MCAT - Biochemistry - DNA & Biotechnology
1/90
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
91 Terms
eukaryotic cells contain
nucleic acids
what are the nucleic acids
dna and rna
dna and RNA are
polymers
DNA
- A polydeoxyribomucleotide that is composed of many monodeoxyribonucleotides linked together.
Nucleosides
- Composed of a five-carbon sugar (pentose) bonded to a nitrogenous base and are formed by covalently linking the base to C-1' of the sugar.
- The carbon atoms in the sugar are labeled with a prime symbol to distinguish them from the carbon atoms in the nitrogenous base.
Adenosine
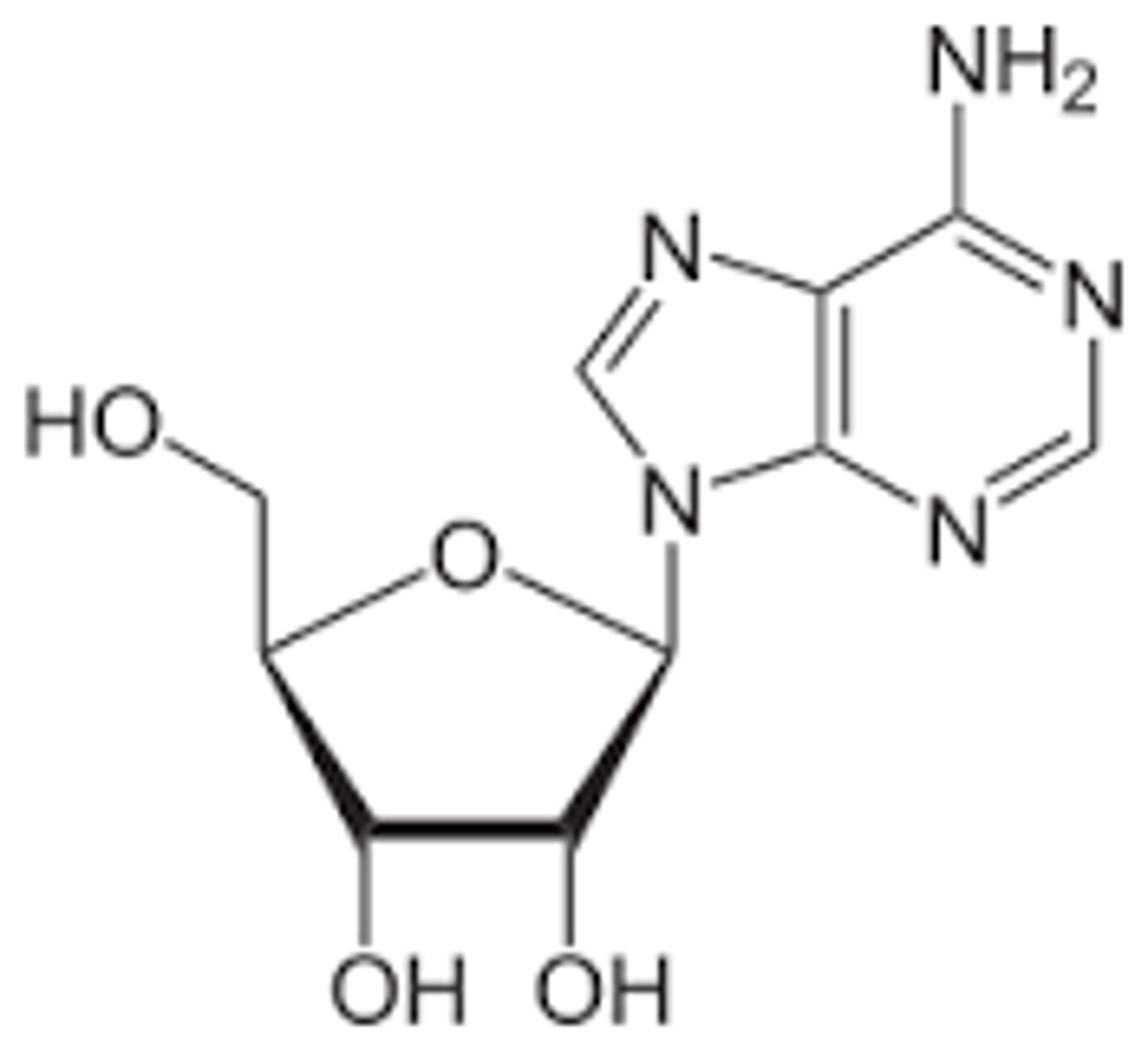
Nucleotides
- Formed when one or more phosphate groups are attached to C-5' of a nucleoside.
- Often these molecules are named according to the number of phosphates present.
- The building blocks of DNA and RNA
nucleoTide has phophate— think TP— like toilet paper
what is true about nucleotides
Adenosine di- and triphosphate (ADP and ATP), for example, gain their names from the number of phosphate groups attached to the nucleoside adenosine. These are high-energy compounds because of the energy associated with the repulsion between closely associated negative charges on the phosphate groups
High-Energy Bonds in ATP
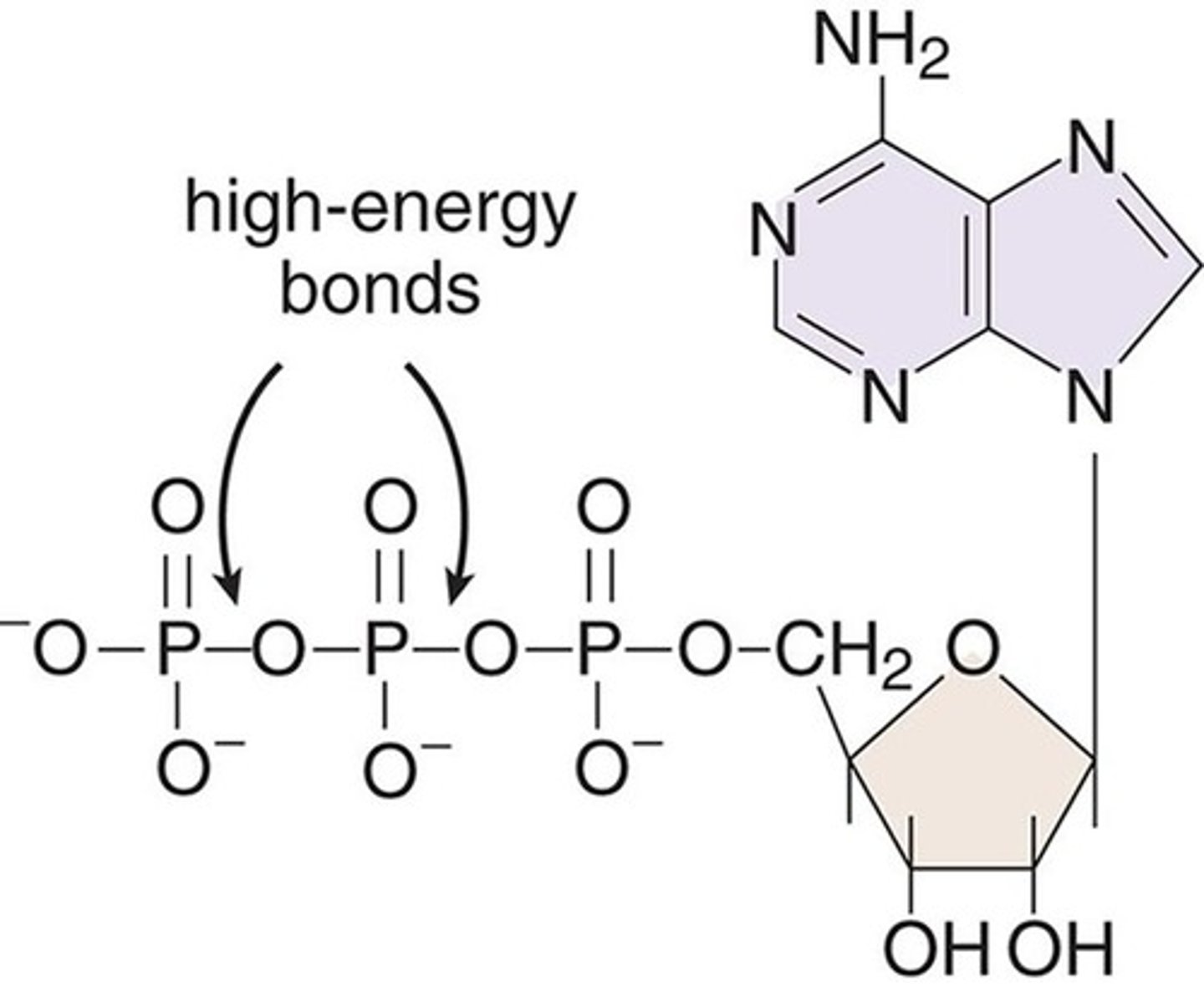
what is true about energy as it relates to ATP, adp, etc
in Chapter 3 of MCAT General Chemistry Review, we learned that bond breaking is usually endothermic and bond making is usually exothermic. ATP offers a biologically relevant—and MCAT tested—exception to this rule. Due to all the negative charges in close proximity, removing the terminal phosphate from ATP actually releases energy, which powers our cells.
how are nucleic acids classified
- Classified according to the pentose they contain.
- If the pentose is ribose, the nucleic acid is RNA; if the pentose is deoxyribose (ribose with the 2'-OH group replaced by -H), then it is DNA.
Ribose
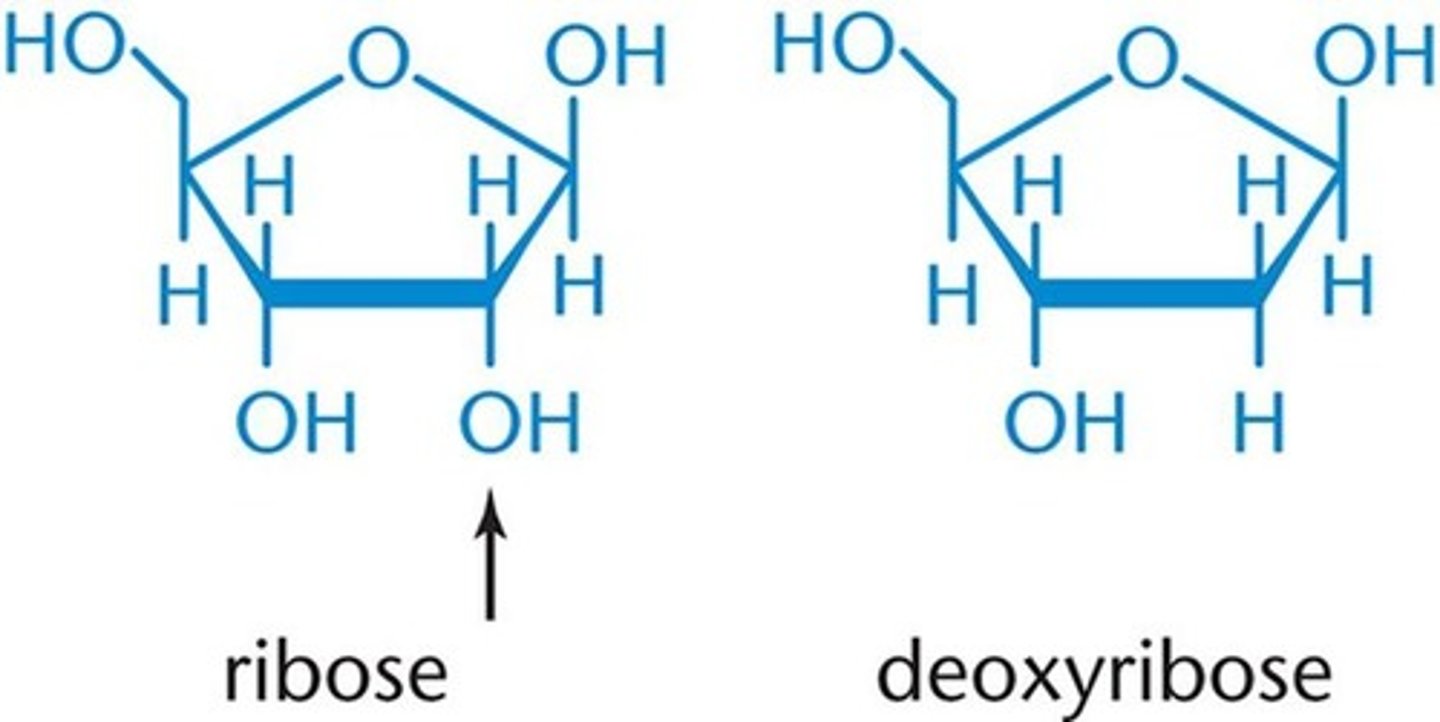
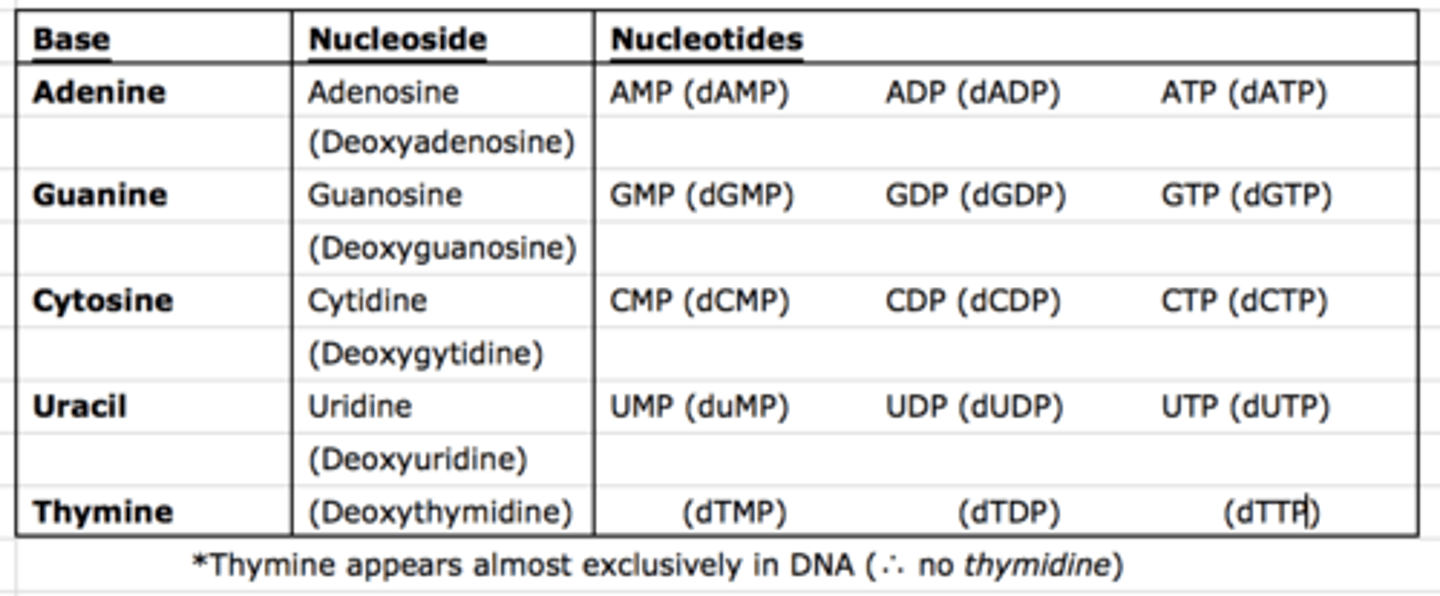
Nomenclature of Important Bases, Nucleosides and Nucleotides
Sugar-Phosphate Backbone
what makes up the backbone
what is the directionality of the backbone
what bonds are present
what direction are the bonds
overall charge of DNA and why
- The backbone of DNA is composed of alternating sugar and phosphate groups; it determines the directionality of the DNA and is always read from 5' to 3'.
- Formed as nucleotides are joined by 3'-5' phosphodiester bonds. That is, a phosphate group links the 3' carbon of one sugar to the 5' phosphate group of the next incoming sugar in the chain.
- Phosphates carry a negative charge; thus, DNA and RNA strands have an overall negative charge.
DNA Strand Polarity Diagram
what does the 5’ end have
what does the 3’ end have
Each strand of DNA has distinct 5′ and 3′ ends, creating polarity within the backbone, as shown in Figure 6.4. The 5′ end of DNA, for instance, will have an –OH or phosphate group bonded to C-5′ of the sugar, while the 3′ end has a free –OH on C-3′ of the sugar. The base sequence of a nucleic acid strand is both written and read in the 5′ to 3′ direction
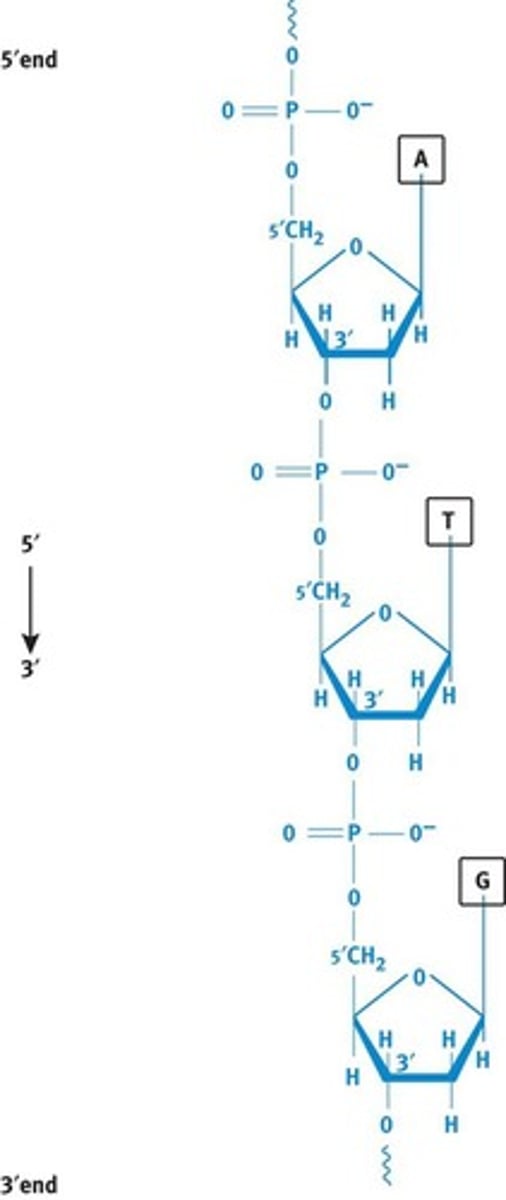
DNA Strand Polarity
- Each strand of DNA has distinct 5' and 3' ends, creating polarity within the backbone.
- The 5' end of DNA will have an -OH or phosphate group bonded to C-5' of the sugar, while the 3' end has a free -OH on C-3' of the sugar.
- The base sequence of a nucleic strand is both written and read in the 5' to 3' direction.
Purines
what are purines
which ones are purines
memory device
- nitrogenous bases that Contain two rings in their structure.
- The two purines found in nucleic acids are adenine (A) and guanine (G); both are found in DNA and RNA.
- PURe As Gold - A and G are purines. It takes two gold rings at a wedding, just like purines have two rings in their structure.
or requires 2 ppl to practice purity
Pyrimidines
what are pyrimidines
which are pyrimidines
memory device:
- nitrogenous bases that contain only one ring in their structure.
- Cytosine, thymine and uracil. Cytosine is found in both DNA and RNA, thymine is only found in DNA and uracil is only found in RNA .
- CUT the PYe. Pie has only one ring of crust, and pyrimidines only have one ring in their structure.
Purines & Pyrimidines are classified as what by structure
- Biological aromatic heterocycles.
Aromatic requirements
- Describes any unusually stable ring system that adheres to the following four specific rules:
1. The compound is cyclic.
2. The compound is planar.
3. The compound is conjugated (has alternating single and multiple bonds, or lone pairs, creating at least one unhybridized p-orbital for each atom in the ring).
4. The compound has 4n+2 (where n is any integer) pi electrons. This is called Huckel's rule. ( how do I know which lone pairs count as conjugates/ are pi electrons? look this up)
Aromatic Stability
- The extra stability is due to the delocalized pi electrons, which can travel throughout the entire compound using available molecular orbitals.
Heterocycles
- Ring structures that contain at least two different elements in the ring.
Watson-Crick Model of DNA
direction of DNA
The two strands of DNA are antiparallel; that is, the strands are oriented in opposite directions. When one strand has polarity 5′ to 3′ down the page, the other strand has 5′ to 3′ polarity up the page
watson-crick model of DNA
where is the backbone, whats on the inside
The sugar–phosphate backbone is on the outside of the helix with the nitrogenous bases on the inside.
watson-crick model of DNA
how does base pairing work, what pairs with what
what holds them together
how many bonds hold each together
There are specific base-pairing rules, often referred to as complementary base-pairing, as shown in Figure 6.7. An adenine (A) is always base-paired with a thymine (T) via two hydrogen bonds. A guanine (G) always pairs with cytosine (C) via three hydrogen bonds. The three hydrogen bonds make the G–C base pair interaction stronger. These hydrogen bonds, and the hydrophobic interactions between bases, provide stability t
Watson-Crick Model of DNA
amount of purine and pyrimidine
Because of the specific base-pairing, the amount of A equals the amount of T, and the amount of G equals the amount of C. Thus, total purines will be equal to total pyrimidines overall. These properties are known as Chargaff's rules.
types of DNA
B and Z DNA
B-DNA
direction of helix
contains what feature and for what purpose
- The double helix of most DNA is a right-handed helix, forming what is called B-DNA.
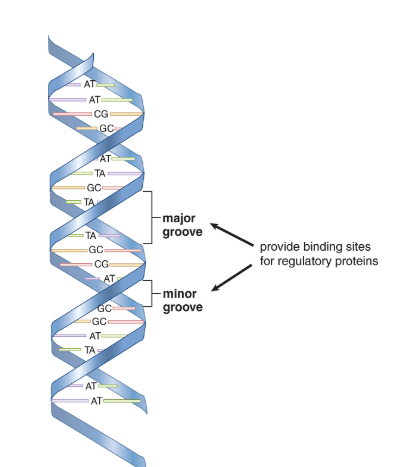
- The helix in B-DNA makes a turn every 3.4nm and contains about 10 bases within that span.
- Major and minor grooves can be identified between the interlocking strands and are often the site of protein binding.
Z-DNA
direction of helix
- Form of DNA named for its zigzag appearance.
- A left-handed helix that has a turn every 4.6nm and contains 12 bases within each turn.
- A high GC-content or a high salt concentrate may contribute to the formation of this form of DNA.
- No biological activity has been attributed to Z-DNA partly because it is unstable and difficult to research.
Denaturation
what occurs here
why is it necessary
how does It occur like what agents cause it
- During processes such as replication and transcription, it is necessary to gain access to the DNA. The double helical nature of DNA can be denatured by conditions that disrupt hydrogen bonding and base-pairing, resulting in the "melting" of the double helix into two single strands that have separated from each other.
- None of the covalent links between the nucleotides in the backbone of the DNA break during this process.
- Heat, alkaline pH, and chemicals like formaldehyde and urea are commonly used to denature DNA.
Reannealing
what is reannealing
how does it happen
Denatured, single-stranded DNA can be reannealed (brought back together) if the denaturing condition is slowly removed. If a solution of heat-denatured DNA is slowly cooled, for example, then the two complementary strands can become paired again,
describe a process that uses annealing of complimentary dna bases
annealing of complementary DNA strands is an important step in many laboratory processes, such as polymerase chain reactions (PCR) and in the detection of specific DNA sequences.
In these techniques, a well-characterized probe DNA (DNA with known sequence) is added to a mixture of target DNA sequences.
When probe DNA binds to target DNA sequences, this may provide evidence of the presence of a gene of interest. This binding process is called hybridization and is described in further detail later in this chapter
describe structure and storage of DNA
chromosomes made up of chromatin made up of nucleosomes
what is chromatin
he DNA that makes up a chromosome is wound around a group of small basic proteins called histones, forming chromati
Histones type and function
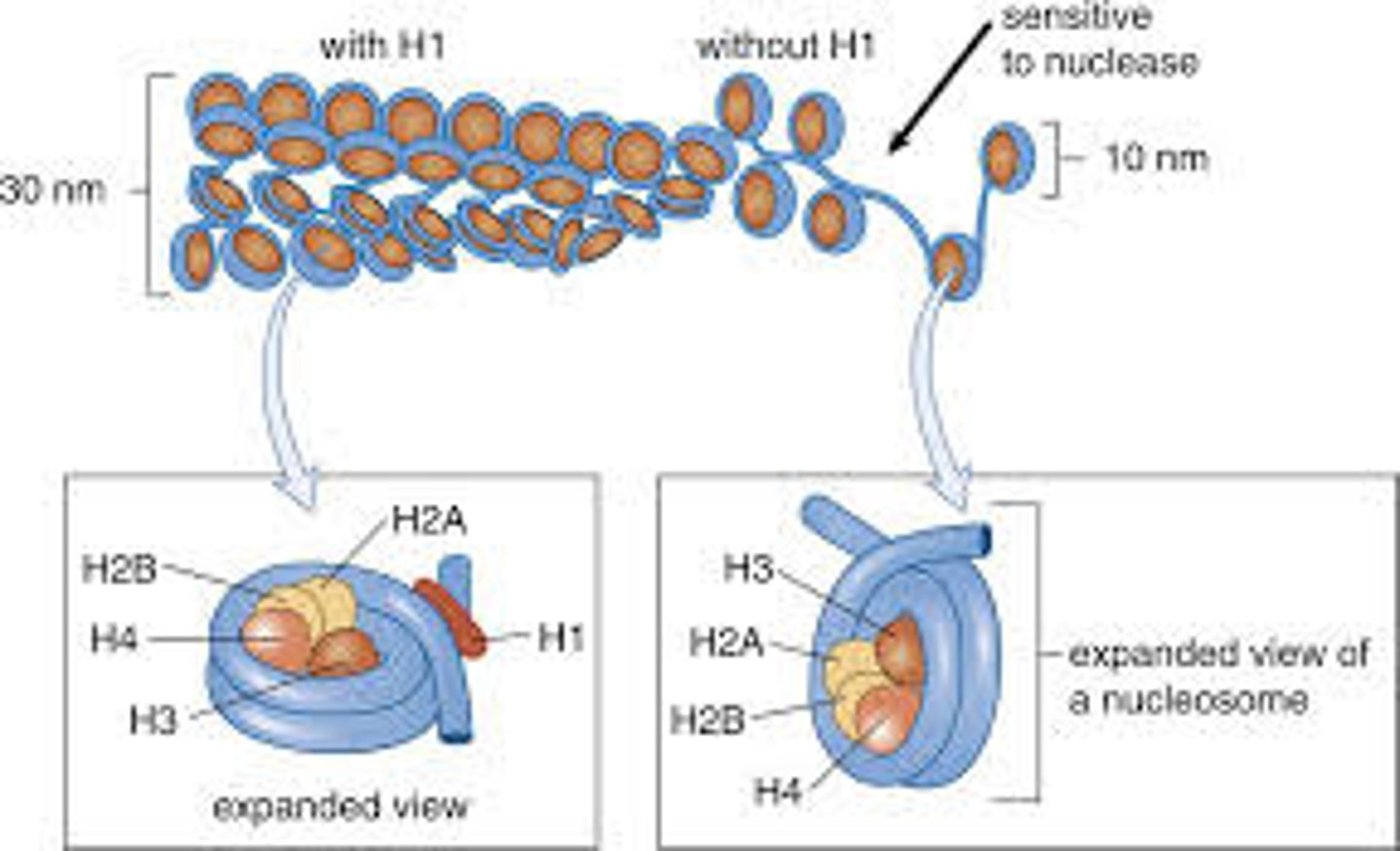
- There are five histone proteins found in eukaryotic cells.
- Two copies each of the histone proteins H2A, H2B, H3 and H4 form a histone core and about 200 base pairs of DNA are wrapped around this protein complex, forming a nucleosome. The last histone, H1, seals off the DNA as it enters and leaves the nucleosome, adding stability to the structure. Together, the nucleosomes create a much more organized and compacted DNA.
- Histones are one example of nucleoproteins.
Nucleosome = one bead
Chromatin = the whole necklace
Nucleosome Structure
types of chromatin
heterochromatin and euchromatin
Heterochromatin
- A small percentage of the chromatin that remains compacted during interphase.
- Appears dark under light microscopy and is transcriptionally silent.
- Often consists of DNA with highly repetitive sequences.
hetero—-thinnk boring, straight
Euchromatin
- The dispersed chromatin of interphase.
- Appears light under light microscopy.
- Contains genetically active DNA.
Telomere
why do we need telomeres
what exactly is a telomere
what protein is involved and why
additional function of telomeres
- DNA replication cannot extend all the way to the end of a chromosome. This will result in losing sequences and information with each round of replication. The solution for our cells is a simple repeating unit at the end of the DNA, forming a telomere.
- Some of the sequence is lost in each round of replication and can be replaced by the enzyme telomerase. Telomerase is more highly expressed in rapidly dividing cells. Studies indicate that there are a set number of replications possible, and that the progressive shortening of telomeres contributes to aging.
- Telomeres also serve a second function: their high GC content creates exceptionally strong strand attracting at the end of chromosomes to prevent unraveling.
Centromeres
- A region of DNA found in the centre of chromosomes.
- Often referred to as sites of constriction because they form noticeable indications.
- This part of the chromosome is composed of heterochromatin, which is in turn composed of tandem repeat sequences that also contain high GC-content. During cell division, the two sister chromatids can therefore remain connected at the centromere until microtubules separate the chromatids during anaphase.
Replisome/Replication Complex
- A set of specialized proteins that assist the DNA polymerases.
what begins the process of replication
DNA unwinds at the origins of replication
replication forks
The generation of new DNA proceeds in both directions, creating replication forks on both sides of the origin
Origins of Replication
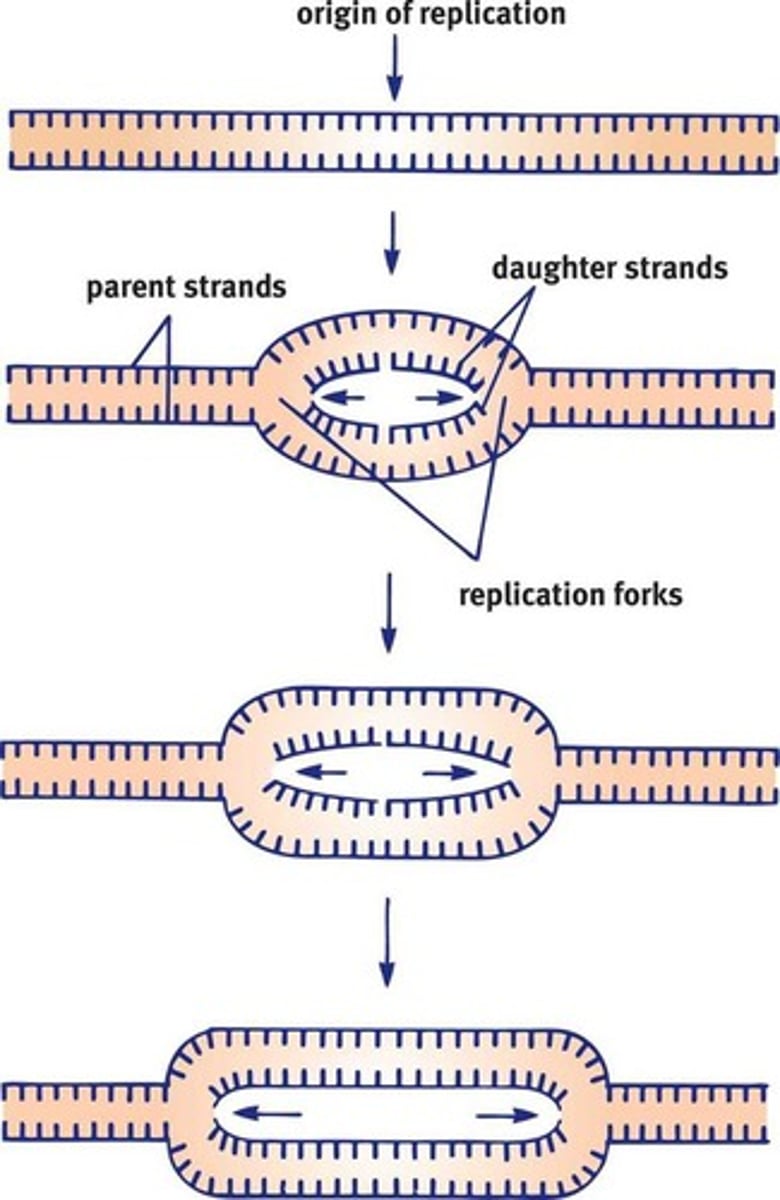
Bacterial Replication
- The bacterial chromosome is a closed, double-stranded circular DNA molecule with a single origin of replication. Thus, there are two replication forks that move away from each other in opposite directions around the circle.
Eukaryotic Replication
has what compared to prokaryotes in terms of origins of replication
- Must copy many more bases compared to prokaryotes and is a slower process.
- In order to duplicate all of the chromosomes efficiently, each eukaryotic chromosome contains one linear molecule of double-stranded DNA having multiple origins of replication. As the replication forks move toward each other and sister chromatids are created, the chromatids will remain connected at the centromere.
Helicase
- The enzyme responsible for unwinding the DNA, generating two single-stranded template strands ahead of the polymerase.
Single-stranded DNA-binding proteins
- Once opened, the unpaired strands of DNA are very sticky, in a molecular sense. The free purines and pyrimidines seek out other molecules with which to hydrogen bond.
- Proteins are therefore required to hold the strands apart. Single-stranded DNA-binding proteins will bind to the unraveled strand, preventing both the reassociation of the DNA strands and the degradation of DNA by nucleases.
Supercoiling & DNA topoisomerases
- As the helices unwinds the DNA, it will cause positive supercoiling that strains the DNA helix.
- Supercoiling is a wrapping of DNA on itself as its helical structure is pushed ever further toward the telomeres during replication.
- To alleviate this torsional stress and reduce the risk of strand breakage, DNA topoisomerases introduces negative supercoils. They do so by working ahead of helices, nicking one or both strands, allowing relaxation of the torsional pressure and then resealing the cut strands.
Semiconservative Replication
- During replication, these parental strands will serve as templates for the generation of new daughter strands.
- The replication process is termed semiconservative because one parental strand is retained in each of the two resulting identical double stranded DNA molecules.
DNA Polymerases
what does it do
reads in what direction
synthesizes in what direction
- Responsible for reading the DNA template, or parental strand, and synthesizing the new daughter strand.
- The DNA polymerase can read the template strand in a 3' to 5' direction while synthesizing the complementary strand in the 5' to 3' direction. This will result in a new double helix of DNA that has the required antiparallel orientation.
- Due to this directionality of the DNA polymerase, certain constraints arise. The two separated parental strands of the helix are also antiparallel to each other. Thus, at each replication fork, one strand is oriented in the correct direction for DNA polymerase; the other strand is antiparallel.
Leading Strand
- The strand in the replication fork that is copied in a continuous fashion, in the same direction as the advancing replication fork.
- This parental strand will be read 3' to 5' and its complement will be synthesized in a 5' to 3' manner.
Lagging Strand
- The strand that is copied in a direction opposite the direction of the replication fork.
- On this side of the replication fork, the parental strand has 5' to 3' polarity.
- DNA polymerase cannot simply read and synthesize on this strand.
Okazaki Fragments
- Because DNA polymerase can only synthesize in the 5' to 3' direction from a 3' to 5' template, small strands called Okazaki fragments are produced. As the replication fork continues to move forward, it clears additional space that DNA polymerize must fill in. Each time DNA polymerase completed an Okazaki fragment, it turns around to find another gap that needs to be filled in .
Enzymes of DNA Replication
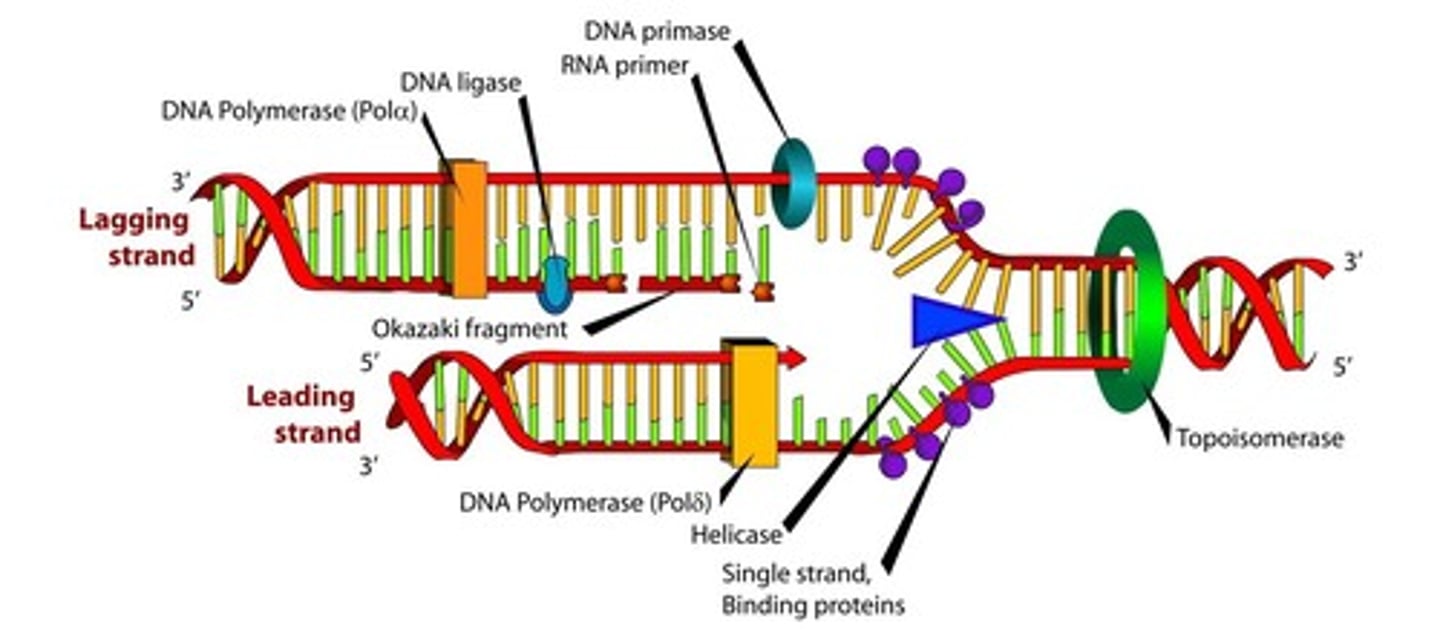
Primase
what is it and why is it necessary
- The first step in the replication of DNA is actually to lay down an RNA primer. DNA cannot be synthesized de novo; that is, it needs another molecule to "hook on" to. RNA on the other hand, can be directly paired with the parent strand. Thus, primase synthesizes a short primer (roughly 10 nucleotides) in the 5' to 3' direction to start replication on each strand.
- These short RNA sequences are constantly being added to the lagging strand because each Okazaki fragment must start with a new primer. In contrast, the leading strand requires only one, in theory.
DNA Polymerase III (Prokaryotes)
DNA polymerases α, δ, and ε (eukaryotes)
- Synthesizes the daughter strand of DNA in the 5' to 3' manner.
- The incoming nucleotides are 5' deoxyribonucleotide triphosphates: dATP, dCTP, dGTP and dTTP. As the new phosphodiester bond is made, a free pyrophosphate (PPi) is released.
DNA polymerase I (prokaryotes)/RNase H (eukaryotes)
- Removes the RNA to maintain sanctity of the genome.
DNA polymerase I (prokaryotes)/DNA polymerase sigma (eukaryotes)
- Adds DNA nucleotides where the RNA primer had been.
DNA ligase
Function: It seals the nicks (gaps) in the sugar-phosphate backbone by forming a phosphodiester bond between adjacent fragments.
Five classic DNA polymerases in eukaryotic cells:
1. DNA polymerases alpha and sigma: work together to synthesize both the leading and lagging strands; DNA polymerase sigma also fills in the gaps left behind when RNA primers are removed.
2. DNA polymerase gamma: replicated mitochondrial DNA.
3. DNA polymerases beta and epsilon: thought to participate mostly in DNA repair.
4. DNA polymerases sigma and epsilon: assisted by the PCNA protein, which assembles into a trimer to form the sliding clam. The clamp helps to strengthen the interaction between these DNA polymerases and the template strand.
sliding clamp
what is it
what synthesizes it
DNA polymerases δ and ε are assisted by the PCNA protein, which assembles into a trimer to form the sliding clamp. The clamp helps to strengthen the interaction between these DNA polymerases and the template strand
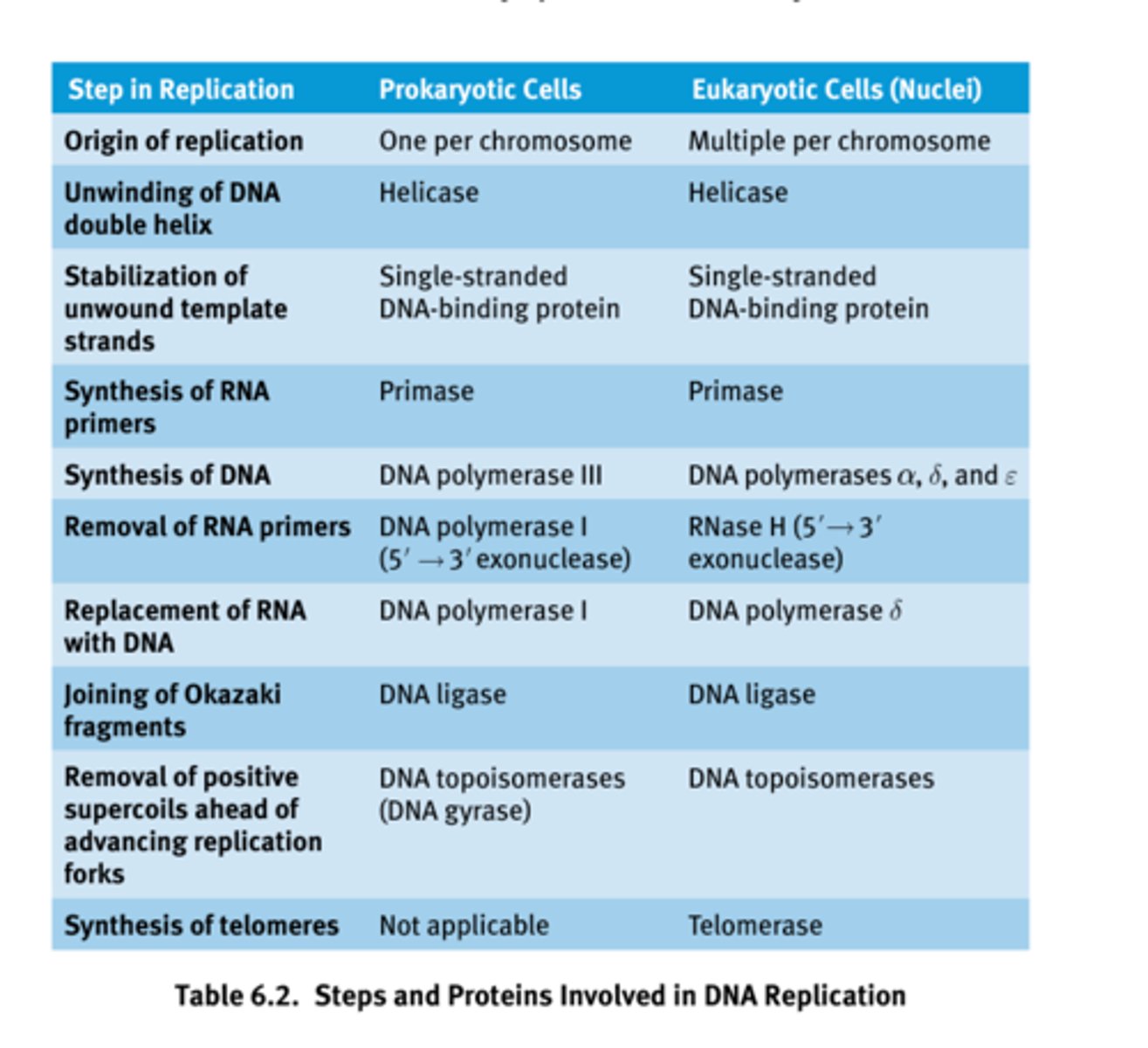
Steps and Proteins Involved in DNA Replication

Oncogenes
what are they
what are they before they are mutated
- Mutated genes that cause cancer.
- Primarily encode cell cycle-related proteins.
- Before these genes are mutated, they are often referred to as pro to-oncogenes.
- The abnormal alleles encode proteins that are more active than normal proteins, promoting rapid cell cycle advancement.
- Typically, a mutation in only one copy is sufficient to promote timor growth and is therefore considered dominant.
Tumor Suppressor Genes
- Encode proteins that inhibit the cell cycle or participate in DNA repair processes. They normally function to stop tumor progression and are sometimes called antioncogenes.
- Mutations of these genes result in the loss of tumor suppression activity, and therefore promote cancer.
- Inactivation of both alleles is necessary for the loss of function because, in most cases, even one copy of the normal protein can function to inhibit tumor formation.
Proofreading
- During synthesis, the two double-stranded DNA molecules will pass through a part of the DNA polymerase enzyme for proofreading. When the complementary strands have incorrectly paired bases, the hydrogen bonds between the strands can be unstable, and this lack of stability is detected as the DNA passes through this part of the polymerase. The incorrect base is excised and can be replaced with the correct one.
- DNA ligase, which closed the gaps between Okazaki fragments, lacks proofreading ability. Thus, the likelihood of mutations in the lagging strand is considerably higher than the leading strand.
If both the parent and daughter strands are simply DNA, how does the enzyme discriminate which is the template strand, and which is the incorrectly paired daughter strand?
- It looks at the level of methylation: the template strand has existed in the cell for a longer period of time, and therefore is more heavily methylated.
Mismatch Repair
- Cells also have machinery in the G2 phase of the cell cycle for mismatch repair; these enzymes are encoded by genes MSH2 and MLH1, which detect and remove errors introduced in replication that were missed during the S phase of the cell cycle. These enzymes are homologues of MutS and MutL in prokaryotes, which serve a similar function.
Nucleotide Excision Repair
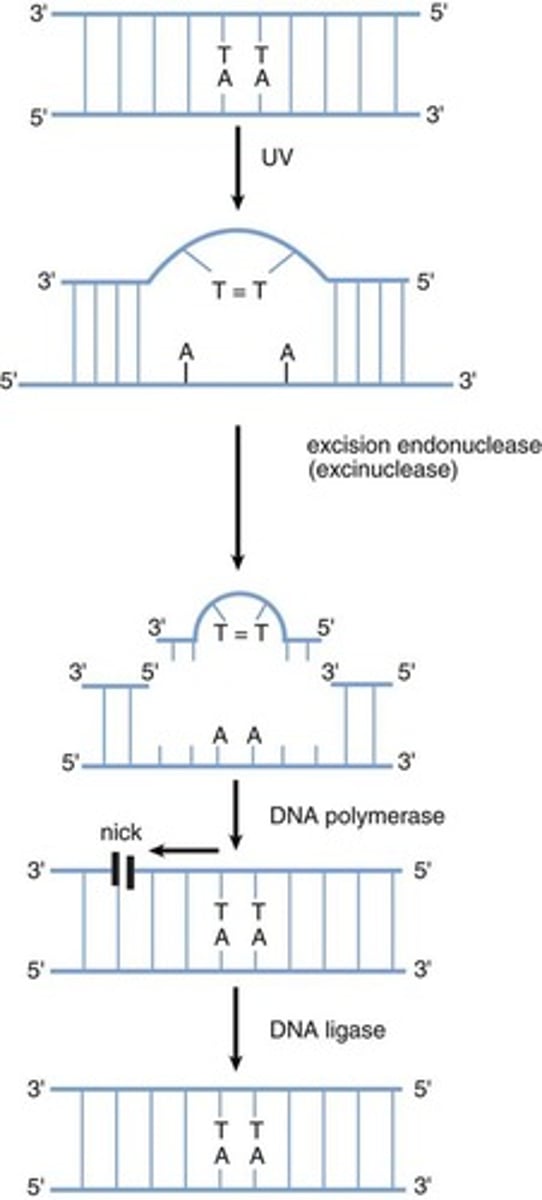
- UV light induces the formation of dimers between adjacent thymine residues in DNA. The formation of thymine dimers interferes with DNA replication and normal gene expression, and distorts the shape of the double helix.
- Thymine dimers are eliminated from DNA by a nucleotide excision repair (NER) mechanism, which is a cut-and-patch process. First, specific proteins scan the DNA molecule and recognize the lesion because of a bulge in the strand. An excision endonuclease then makes nicks in the phosphodiester backbone of the damaged strand on both sides of the thymine dimer and removes the defective oligonucleotide. DNA polymerase can then fill in the gap by synthesizing DNA in the 5' to 3' direction, using the undamaged strand as a template. Finally, the nick in the strand is sealed by DNA ligase.
Thymine Dimer Formation and Nucleotide Excision Repair
Base Excision Repair
- Alteration to bases can occur with other cellular insults. For example, thermal energy can be absorbed by DNA and may lead to cytosine reanimation. This is the loss of an amino group from cytosine and results in the conversion of cytosine and results in the conversion of cytosine to uracil. Uracil should not be found in a DNA molecule and is thus easily detected as an error; however, detection systems exist for small non-helix-distorting mutations in other bases as well.
- These are repaid by base excision repair. First, the affected base is recognized and removed by a glycosylase enzyme, leaving behind an apurinic/apyrimidic (AP) site, also called an abasic site. The AP site is recognized by an AP endonuclease that removes the damaged sequence from the DNA. DNA polymerase and DNA ligase can then fill in the gap and seal the strand.
Recombinant DNA Technology
- Allows a DNA fragment from any source to be multiplied by either gene cloning or polymerase chain reaction (PCR). This provides a means of analyzing and altering genes and proteins. It also provides the reagents necessary for genetic testing, such as carrier detection (detaching heterozygote status for a particular disease) and prenatal diagnosis of genetic disease; it is also useful for gene therapy.
- Additionally, this technology can provide a source of a specific protein, such as recombinant human insulin, in almost unlimited quantities.
Cloning Recombinant DNA
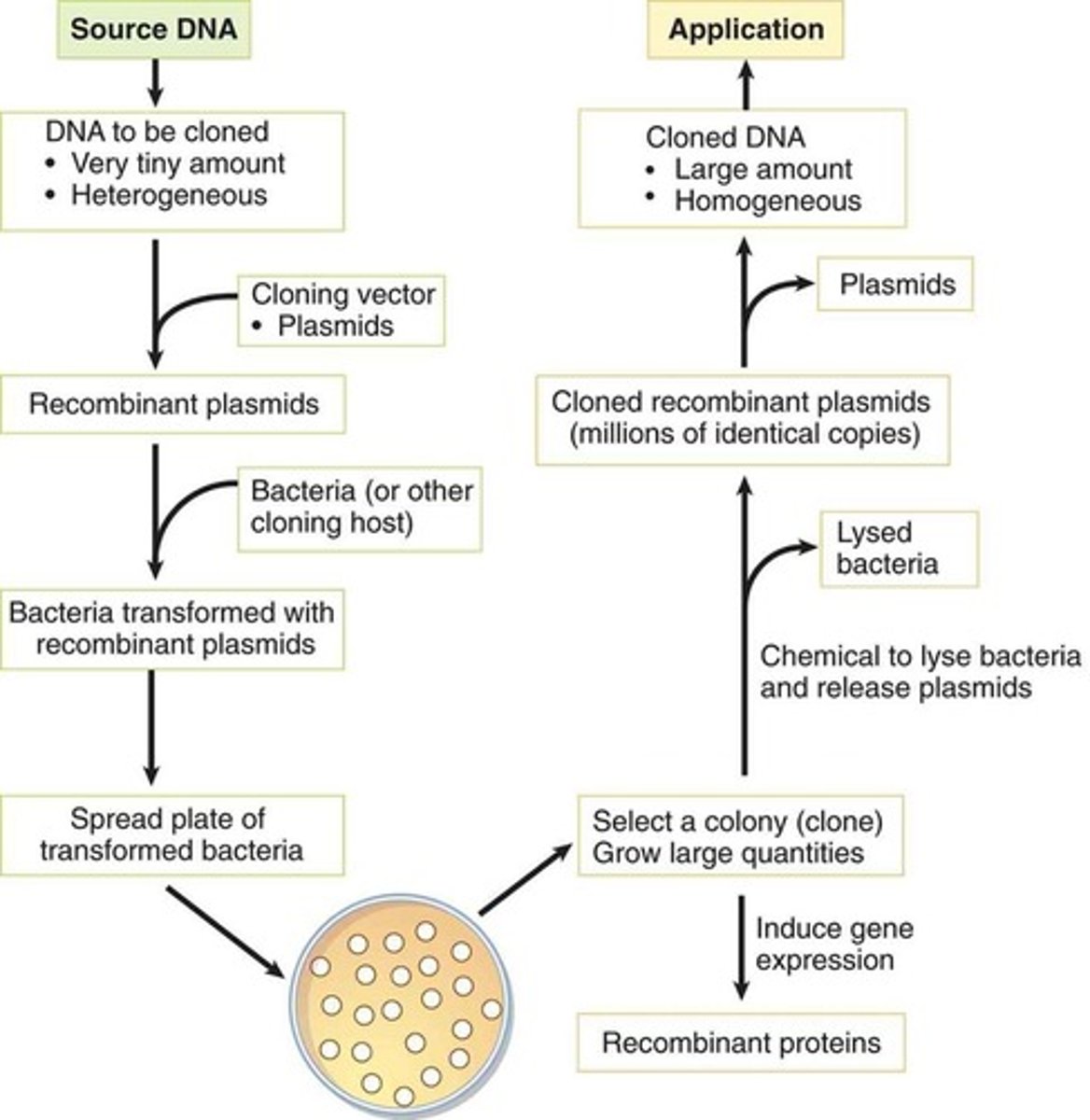
DNA Cloning
- A technique that can produce large amounts of a desired sequence.
- Often the DNA to be cloned is present in a small quantity and is part of a heterogeneous mixture containing other DNA sequences. The goal is to produce a large quantity of homogenous DNA for other applications.
- Cloning requires that the investigator ligate the DNA of interest into a piece of nucleic acid referred to as a vector, forming a recombinant vector. Vectors are usually bacterial or viral plasmids that can be transferred to a host bacterium after insertion of the DNA of interest.
- The bacteria are then grown in colonies, and a colony containing the recombinant vector is isolated. This can be accomplished by ensuring that the recombinant vector also includes a gene for antibiotic resistance; antibiotics can then kill off all of the colonies that do not contain the recombinant vector. The resulting colony can then be grown in large quantities.
- Depending on the investigator's goal, the bacteria can then be made to express the gene of interest (generating large quantities of recombinant protein), or can be lysed to resolute the replicated recombinant vectors (which can be processed by restriction enzymes to release the cloned DNA from the vector).
Restriction Enzymes
- Enzymes that recognize double-stranded DNA sequences. These sequences are palindromic, meaning that the 5' to 3' sequence of one strand is identical to the 5' to 3' sequence of the other strand (in antiparallel orientation).
- Isolated from bacteria, which are their natural source. In bacteria, they act as part of a restriction and modification system that protects the bacteria from infection by DNA viruses.
- Once a specific sequence has been identified, the restriction enzyme can cut through the backbones of the double helix.
Sticky Ends
- Some restriction enzymes produce offset cuts, yielding stick ends on the fragments.
- Sticky ends are advantageous in facilitating the recombination of a restriction fragment with the vector DNA. The vector of choice can also be cut with the same restriction enzyme, allowing for the fragments to be directly inserted into the vector.
DNA Vectors
- Contain at least one sequence, if not many, recognized by restriction enzymes.
- A vector also requires an origin of replication and at least one gene for antibiotic resistance to allow for selection of colonies with recombinant plasmids.
Formation of a Recombinant Plasmid Vector
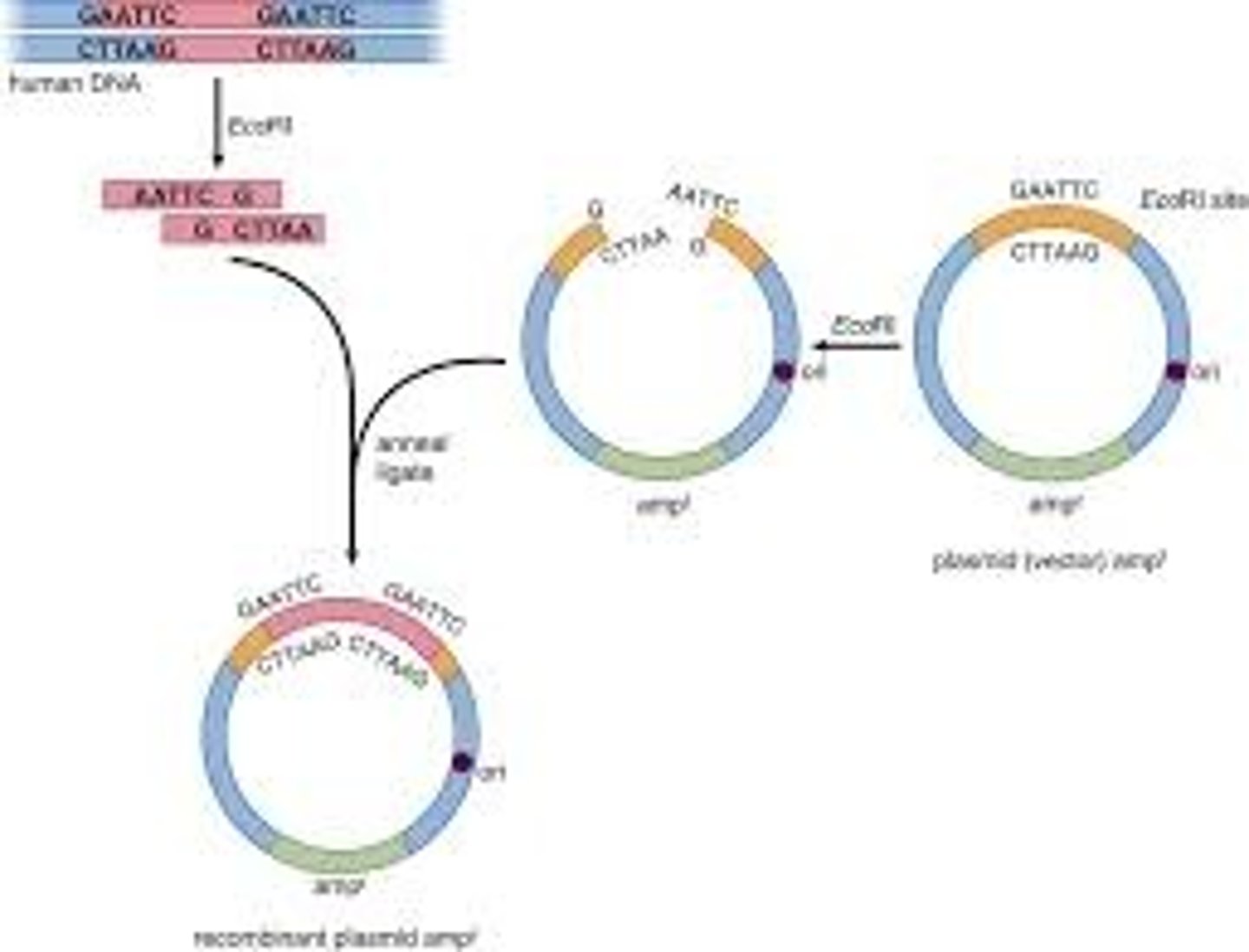
DNA Libraries
- Large collections of known DNA sequences; in sum, these sequences could equate to the genome of an organism.
- To make a DNA library, DNA fragments, often digested randomly, are cloned into vectors and can be utilized for further study.
- Libraries can consist of either genomic DNA or cDNA.
Genomic Libraries
- Contain large fragments of DNA, and include both coding (exon) and noncoding (intron) regions of the genome.
cDNA (complementary DNA) Libraries
- Constructed by reverse-transcribing processed mRNA. As such, cDNA lacks noncoding regions, such as introns, and only includes genes that are expressed in the tissue from which the mRNA was isolated. Thus, these libraries are sometimes called expression libraries.
- While genomic libraries contain the entire genome of an organism, genes may by chance be split into multiple vectors. Therefore, only cDNA libraries can be used to reliably sequence specific genes and identify disease-causing mutations, produce recombinant proteins (such as insulin, clotting factors or vaccines), or produce transgenic animals.
Cloning Expressed Genes by Producing cDNA
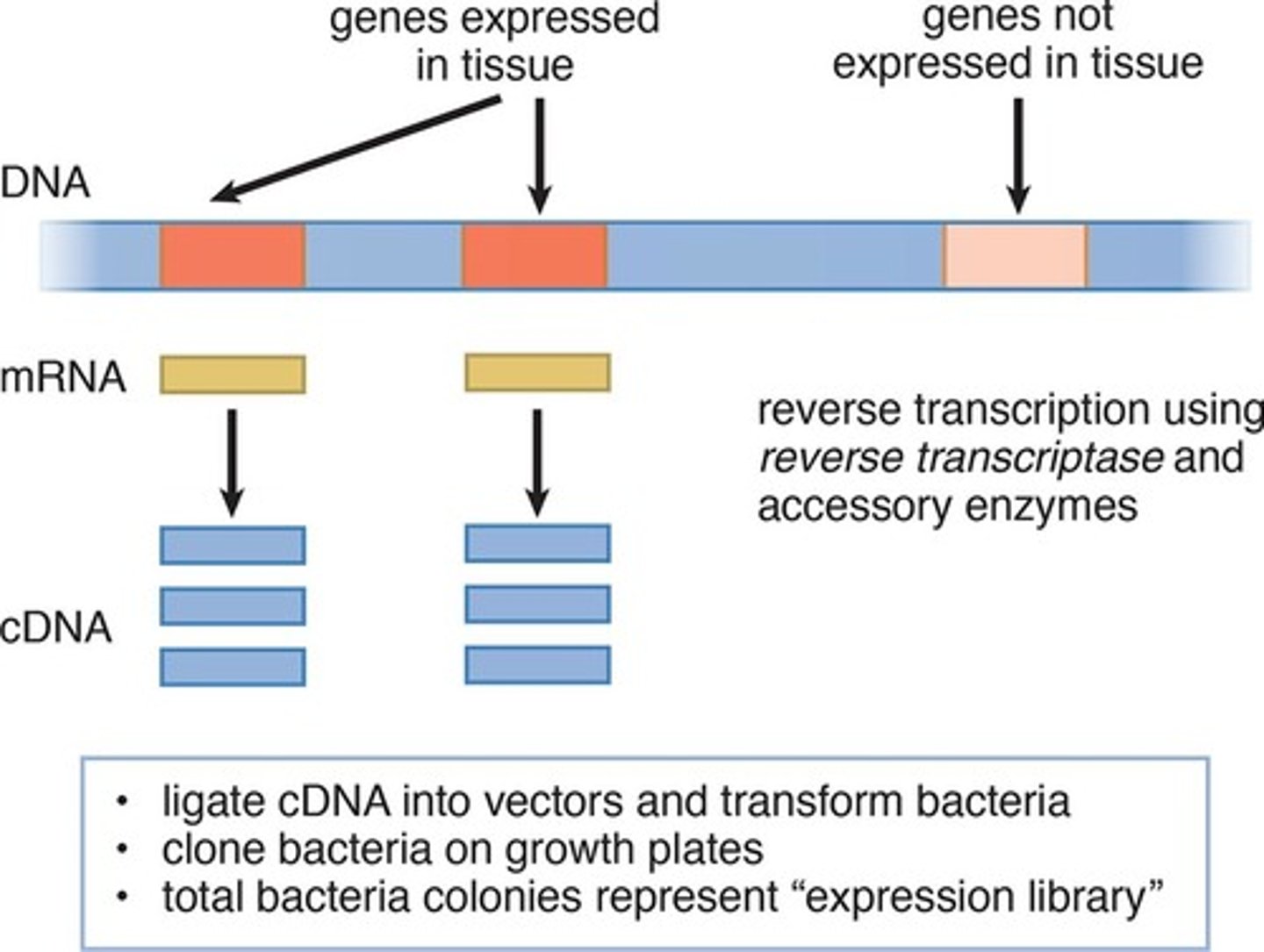
Hybridization
- The joining of complementary base pair sequences. This can be DNA-DNA recognition or DNA-RNA recognition.
- Uses two single-stranded sequences and is a vital part of PCR and Southern blotting.
Polymerase Chain Reaction
- An automated process that can produce millions of copies of a DNA sequence without amplifying the DNA in bacteria.
- Knowing the sequences that flank the desired region of DNA allows for the amplification of the sequence win between.
- Requires primers that are complementary to the DNA that flanks the region of interest, nucleotides and DNA polymerase. The reaction also needs heat to cause the DNA double helix to denature. Unfortunately, the DNA polymerase found in the human body does not work at high temps. Thus, the DNA polymerase from Thermus aquatics, a thermophilic bacteria is used instead.
- During PCR, the DNA of interest is denatured, replicated and then cooled to allow reannealing of the daughter strands with the parent strands. This process is repeated several times, doubling the amount of DNA with each cycle, until enough copies of the DNA sequence are available for further testing.
Gel Electrophoresis
- A technique used to separate macromolecules, such as DNA and proteins, by size and change.
- All molecules of DNA are negatively charged because of the phosphate groups in the backbone of the molecule, so all DNA strands will migrate toward the anode of an electrochemical cell.
- The preferred gel for DNA electrophoresis is agarose gel, and the longer the DNA strand, the slower it will migrate in the gel.
Southern blot
- Used to detect the presence and quantity of various DNA strands in a sample.
- DNA is cut by restriction enzymes and then separated by gel electrophoresis. The DNA fragments are then carefully transferred to a membrane, retaining their separation. The membrane is then probed with many copies of a single stranded DNA sequence. The probe will bind to its complementary sequence and form double-stranded DNA. Probes are labeled with radioisotopes or indicator proteins, both of which can be used to indicate the presence of a desired sequence.
DNA Sequencing
- A basic sequencing reaction contains the main players from replication, including template DNA, primers, an appropriate DNA polymerase, and all four deoxyribonucleotide triphosphates. In addition, a modified base called a dideoxyribonucleotide is added in lower concentrations. Dideoxyribonucleotides contain a hydrogen at c-3', rather than a hydroxyl group; thus, once one of these modified bases has been incorporated, the polymerase can no longer add to the chain. Eventually, the sample will contain many fragments (as many as the number of nucleotides in the desired sequence), each one of which terminates with one of the modified bases. These fragments are then separated by size using gel electrophoresis. The last base for each fragment can be read, and because gel electrophoresis separates the strands by size, the bases can easily be read in order.
Gene Therapy
- Intended for diseases in which a given gene is mutated or inactive, giving rise to pathology. By transferring a normal copy of the gene into the affected tissues, the pathology should be fixed, essentially curing the individual.
- For gene replacement therapy to be a realistic possibility, efficient gene delivery vectors must be used to transfer the cloned gene into the target cells' DNA. Because viruses naturally infect cells to insert their own genetic material, most gene delivery vectors in use are modified viruses. A portion of the viral genome is replaced with the cloned gene such that the virus can infect but not complete its replication cycle.
- Randomly integrated DNA poses a risk of integrating near and activating a host oncogene.
Transgenic Mice
- Once DNA has been isolated, it can be introduced into eukaryotic cells. Transgenic mice are altered at their germ line by introducing a cloned gene into fertilized ova or into embryonic stem cells. The cloned gene that is introduced is referred to as a transgene. If the transgene is a disease-producing allele, the transgenic mice can be used to study the disease process from early embryonic development through adulthood.
Knockout Mice
- A gene has been intentionally deleted.