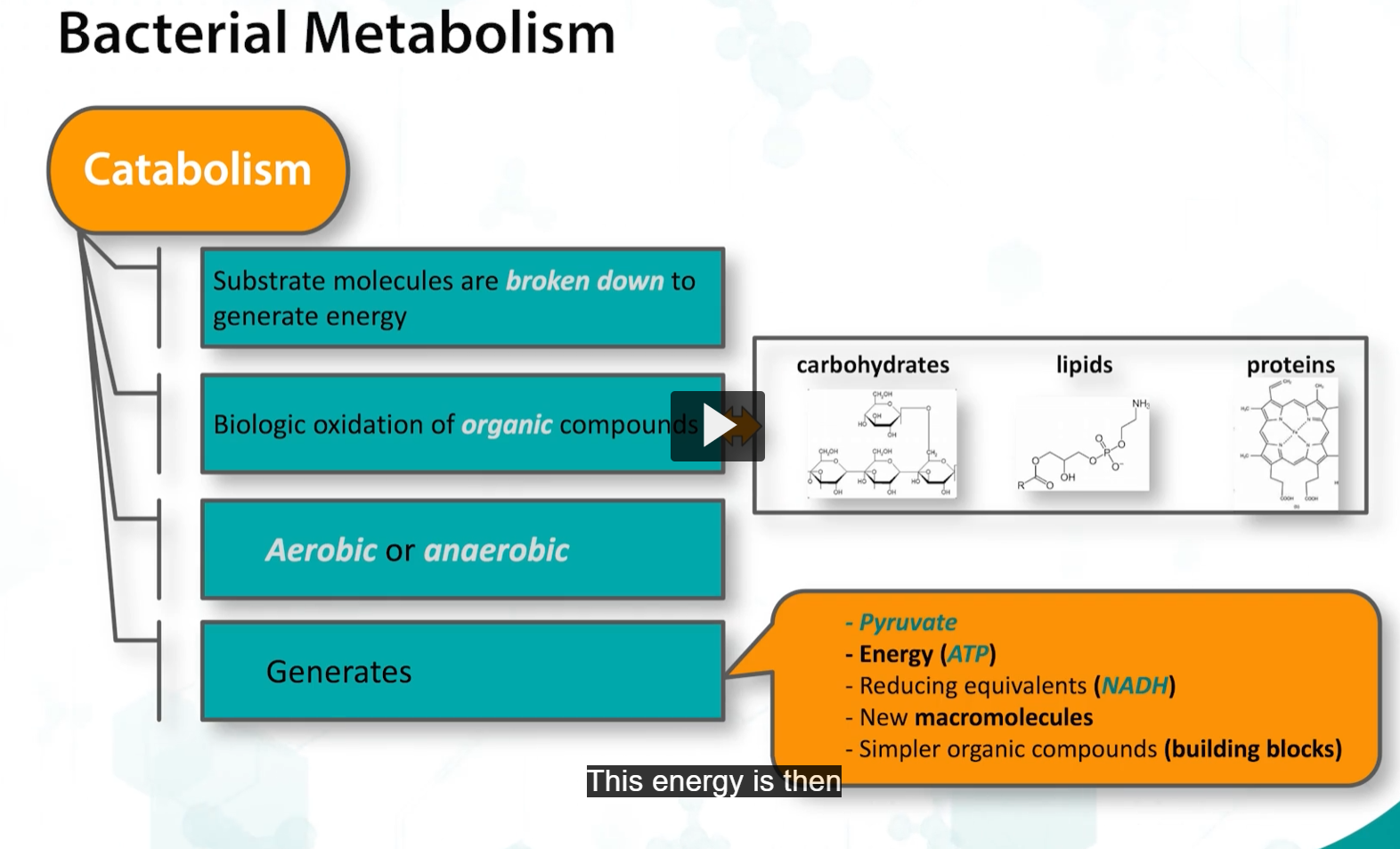
This energy (from catabolism) is utilized during anabolism, which refers to the synthesis of macromolecules need for growth and proliferation of bacteria.
This process requires a variety of precursors, building blocks, and energy compounds.
Before utilizing sugars in the various metabolic pathways, these (referring to the diagram), must be transported into the cell. For that (transportation), bacteria is able to use various mechanisms

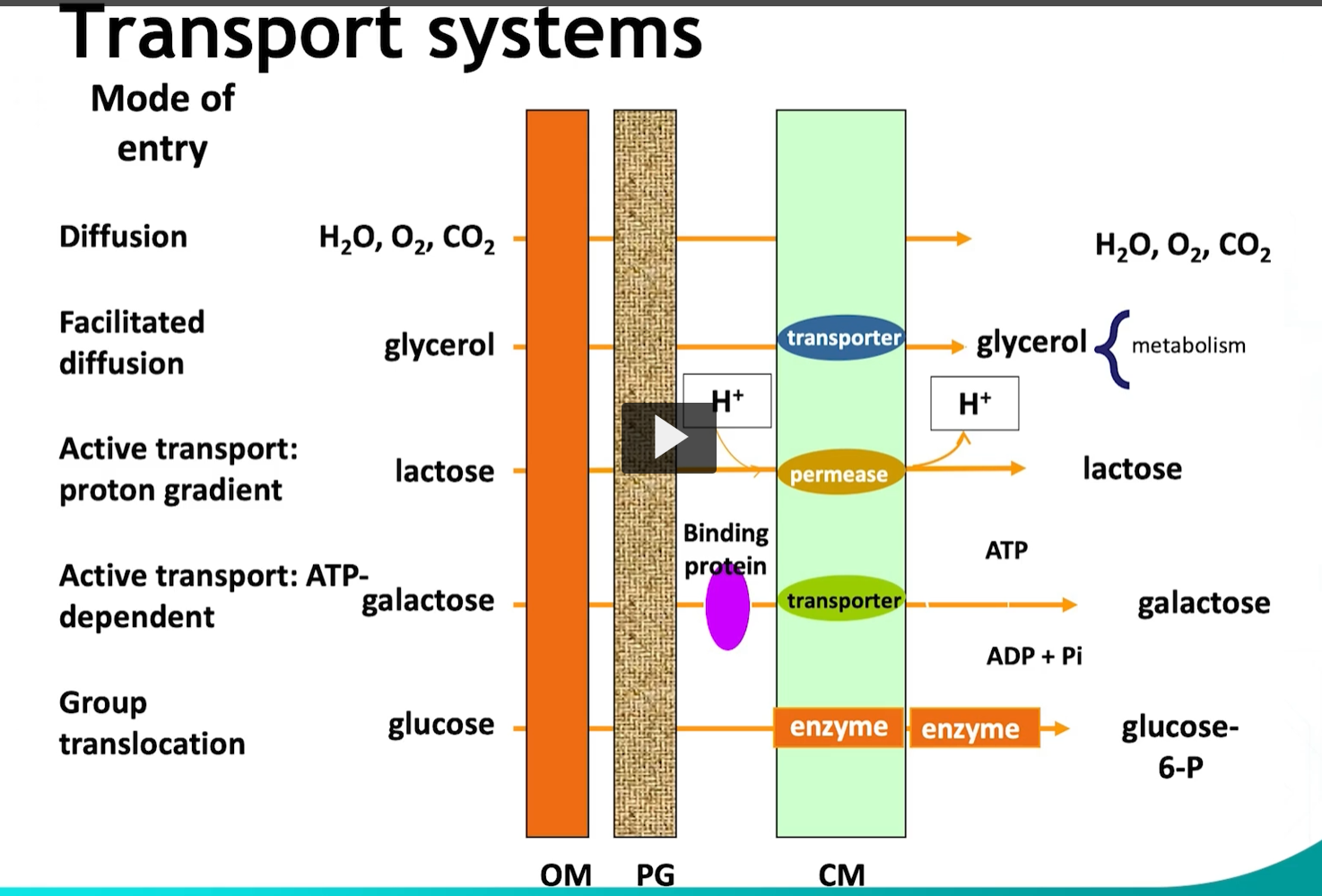
The first transport mechanism is facilitated diffusion. For example, glycerol.
Glycerol is transported without spending any energy, glycerol then enters immediately into the metabolic pathway and then is phosphorylated by phosphoenolpyruvate. (CHATGPT says Glycerol-kinase).
During active transport, energy is used to move the substrate across the membrane.
One example of this will be lactose.
Energy can come from ATP hydrolysis, or the protonmotor force is used to alter the conformation of a protein carrier, and phosphorylate the substrate during transport.


The last transport mechanism is group translocation, for example, glucose transport in bacteria. In this process, a series of proteins (including a permease) chemically modifies the substrate during transport by phosphorylation, using energy derived from phosphoenolpyruvate (PEP).
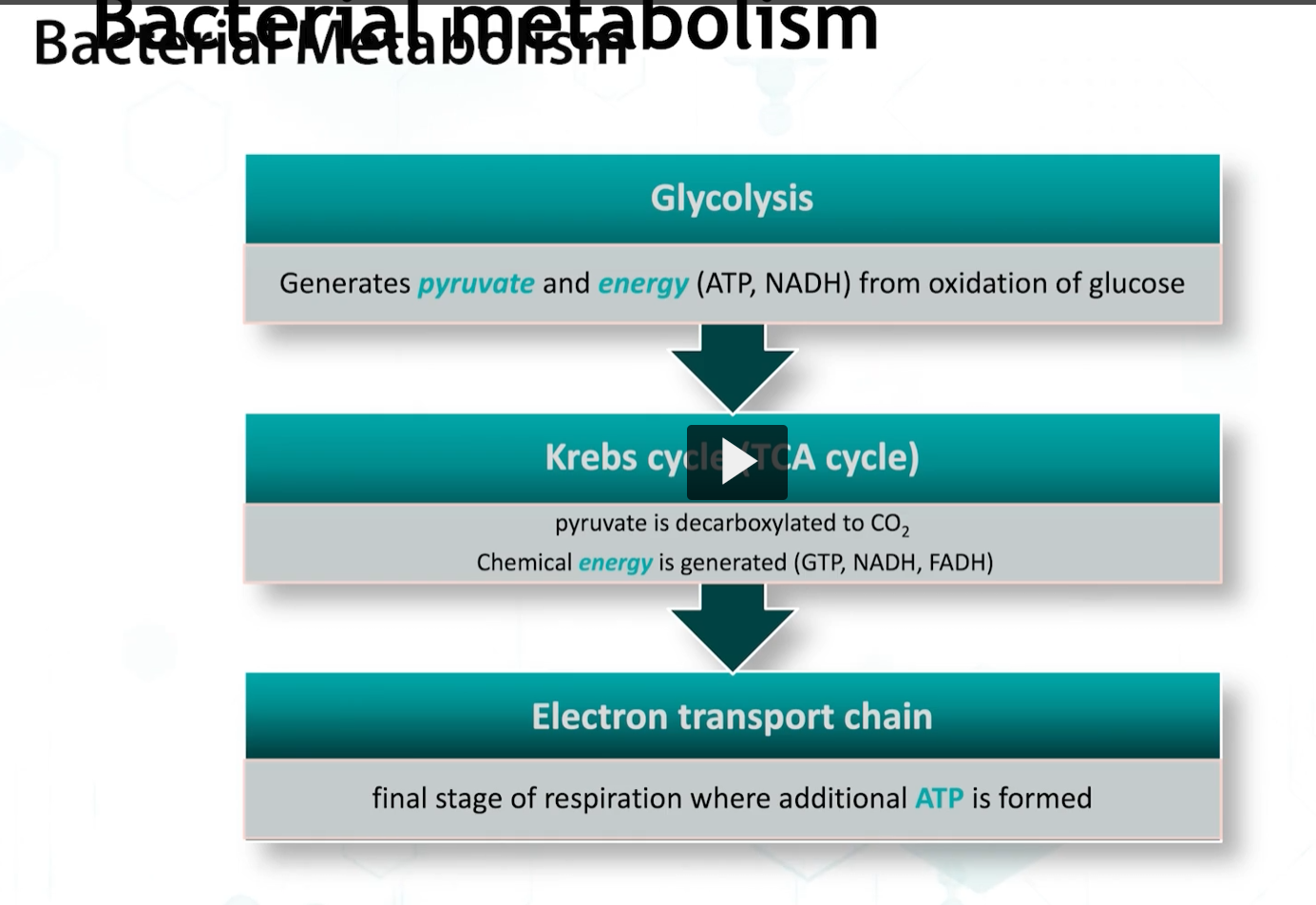
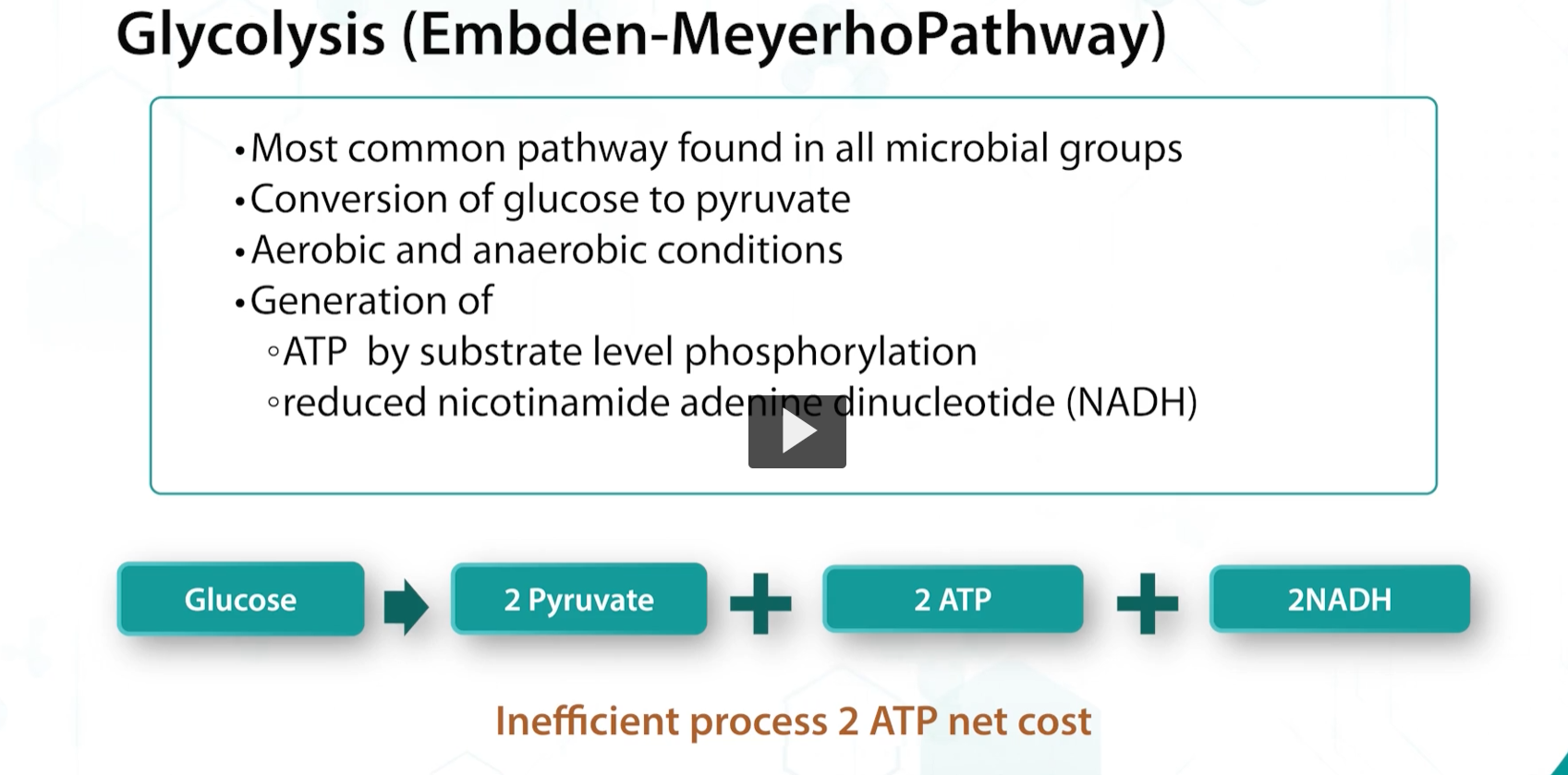
the main pathway for glucose metabolism is glycolysis or the embden-meyerhof cycle, in which glucose is metabolized to 2 pyruvate molecules and ATP.
This is followed by the Kreb’s Cycle, which is called the tricarboxylic acid cycle (TCA).
Kreb’s produces energy by substrate level phosphorylation, and also, NADH, and FADH, that are converted into energy by the electron transport chain.
embden-meyerhof pathway
pentose-phosphate shunt
entner-doudoff pathway
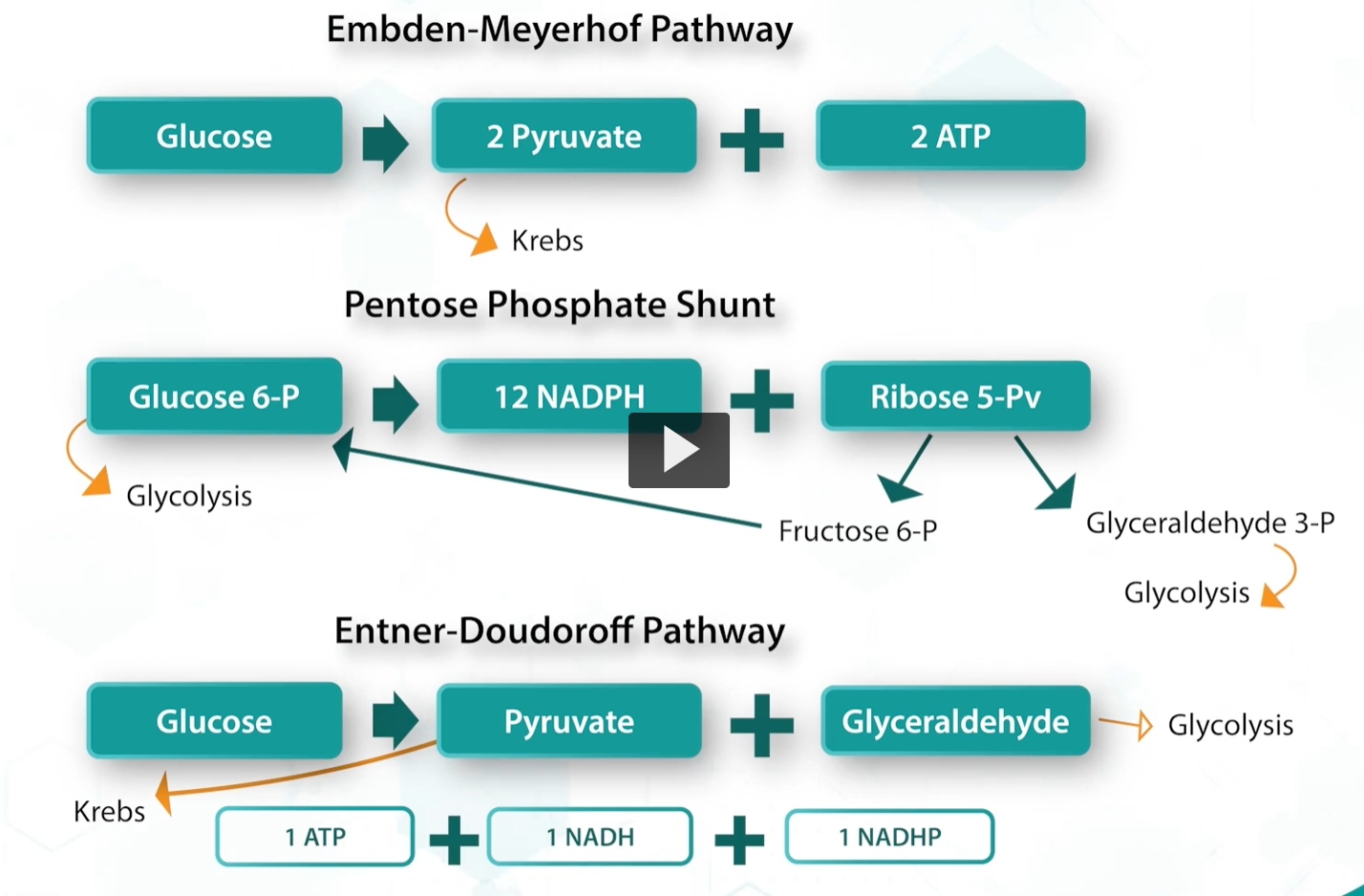
depending on the needs, bacteria can shift to other sugar metabolic pathways.
The bacteria needs 5-carbon sugars, for example, ribose, for the synthesis of nucleotides, they can shuffle glucose into the pentose phosphate shunt. If the enzymes of glycolysis are missing, such as phosphofructokinase, bacteria can use the entner-doudoff pathway.
In bacteria, the pyruvate produced by the Entner-Doudoroff pathway can enter the Krebs cycle. Glyceraldehyde-3-phosphate enters glycolysis, where it is further metabolized to pyruvate, which can then also enter the Krebs cycle.
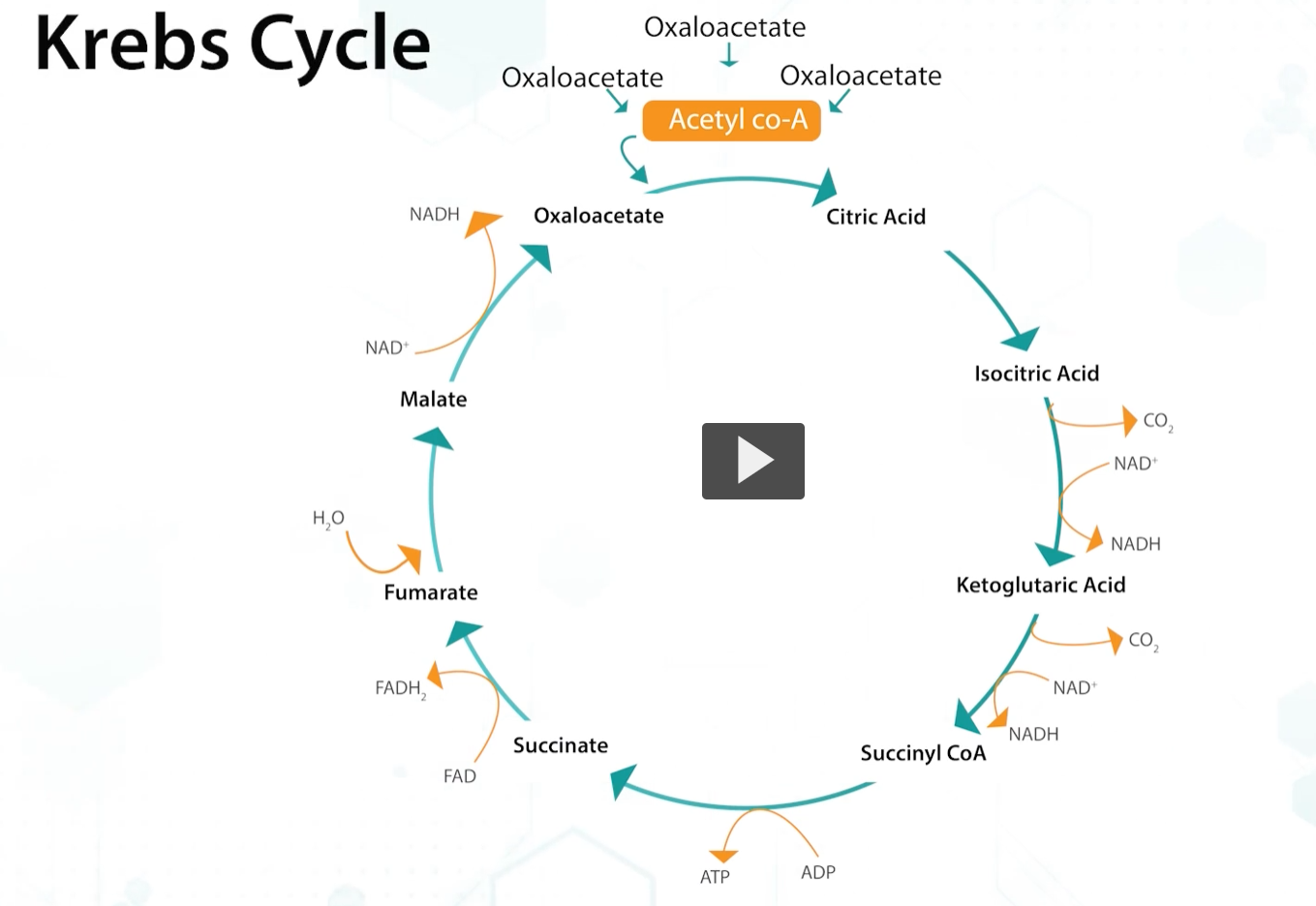
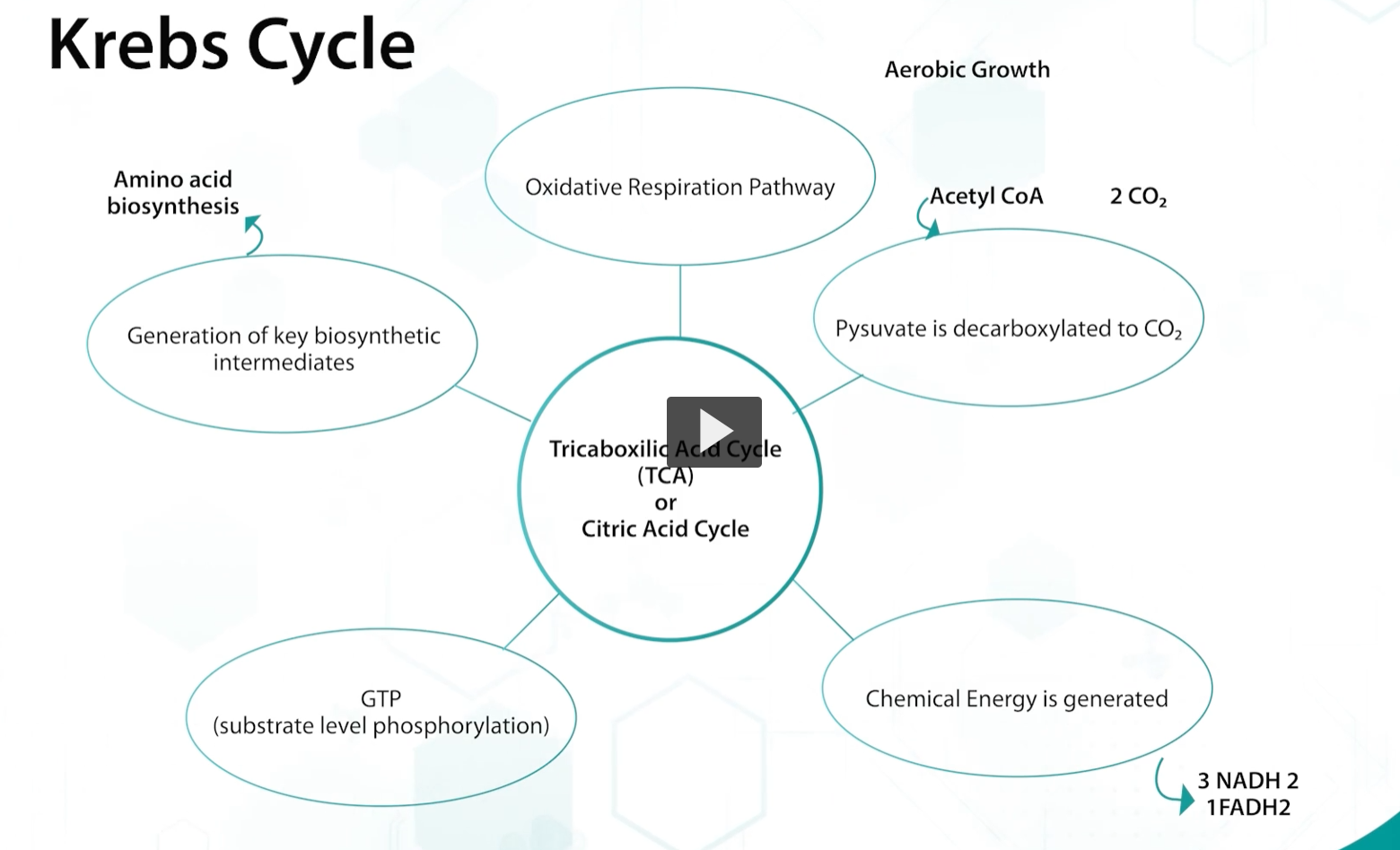
The Kreb’s, or TCA cycle, is part of the oxidative respiration pathway used by AEROBIC bacteria.
Kreb’s is the important focal point of central where acetyl-CoA, that is produced by different pathways, completes the process to generate energy.
during krebs, pyruvate is complete decarboxylated to CO2, generating chemical energy in the form of NADH, FADH, or GTP, through substrate level phosphorylation.
and also, key biosynthetic intermediates are produced.
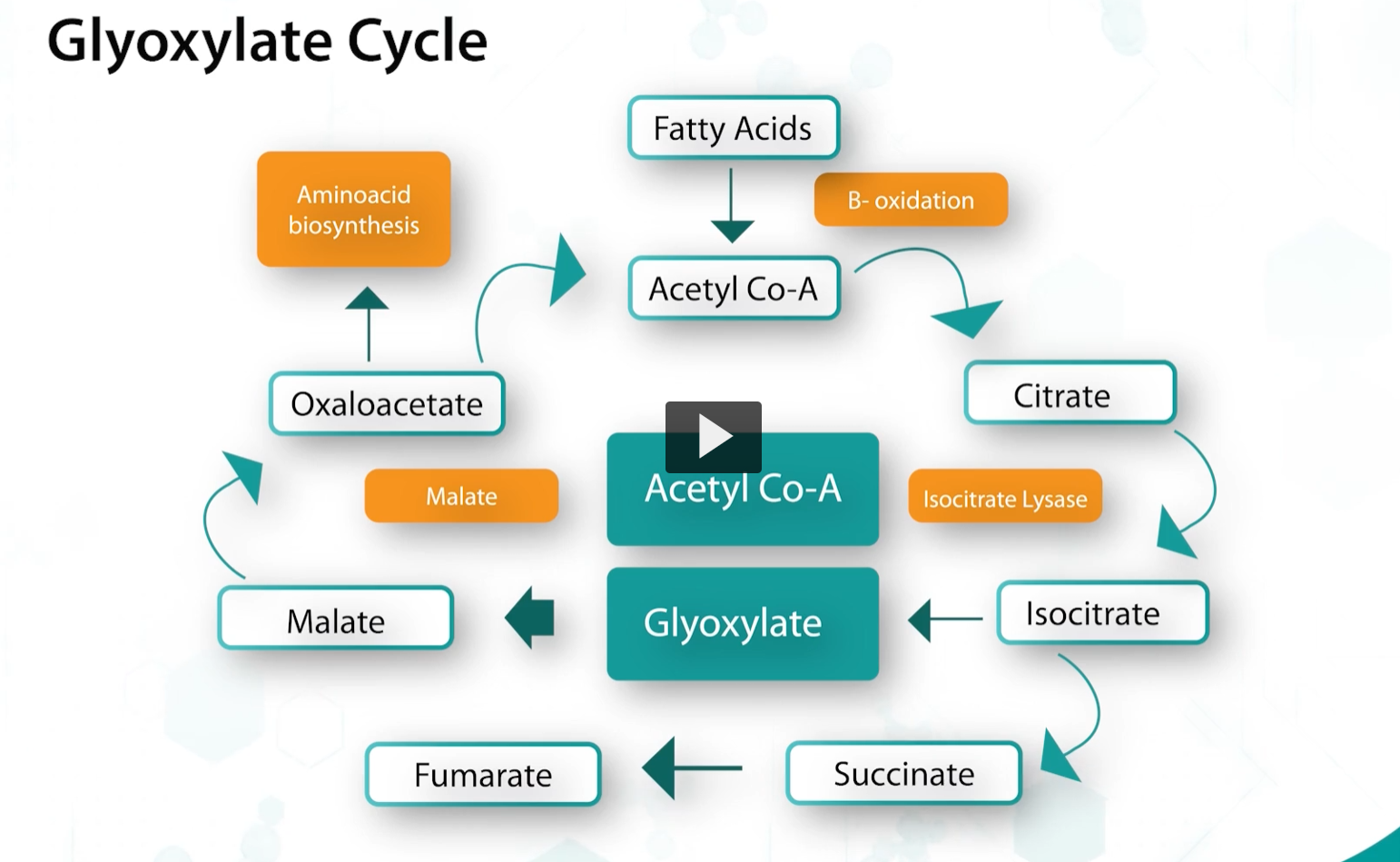
glyoxylate cycle
bacteria can also use a modification of the kreb’s cycle called a glyoxylgate cycle, and it’s used to catabolize acetyl-CoA.
This cycle is useful to bacteria because it replenishes key TCA intermediates that are being used in other biosynthetic reactions, this is why this cycle is referred to as an anapleurotic cycle.
Depletion of these metabolites will bring the TCA cycle to halt, but bacteria can solve this problem through the beta-oxidation fatty acid production of acetyl-CoA, which is subsequently combined with glyoxylate to produce malate. And this allows the TCA cycle to continue.
Respiration refers to the complete dissimulation of sugars resulting in generation of 38 ATPs per molecule, 6 CO2 molecules, and water.
This is more efficient in eukaryotes, where 2 ATPs are used in a shuttle, that transports NADH into the mitochondria.
The final step in respiration is the electron transfer system, where oxidation reduction and electron transfer reactions occur
In bacteria, respiration can occur in BOTH aerobic and anaerobic conditions, the difference being who is the final electron acceptor.
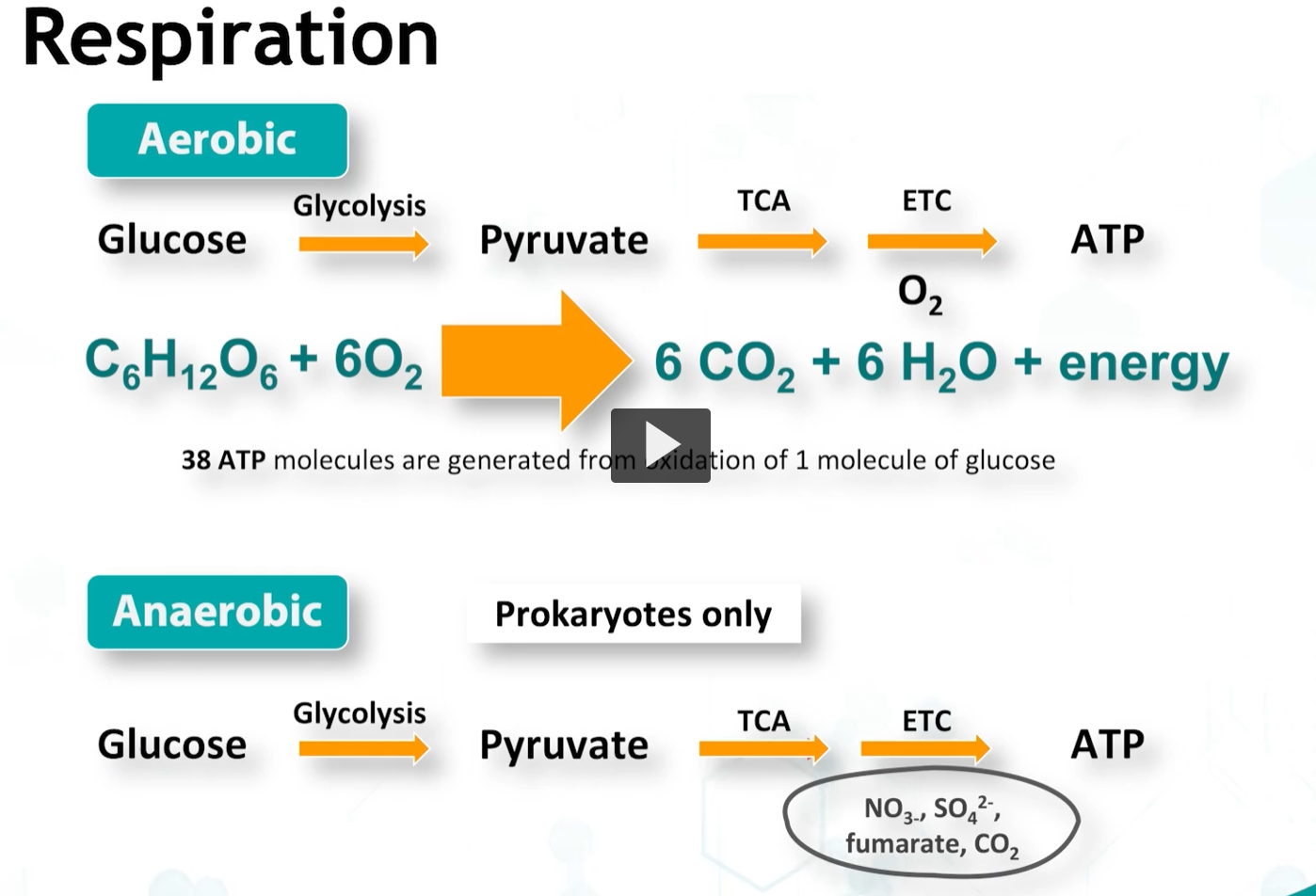
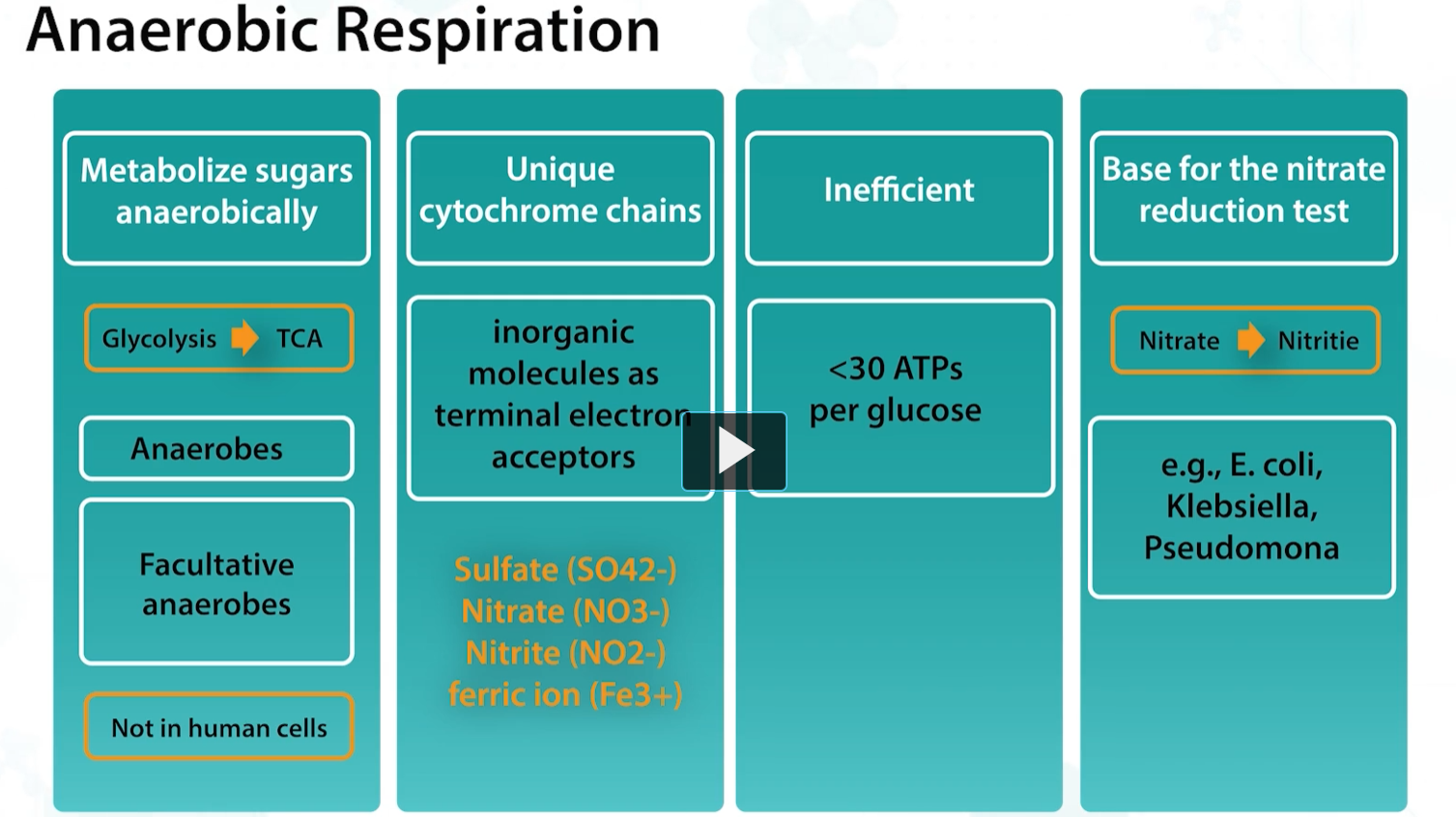
anaerobic respiration is used by anerobes and faculating anaerobes to metabolize sugars in the absence of oxygens, and this allows the production of additional ATPs than possible by using only glycolysis, although, less efficiently, with only 30 ATPs vs 38 ATPs formed.
It uses unique cytochromes and inorganic molecules such as sulfate, nitrate, nitrite, as terminal electron acceptors.
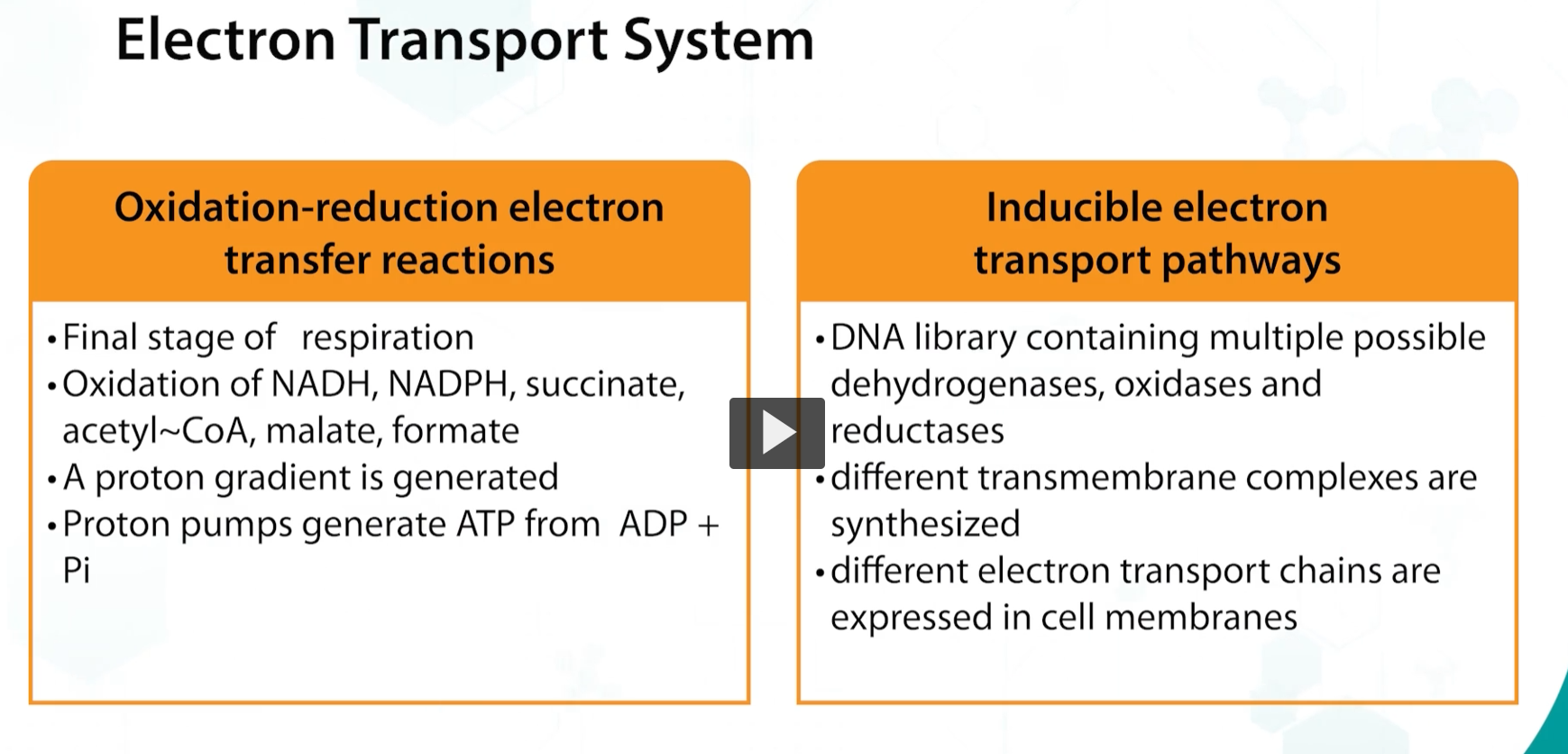
a key aspect of electron transport systems is that it is the final stage of respiration, consisting of oxidation and reduction reactions. The net result is a creation of a transmembrane proton gradient, known as the protonmotive force. and the production of ATP, every time that a hydrogen ion is transported by the proton pumps
This system is inducible, and depending on the environment in which the bacteria is growing.
Various dehydrogenases, oxidases, and radicases can be expressed.
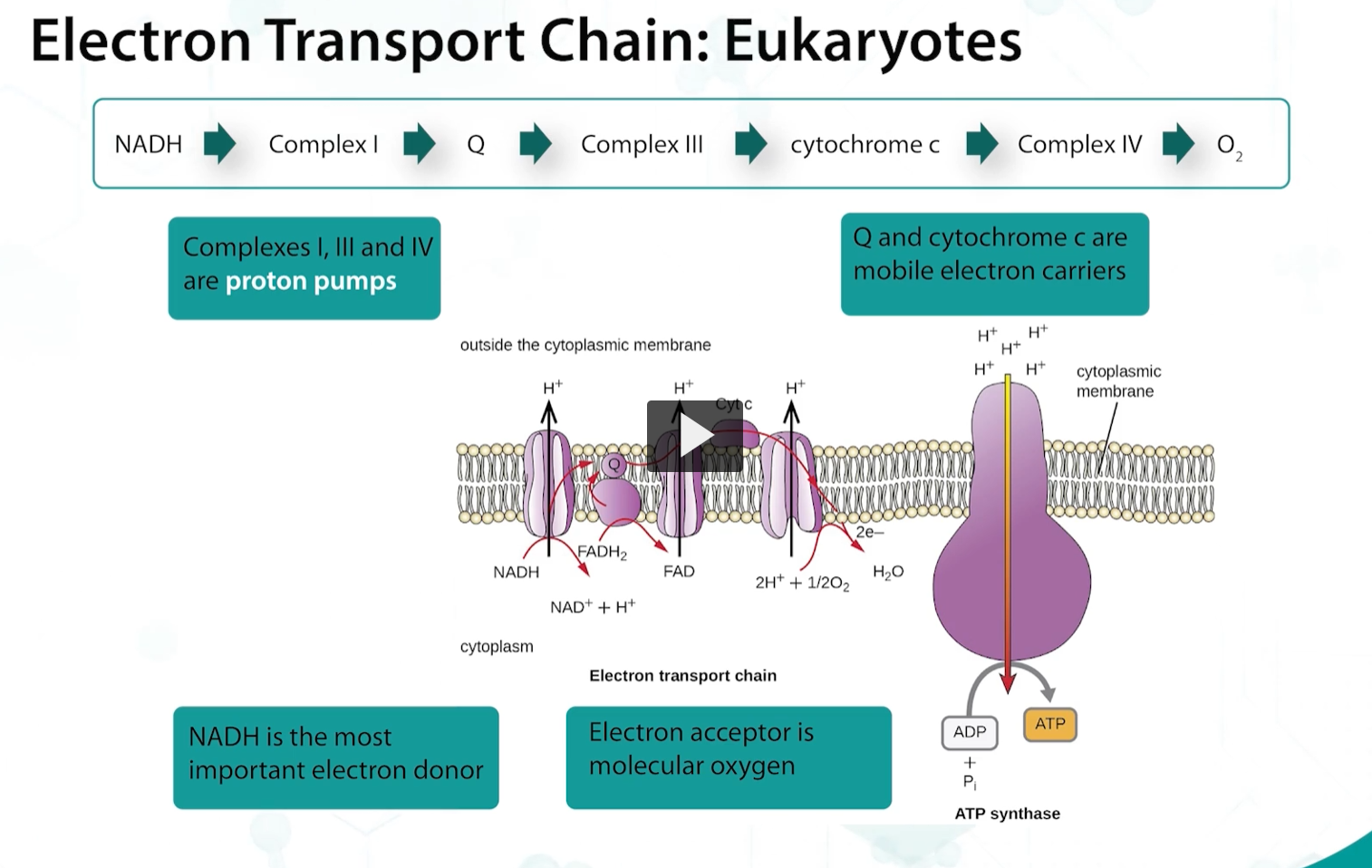
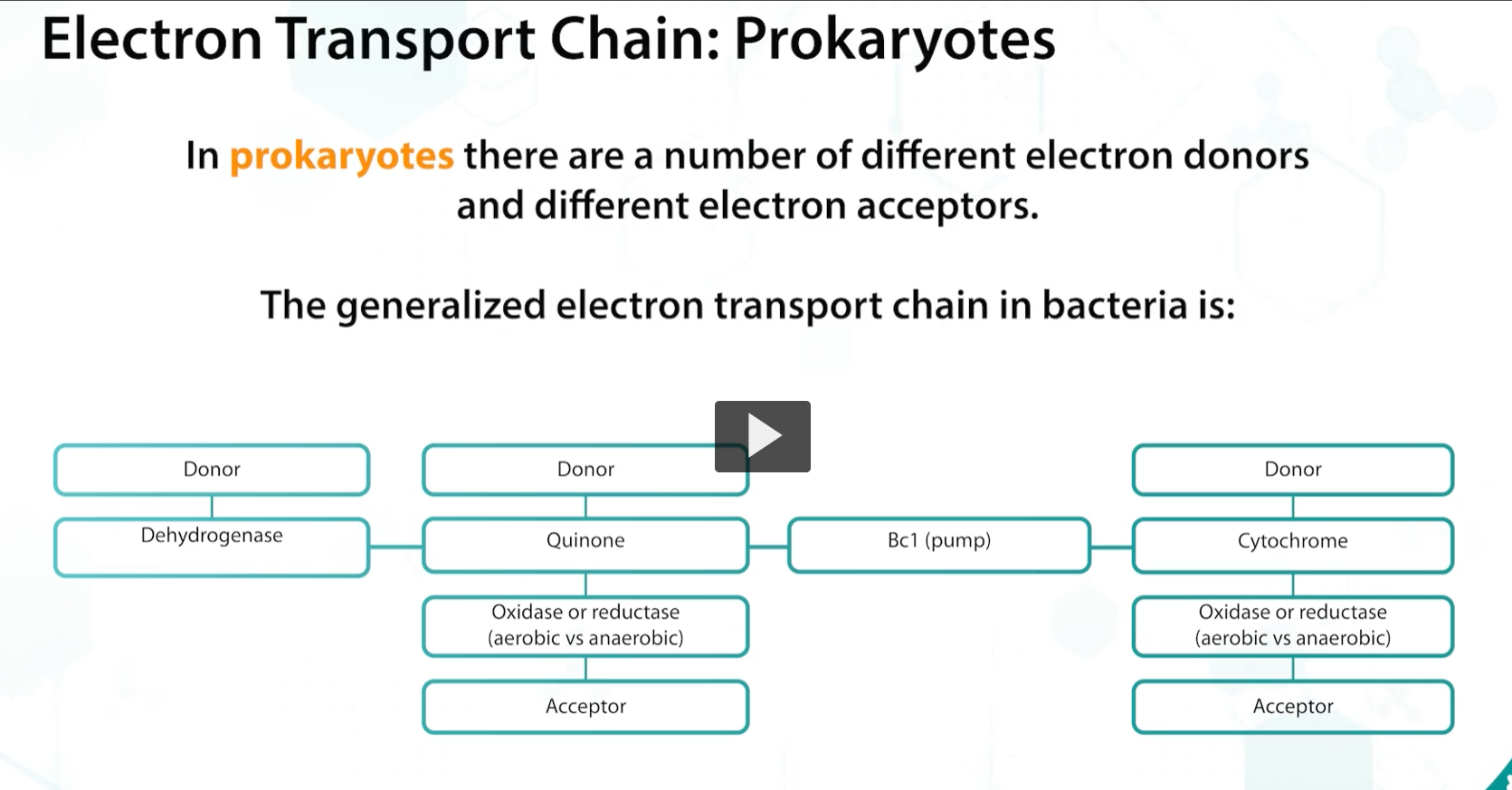
The sequence of events of the electron transport chain of bacteria is similar to eukaryotes, in which there are proton pumps, electron carriers such as cytochromes, and electron donors such as NADH.
but, PROKARYOTIC ELECTRON DONORS use a variety of electron donors and acceptors, and the choice of dehydrogenases, oxidases and reductases varies, allowing the bacteria to use different inorganic molecules that may be available as terminal electron acceptors.

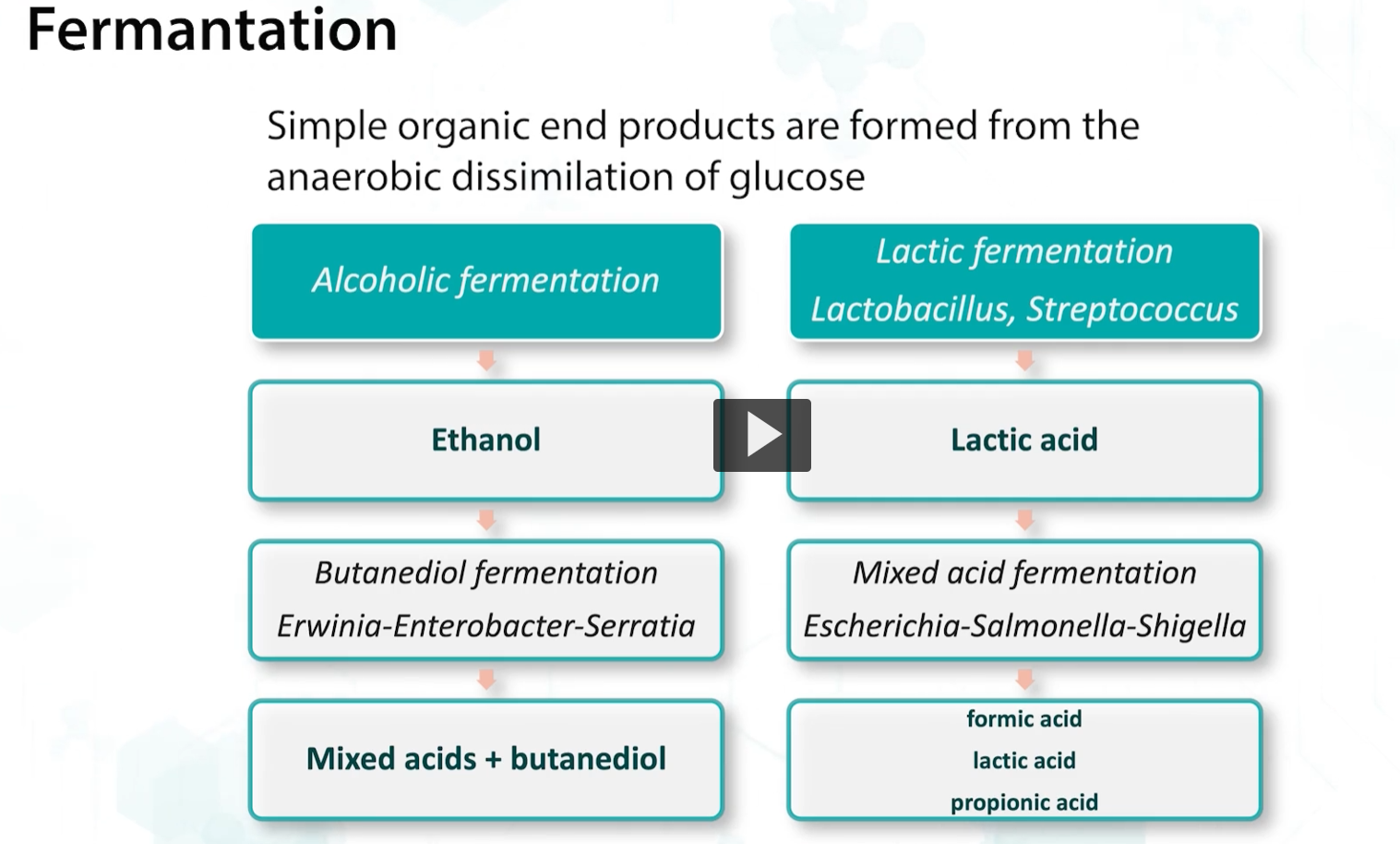
lastly we will discuss fermentation, defined as the metabolic process that releases energy from a sugar or other compounds that does not require molecular oxygen and uses an organic compound as the final electron acceptor.
fermentation produces 2 ATPs per molecule of glucose + 2 molecules of lactate, although the efficiency is low compared to to respiration that yields 38 ATPs, it is valuable as it allows catabolization of sugars to continue in the absence of oxygen.
during fermentation, simple organic and other products are formed, including alcohols and acids which are useful in food.
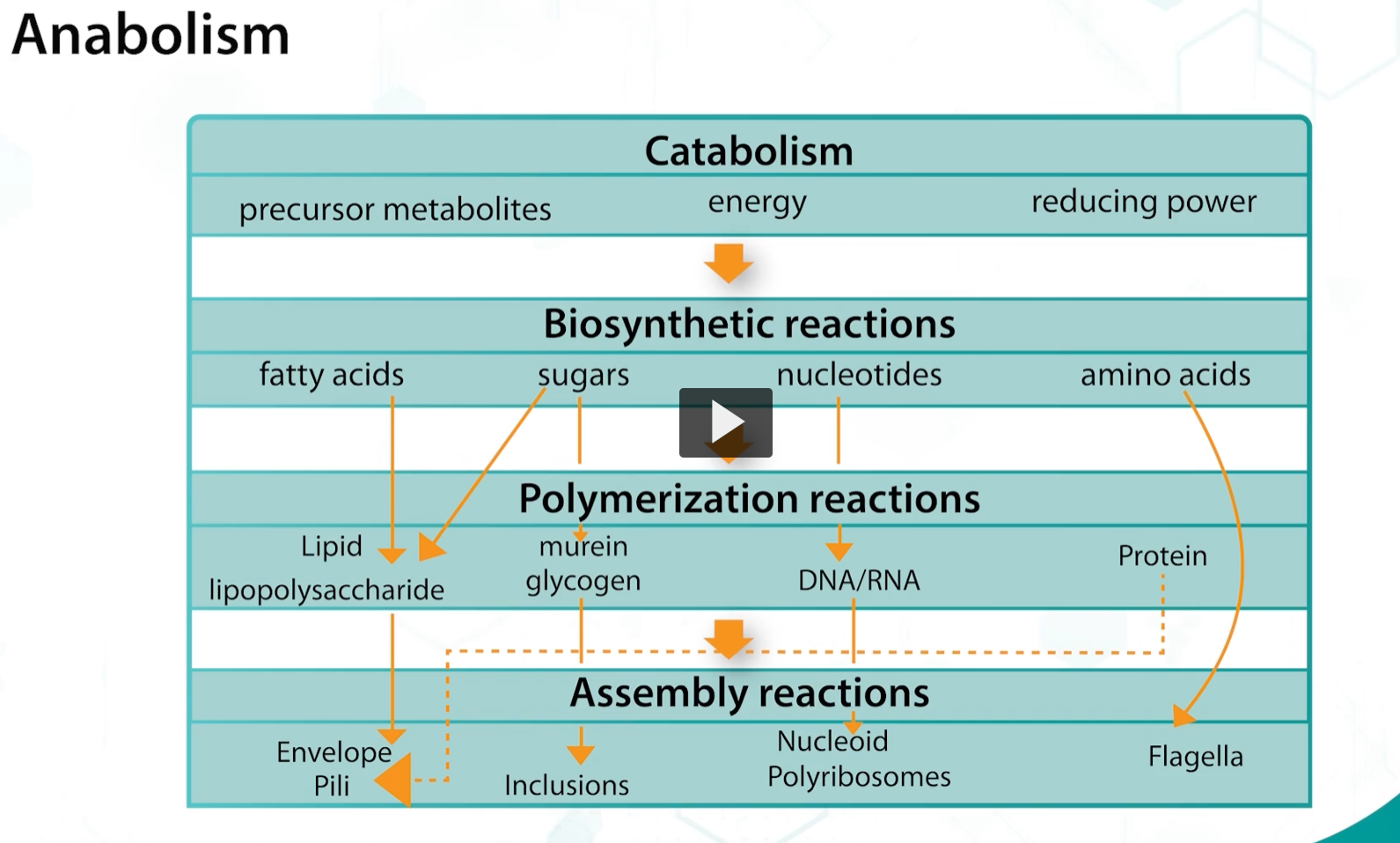
To summarize, during catabolism, organic substrates are broken down to produce energy, which is subsequently used in biosynthetic reactions to synthesize the building blocks necessary for the synthesis for the macromolecules that form the bacterial cell structures.