Cell Biology Exam 3
1/101
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
102 Terms
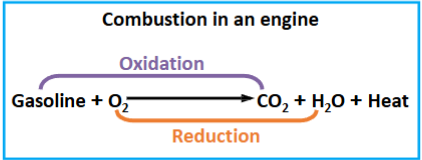
Combustion
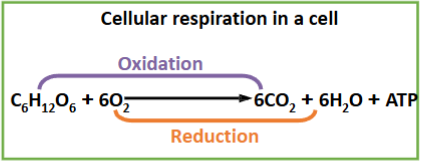
Cellular respiration
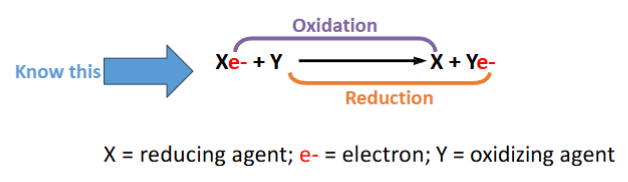
What does it mean if a molecule is oxidized or reduced?
Oxidized – when electrons are taken away from a molecule it is oxidized
Reduced – when a molecule accepts electrons it is reduced
What is cellular respiration?
the process of oxidation of fuel molecules coupled with the production of ATP in the cell.
3 major steps:
Glycolysis – in the cytoplasm
The citric acid cycle (Krebs cycle) - in the mitochondrial matrix
The electron transport and ATP synthesis – in the inner mitochondrial membrane
Max of 38 ATP molecules can be produced per glucose molecule
What is the location, starting material, and net result for the first step of cellular respiration?
Glycolysis
Occurs in they cytoplasm
Net result: Glucose = 2 Pyruvate + 2 ATP + 2 NADH
Glycolysis is anaerobic = meaning no oxygen is required
What is the location, starting material, and net result for the second step of cellular respiration?
Citric Acid Cycle
Occurs in the mitochondrial matrix
Acetyl CoA is completely oxidized to CO2 through 8 reactions in mitochondrial matrix
Electrons derived from these reactions are carried by NADH and FADH2
This can produce 2 ATP molecules
Net result: One turn of the cycle produces three NADH, one GTP, and one FADH2, and releases two molecules of CO2
What is the location, starting material, and net result for the third step of cellular respiration?
The Electron transport chain (ETC)
Occurs in the inner mitochondrial membrane
The electrons carried by NADH and FADH2 are passed to the ETC, which contains more than 40 components.
Most of the ETC components are membrane proteins embedded in the inner membrane of the mitochondria
Electrons from NADH and FADH2 are transported from higher energy level to lower energy level components.
Define anaerobic and aerobic
Anaerobic respiration:
Pyruvate is converted to lactic acid (in muscle cells) or ethanol (in yeast)
Yields Lactic acid or Ethanol
-O2 Fermentation
Aerobic respiration:
Pyruvate generated from glycolysis is transported to mitochondria and converted to acetyl CoA
Yields Acetyl CoA (enters TCA)
+O2 Citric acid Cycle
Oxidative phosphorylation
The energy released from electron transport is used to create H+ gradient between the matrix and inner membrane space.
The H+ gradient drives ATP synthesis
~34 ATP molecules are produced
Oxidative Phosphorylation – ATP synthase
ATP synthase is a huge protein complex located in the inner membrane of mitochondria. It creates a passage for H+ and converts ADP to ATP
ATP is produced by chemiosmosis:
Chemiosmosis – the movement of ions across a semipermeable membrane bound structure, down their electrochemical gradient.
How does ATP synthase compare to ATPase (lecture 11)?
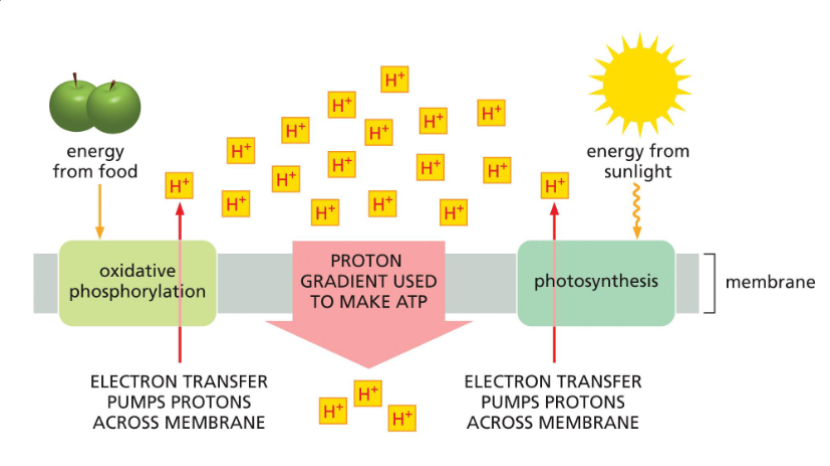
1. Energy from electrons released from an electron transport chain give energy to membrane proton pumps
2. Proton pumps move protons across the membrane. This creates a proton (H+) gradient (high proton environment on one side of the membrane and a low proton environment on the other side).
3. ATP synthase carries protons across the membrane, and uses the energy released from this to make ATP
Understand and describe membrane-based energy production
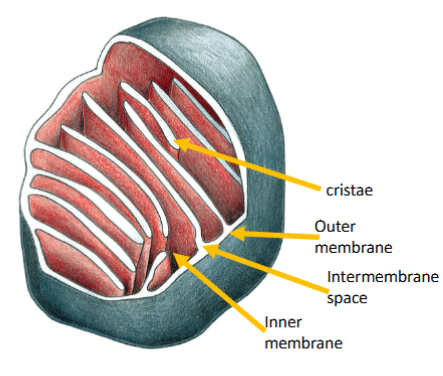
Describe the structure and function of the mitochondria (including be able to label mitochondria structures)
A mitochondrion has outer and inner membranes separated by an inter-membrane space.
The inner membrane folds into cristae
The space enclosed by the inner membrane is the matrix.
The major function of mitochondria is to produce ATP through oxidation of fuel molecules.
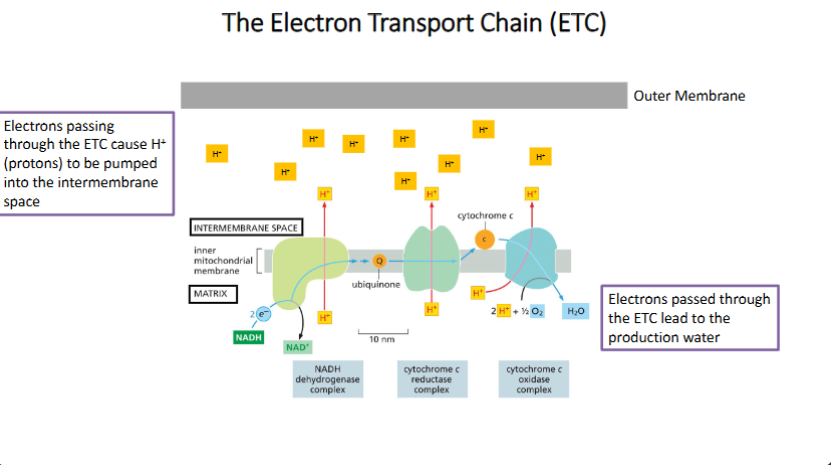
Know, be able to draw, and apply to situations how the mitochondrial electron transport chain works (including H+ flow direction and electron movement)
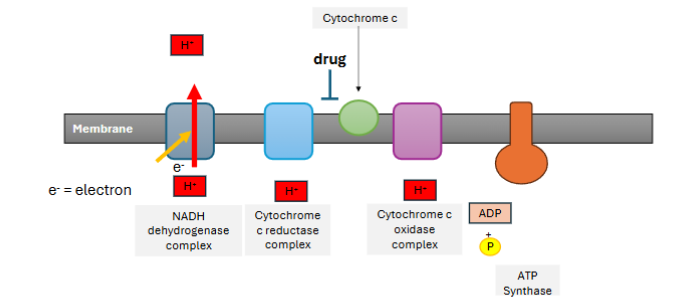
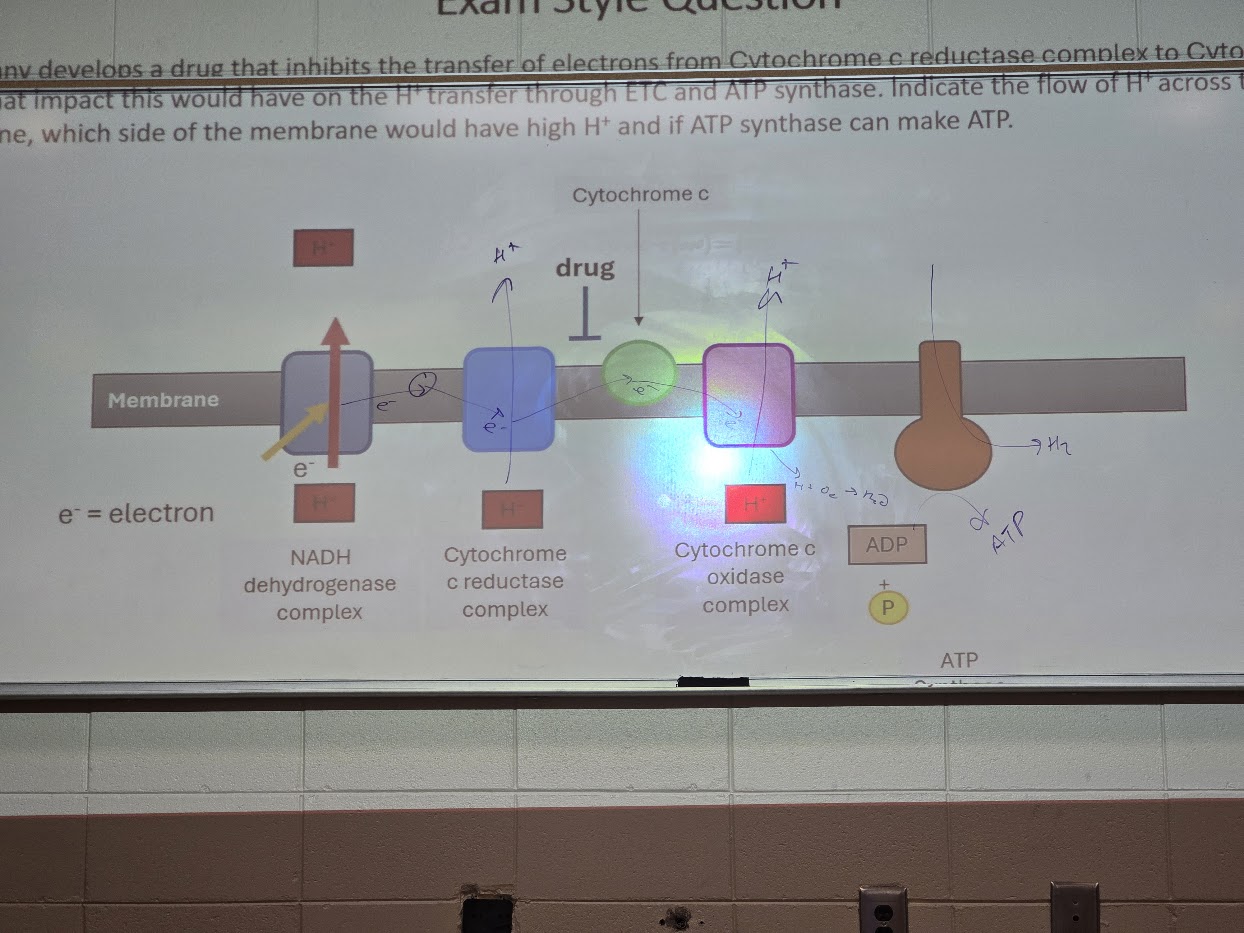
A company develops a drug that inhibits the transfer of electrons from Cytochrome c reductase complex to Cytochrome C. Draw what impact this would have on the H+ transfer through ETC and ATP synthase. Indicate the flow of H+ across the membrane, which side of the membrane would have high H+ and if ATP synthase can make ATP.
Redox reactions
Chemical reactions involve reduction and oxidation.
Describe the structure of mitochondrion. Why is it important to have intermembrane space and cristae?
Cristae - extensions of the inner membrane
Contain and organize the electron transport chain and the ATP synthase.
Increase the surface area of the inner membrane.
Separate the matrix from the intermembrane space, which stores H+ and allows H+ gradient formation.
List 3 major differences in Energy Harvest in an Engine vs in a Cell
1. The Method of Release
Engine: Combustion. A single, rapid, high-temperature reaction (a "burn") that releases energy all at once.
Cell: Stepwise Oxidation. A series of small, enzyme-controlled steps at room/body temperature that releases energy gradually.
2. Form of Energy Used
Engine: Heat/Expansion. Converts chemical energy into heat, which expands gas to push a piston (mechanical work).
Cell: Chemical Coupling. Converts chemical energy from food into a Proton Gradient (electrochemical energy) to produce ATP.
3. Storage and Transport
Engine: Direct/Immediate. Energy must be used the moment it’s released; there is no "mobile battery" molecule.
Cell: Portable (ATP). Energy is captured in ATP molecules, which can travel to different parts of the cell to power specific reactions later.
Describe how cells obtain energy from food in 3 stages (where does each stage take place?)
Stage 1: Glycolysis
Location: Cytosol (the fluid inside the cell, outside the mitochondria).
The Process: A single molecule of glucose (6 carbons) is broken down into two molecules of pyruvate (3 carbons each).
Energy Yield: A small amount of ATP is made directly, and some high-energy electrons are captured by NADH.
Note: This is the only stage that doesn't require oxygen (anaerobic).
Stage 2: The Citric Acid Cycle (Krebs Cycle)
Location: Mitochondrial Matrix (the innermost compartment of the mitochondria).
The Process: Pyruvate is converted into Acetyl-CoA and enters the cycle. It is completely broken down into $CO_2$, which you eventually breathe out.
Energy Yield: More NADH and another electron carrier, $FADH_2$, are loaded up with "excited" electrons to be used in the final stage.
Stage 3: Oxidative Phosphorylation
Location: Inner Mitochondrial Membrane (specifically on the cristae or folds).
The Process: This is the "big harvest." High-energy electrons from NADH and $FADH_2$ are passed down the Electron Transport Chain. This flow of electrons pumps protons across the membrane to create a gradient.
Energy Yield: Protons rush back through a protein called ATP Synthase (like water through a turbine), generating the vast majority of the cell's ATP.
Note: Oxygen is the "final electron acceptor" here; it grabs the electrons at the end to form water ($H_2O$).
Lactic Acid Fermentation
This is common in animal muscle cells (like yours during a heavy workout) and certain bacteria (like those used to make yogurt).
The Process: Pyruvate is reduced directly by $NADH$ to form Lactate (Lactic Acid). There is no release of $CO_2$.
The Goal: By turning Pyruvate into Lactate, the cell turns $NADH$ back into $NAD^+$. This $NAD^+$ goes back to the beginning of Glycolysis to keep making a tiny bit of ATP.
Chemical Logic:
\text{Pyruvate} + \text{NADH} \rightarrow \text{Lactate} + \text{NAD}^+
Alcohol Fermentation
This occurs in yeast and many types of bacteria. This process is the foundation of baking and brewing.
The Process: This happens in two steps:
Decarboxylation: $CO_2$ is removed from Pyruvate, converting it into a 2-carbon molecule called Acetaldehyde.
Reduction: $NADH$ reduces Acetaldehyde into Ethanol (Ethyl Alcohol).
The Goal: Just like in lactic acid fermentation, the main point is to regenerate $NAD^+$ so Glycolysis can continue.
Chemical Logic:
\text{Pyruvate} \rightarrow \text{CO}_2 + \text{Acetaldehyde} \xrightarrow{\text{NADH}} \text{Ethanol} + \text{NAD}^+
Know the locations of protein synthesis and understand how signaling patches/sequences control protein localization.
Proteins are directed to their destinations by signal sequences or signal patches (like zip code)
The information required to transport a protein from cytosol to an organelle is contained in a signal sequence or signal patch.
signal sequence – a short sequence of amino acids (15-60 aa) – signal sequence
signal patch – formed from amino acids in different segments of the protein.
Cytosolic proteins
are synthesized by free ribosomes. They stay in the cytosol for their functions.
Organelle proteins
are synthesized by free ribosomes in cytosol, then transported to the specific organelles.
Secreted and membrane proteins
are synthesized by ribosomes on the ER, then, transported to the cell membrane, or secreted out of the cell.
Gated transport
between the cytosol and the nucleus
The transport of molecules between the nucleus and the cytosol through nuclear pores.
All nuclear proteins (example - histones) are synthesized in the cytosol and transported into the nucleus through nuclear pores.
The molecules produced in the nucleus (example - mRNA) are exported to the cytosol also through nuclear pores.
The transport of molecules between the nucleus and the cytosol is bi-directional.
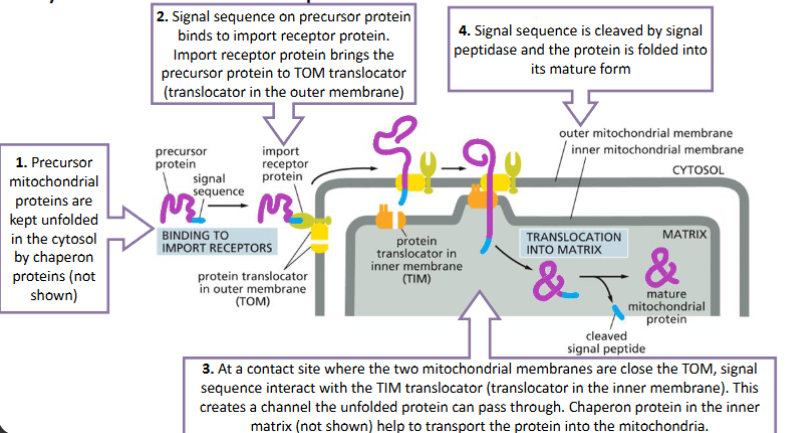
Transmembrane transport - mitochondria and chloroplast
from cytosol to mitochondria, chloroplasts, peroxisomes, and the ER
Transporting proteins across the membrane need a group of proteins, called translocators, on the membrane.
The transported proteins are kept unfolded to go through the membrane (different from nuclear pore transport).
Depending on their destinations, transported proteins may need more than one signal sequences.
Transport proteins to mitochondria and chloroplasts follows the similar mechanism
The signal sequence required to import a protein to mitochondria or chloroplasts is usually at the N-terminus.
The protein to be imported is called precursor protein since the signal sequence will be removed by protease
Transmembrane transport - Proteins into the Mitochondrial Matrix
Two sequences needed – signal sequence and stop-transfer sequence
Protein translocation is initiated by the same way as for mitochondrial matrix proteins
At the inner membrane, the protein translocation is blocked by the stop-transfer sequence. The proteins is then anchored to the inner membrane with the stop-transfer sequence (predict the property of this sequence).
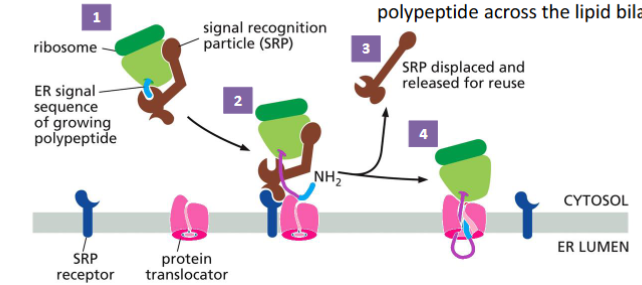
Vesicular transport
transport between components (the ER, Golgi, and lysosomes) and secretion out of the cell
ER is a network of interconnected internal membranes.
ER consists of closed, flattened membrane sacs called cisternae
Rough ER - has bound ribosomes and produces cell membrane and secretory proteins.
Smooth ER - lacks ribosomes; it synthesizes lipid and steroids. Import of proteins to ER is co-translational translocation (translation and translocation occurs simultaneously)
The ER signal sequence is a short sequence at the N-terminus that contain a stretch of hydrophobic amino acid residues.
The signal sequence is cleaved while the protein is still growing on the ribosome.
After translation is completed, the proteins are modified (such as glycosylation) and delivered to different locations.
The signal recognition particle (SRP) binds to the ER signal and the ribosome. This slows translation.
The SRP-ribosome complex binds to the SRP receptor in the ER membrane.
The SRP is released, and the ribosome passes from the SRP receptor to a protein translocator in the ER membrane.
Protein synthesis resumes, and the translocator starts to transfer the growing polypeptide across the lipid bilayer.
The signal sequence is removed by a signal peptidase – an integral membrane protein that cleaves the signal peptide. The protein is released to the ER lumen.
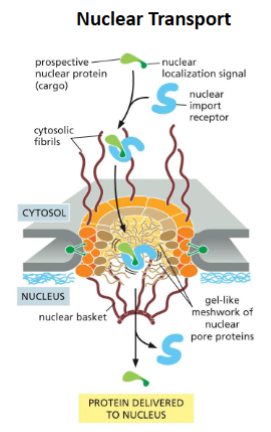
Protein transport to the nucleus
Most nuclear proteins are too large to passively enter the nucleus
Nuclear proteins contain a nuclear localization signal (NLS)
NLS features:
Can be anywhere in the protein
Can be a single sequence or two sequences (bipartite)
Nuclear protein with NLS (cargo) is recognized and interacts with the nuclear import receptor
Receptor interacts with nuclear pore cytosolic fibrils
Receptor-cargo complex move through gel-like nuclear pore
Entry into the nucleus triggers receptor to release cargo
Cargo protein is transported into nucleus fully folded and NLS is not cleaved from protein
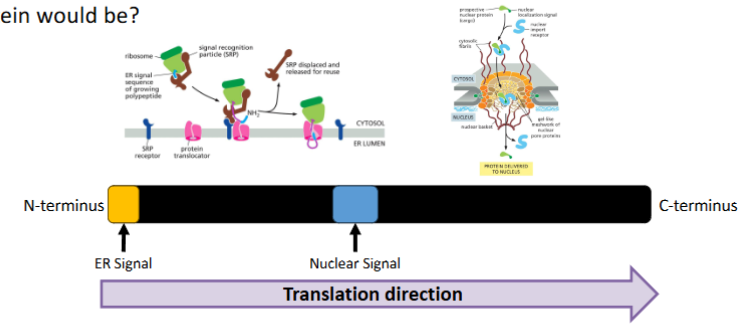
Consider a protein that contains an ER signal sequence at is N-terminus and a nuclear localization sequence in its middle. What do you think the fate of this protein would be?
The Rule: ER signals are usually co-translational (happen during synthesis). Nuclear signals are post-translational (happen after synthesis).
The Winner: The ER signal "wins" because it is synthesized first and moves the protein into a membrane-bound compartment before the rest of the protein is even finished.
Nucleus protein transport
Site of Synthesis: Free ribosomes in the cytosol
Location of Signal: Usually internal (anywhere in chain)
Timing of Transport: Post-translational (after synthesis)
Protein State: Folded (can pass through large pores)
Signal Fate: Retained (not removed)
Mitochondria/chloroplast protein transport
Site of Synthesis: Free ribosomes in the cytosol
Location of Signal: N-terminus
Timing of Transport: Post-translational (after synthesis)
Protein State: Unfolded (must be "threaded" through translocators)
Signal Fate: Cleaved (removed by signal peptidase)
Endoplasmic reticulum protein transport
Site of Synthesis: Ribosomes attached to the Rough ER
Location of Signal: N-terminus
Timing of Transport: Co-translational (during synthesis)
Protein State: Unfolded (threaded through as it's being made)
Signal Fate: Cleaved (removed by signal peptidase)
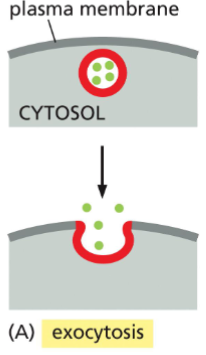
Exocytosis
vesicles containing proteins or other molecules fuse with the plasma membrane, causing vesicle contents to be secreted from cell
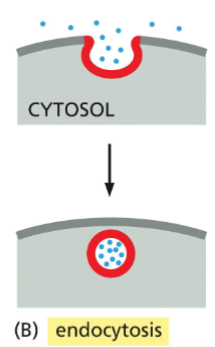
Endocytosis
internalization of proteins/molecules from outside the cell by plasma membrane budding, creating vesicles.
Transporting proteins to the cell exterior: Exocytosis (secretion)
Proteins travel through:
The ER – proteins are covalently modified (e.g., disulfide bonds are formed, proteins are glycosylated).
Golgi apparatus – proteins further modified, and sorted for transport to final destination (e.g., plasma membrane or lysosome).
Secretory vesicles – travel to the plasma membrane for exocytosis (and secretion)
(know the labels and arrows in the attached image)
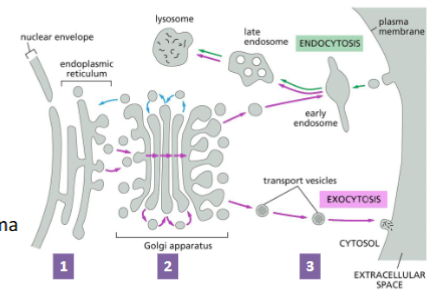
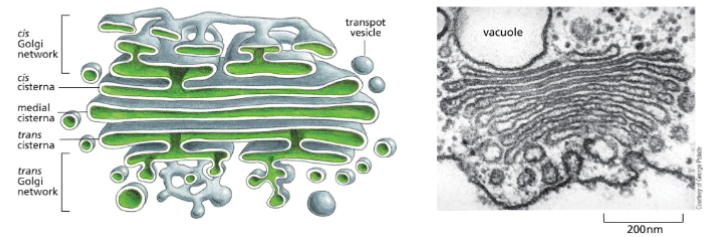
The Golgi Apparatus Structure and Function
A stack of flattened membrane-enclosed structures called cisternae
Has 3 defined regions – the cis (adjacent to ER), medial (in middle) and trans region (pointing towards plasma membrane).
The Golgi processes, modifies (adds more sugars to glycoproteins), and sorts secreted proteins and membrane proteins.
Exocytosis – Constitutive Secretion
Constitutive Secretion:
occurs in all eukaryote cells
continually releases soluble proteins from the cell
supplies the plasma membrane with newly synthesized lipids and proteins
transport vesicles fuse with plasma membrane when it arrives there
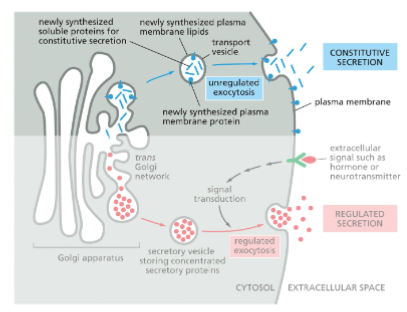
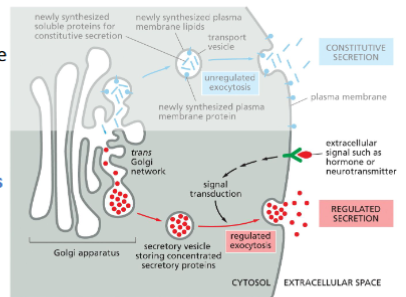
Exocytosis – Regulated Secretion
Regulated Secretion
Specialized secretory cells (e.g., β cells in the pancreas, neuronal cells)
Proteins are concentrated and stored in secretory vesicles
Secretory vesicles are stored near plasma membrane and only fuse when cell receives signals, such as hormones and neuron transmitters
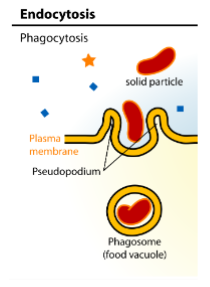
Endocytosis - Phagocytosis
internalization of bacteria, damaged cells, or other large particles by certain cells, such as macrophages and neutrophils
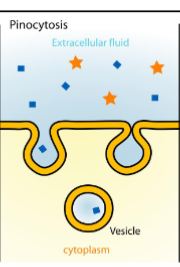
Endocytosis - Pinocytosis
internalization of small droplets of liquid materials.
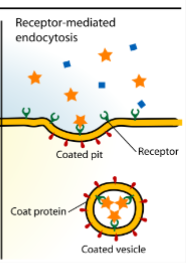
Endocytosis - Receptor-mediated
receptor proteins on the cell surface are used to capture a specific target molecule
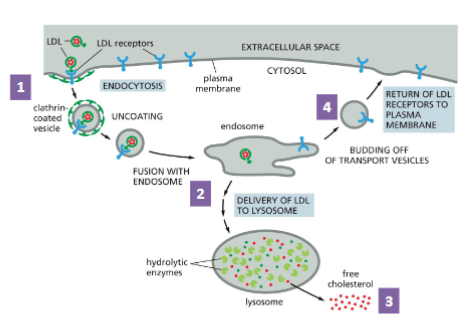
Receptor-mediated Endocytosis of LDL
Low density lipoprotein(LDL) is a carrier of blood cholesterol
An LDL particle is composed of a phospholipid-protein shell and cholesterol core.
It carries cholesterol to target cells (e.g., gonads)
LDL is recognized by its receptor at the cell surface and internalized in the form of coated vesicle
The vesicle fuses with an endosome, which becomes lysosome
Free cholesterol is released to the cell
The receptor is recycled to the cell membrane
Why do we have LDL?
LDL carries cholesterols to target cells.
Example – Brings cholesterol to the ovary. Granulosa cells in the ovary use cholesterol to make hormones like estradiol
HDL on the other hand brings cholesterol to the liver.
The liver is able to “flush” cholesterol out of the body
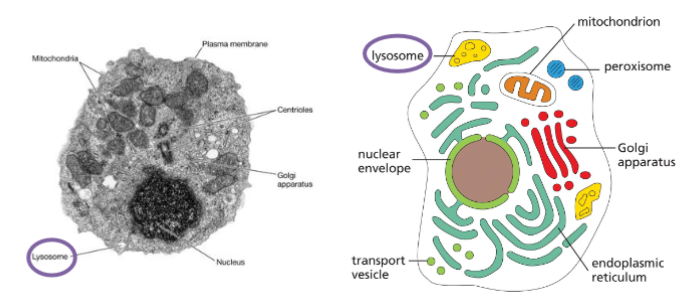
Lysosomes – the sites for intracellular digestion
Single membrane bound small organelles
Only found in animal cells
Filled with hydrolytic enzymes (hydrolases) that function at pH 4-5
lysosomes are “garbage disposal system” of the cell
Degrade the endocytic materials, such as bacteria.
Process the damaged internal organelles, such as mitochondria.
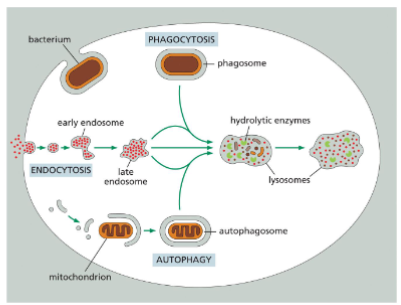
Three Steps of Vesicular Traffic
bud from the donor membranes
travel
fuse with the target membranes.
First step of Vesicular Traffic
Budding
Cargo binds to membrane embedded cargo receptor (on extracellular side). Adaptin bind cargo receptor on cytosol side.
Clathrin binds to adaptin on the cytosolic surface and assemble into a basket like shape. Forming a bud.
Dynamin proteins assemble around the neck of the budding vesicle leading to pinching off the vesicle.
Clathrin-coat is removed from vesicle once budding is complete
Third step of Vesicular Traffic
Fusion
Tethering: Tethering protein (on membrane surface) binds to Rab protein on vesicle. This brings the vesicle to its target membrane
Docking: v-SNARE in vesicle binds to t-SNARE on target membrane
Fusion: v-SNARE and t-SNARE catalyze the fusion of the two membranes. Ultimately delivering the cargo protein
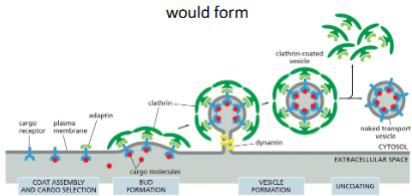
The budding of clathrin-coated vesicles from eukaryotic plasma membrane fragments can be observed when adaptins, clathrin, and dynamin are added to the membrane preparation. What would you observe if you omitted (A) adaptins, (B) clathrin, or (C) dynamin?
Adaptins?
Binds to the receptor and links clathrin and the membrane.
Without adaptins no clathrin could bind, so no bud could form
Clathrin?
Clathrin creates a basket that causes bud formation.
Without clathrin, adaptin would bind to receptor but no bud would form
Dynamin?
Dynamin pinches of the bud to create a vesicle.
Without dynamin a clathrin coated bud would form, but would not be released from membrane
Consider the v-SNAREs that direct transport vesicles from the tran Golgi network to the plasma membrane. They, like all other v-SNARES, are membrane proteins that are integrated into the membrane of the ER during their biosynthesis and are then carried by transport vesicles to their destination. Thus, transport vesicles budding from the ER contain at least two kinds of v- SNAREs - those that target the vesicles to the cis Golgi cisternae, and those that are in transit to the trans network to be packaged in different transport vesicles destined for the plasma membrane. (A) why might this be a problem? (B) Suggest possible ways in which the cell might solve it.
The Problem: Mixed signals lead to docking at the wrong organelle.
The Solution: Use Rab proteins as a secondary "zip code" and keep "in-transit" SNAREs inactive or masked until they reach their correct staging area (the Trans-Golgi).
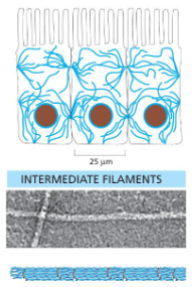
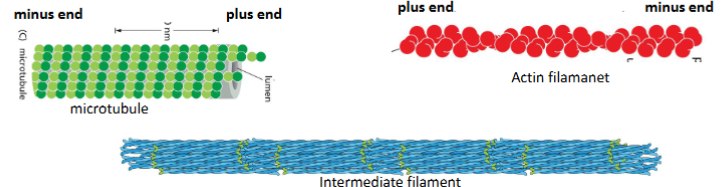
Intermediate filaments
Ropelike fibers
• 10nm diameter
• Made of fibrous intermediate filament proteins
• Are flexible and have great tensile strength
• Have roles in mechanical strength of cells and nuclear membrane structure
Actin Filaments
Helical polymers of actin proteins
• Diameter of 7nm
• Are flexible and can organize into a variety of liner bundles or 3D meshes
• Highly concentrated in cortex (layer of cytoplasm just beneath the plasma
membrane)
Microtubules
Hollow cylinder made of alpha and beta tubulin proteins
• Diameter of 25nm
• More rigid than actin filaments or intermediate filaments
• Rupture when stretched
Cytoskeletal assembly and disassembly – general principles
Cytoskeletal filaments are very dynamic structures (they assemble and disassemble fast).
Assembly and disassembly regulated via accessory proteins
Dynamic nature a result of filaments being made of small subunits held together via noncovalent interactions
Why are smaller cytoskeletal subunits necessary?
Small subunits can quickly get to their cellular location
• Noncovalent interaction among subunits allows quick assembly and disassembly of the cytoskeleton, thus a rapid change of cellular function
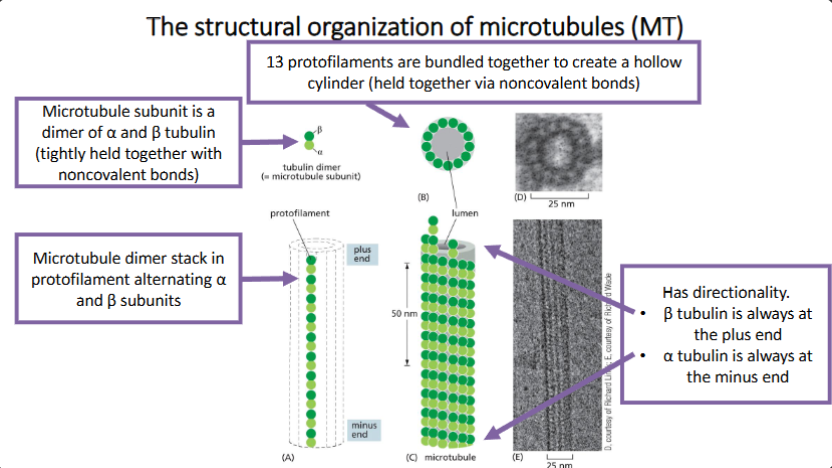
Microtubule (MT) Properties
MT are tube-like structures assembled from tubulin subunits, which is a heterodimer of α tubulin and β tubulin.
MT have structural polarity and are dynamic.
The organization and the stability of MT are regulated by MT associated proteins (MAP).
MT regulate:
cell movement
mitosis
organelle orientation
cellular transportation.
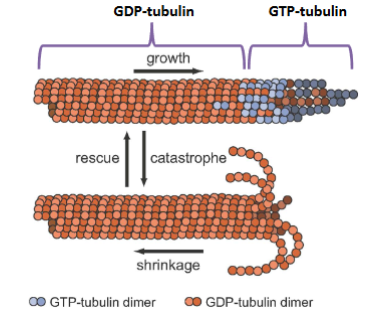
Microtubule (MT) Assembly and Disassembly
Preferentially takes place at the plus (+) end (β tubulin end).
• GTP bound tubulins are at the + end, forming a stabilizing GTP-cap.
• GDP bound tubulins are at the minus (–) making MT unstable causing them to
depolymerize
GTP to GDP: Protein Function
GTP binds to protein (this binding is fast) causing protein activation
• GTP is hydrolyzed to GDP causing protein inactivation
• GDP dissociates from protein (this dissociation is slow)
• Protein without GDP can bind new GTP
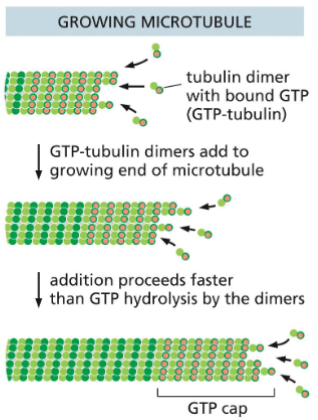
Microtubule (MT) Assembly (Growth)
GTP bound tubulin dimer binds to the MT plus end
Addition of GTP bound tubulin dimer is faster then GTP hydrolysis
GTP cap is formed, and MT grows
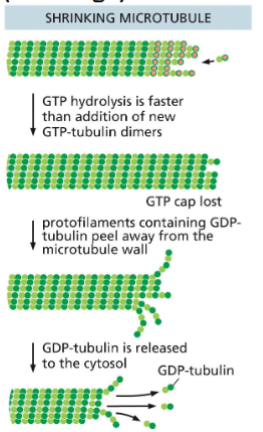
Microtubule (MT) Disassembly (Shrinkage)
GTP hydrolysis is faster the GTP-tubulin
dimer addition
• GTP cap is lost
• GDP-tubulin protofilaments peel away
• GDP-tubulin is released into the cytosol
• MT shrinks in size
Microtubule Dynamics: Dynamic Instability
MT can grow and shrink. The oscillation
between growth and shrinking of MT is
called dynamic instability.
• Switching back and forth between
polymerization and depolymerization
• Allows microtubules to rapidly remodel
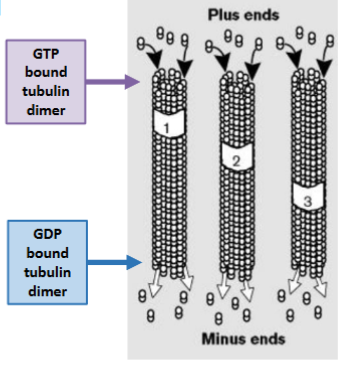
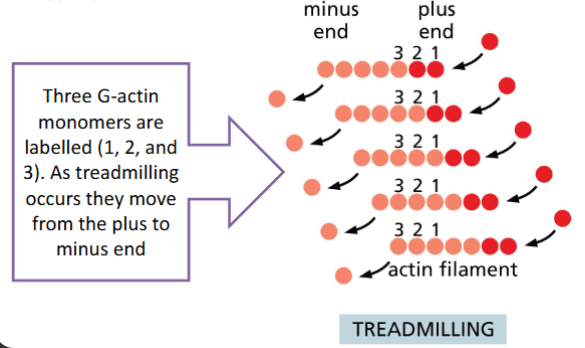
Treadmilling
The rate of monomer loss from one end
equals monomer addition at the other
end so the filament remains the same
length
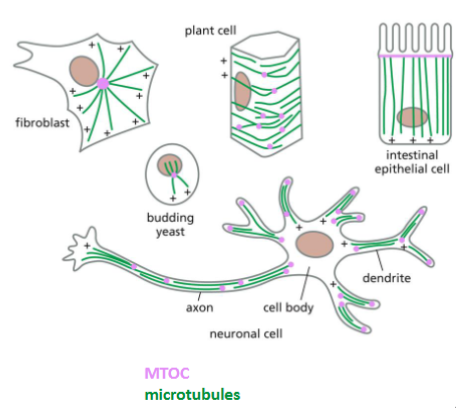
Microtubule Organizing Centers (MTOC)
A MTOC is a protein complex
made of γ tubulin and other
types of proteins.
• The major MTOC in mitotic cells
is called centrosome, which
contains a pair of centrioles in
animal cells.
• In some eukaryotic cells, MTOC
can also be found in basal
bodies that are associated with
cilium or flagellum
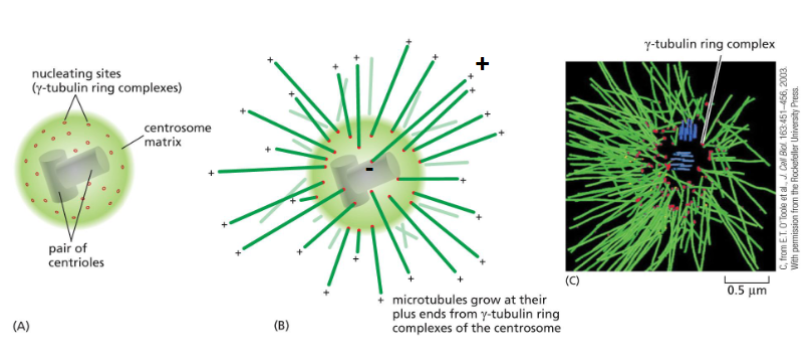
Centrosome Structure
A MTOC initiates the assembly and orientation of MT.
• MT are arranged in a way that all the (-) ends located at the MTOC.
Each microtubule grows and shrinks independently.
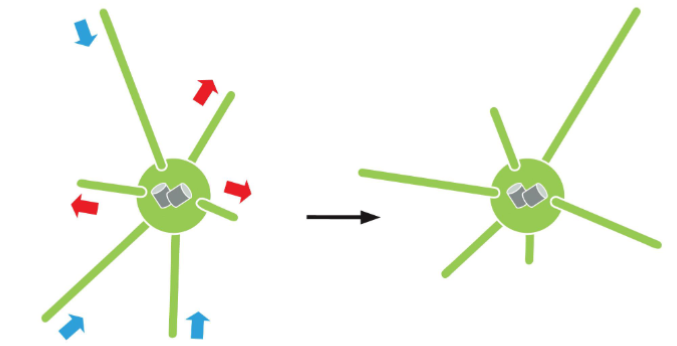
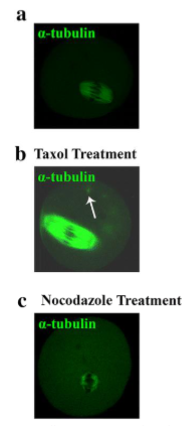
Microtubule (MT) as Anti-Cancer Drug Treatment
Microtubule-specific Drug; Action
Taxol: Binds to filaments and prevents depolymerization
Colchicine: Forms a complex with tubulin dimers that binds to the end of a microtubule, preventing further polymerization
Nocodazole: Binds tubulin dimers and prevents their polymerization
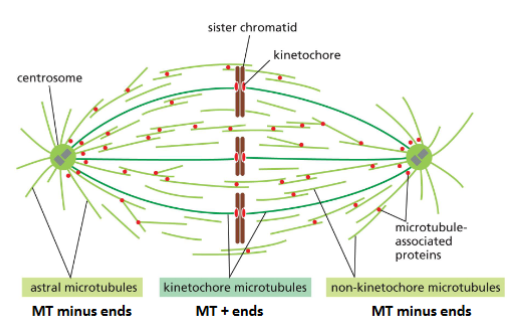
Microtubule Functions: Mitotic Spindle
Microtubules form mitotic apparatus to separate chromosomes during mitosis
• Combined action of MT dynamics and motor proteins is responsible for
chromosome separation during mitosis
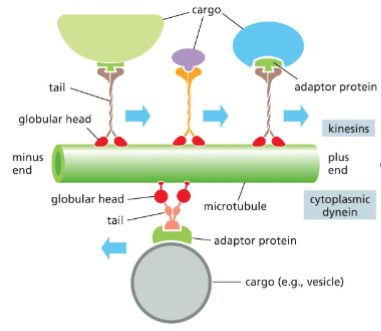
Microtubule Functions: Intracellular Transport and Motor Proteins
MT provide intracellular highway systems for the movement of organelles and vesicles
• Two families – kinesins and dynein
Kinesins - Most move the cargoes toward the (+) end
Dynein - move the cargoes toward the (-) end
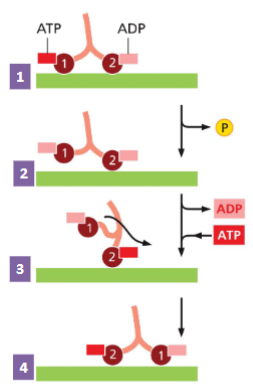
How Motor Proteins Move
1) The two heads of the motor protein are
attached to the MT. Head 1 has an ATP
bound, Head 2 has an ADP bound.
2) ATP hydrolysis (ATP to ADP) loosens
attachment of Head 1.
3) ADP is replaced by ATP on Head 2,
causing a confirmation change pulling
Head 1 forward in a “hand-over-hand”.
4) The motor protein has stepped forward.
This process continues to move motor
protein in “hand-over-hand along MT.
1
Motor proteins are enzymes with ATP
hydrolysis (ATPase) activity.
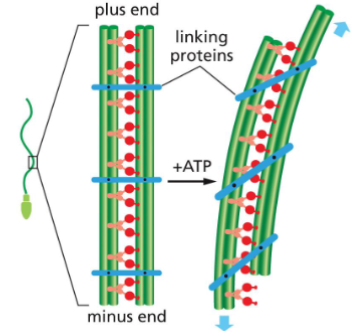
Microtubule Functions: Cell movement (Flagella)
Flagella are microtubule whip-like structures that are used to propel the cells
• Found in prokaryotes and some eukaryotic cells
How do flagella move?
• Dynein tail is attached to outer MT
• Dynein heads interact with adjacent MT
• Linker Proteins hold the two MT together,
causing MT bending
• MT bending leads to flagella movement
• Similar action happens in cilia
(not discussed here)
Actin filaments (F-Actin) Properties
Are polymers of actin subunits, one of the most abundant proteins in cells.
• Are essential for maintaining cell shape and movement of the cells outer membrane.
• Are dynamic and interactive with a large number of actin binding proteins can increase or decrease actin filament dynamics.
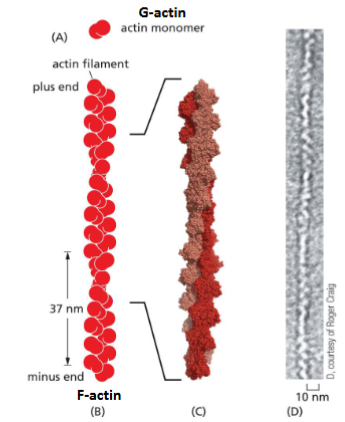
Actin Filament (F-actin) Structure
Made of actin subunits (monomer), a
globular protein called G-actin.
• G-actins polymerize to form actin
filaments (F-actin)
• F-actin has polarity - all G-actin are
oriented in the same direction (head
to tail)
• The plus end is more dynamic (grow
and shrink) than the minus end
• Each filament is a two-stranded helix.
Multiple, lateral interactions between
the two strands prevent the strands
from separating
F-actin Treadmilling
When G-actin addition and loss from F-actin is equal.
• F-actin stays the same size, but the incorporated monomers are not the
same
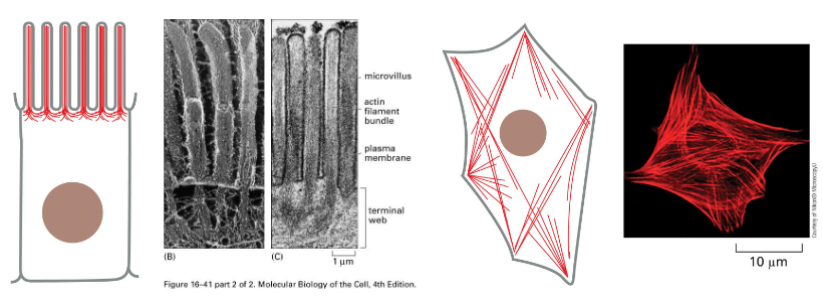
Actin Filament Functions: Maintaining cell shape and polarity
In many cells actin meshwork is found in the cortex (the area just under the plasma membrane.
• This actin meshwork and filaments allows the cell to hold its shape or move specialized cell features (e.g., microvilli)
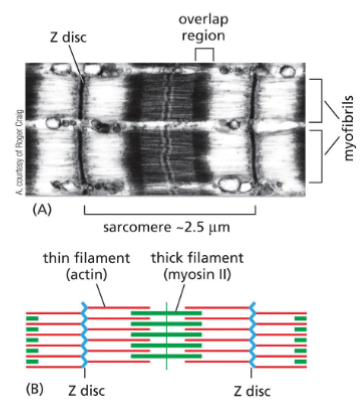
Actin Filament Functions: Muscle contraction
Sarcomeres are the functional contractile until of muscles
• Made of three regions:
1) Actin filaments
2) Thick filament (made of myosin II)
3) Z disc (attachment point for actin
filament
In sarcomeres this myosin filament mediated actin filament movement
causes muscle contraction.
• The size of the filaments remains the same, but the sarcomere gets smaller
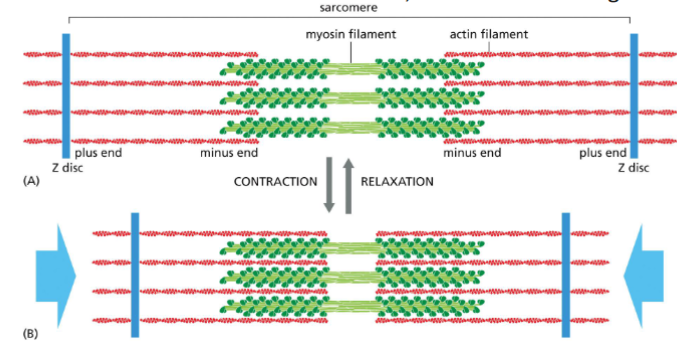
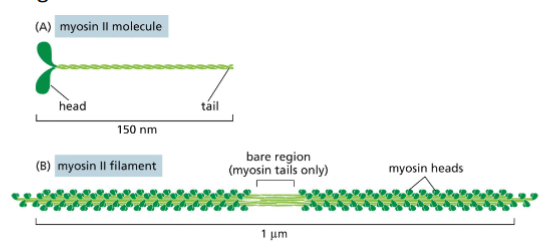
Myosin II and Thick Filaments
Myosin II is a motor protein and an actin binding protein with actin
binding head and tail.
• Myosin II tails can associate with one another to form a bipolar myosin
filament.
• heads project outward from the middle in opposite directions. The
bare region in the middle of the filament consists of tails only.
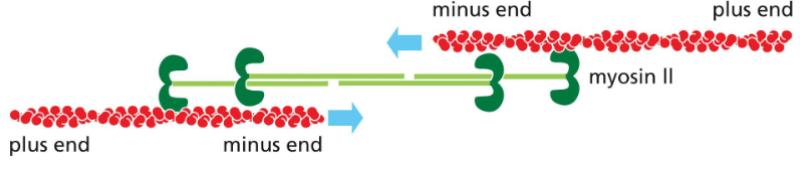
Myosin II filaments can interact with two actin filaments and move them in
opposite directions.
• The myosin heads walk towards the plus end of the actin filament, causing
contraction.
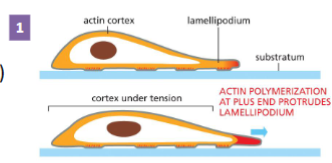
Actin Filament Functions: Cell Migration step 1
Cell sends out protrusions at the leading
edge
Actin filament plus end (where actin is most dynamic)
is right under the plasma membrane.
• Actin polymerizes forming the cell protrusion (e.g.,
lamellipodium)
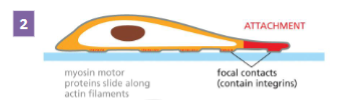
Actin Filament Functions: Cell Migration step 2
Protrusions adhere to the surface cell is
crawling over
Integrins are transmembrane proteins
• Extracellularly they bind to the surface the cell is
crawling over
• Intracellularly they bind to actin filaments
• This makes contact points for attachment
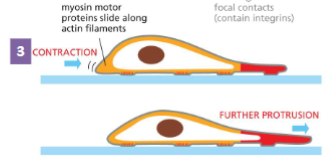
Actin Filament Functions: Cell Migration step 3
Rest of the cell drags itself forward using
the attachment points for traction
Myosin I motor attached to the plasma
membrane moves along actin filaments. This
brings the tail end of the cell forward
• Cell contraction
Myosin II vs. Myosin I
Myosin I
• Has a single F-actin binding head
• Tail bound to membrane
Myosin II
• Has two F-actin binding heads
• Forms thick filaments in muscle
Intermediate filaments (IF) Properties
• Exist in a various forms and expressed in different cells
• Different Rope-like filaments assembled from different protein subunits (monomers) by lateral binding and twisting
• Organized into bundles and meshworks by various IF associated proteins.
• Relatively stable compare with microtubules and actin filaments, but they are also dynamic.
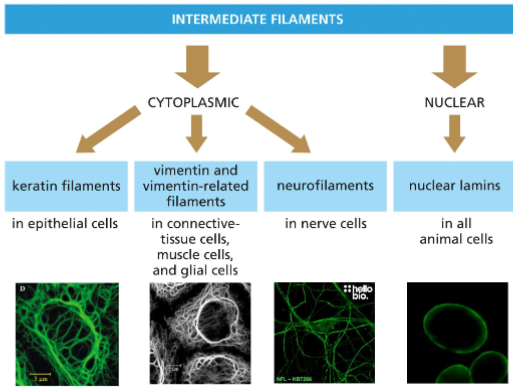
Intermediate filaments (IF) Classification
Are classified by cell type and subcellular localization
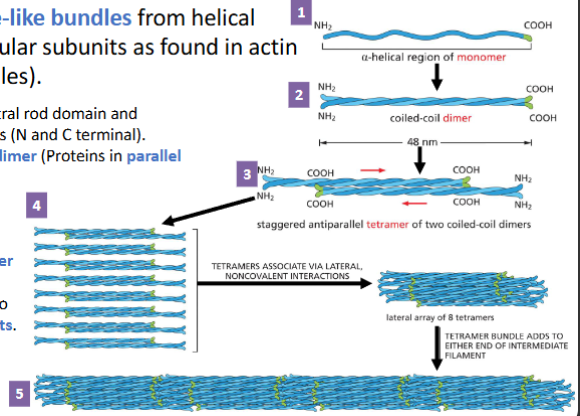
Intermediate filaments (IF) Assembly
IF are assembled to rope-like bundles from helical proteins (instead of globular subunits as found in actin filaments and microtubules).
1. IF monomer has α-helical central rod domain and unstructured terminal domains (N and C terminal).
2. Monomers associate to form dimer (Proteins in parallel direction).
3. Two dimers form a staggered antiparallel tetramer.
4. Eight tetramers pack into a helical array creating a tetramer strand
5. Tetramer strands assemble into ropelike intermediate filaments. Addition of tetramer array on either end causes filament to grow.
Intermediate filaments:
• Have no polarity
• Are easy to bend but not break
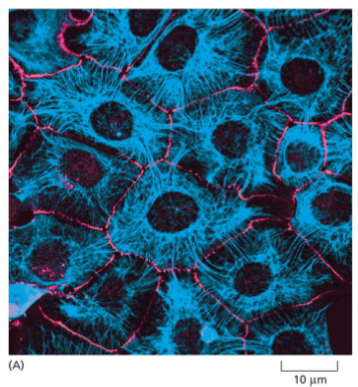
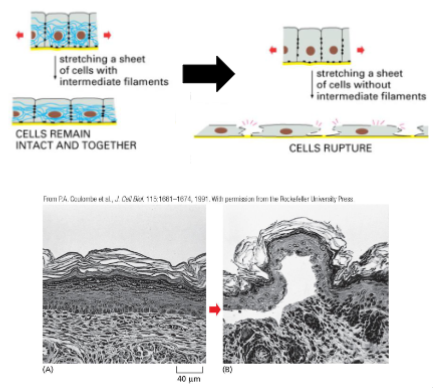
Intermediate filament function: Mechanical Strength in Epithelial Tissue
Keratin increases the mechanical strength in epithelial tissue
• Keratin binds to Cell-Cell Adhesions and Cell-Matrix adhesions (next lecture).
• This allows the cells in the tissue to withstand stretching, and remain as an intact tissue
• Loss of keratin (or keratin adhesion interaction) causes the cell layer to rupture when stretched.
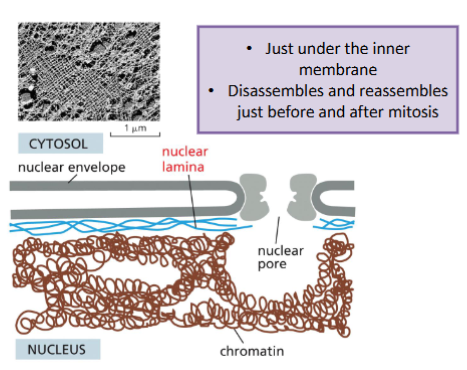
Intermediate filament function: Nucleus Shape and Gene Regulation
Lamin creates a mesh (nuclear lamina) work the supports the nuclear membrane
Lamin can also regulate gene expression
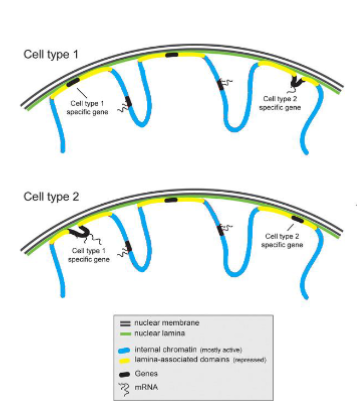
• Regions of DNA (as chromatin) is associated with the nuclear lamina (called lamina-associated domains).
• This association represses gene expression of genes in the lamina-associated domains.
• Different cells types can have different genes repressed via the nuclear lamina
• This is one of the ways different cells types have different gene expression profiles.
Why do you suppose it is much easier to add tubulin to existing microtubules than to start a new microtubule from scratch? Explain how the centrosome helps to overcome this hurdle.
Why is "scratch" hard? Small clusters of tubulin are unstable and fall apart before they can form a tube.
How does the centrosome help? It provides the $\gamma$-tubulin ring complex ($\gamma$-TuRC), which acts as a permanent, stable template/base.
Result: It lowers the concentration of tubulin required for a microtubule to start growing.
The drug Taxol, extracted from the bark of yew trees, has an opposite effect
to the drug colchincine, an alkaloid from autumn crocus. Taxol binds tightly
to microtubules and stabilizes them; when added to cells, it causes much of
the free tubulin to assemble into microtubules. In contrast, colchincine
prevents microtubule formation. Taxol is just as pernicious to dividing cells
as colchicine, and both are used as anticancer drugs. Based on your
knowledge of microtubule dynamics, suggest why both drugs are toxic to
dividing cells despite their opposite actions.
1. Colchicine: The "No-Growth" Problem
Colchicine binds to free tubulin dimers and prevents them from polymerizing.
The Effect: Existing microtubules eventually shrink and disappear because they can't be replaced.
Impact on Mitosis: The cell cannot build the mitotic spindle. Without a spindle, the chromosomes cannot be organized or moved to the center of the cell.
Result: The cell gets stuck in prophase/prometaphase and eventually undergoes programmed cell death (apoptosis).
2. Taxol: The "No-Shrink" Problem
Taxol does the opposite: it binds to the microtubule polymer and locks the subunits in place, preventing them from falling apart.
The Effect: Microtubules become "frozen" and incredibly stable. They cannot undergo dynamic instability (the rapid growing and shrinking).
Impact on Mitosis: During anaphase, the cell must shorten the spindle fibers to pull the sister chromatids toward opposite poles. If Taxol has "locked" the fibers, they cannot shorten.
Result: The chromosomes stay pinned in the middle of the cell. Because the cell cannot complete the division process, it triggers a "mitotic checkpoint" and dies.
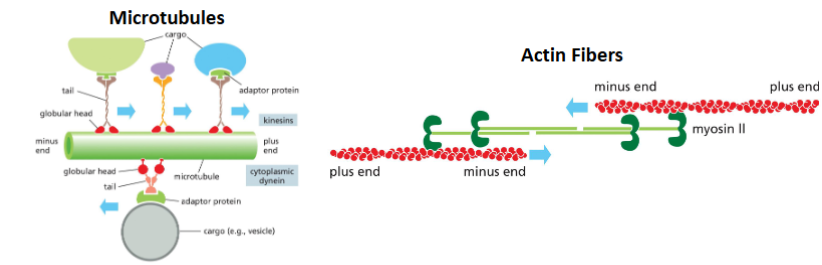
There are no known motor proteins that move on intermediate filaments.
Suggest an explanation for this.
1) What cytoskeleton proteins have known motor proteins?
Microtubules & Actin Fibers
2) What do structure does microtubules and actin fibers have in common that intermediate filaments do not?
Microtubules and actin filaments have directionality, intermediate filaments don’t
3) How motor proteins work? They move along cytoskeleton from one end to another (i.e., they uses the cytoskeleton directionality)
Intermediate filaments have no polarity; their ends are chemically indistinguishable. It would therefore be difficult to envision how a hypothetical motor protein that bound to the middle of the filament could sense a defined direction. Such a motor protein would be equally likely to attach to the filament facing one end or the other.
If both actin and myosin filaments of muscle are made up of subunits held together by weak noncovalent bonds, how is it possible for a human being to lift heavy objects?
Summation: Thousands of weak bonds added together create a very strong, stable polymer.
Distribution: The heavy load is shared across millions of myosin heads (cross-bridges) simultaneously.
Organization: The sarcomere structure anchors the filaments, allowing them to translate molecular "pulling" into macroscopic "lifting."
Actin filaments
Cell Shape and Surface Strength: Forms the cortex (a dense network just under the plasma membrane) that provides mechanical strength to the cell surface.
Cell Motility: Powers "crawling" through the formation of lamellipodia and filopodia.
Muscle Contraction: Interacts with myosin motor proteins to shorten sarcomeres.
Cytokinesis: Forms the contractile ring that pinches a dividing cell into two daughter cells.
Microtubules
Intracellular Transport: Acts as "railroad tracks" for motor proteins (kinesin and dynein) to move organelles and vesicles throughout the cell.
Cell Polarity and Organization: Originating from the centrosome, they establish the internal layout of the cell.
Mitosis: Forms the mitotic spindle required to physically pull duplicated chromosomes apart.
Cilia and Flagella: Provide the structural core (axoneme) that allows cells to swim or move fluid over their surface.
Intermediate Filaments
Mechanical Strength (Tensile Strength): Acts like a "rope" or "cable" that prevents cells from tearing when stretched or under physical stress.
Nuclear Integrity: Forms the nuclear lamina, a meshwork that supports the nuclear envelope and protects the DNA.
Cell-to-Cell Anchoring: Connects to specialized junctions like desmosomes, linking the cytoskeletons of adjacent cells together to form a continuous mechanical sheet (like in your skin).
Cytoskeletal elements summary - Actin
Subunit: G-actin monomers
Main job: Movement and Surface
Motors: Myosin
Structure: Flexible 2-strand helix
Cytoskeletal elements summary - Microtubules
Subunit: aB-tubulin dimers
Main job: transport and mitosis
Motors: Kinesin and Dynein
Structure: Hollow, rigid tube
Cytoskeletal elements summary - Intermediate filaments
Subunit: Fibrous proteins (keratin)
Main job: Strength and Durability
Motors: none
Structure: Tough, rope-like cable
What protein(s) are not involved in membrane fusion
clathrin
What structure does the protein have as it is transported into the organelle?
Endoplasmic Reticulum: co-translational translocation
Nucleus: Fully folded mature protein
Mitochondria: Unfolded precursor protein
You have two proteins - Protein A is a cytosolic protein and Protein B is a mitochondrial protein. In the lab you remove the mitochondrial localization signal from Protein B and add it to Protein A. What happens to the localization of these proteins.
Protein B becomes cytosolic and Protein A becomes mitochondrial
Once a protein is transported into certain organelles the sequencing signal is cleaved. Match the organelle transported into and what happens to the protein.
Mitochondria: Protein is cleaved, peptide stays in organelle
Nucleaus: Protein is not modified (no cleavage)
ER: Protein is cleaved, peptide stays in organelle membrane
A protein that is synthesized in the cell to be exocytosised to outside the cell travels through the endomembrane system from one membrane bound structure to another. What membrane bound structures do they go through, and in what order from site of protein synthesis to cell surface.
Protein Synthesis
Endoplasmic Reticulum
Golgi Apparatus
Secretory vesicles
Plasma membrane
Cell Surface