BIOC 4331 Lecture 11
1/45
Earn XP
Description and Tags
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
46 Terms
What is protein folding?
A polypeptide adopts its native, compact, functional 3D structure.
What determines a protein’s final (tertiary) structure?
The amino acid sequence (primary structure).
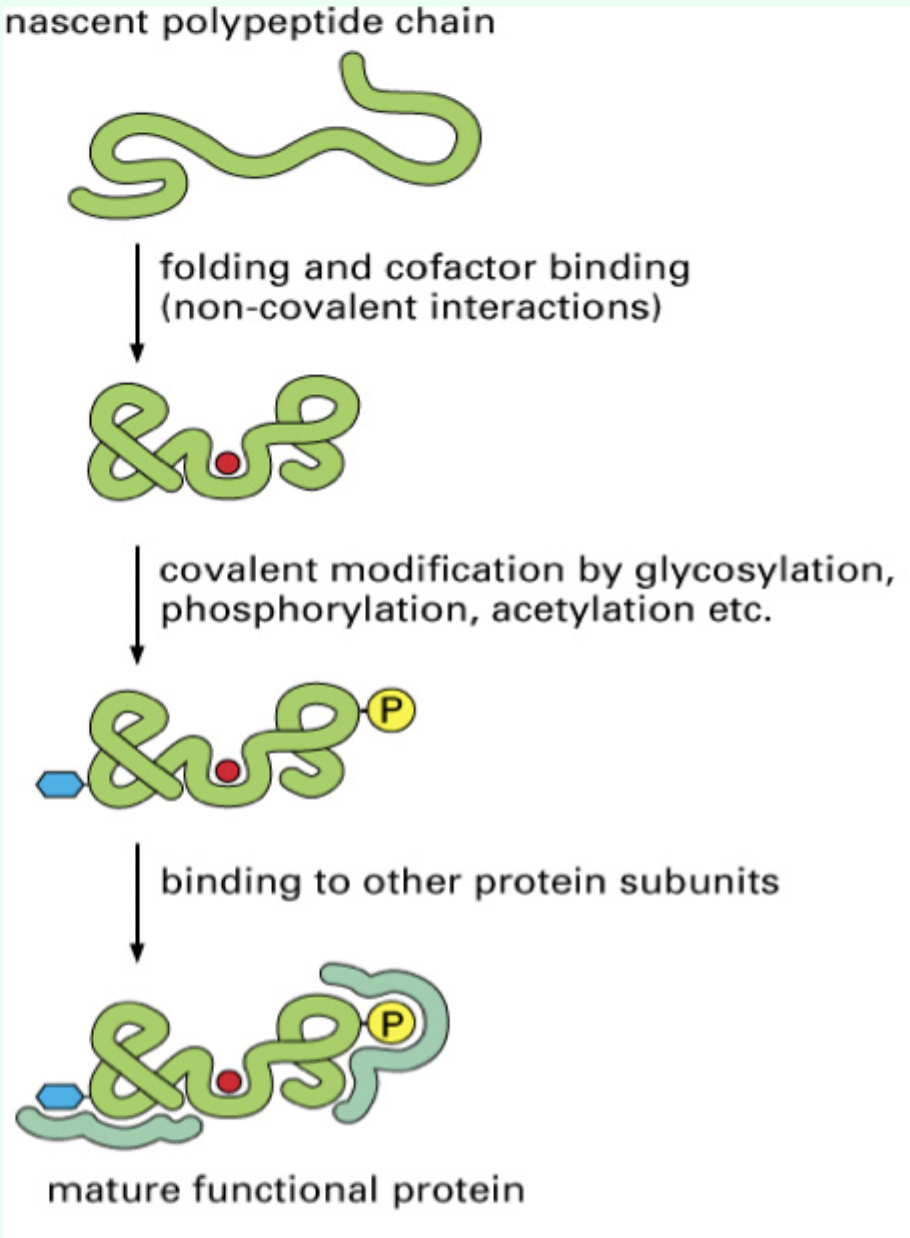
What are the major steps to making a functional protein?
Folding
Cofactor binding
Covalent modification
Translocation
Assembly of multi-subunit complexes
When will a protein fold spontaneously?
When ΔG < 0 (folded state has lower free energy than unfolded).
What thermodynamic equation governs protein folding?
ΔG = ΔH − TΔS
Folding decreases the entropy of the polypeptide chain — so why is folding spontaneous?
Folding releases ordered water molecules, increasing solvent entropy and compensating for the loss of protein entropy.
Is protein folding mostly enthalpically or entropically driven?
Mostly entropically driven.
Why is ΔH often close to zero during folding?
Favorable intramolecular interactions are often offset by loss of favorable protein-water interactions.
What is the global free energy minimum?
The native folded state.
What forms first in the hierarchical model of folding?
Local secondary structures (α-helices and β-sheets).
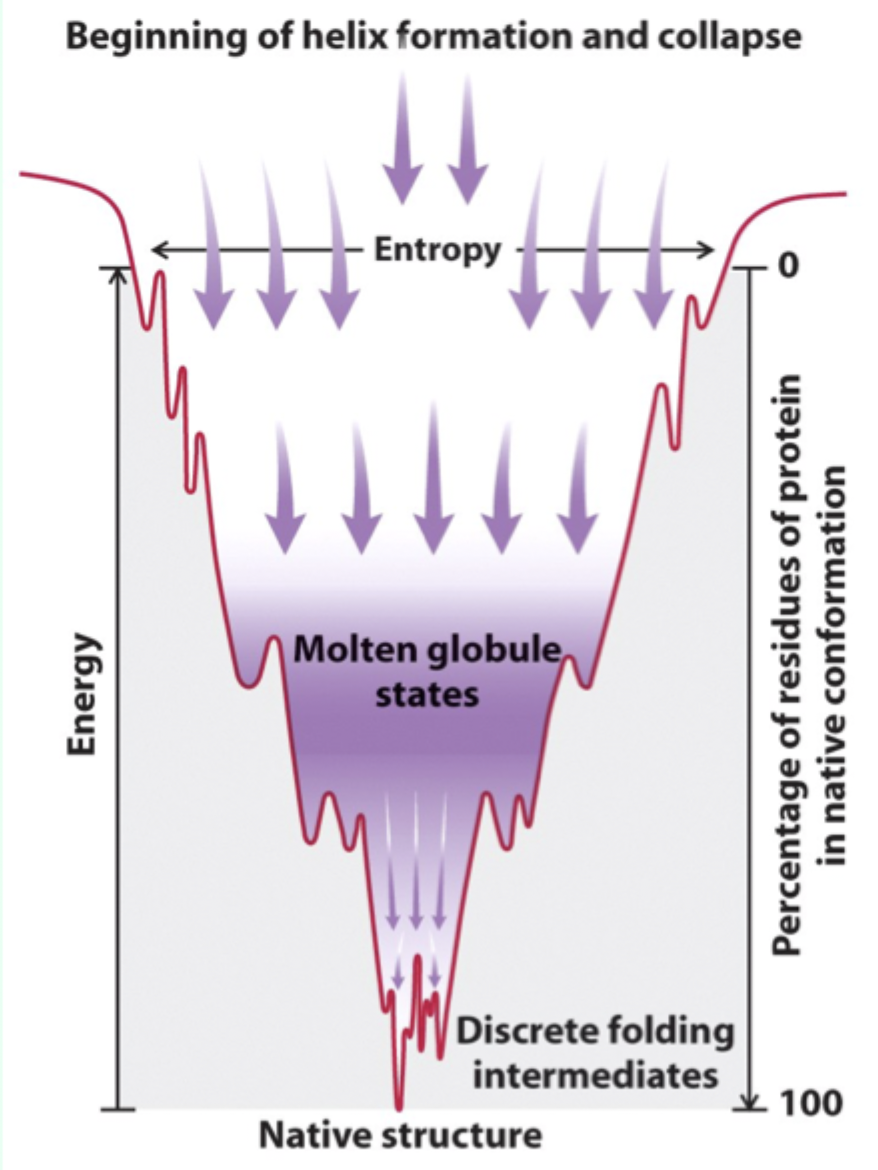
What is a molten globule?
A collapsed intermediate state with much of the secondary and tertiary structure formed.
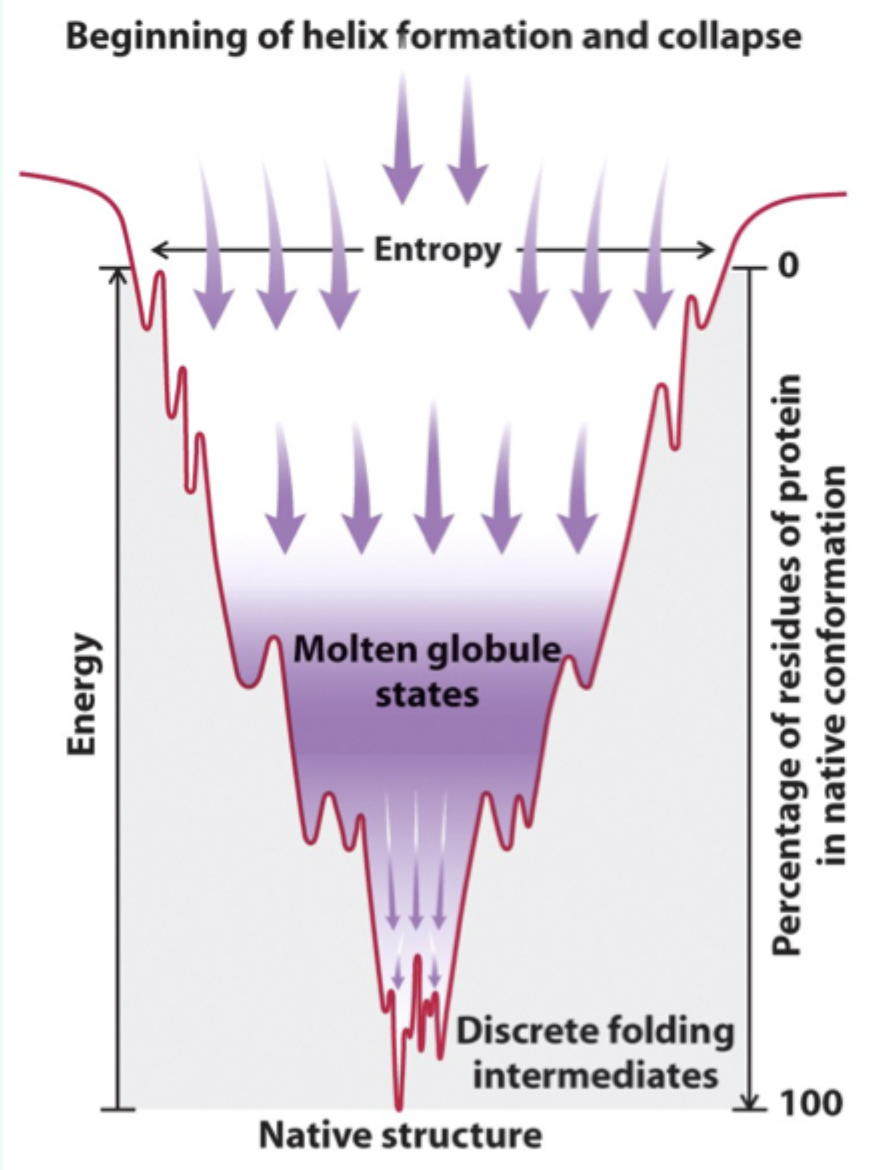
What drives formation of the molten globule?
Rapid hydrophobic collapse to separate hydrophobic residues from water.
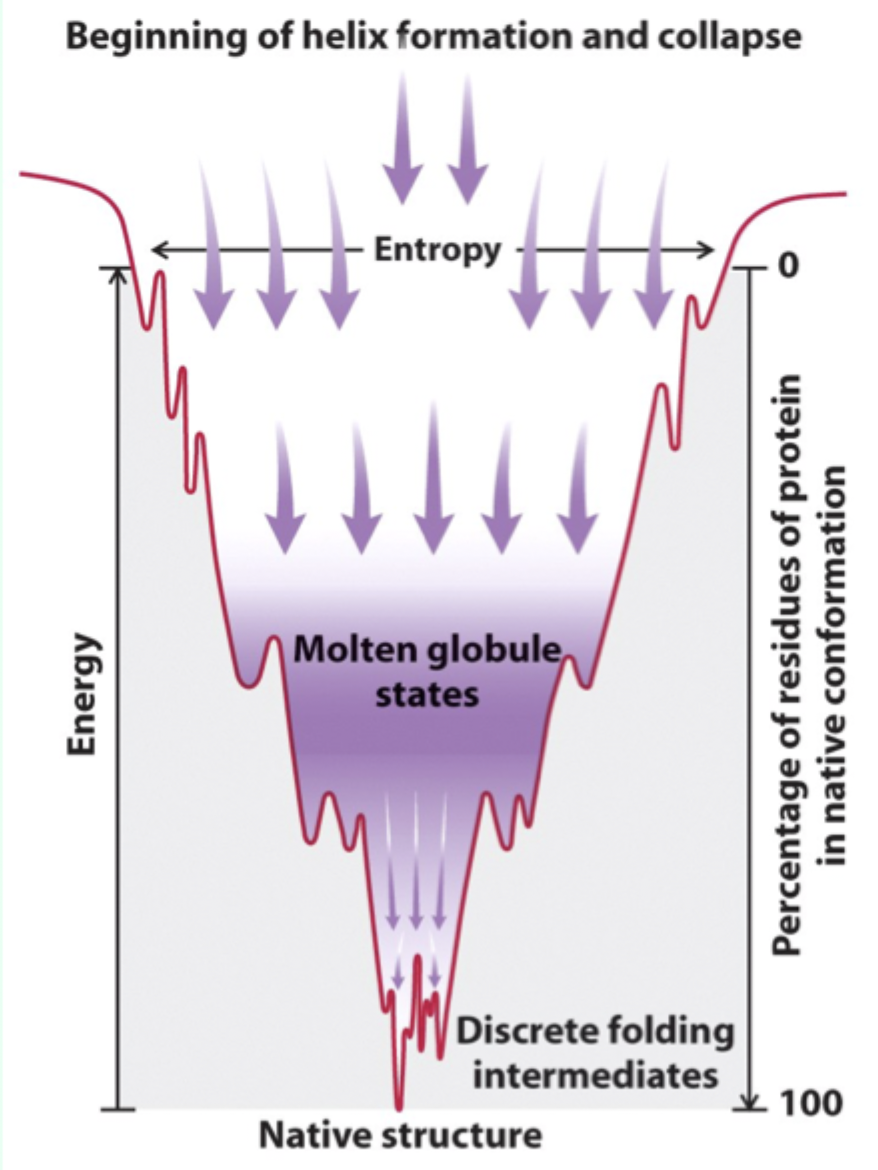
What is another name for the fully folded, functional protein?
The native state.
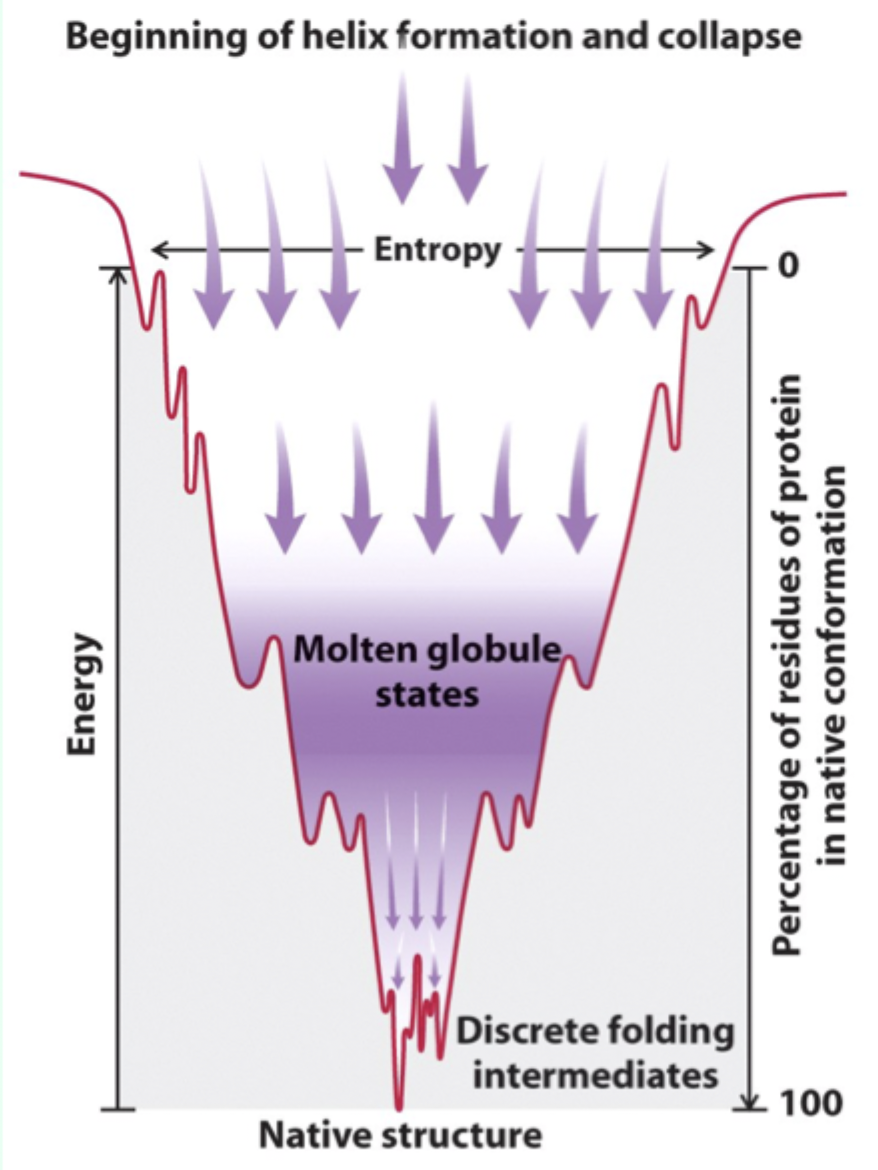
Why do some proteins get “trapped” during folding?
They become stuck in a local free energy minimum and can’t overcome the energy barrier alone.
What do molecular chaperones do?
Assist proteins in folding by helping them overcome energy barriers.
Can most proteins fold on their own?
Only small, soluble globular proteins tend to fold independently.
Do chaperones change the final structure of a protein?
No — they do not dictate structure, they only assist folding.
What are the two major families of molecular chaperones?
Hsp70 and Hsp60
What does Hsp stand for?
Heat shock protein
Why are heat shock proteins upregulated at high temperatures?
Heat denatures (unfolds) proteins, so HSPs help with protein rescue and refolding.
What does Hsp70 bind to?
Hydrophobic patches on nascent polypeptides.
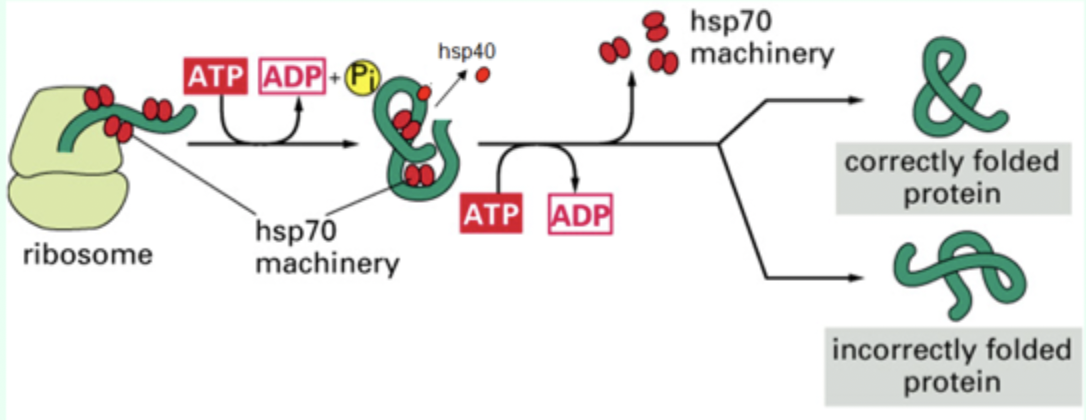
When does Hsp70 bind proteins?
As proteins emerge from the ribosome.
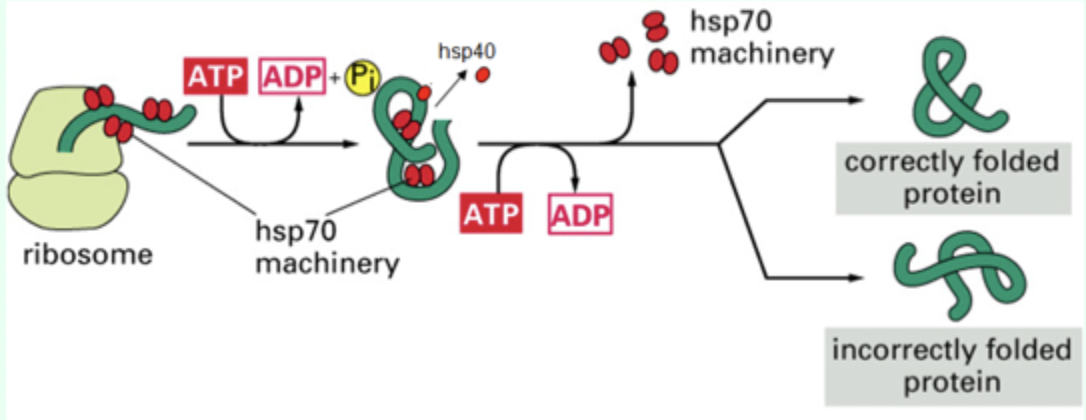
What small protein assists Hsp70?
Hsp40 (DnaJ)
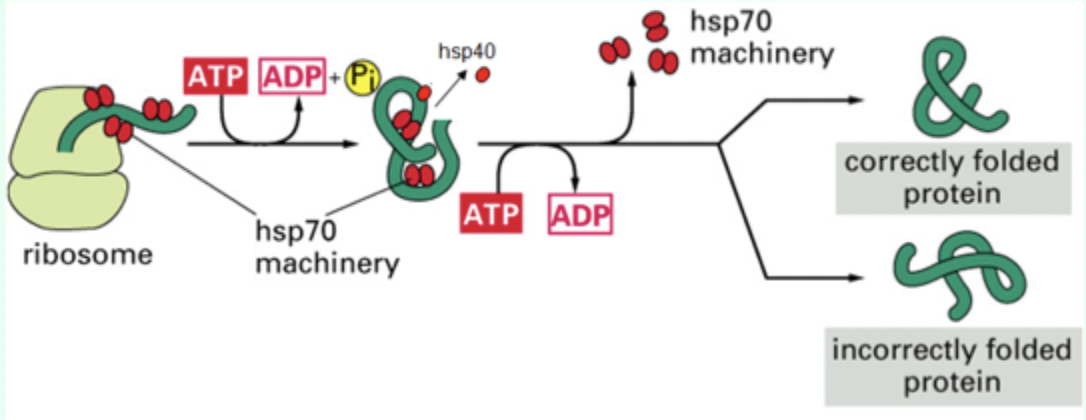
What nucleotide is required for Hsp70 function?
ATP
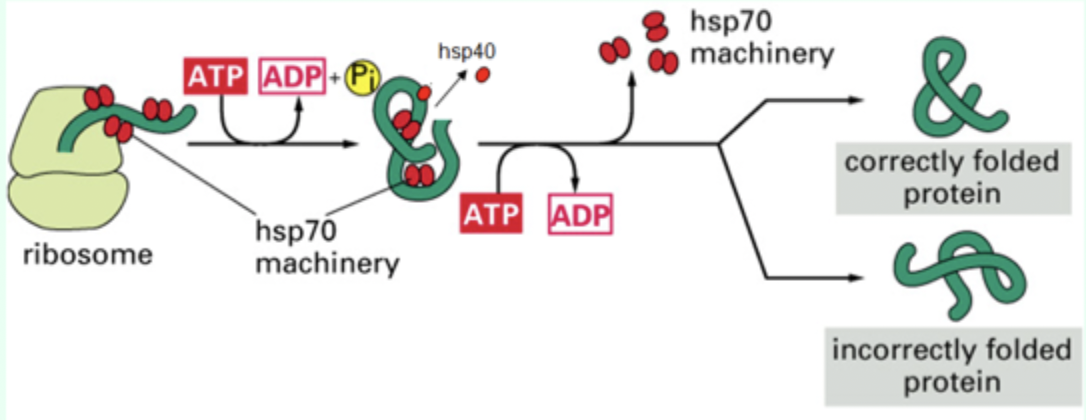
What happens after ATP hydrolysis in Hsp70?
Conformational change locks Hsp70 onto the polypeptide.
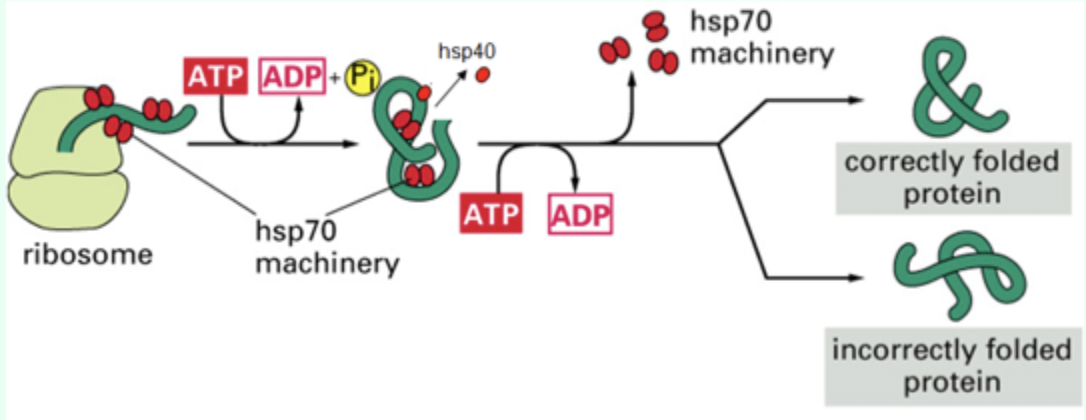
How does Hsp70 release the protein?
Binding of a new ATP molecule.
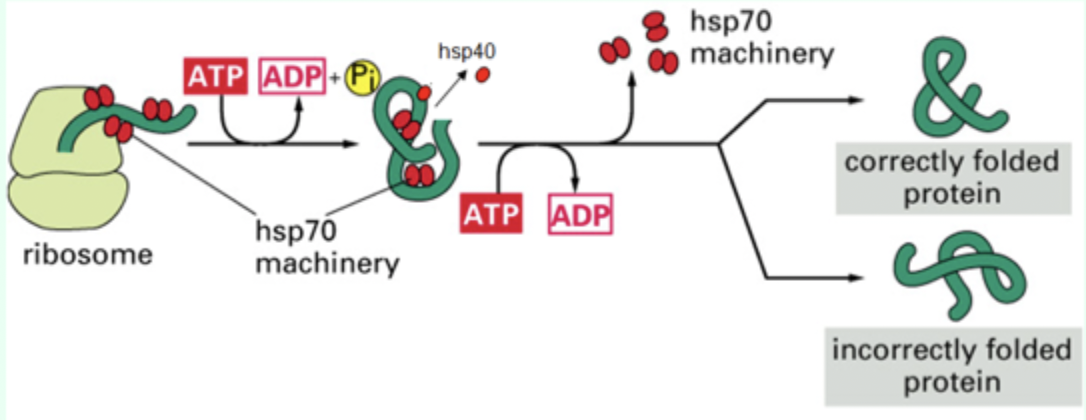
How many binding/release cycles occur?
Multiple cycles at multiple sites.
What type of proteins does Hsp60 act on?
Fully synthesized polypeptides
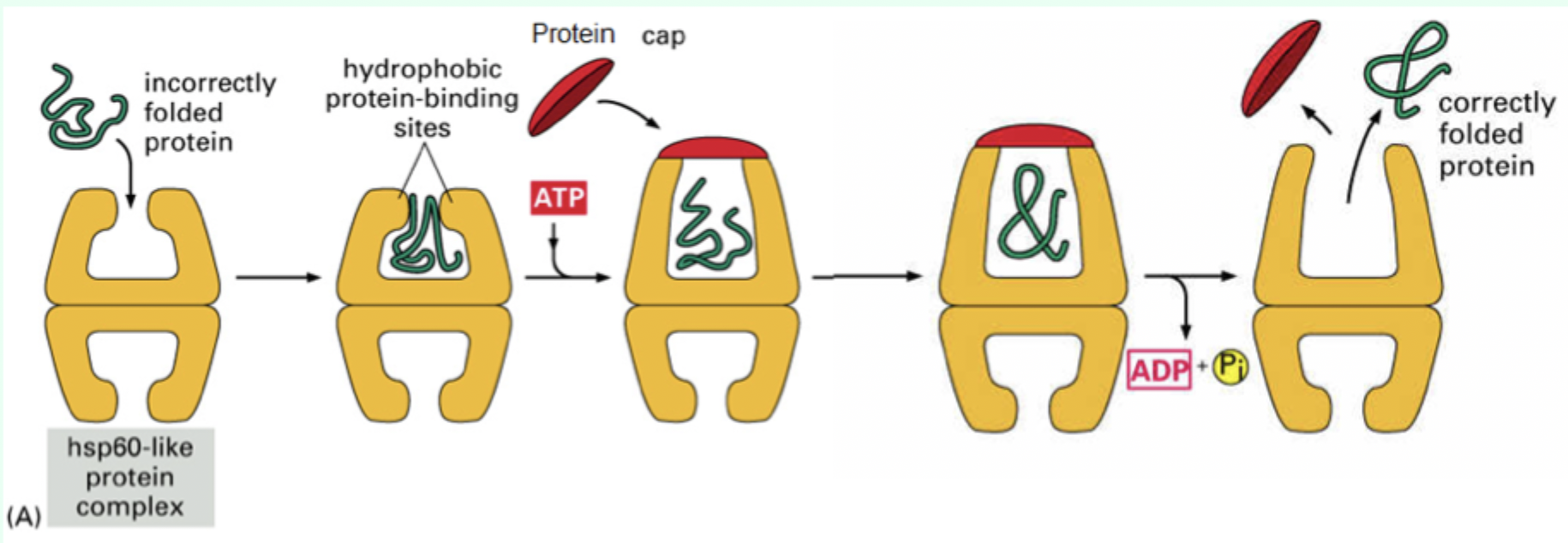
Where does the polypeptide initially bind in Hsp60?
Hydrophobic regions around the rim
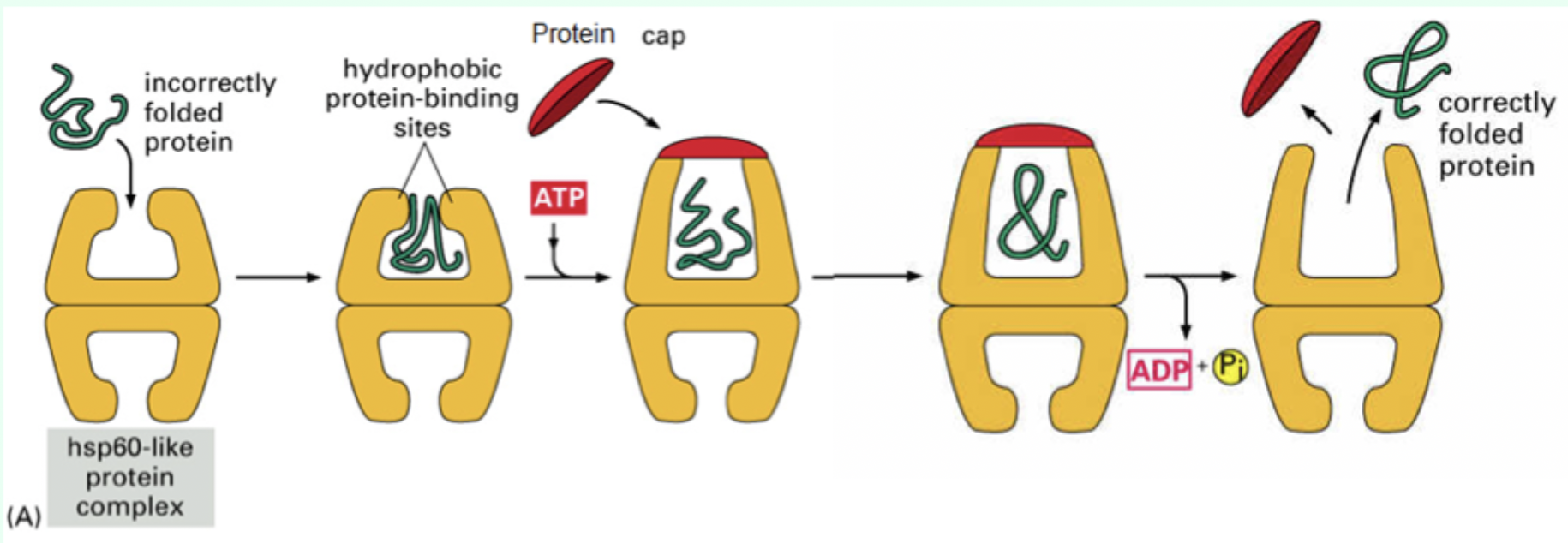
How many ATP molecules bind Hsp60?
7 ATP molecules
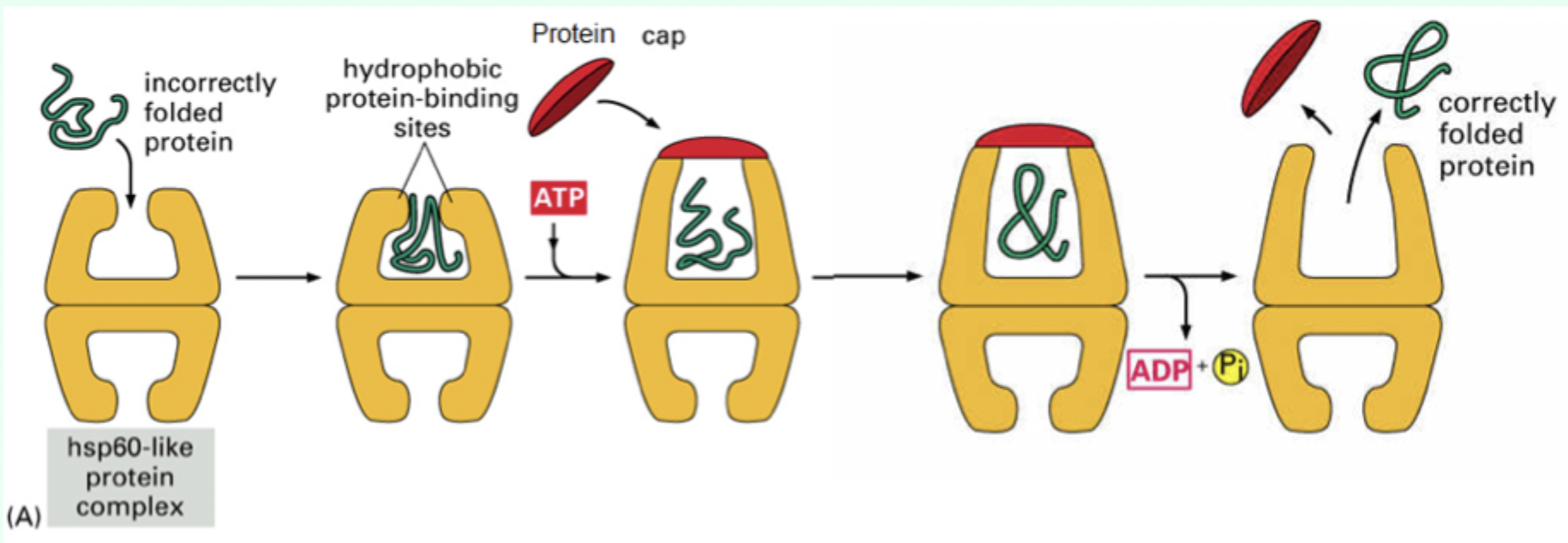
What is the role of the protein “cap”?
Confines the protein inside the chamber to allow folding.
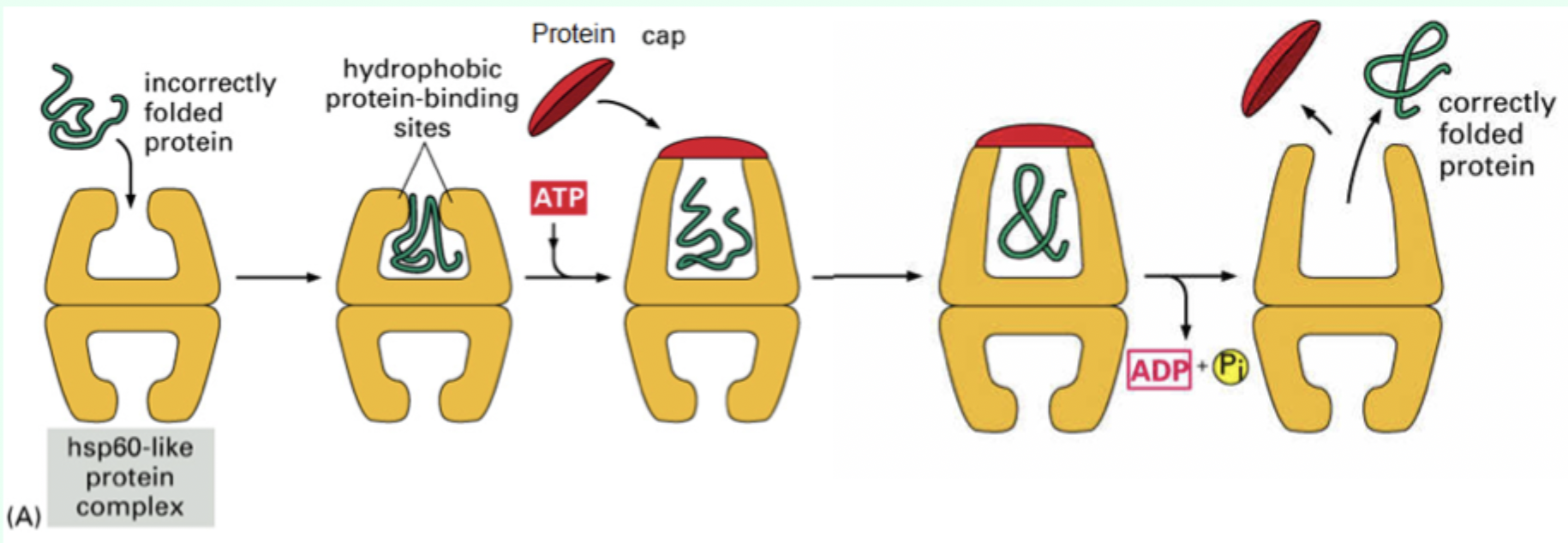
What environment does the Hsp60 chamber provide?
A mixed hydrophobic/hydrophilic environment.
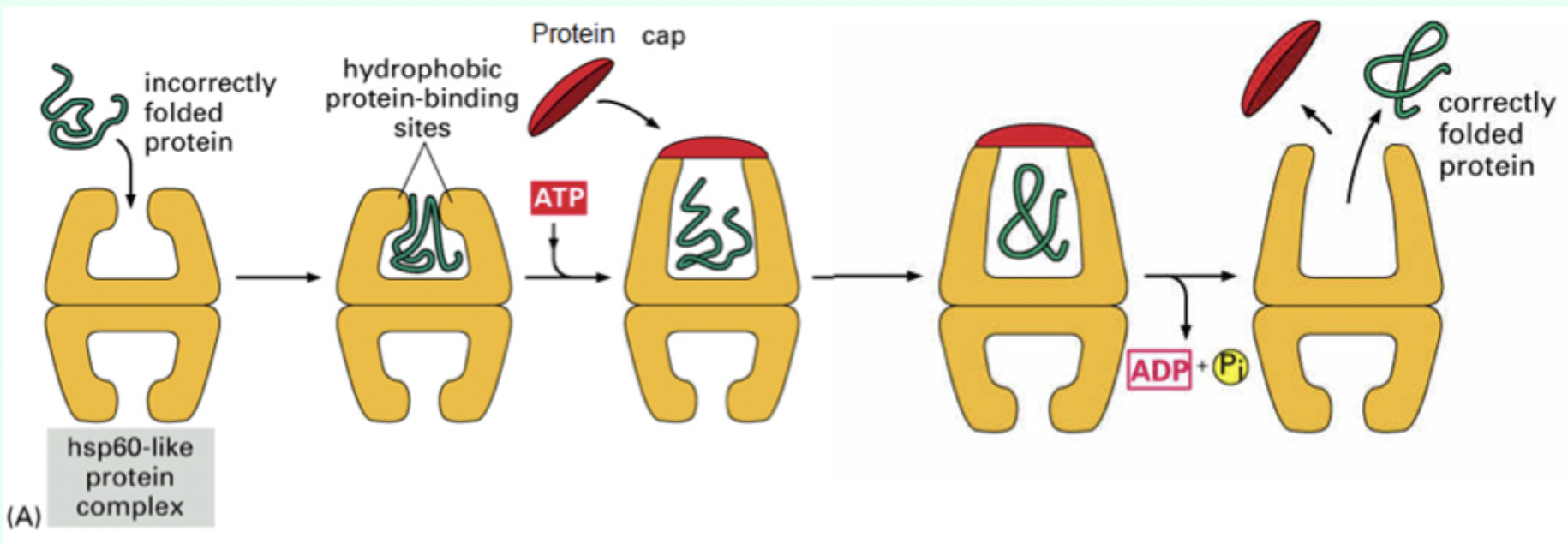
After ATP hydrolysis, what happens to the protein?
It is released whether fully folded or not.
Maximum protein size that Hsp60 can accommodate?
~57 kDa (~520 residues)
What does protein disulfide isomerase (PDI) do?
Shuffles incorrect disulfide bonds until the correct ones form.
What does peptidyl-prolyl cis-trans isomerase (PPI) do?
Catalyzes cis-trans isomerization of proline residues.
What do proteases remove during protein maturation?
Signal sequences
Prosequences
Secretion signals
What is protein denaturation?
Loss of compact 3D structure → loss of function.
How does heat denature proteins?
Disrupts non-covalent interactions.
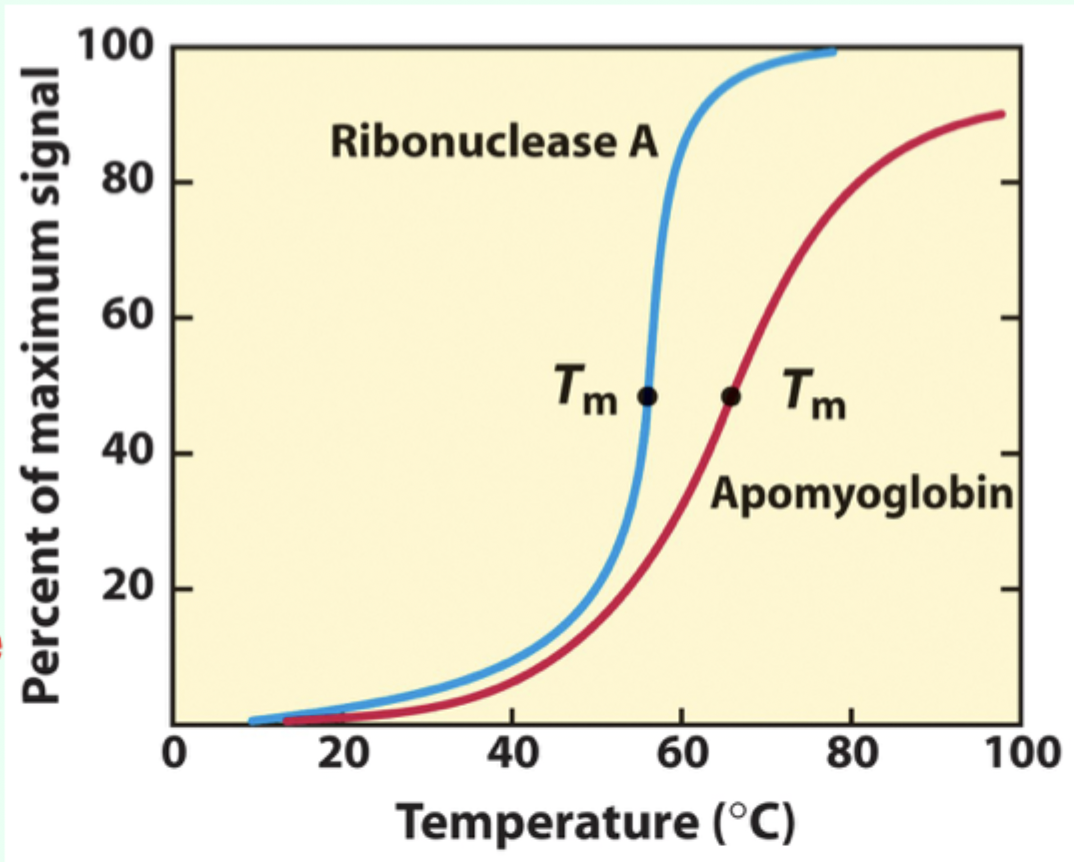
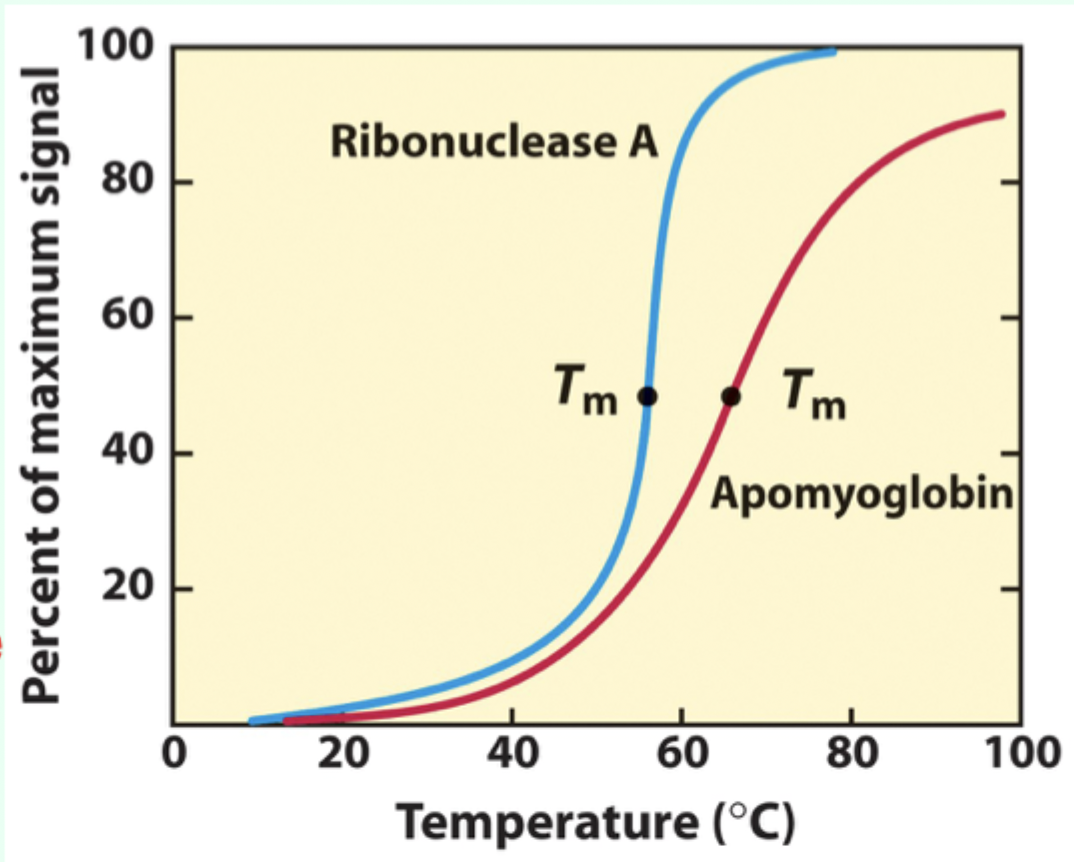
What does a sharp unfolding transition indicate?
Cooperative unfolding
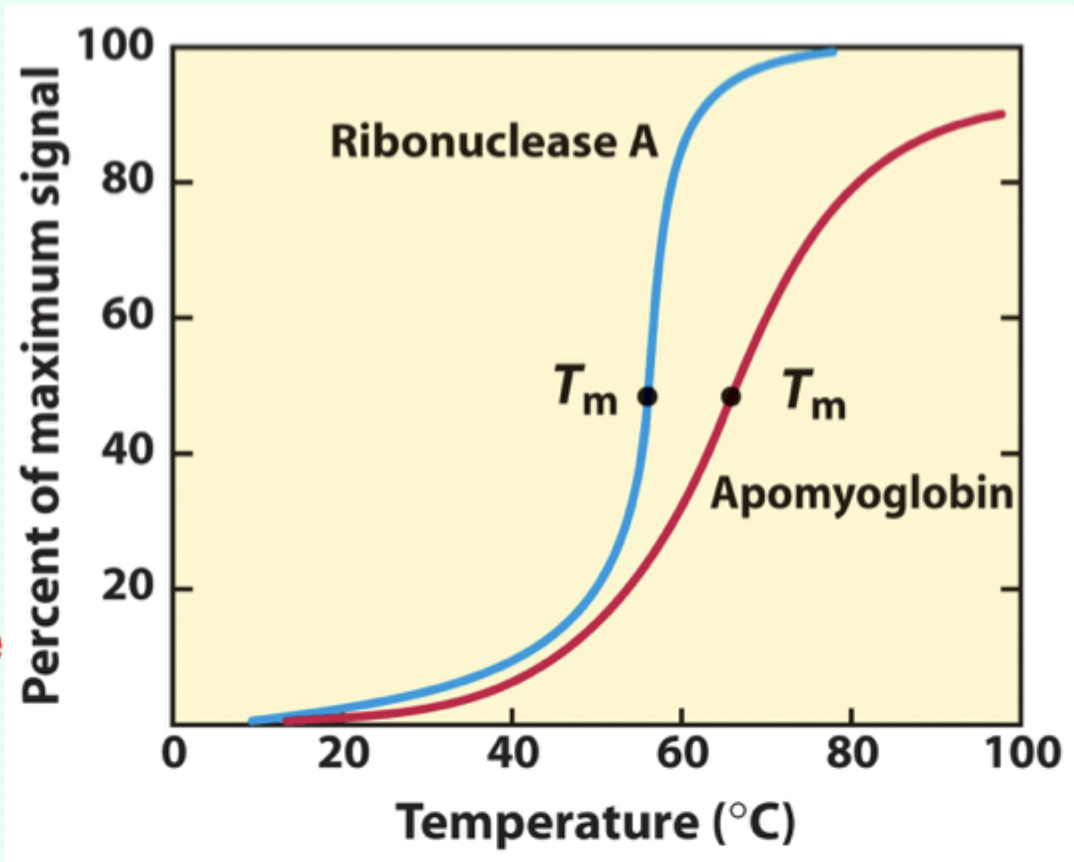
What is Tₘ (melting temperature)?
The midpoint temperature where denaturation occurs.
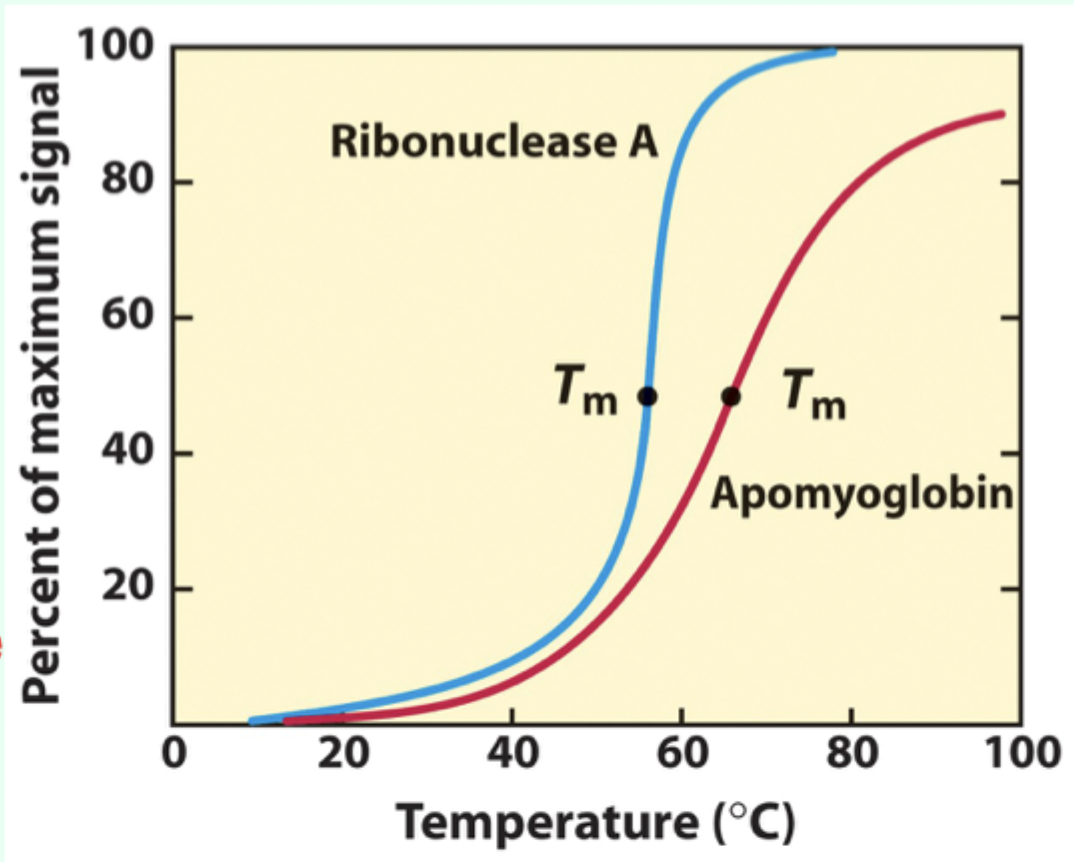
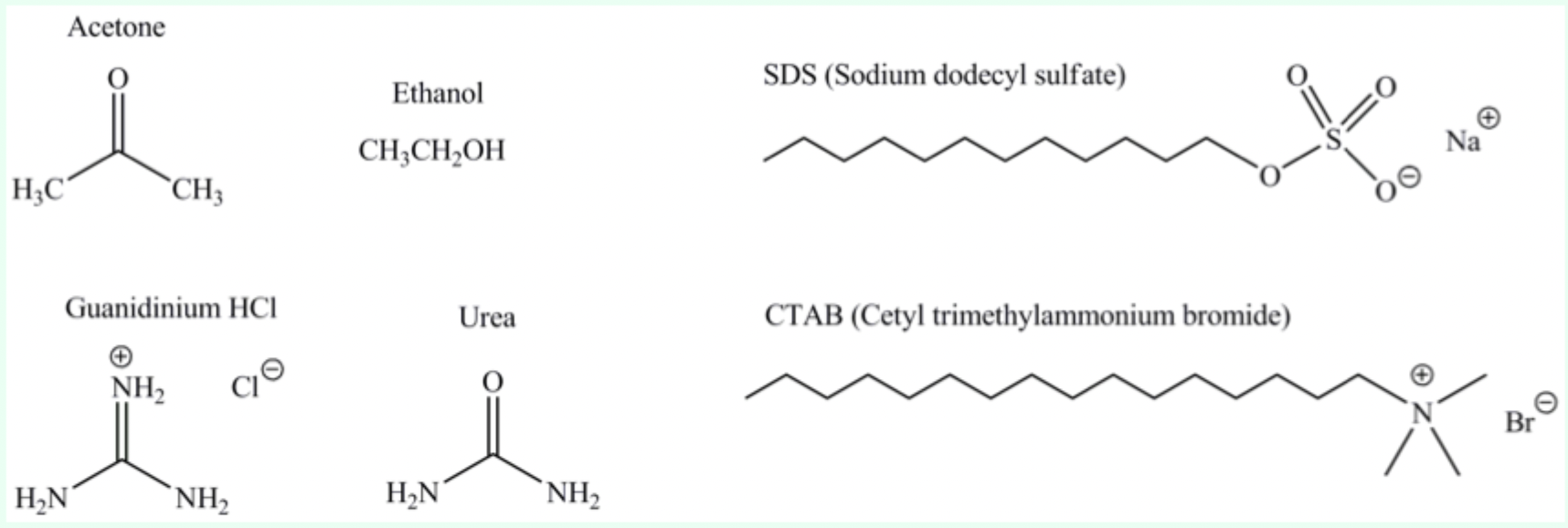
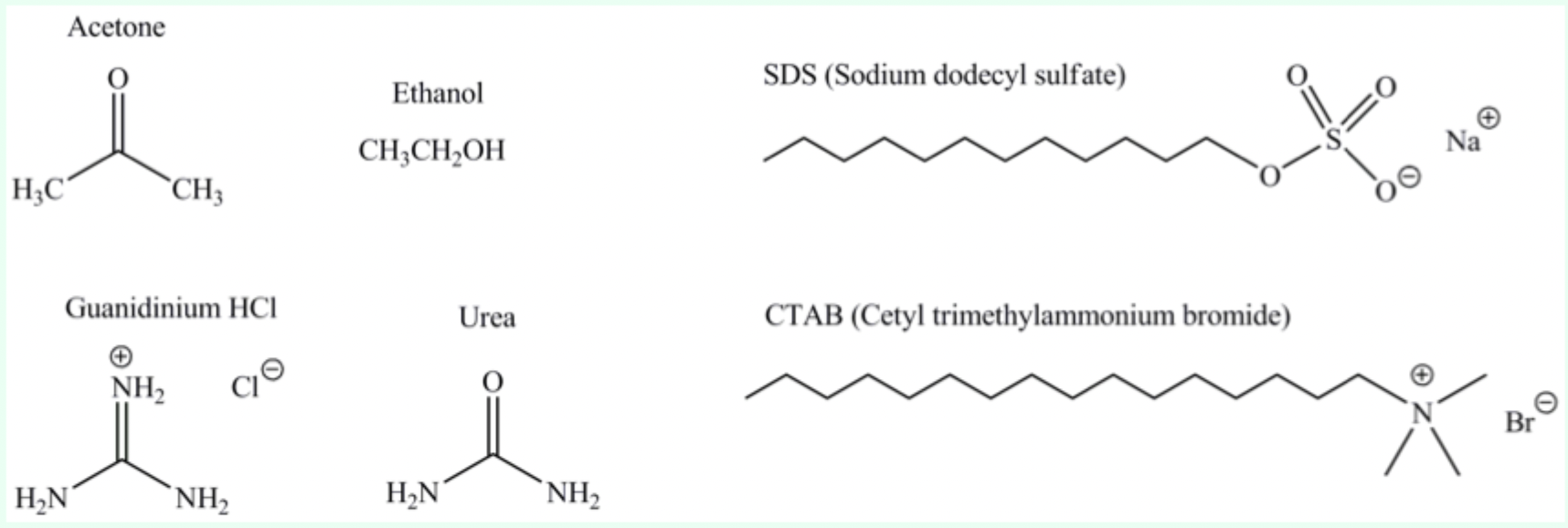
What do urea, detergents, and organic solvents disrupt?
The hydrophobic core of proteins.
How does pH denature proteins?
Changes net charge → electrostatic repulsion + hydrogen bond disruption.
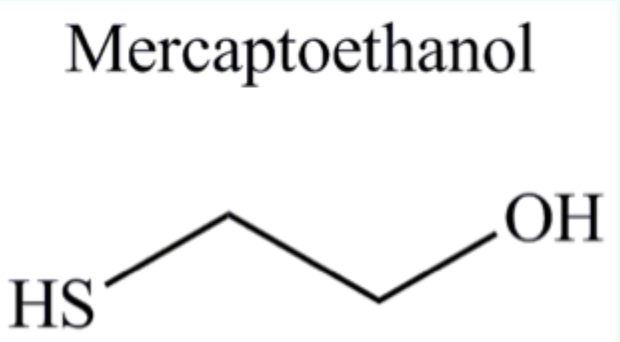
What does β-mercaptoethanol do?
Reduces disulfide bonds between cysteine residues.
Can proteins refold after removal of urea and mercaptoethanol?
Yes — small proteins can renature.
What is the approximate weight of a 100-residue protein?
Equal to the number of residues × 110