Bacterial Gene Expression
1/48
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
49 Terms
The Central Dogma is the flow of genetic information
DNA {transcription}→ RNA {translation}→ protein
DNA= instructions (recipe)
RNA= messenger (copy of recipe)
Protein= function (actual dish)
DNA
Stores genetic info
determines genotype
RNA
Intermediate carrier
Transfers info from DNA
Protein
Does the work (enzymes, structure, transport, function)
determines phenotype
Transcription
Making RNA from DNA
The process of when genetic info in DNA is copied into RNA
Monocistronic and Polycistronic
Monocistronic
1 mRNA codes for 1 protein
Polycistronic
1 mRNA codes for multiple proteins
Translation
The process where ribosomes read mRNA and use it to build a protein
Transcription and Translation coupled occur where in a bacterial cell?
Cytoplasm
RNA Polymerase
An enzyme that reads a DNA template and builds a complementary RNA strand during transcription
Synthesizes in the 5’-3’ direction (like DNA)
Can start on its own, NO PRIMER
Sigma Factor (σ)
A protein that helps RNA polymerase find the promoter, bind to it, and start transcription in bacteria
Will fall off after the core puts on about 10 bases
Recognizes -35 and -10 regions
Sigma + Core =
Holoenzyme
Promotor
Transcription start site
Steps to Transcription
Initiation
Elongation
Termination
Prokaryotes vs Eukaryotes
Bacteria: transcription and translation happen at the same time and in the cytoplasm, operons are multiple genes together, no introns or exons they just transcribe what they have
Eukaryotes: have introns and exons, need splicing, introns removed and mRNA is formed
Bacteria
Simple, fast, together
Eukaryotes
Complex and separated
Coding Strand
DNA strand that matches the RNA sequence (except T replaces U)
Template strand
The DNA strand used as a guide to make RNA during transcription
-Inverse of the mRNA
Transcription Initiation
The first stage of transcription where RNA polymerase binds to the promotor (with help from sigma factor in bacteria) and begins RNA synthesis
Transcription Elongation
The stage in transcription where RNA polymerase moves along the DNA template and adds nucleotides to lengthen the RNA strand
Transcription Termination
The stage in transcription where RNA polymerase reaches a termination signal and releases the newly made RNA transcript
Two ways:
-Rho-dependent
-Rho-independent
Rho-dependent
Type of transcription termination in bacteria that requires the Rho protein to unwind the RNA-DNA hybrid and release the RNA transcript
Uses Rho protein
A protein called Rho binds to the newly made RNA strand
Rho moves along the RNA toward RNA polymerase using energy (ATP)
RNA polymerase pauses at a termination region on the DNA
Rho catches up to RNA polymerase
Rho helps separate the RNA from the DNA template, stopping transcription
Rho-independent
RNA polymerase transcribes a DNA sequence that forms a GC-rich hairpin loop in the RNA
Right after the hairpin, there is a stretch of Us in the RNA
The hairpin causes RNA polymerase to pause
The weak A-U bonds (between RNA and DNA) break easily
The RNA strand detaches, ending transcription
There are 3 types of RNA
mRNA
rRNA
tRNA
mRNA
Carries genetic information to ribosome that encodes for a protein
rRNA
RNA associated with the ribosome
5s rRNA (associated with the large subunit)
16s rRNA orients/positions the ribosome on the mRNA (associated with the large subunit)
23s rRNA peptidyl transferase activity (associated with the small subunit)
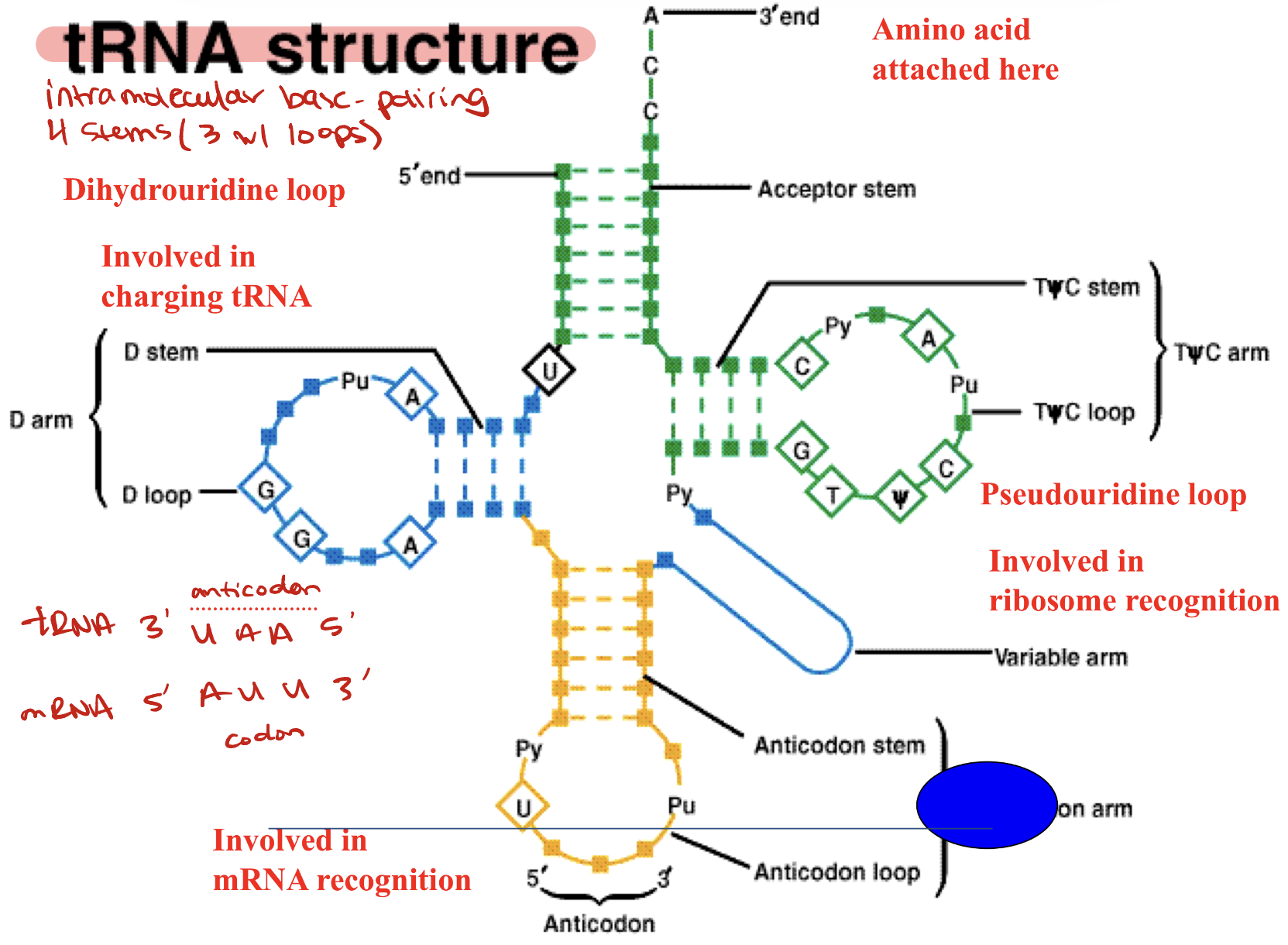
tRNA
Brings amino acids to the ribosome during protein synthesis
-Dihydrourine loop → involved in charging tRNA
-Pseudouridine loop → involved in ribosome recognition
-Anticodon loop → involved in mRNA recognition; recognizes codons in mRNA 3 bases at a time
-3’ end → amino acid attached here to the rRNA
Aminoacyl-tRNA Synthetases
Enzymes that catalyze the attachment of amino acids to tRNA
-There is a specific enzyme for each amino acid
No proofreading in the ribosome, so this is the step that adds specificity
less specific for tRNA (can recognize tRNA for each amino acid)
Genetic Code
Triplet code (codon = 3 bases on mRNA)
Anticodon = matches on tRNA
Start codon
Code is degenerate (specific to certain amino acids)
Base wobble
Start Codon
AUG aka met AKA fMet in bacteria! (N-formyl methionine, first amino acid)
different than other AUG
Bacteria are more flexible than eukaryotes and other start codons sometimes
Base Wobble
3rd base is flexible/wobbly!
A single tRNA can recognize multiple codons → less tRNA needed in the cell
Allows for more wiggle room
Each codon codes for a specific amino acid, but some amino acids can have multiple codons for one amino acid
Steps to Translation
Ribsosome binds mRNA
tRNA brings amino acids
Peptide bonds form
Protein grows
ALSO HAS Initiation, Elongation, and Termination
Shine-Dalgarno Sequence
A sequence that is rich in A and G that comes before the actual start codon
Helps it line up with the chromosome (aligns the ribosome with the start codon)
16s rRNA is crucial for this alignment
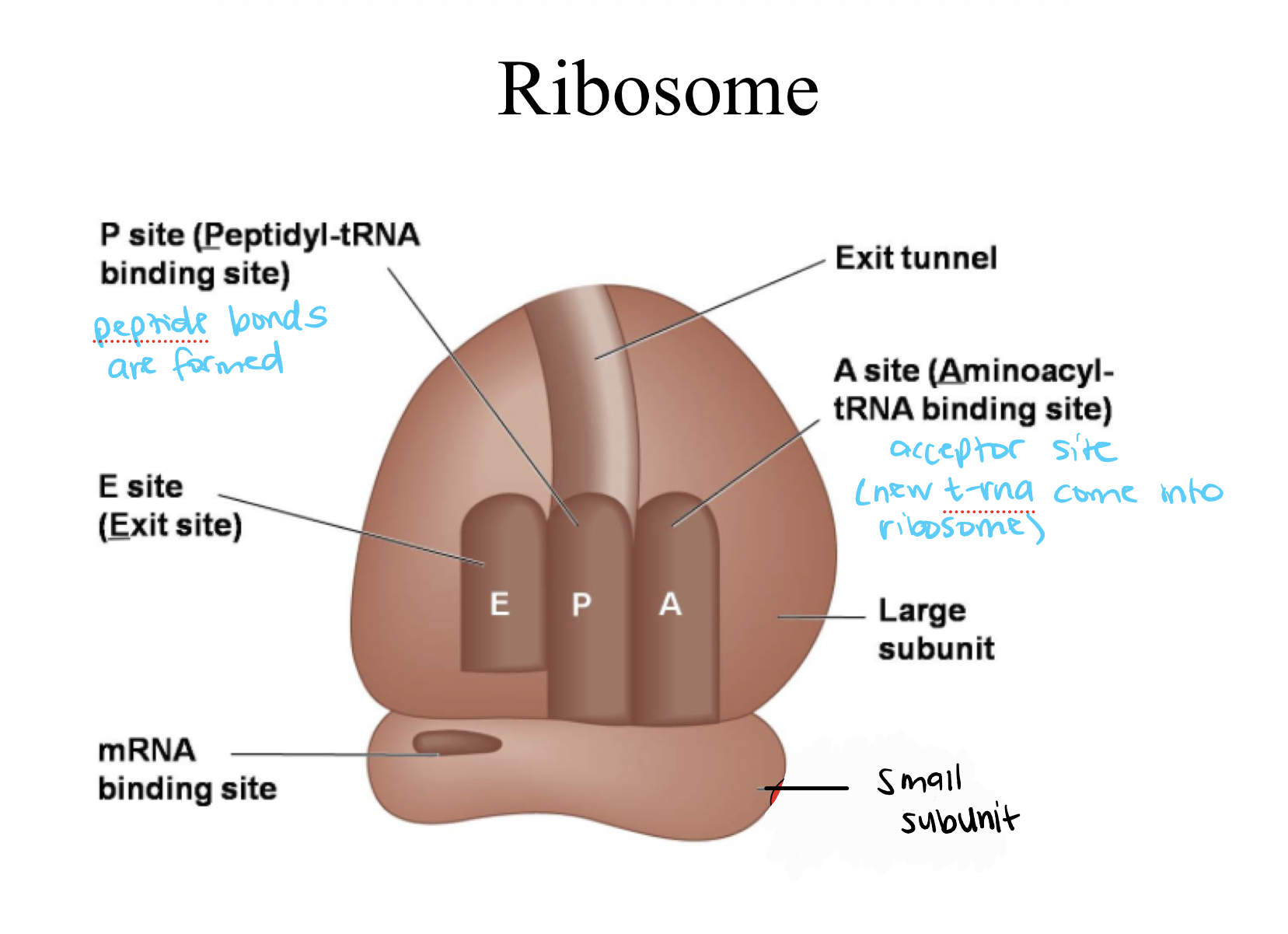
Ribosome
A cellular structure that reads mRNA and builds proteins by linking amino acids together
E-site (Exit site)
Where tRNA leaves after donating its amino acid
P-site (peptidyl-tRNA binding site)
Where peptide bonds are formed in the ribosome
Where Shine-Dalgarno Sequence binds to SSU
A site (Aminoacyl-tRNA binding site)
Accepts new tRNA
Small subunit
Part of the ribosome that binds to mRNA and helps it correctly position the start codon for translation
Large subunit
The part of the ribosome that forms peptide bonds between amino acids during protein synthesis
G proteins
Other proteins that help other than the ribosome (helper proteins)
-Initiation and Elongation factors
-GTP switches from active to inactive states
Translation Initiation
First stage of translation where the ribosome assembles on mRNA and the first tRNA binds to the start codon
Ribosome binds to mRNA and looks for Shine-Dalgarno Sequence so it can “Start” and line up with the ribosome
Finds the start codon
First tRNA enters, methionine enters with tRNA and binds to AUG
tRNA sits in the P site, GTP hydrolyses
IF3 gets bumped off by IF-1
Large subunit comes in and IF-1 leaves
Complex is complete and is ready to build a protein
Translation Elongation
Stage of translation where the ribosome moves along the mRNA, and tRNAs bring amino acids that are added to the growing protein chain
Goal: add amino acids 1 by 1
brings appropriate tRNA to the A site (via EFU-TU complex)
Peptide bond formation → created by 23s ribosomal RNA
Translocation → ribosome moves 3 bases toward 3’ end (A to P, P to E, E to outside)
Chloramphenicol
Inhibits peptide bond formation
Erythromycin
Inhibits translocation
Translation Termination
Final stage in translation where a stop codon is reached and the completed protein is released from the ribosome
Ribosome hits a STOP codon
Release factor binds
Protein is released
Ribosome falls apart (subunits separate and mRNA is released) IF3 attaches to the small subunit and triggers the separation of the subunits
“Stop, release, disassemble”
-different release factors recognize different stop codons
-peptide gets cleaved off from tRNA in p-site
No tRNA recognizes stop codons…
They recognize release factors!
IF-3
G-protein in bacteria that prevents small and large subunits from prematurely joining, ensuring correct initiation of translation
EFU-TU complex
Bacterial protein that delivers aminoacyl-tRNA to the ribosome during translation elongation