SMU Genetics final (Practice Questions and Vocabulary)
1/88
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
89 Terms
1. Which of the following statements about regulation of gene expression is CORRECT?
A) An inducible gene is transcribed when a specific substance is absent.
B) A gene is any DNA sequence that is transcribed into an mRNA molecule only.
C) All genes are transcribed at all times as long as they have a functional promoter.
D) The regulation of gene expression is the same in both eukaryotes and prokaryotes.
E) The regulation of gene expression is critical for the control of life processes in all organisms.
E) The regulation of gene expression is critical for the control of life processes in all organisms.
2. Which of the following statements about gene regulation concerning operons is INCORRECT?
A) A negative repressible gene is controlled by a regulatory protein that inhibits transcription.
B) For a gene under negative repressible control, a small molecule is required to prevent the gene's repressor from binding to DNA.
C) For a gene under positive repressible control, the normal state is transcription of a gene, stimulated by a transcriptional activator.
D) A regulator gene has its own promoter and is transcribed into an independent mRNA.
E) Presence of operons where genes of related functions are clustered is common in bacteria but not in eukaryotes.
B) For a gene under negative repressible control, a small molecule is required to prevent the gene's repressor from binding to DNA. is the INCORRECT STATEMENT
3. E. coli lac operon control by lac I is:
A) negative inducible.
B) negative repressible.
C) positive inducible.
D) positive repressible.
E) attenuation.
A) negative inducible.
4. Transcriptional control that acts by regulating the continuation of transcription is called:
A) riboswitching.
B) antitermination.
C) negative control.
D) operator mutation.
E) attenuation.
E) attenuation.
5. RNA molecules that are complementary to particular sequences on mRNA are called: A) complementary RNA. B) sense RNA. C) antisense RNA. D) riboswitches. E) ribozymes.
C) antisense RNA x
6. How does histone acetylation affect chromatin?
A) It loosens the chromatin and allows increased transcription.
B) It allows DNA to become resistant to damage.
C) It helps the histones have a greater affinity for DNA.
D) It inhibits DNA replication by making it more difficult to separate the DNA strands.
E) It causes the chromatin to become more condensed in preparation for metaphase.
A) It loosens the chromatin and allows increased transcription.
7. Which of the following is an example of an epigenetic change in eukaryotes?
A) a loss of an AT base pair from a gene
B) the addition of methyl groups to cytosines in the promoter region of a gene
C) the substitution of an AT base pair by a GC base pair in a gene as a result of a mistake during DNA replication
D) a deletion that simultaneously removes two genes from the genome
E) None of these examples represents epigenetic changes.
B) the addition of methyl groups to cytosines in the promoter region of a gene x
The agouti locus helps determine coat color in mice, and this phenotype can vary from light to dark between genetically identical individuals. You have discovered a drug that reduces the variation in the agouti phenotype. What is a likely explanation for this drug's mechanism of action?
A) It inhibits DNA polymerases.
B) It inhibits DNA methyl transferases.
C) It activates shelterin proteins.
D) It activates mitochondrial transcription.
E) It causes DNA damage.
B) It inhibits DNA methyl transferases.
Which of the following statements about regulation of eukaryotic gene expression is INCORRECT?
A) The presence of a nuclear membrane separating transcription and translation in eukaryotes led to the evolution of additional mechanisms of gene regulation.
B) In eukaryotes, most structural genes are found within operons.
C) Eukaryotic mRNAs are generally more stable than prokaryotic mRNAs.
D) The rate of degradation of mRNAs is important in regulation in eukaryotes. E) Posttranslational regulation of histones is unique to eukaryotes.
B) In eukaryotes, most structural genes are found within operons.
10. Which of the following statements about histones and gene expression is CORRECT?
A) In a general sense, highly condensed DNA bound with histone proteins represses gene expression.
B) Acetylation involves the addition of acetyl groups to histone proteins, and it usually results in repression of transcription.
C) Addition of methyl groups to the tails of histone proteins always results in activation of transcription.
D) Histone code refers to the modification that takes place on the globular domain of the octamer histone core.
E) Phosphorylation of cytosines generally leads to increase in transcription..
A) In a general sense, highly condensed DNA bound with histone proteins represses gene expression
11. When regions around genes become sensitive to the enzyme DNase I, this is an indication that those regions are:
A) becoming transcriptionally active.
B) becoming more condensed.
C) binding to the single-strand binding proteins.
D) destabilizing and transcriptionally inactive.
E) becoming highly methylated by a methylase.
A) becoming transcriptionally active.
12. A boundary element is also known as a(n):
A) insulator.
B) repressor.
C) enhancer.
D) coactivator.
E) mediator.
A) insulator. X
13. Which of the following statements about response elements is INCORRECT?
A) A single eukaryotic gene is regulated by only one unique response element.
B) Response elements are composed of specific consensus sequences that are unique from one another.
C) Different genes can possess a common regulatory element upstream of their start site.
D) Multiple response elements allow the same gene to be activated by different stimuli.
E) Response elements allow complex biochemical responses in eukaryotic cells.
A) A single eukaryotic gene is regulated by only one unique response element.
14. Consider the regulation of galactose metabolism through GAL4. Which of the following would result from a mutation that allowed GAL3 to bind to GAL80 in the absence of galactose?
A) Transcription of the genes involved in galactose metabolism would occur both in the presence and in the absence of galactose.
B) GAL80 would be able to bind to GAL4, and transcription of the genes involved in galactose metabolism would be repressed.
C) GAL4 would no longer be able to bind to the DNA; thus, transcription of the genes involved in galactose metabolism would occur.
D) GAL80 would no longer be able to stimulate transcription of the genes involved in galactose metabolism.
E) There would be no change in the regulation of galactose metabolism because GAL3 normally binds to GAL80 to cause a conformation change in GAL80.
A) Transcription of the genes involved in galactose metabolism would occur both in the presence and in the absence of galactose.
A fly (Drosophila) with an XY genotype has a mutation in its sex-lethal (Sxl) gene that renders its protein product nonfunctional. Which of the following describes the sex of this fly? A) male B) female C) intersex D) male, but it will be sterile E) female, but it will be sterile
A) male
16. siRNAs and miRNAs are produced by the:
A) cleavage of RISCs by endonucleases.
B) cleavage of functional mRNA within the cytoplasm.
C) cleavage of pre-mRNA in the nucleus.
D) cutting and processing of double-stranded RNA by Dicer enzymes.
E) cutting and processing of double-stranded RNA by Slicer enzymes.
D) cutting and processing of double-stranded RNA by Dicer enzymes. x
17. Which of the following information CANNOT be learned from RNA sequencing?
A) the types and number of RNA molecules produced by transcription
B) the presence of alternatively processed RNA molecules
C) information about the differential expression of the two alleles in a diploid individual
D) different RNA molecules generated by bidirectional or overlapping transcription of DNA sequences
E) the active sites of the enzyme the RNA encodes
E) the active sites of the enzyme the RNA encodes x
18. A linear DNA fragment is cut with a restriction enzyme to yield two fragments. How many restriction sites are there for the enzyme in this fragment?
A) one
B) two
C) three
D) four
E) It cannot be determined with this information.
A) one
19. What is the purpose of Taq polymerase in a PCR reaction?
A) DNA denaturation
B) primer annealing
C) DNA synthesis
D) heating of the reaction
E) heating of the reaction and DNA denaturation
C) DNA synthesis
20. Which of the following DNA sequences are contained within a cDNA library?
A) exons
B) introns
C) promoters
D) enhancers
A)Exons x
Transcription
(genetics) the organic process whereby the DNA sequence in a gene is copied into mRNA
Structural Genes
genes that code for proteins
Regulatory Genes
genes that control gene expression
Cis-Regulatory Genes
DNA sequences next to genes that help regulate transcription.
DNA binding proteins
Contain a domain with one of several kinds of DNA binding motif (motif = a stereotypical tertiary structure).
Operons
genes that coordinate the regulation of gene expression
polycistronic mRNA
A single molecule of messenger RNA that is formed by the transcription of a group of functionally related genes located next to one another along bacterial DNA.
Negative Control
Regulatory protein = Repressor, inhibits transcription whenbound to DNA
Positive Control
Regulatory protein = Activator, stimulates transcription whenbound to DNA
Inducible Operon
Transcription OFF (or low) until inducer turns it ON.
Repressible Operon
Transcription ON until co-repressor turns it OFF.
allosteric regulation
The binding of a regulatory molecule to a protein at one site that affects the function of the protein at a different site.
CAP protein
Activates transcription of the lac operon in the absence of glucose.
cAMP (cyclic AMP)
cAMP serves as a second messenger to activate or inactivate proteins within the cell. Termination of the signal occurs when an enzyme called phosphodiesterase converts cAMP into AMP
catabolite repression
inhibits cells from using carbon sources other than glucose
trp operon
tryptophan binds to the repressor protein and enables it to repress gene transcription.
trpR
regulatory gene or trp operon; codes for trp repressor that is active when effector is bound and attaches to DNA
Attenuation
premature transcription termination
antisense RNAs
Regulation of gene expression
OmpF
a transmembrane protein that permits ion diffusion in/out of cell
92 nt micF antisense RNA
base pairs with ompF mRNA, blocks translation
Riboswitches
Regulatory sequences within mRNAs
Ribozyme
RNA with catalytic activity
Where are prokaryotic genes regulated?
Prokaryotic genes primarily regulated at level of transcription
Where are Eukaryotic Genes regulated?
extensively regulated at multiple levels
Helix-turn-helix, zinc fingers, and leucine zippers, are all examples of what
DNA-binding proteins
What's the rundown on Bacteria sugar utilization
They generally like to turn to Glucose, easy to use and has lots of energy
They will use Lactose if Glucose is not present, if Lactose is the only option. This where the Lac Operon comes in, multiple genes are required to be turned on at once to activate the Lac Operon.
Case Study: The regulator protein is a repressor that binds to the operator and prevents transcription of the structural genes. What type of operon is it?
Negative Inducible Operon
Regulatory Protein: Repressor, transcripton on in the absence of signal
Negative Repressible
Regulatory Protein: Repressor, transcription off in the absence of signal
Negative Inducible
Regulatory Protein: Activator, transcription on in the absence of signal
Positive Repressible
Regulatory Protein: Activator, transcription off in the absence of signal
Positive Inducible
Small molecule binds to one site on protein & changes activity of another site Signaling molecule (e.g., allolactose) binds to lac repressor & changes position of DNA binding domain.
What is this process?
Allosteric regulation of regulatory protein activity
What happens when product of biosynthetic pathway is not present?
Repressor not able to bind to DNA and operon is transcribed.
_______ & ______ needed to (a) bring lactose into cell & then (b) convert it into galactose & glucose.
Permease (lacY), b-galactosidase (lacZ)
_____ actively transports glucose into the cell
Permease
the enzyme _____ breaks down the lactose transported in by permease into glucose and galactose
B-galactidosae
B-galactosidase also converts compound into
allolactose
Histones
protein molecules around which DNA is tightly coiled in chromatin
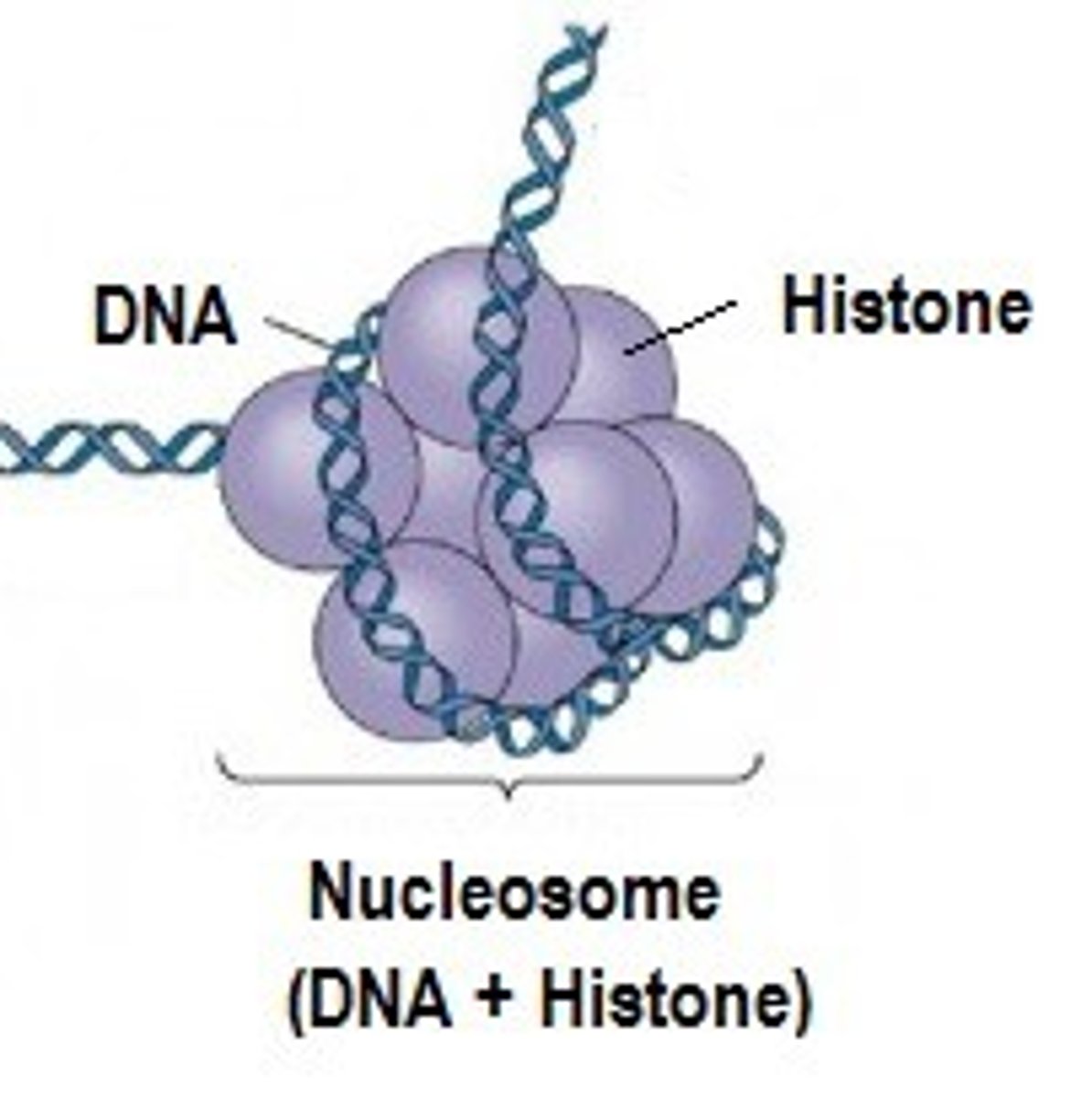
Binding of ___s to promoters require ____ to be removed
TFs, histones
TFIID
the first general transcription factor to bind the promoter, binds to the TATA box through the TATA binding protein (TBP)
basal transcription factors
Proteins, present in all eukaryotic cells, that bind to promoters and help initiate transcription.
RNA polymerase II
a multiprotein complex that transcribes DNA into precursors of messenger RNA (mRNA) and most small nuclear RNA (snRNA) and microRNA.
activator TF
attract histone acetylases(histone tails can't interact with each other--loose form of DNA that can be transcribed); TF indirect control-affect chromatin structure
chromatin remodeling complexes
complex of proteins that alters chromatin structure without acetylating histone proteins
Transcription Pre-initiation Complex
a complex of approximately 100 proteins that is necessary for the transcription of protein-coding genes in eukaryotes and archaea.
DNase I hypersensitive sites
Histone free regions that represent regions that are transcriptionally active
histone code hypothesis
Covalent modifications of tails of core histones play important roles in modulating transcription, by altering the abilities of nonhistone proteins to bind to nucleosomes.
Arabidopsis
An example of how histone acetylation can control flowering
DNA methylation
The addition of methyl groups to bases of DNA after DNA synthesis; may serve as a long-term control of gene expression. Hinders transcription
MeCP proteins
The MECP2 gene provides instructions for making a protein called MeCP2. This protein helps regulate gene activity (expression) by modifying chromatin, the complex of DNA and protein that packages DNA into chromosomes.
CpG islands
DNA methylation at CpG islands represses transcription
CpG
transcriptional repressors, mediating gene silencing via DNA cytosine methylation
Epi-
over, above
Can cause phenotypes without modifying the DNA sequence.
What is this?
Epigenetics
Rats licking their children which affects the patterns of DNA metylation in the offspring
What is this an example of
Epigenetics
Transcriptional Activators Interact with what:
Basal Tfs or Mediator
Regulatory Promoters
TF binding sites upstream of core promoter.
Usually within a few hundred bp of transcription start site.
Enhancers
Clusters of TF binding sites that can be located at variable positions relative to the promoter.
Can be located either close to a promoter or very far away.
Can be located upstream of a promoter, downstream of a gene, or even within introns.
Response elements (a type of cis-element):
help regulate genes in response to a stimulus
Insulators
a type of cis-regulatory element known as a long-range regulatory element. Found in multicellular eukaryotes and working over distances from the promoter element of the target gene, an insulator is typically 300 bp to 2000 bp in length. Insulators contain clustered binding sites for sequence specific DNA-binding proteins and mediate intra- and inter-chromosomal interactions. (sheriffs)
Are Enchancers Cis or Trans Acting
Trans acting
P-TEFb
Overcomes stalling of RNA Polymerase, allows RNA Polymerase II to continue transcription
HSF
heat shock factor (transcription factor) binds to Heat-Shock Element within regulatory promoters of heat shock genes & stimulates RNA polymerase to begin elongation.
Repressor TFs
inhibit transcription initiation or transition from initiation to elongation
GAL genes
genes in yeast responsible for metabolizing galactose
Gal4
DNA binding transcription factor in yeast that regulates galactose metabolism. The Gal4 protein contains a sequence specific DNA binding domain and an activation domain. (Activator)
Gal80
A repressor protein that interacts with Gal4 and prevents transcription of GAL structural genes. (Repressor)
UASg
enhancer with GAL4 binding sites