Genetics test 3 Dr. Seibenhener
1/111
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
112 Terms
what most people think of when they hear the word chromosome?
metaphase chromosome
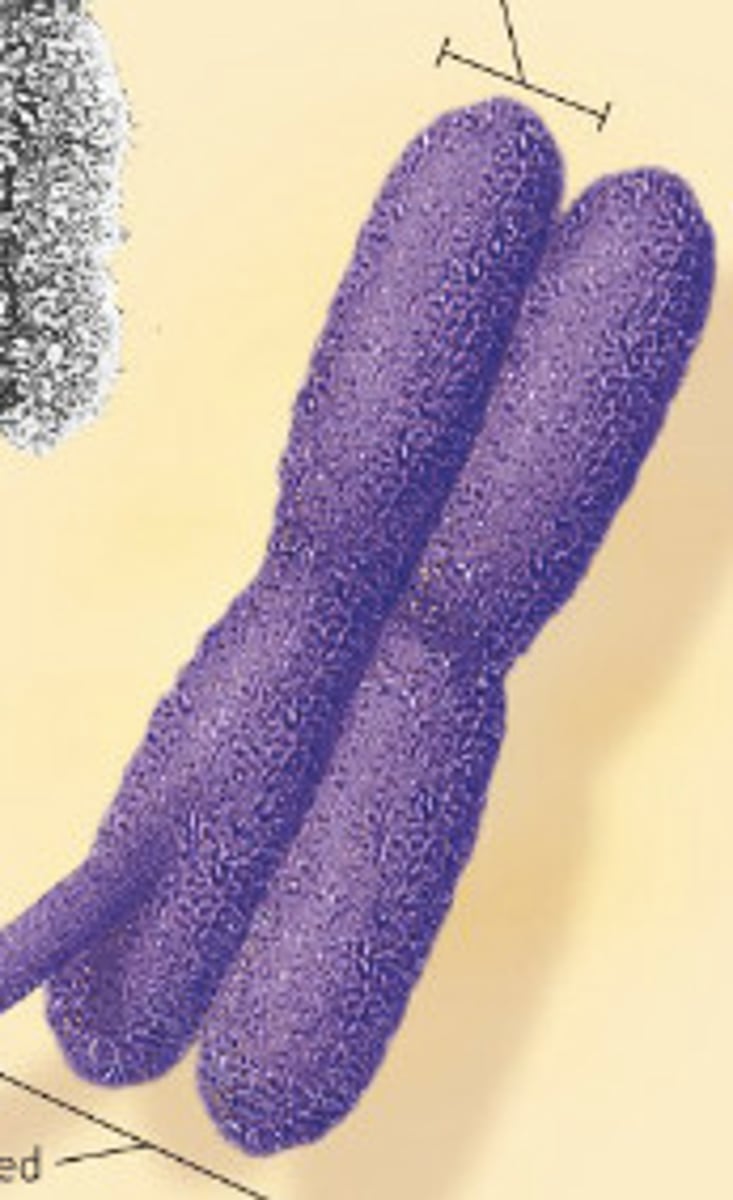
general chromosome infomation
humans have 22 pairs of autosomes and 1 pair of sex chromosomes (allosomes)
for a total of 23 pairs of chromosomes (46 chromosomes)
about 6 billion bp in every cell ~2 meters of DNA in a 10 nm nucleus
chromatin condensation
DNA
Chromatin
Metaphase Chromosome
what is chromatin?
DNA + protein (histones)
2 types of chromatin
euchromatin and heterochromatin
euchromatin (active)
lightly packed chromatin rich in gene concentration and is most often under active transcription
heterochromatin (silent)
tightly packed chromatin consisting mainly of the genetically inactive sequences
examples of heterochromatin
telomeres
centromeres
why are the genes in heterochromatin inactive?
it is bound so tightly that they cannot be accessed by transcription mechanisms
2 types of heterochromatin
1. Constitutive
2. Facultative
constitutive heterochromatin
Regions that are always heterochromatic
Permanently inactive with regard to transcription, as tightly packed as you can get
Usually contain highly repetitive sequences
Example: centromeres
Constitutive heterochromatin is located in
the centromeres of all chromosomes
facultative heterochromatin
"change"
Regions that can interconvert between euchromatin and heterochromatin (less tight to tight)
facultative heterochromatin example
heat shock proteins
-you don't need them all the time, they can remain in facultative until they are needed and then they change to become accessible
every cell in the body has the same
genetic information
models to explain association of DNA and proteins
1. folded fiber model
2. nucleosome model
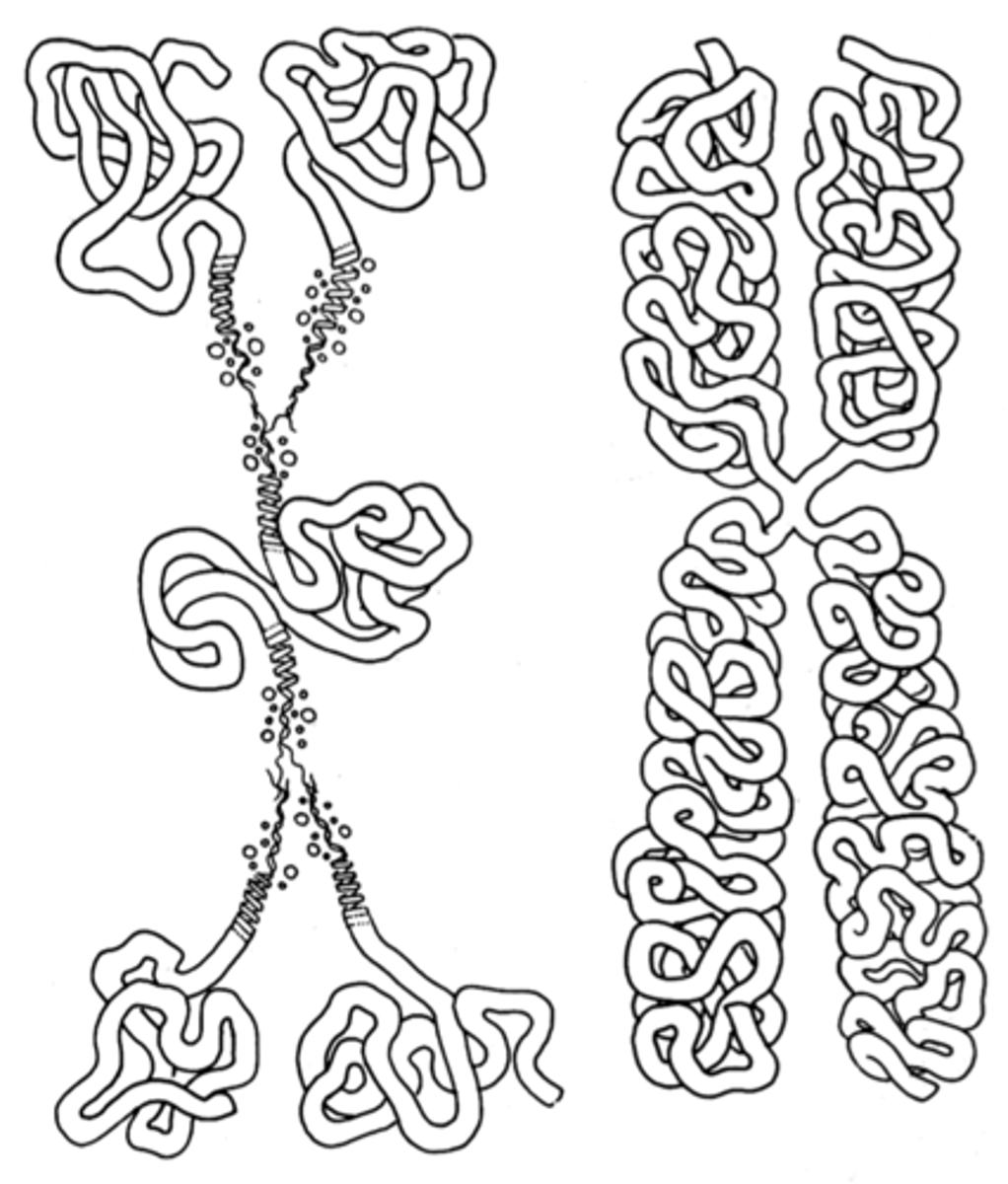
Folded fiber model
A model of eukaryotic chromosome organization in which each sister chromatid consists of a single chromatin fiber composed of double-stranded DNA and proteins wound like a tightly coiled ball of yarn (DNA IS RANDOMLY PACKED)
-whole mounts of human WBC
-found few or no free fiber ends and concluded each chromatid must consist of a single fiber
-0.2 nm double helix of DNA
Type A fiber 1-10 nm
Type B fiber 20-25 nm (extensive type B folding forms a chromatid)
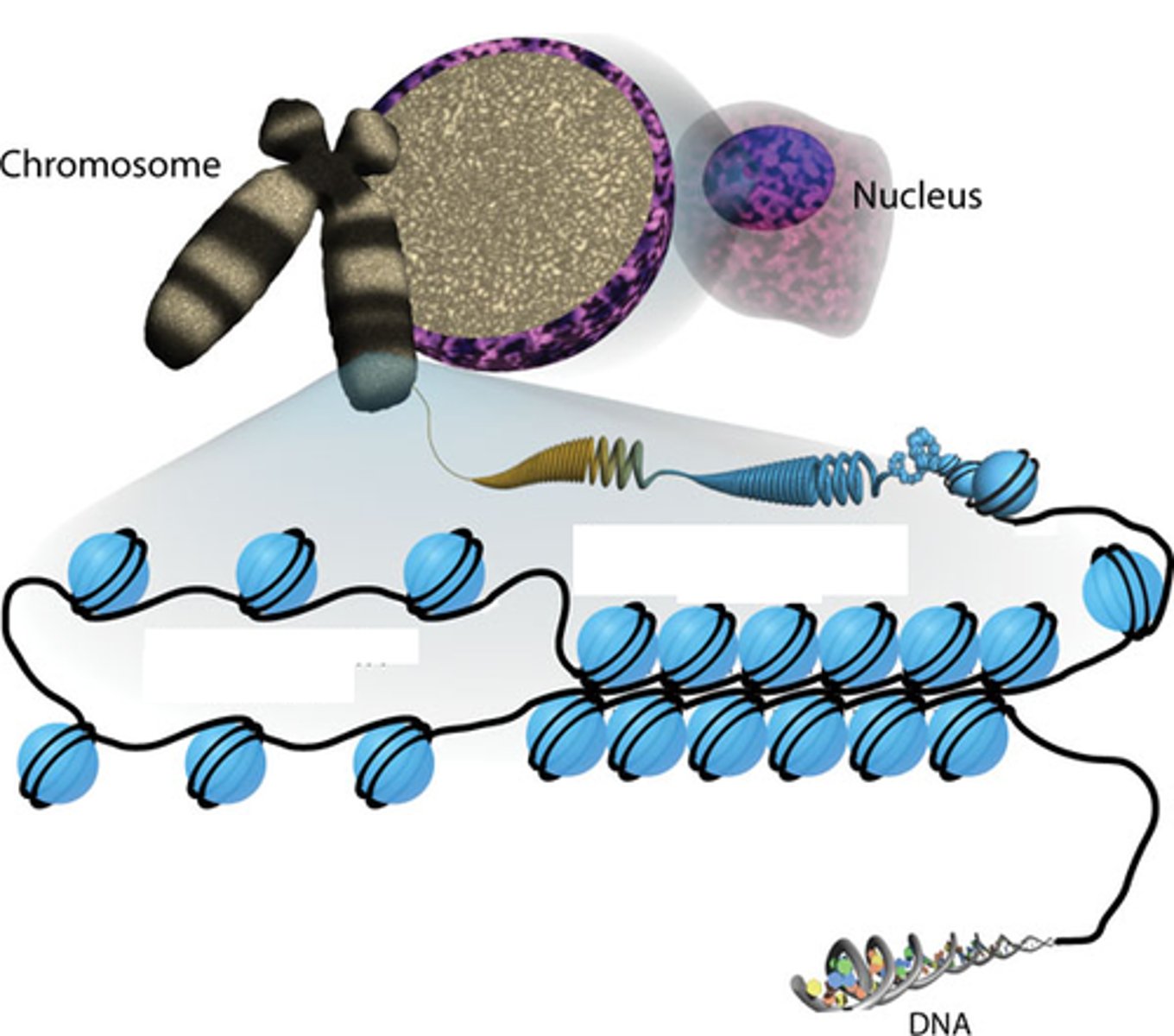
Nucleosome Model
Most commonly accepted model for DNA packaging
This model allows for more packaging than folded fiber, more organized
nucleosome
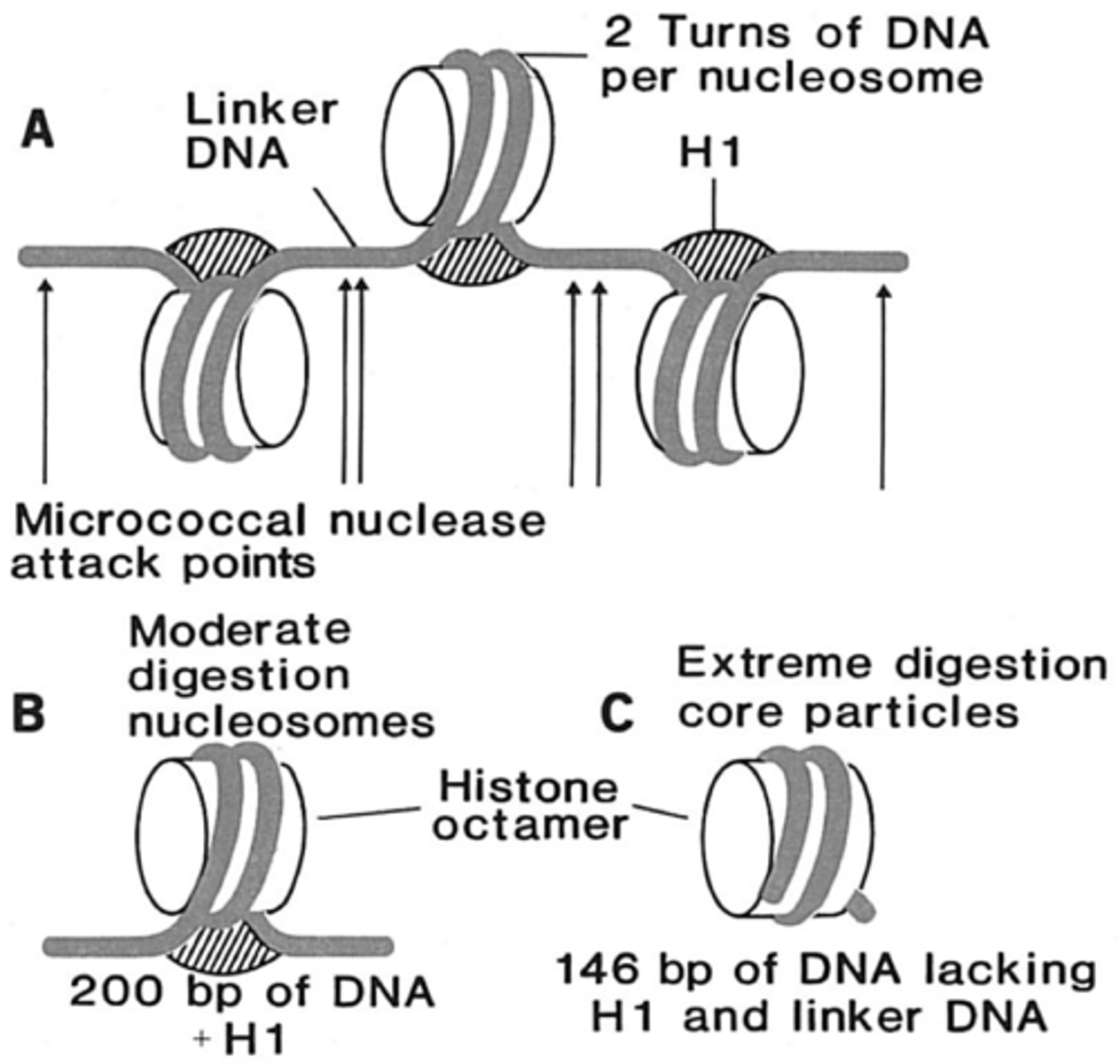
nucleosome
simplest packaging structure of all eukaryotic chromatin
classes of histones
core and linker
core histones
The four histone proteins (H2A, H2B, H3 and H4) that form the octameric core of a nucleosome
consists of approx. 120 amino acids each
highly conserved
in combination form the core particle
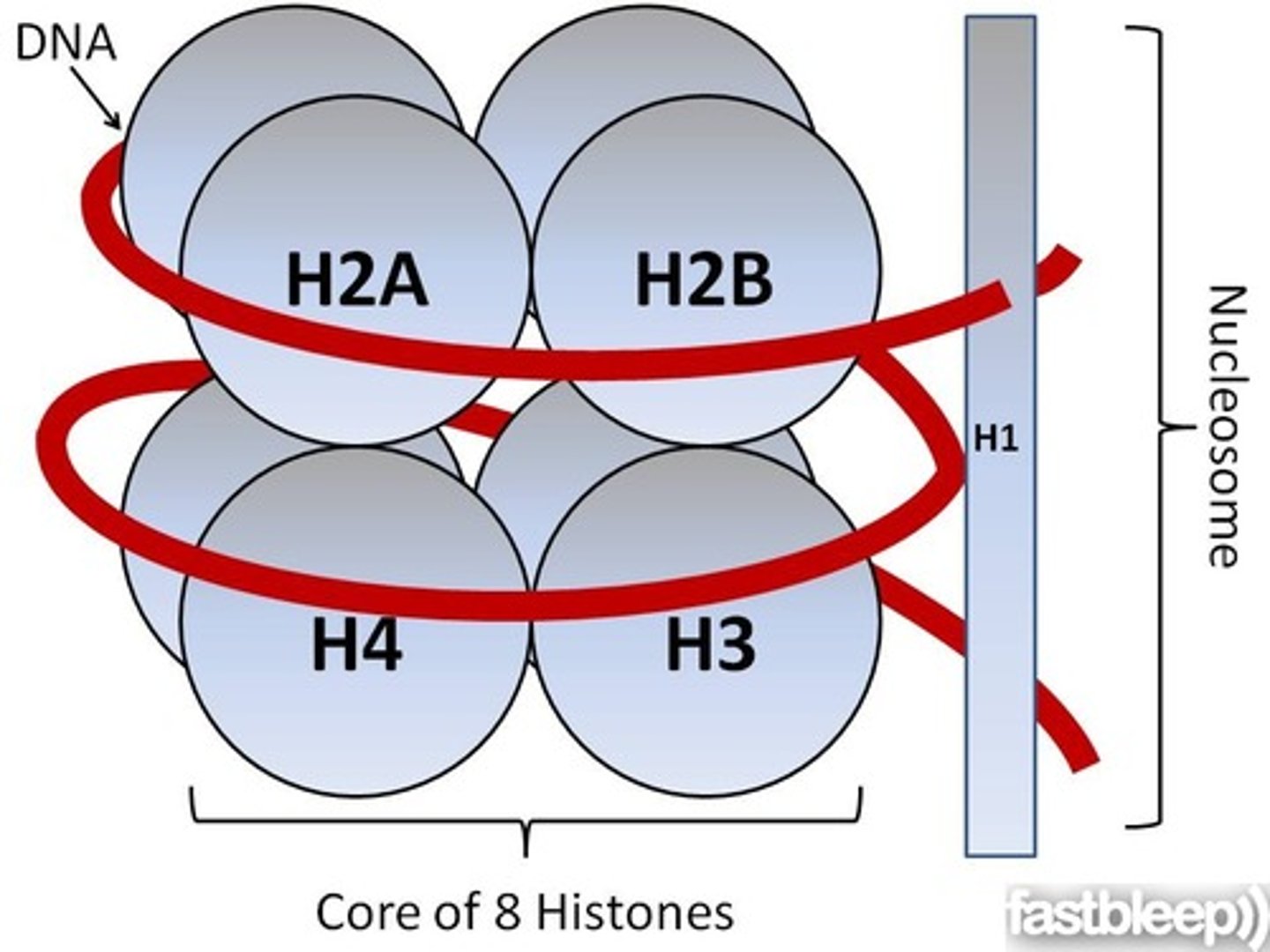
linker histone
The histone protein H1, which binds to the linker DNA adjacent to the nucleosome.
consist of approx. 200 amino acids
tissue specific expression and not highly conserved
loosely associated with core particle
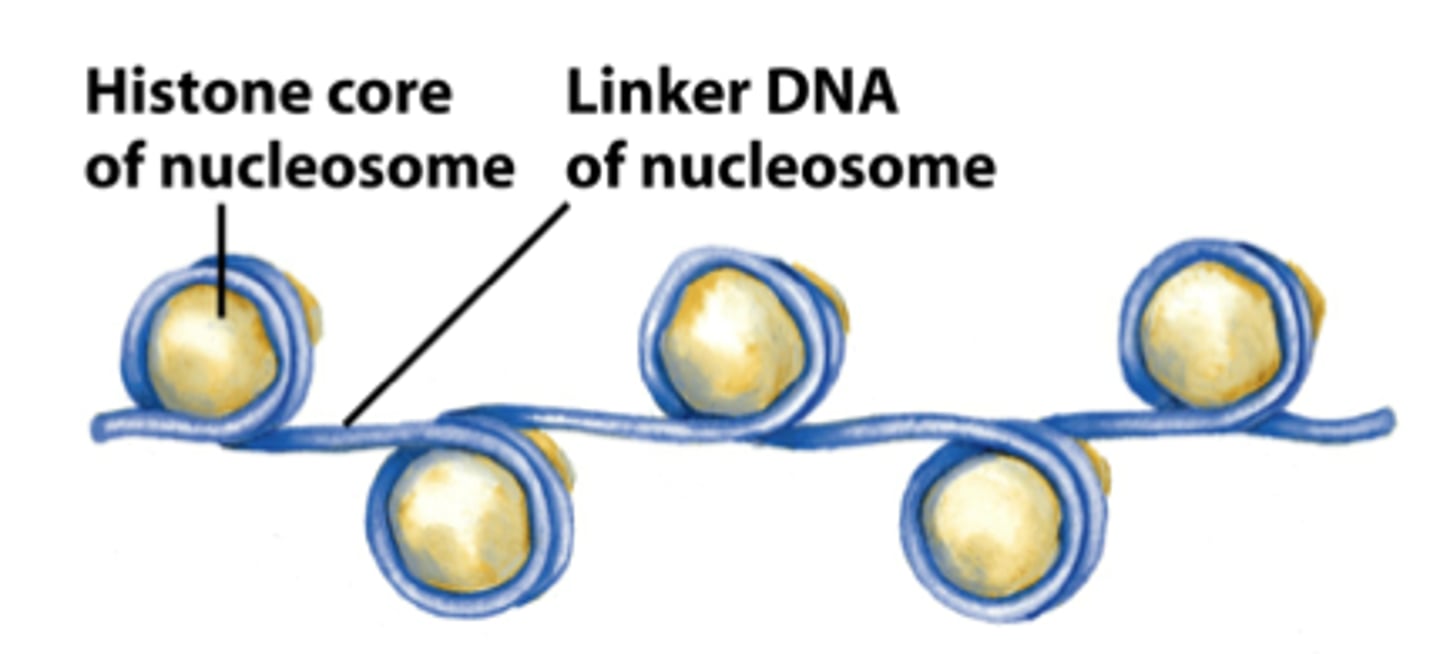
each of the four core histones are __________, leading to the structure of an octamer
dimers
parts of the nucleosome
the octamer (8 histones)
the DNA
and the H1 linker
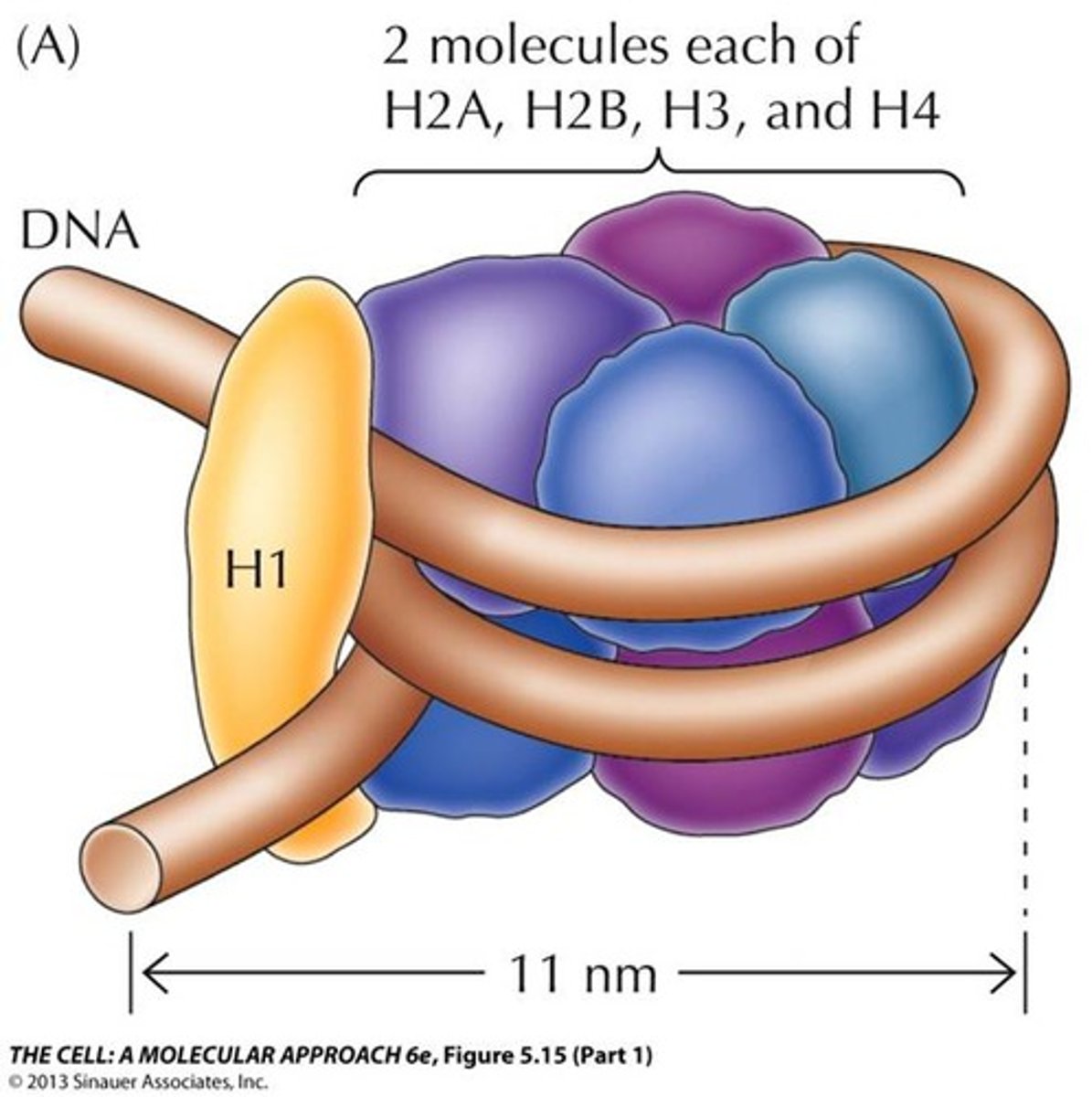
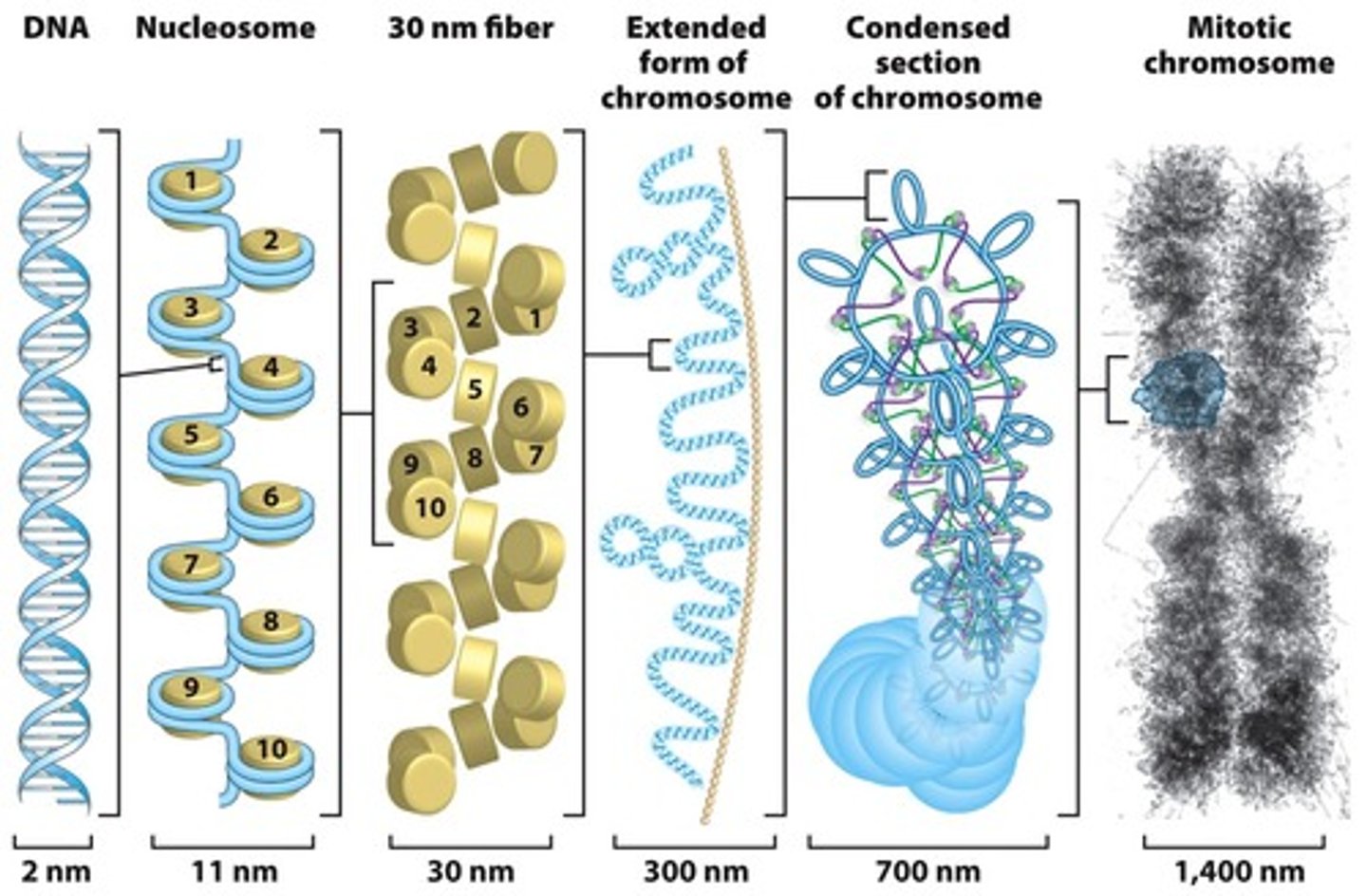
formation of the nucleosome
DNA 2 nm
nucleosomes 10 nm
soleniod, zig zag 30 nm
chromatin loops 300 nm
condensed chromatin loops 700 nm
chromosome 1400 nm
formation of the nucleosome: DNA
200 bp DNA
-146 bp wrap core
-54 bp link to the next nucleosome by linker DNA
reduce DNA length by ~7 times
linear arrangement
10 nm fiber: primary packaging of the chromatin
histone tails
strings of amino acids that protrude from the histone proteins in the nucleosome
are flexible
allows histone protein to grab onto the DNA
"beads on a string"
nucleosome
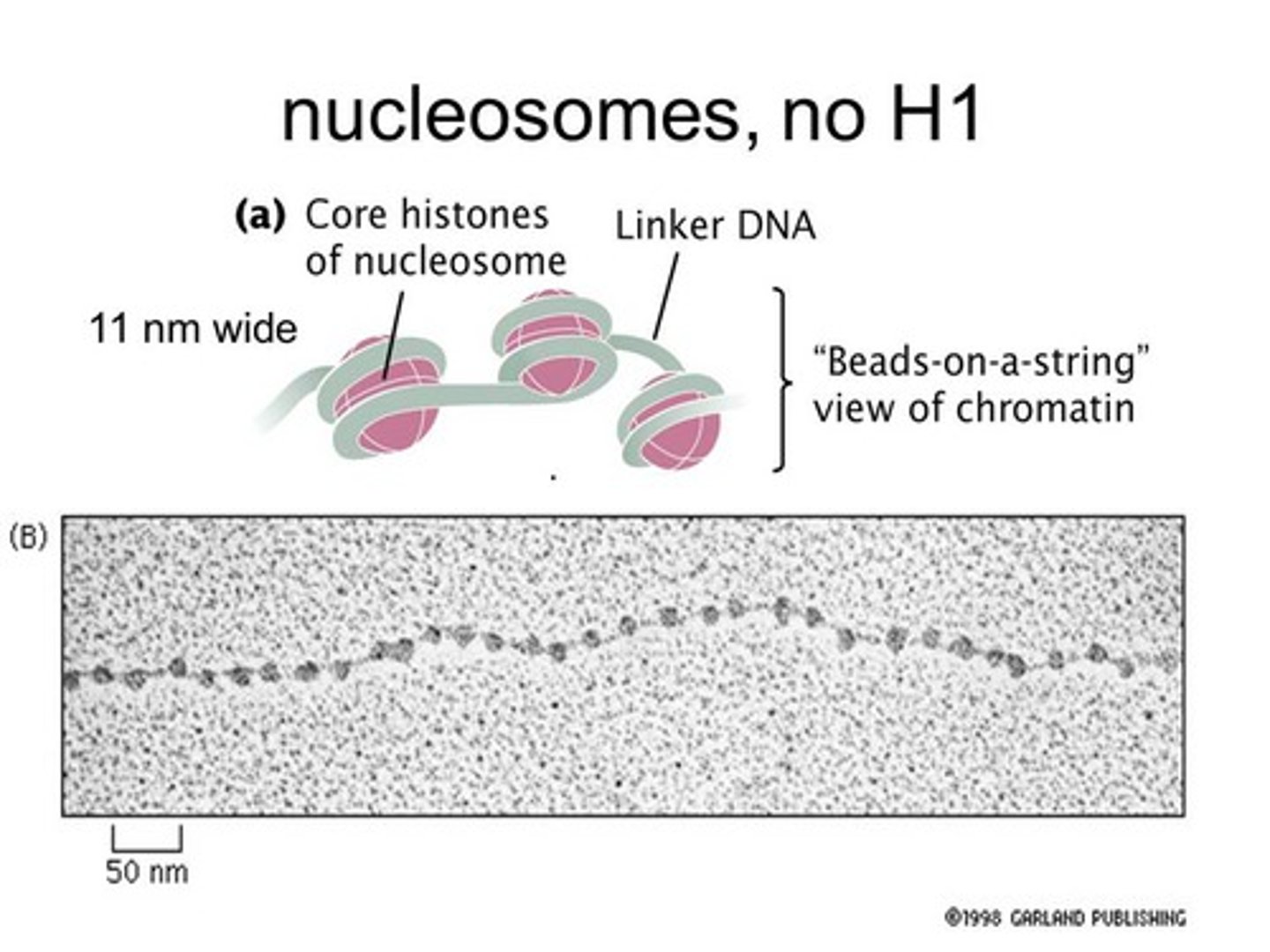
supercoiling of DNA- formation of the solenoid
H1 histone is responsible for packaging. it pulls it all together
Solenoid: helical coiling of 10 nm fibers consisting of 6 nucleosomes
-The 30 nm particle
Supercoiling- reduces the length of DNA by about 7 times
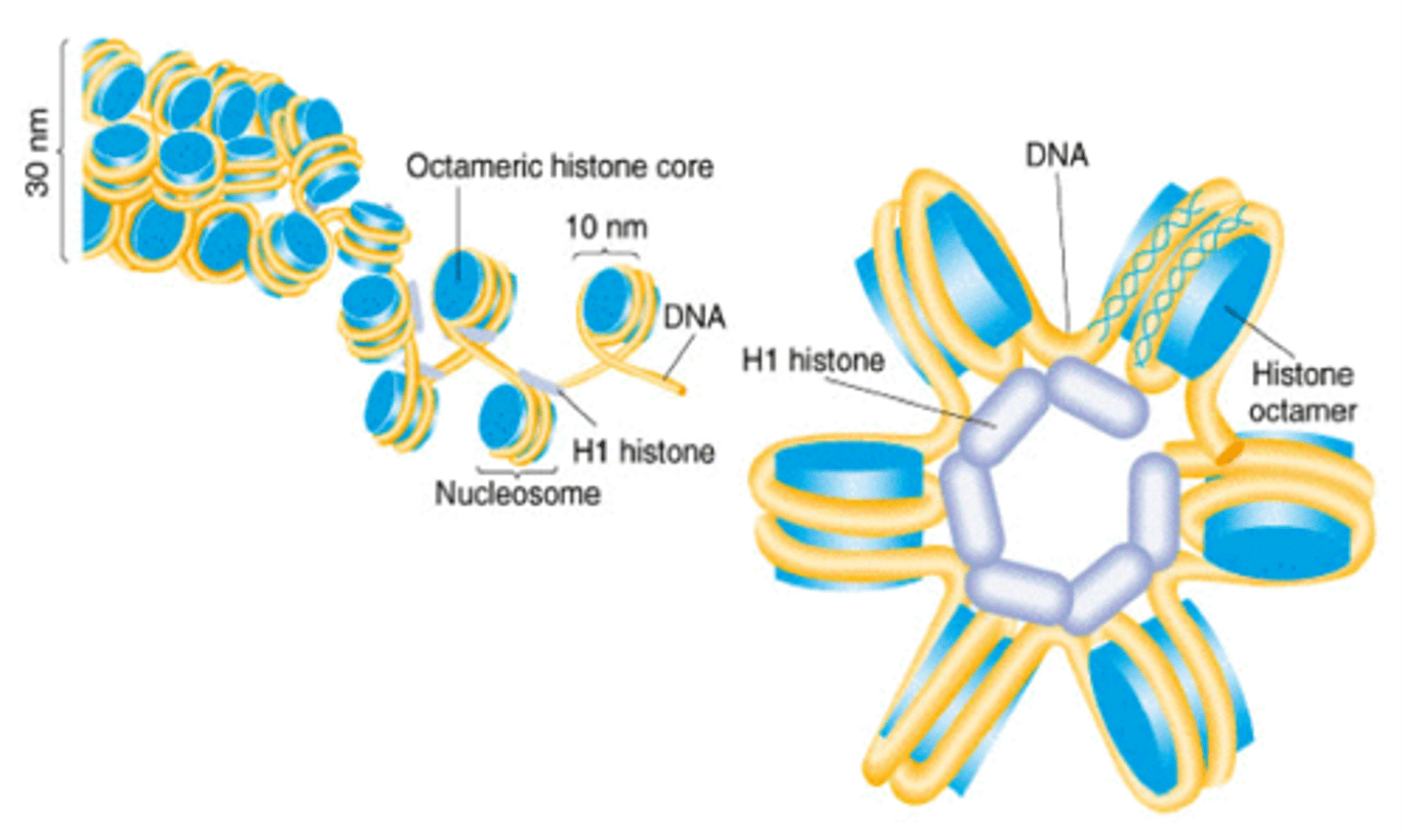
how does the histone H1 work?
it binds 2 distinct region:
1. linker DNA
2. portions of the 146 bp core
40x compaction, induces tighter DNA wrapping around histones, makes the solenoids come together
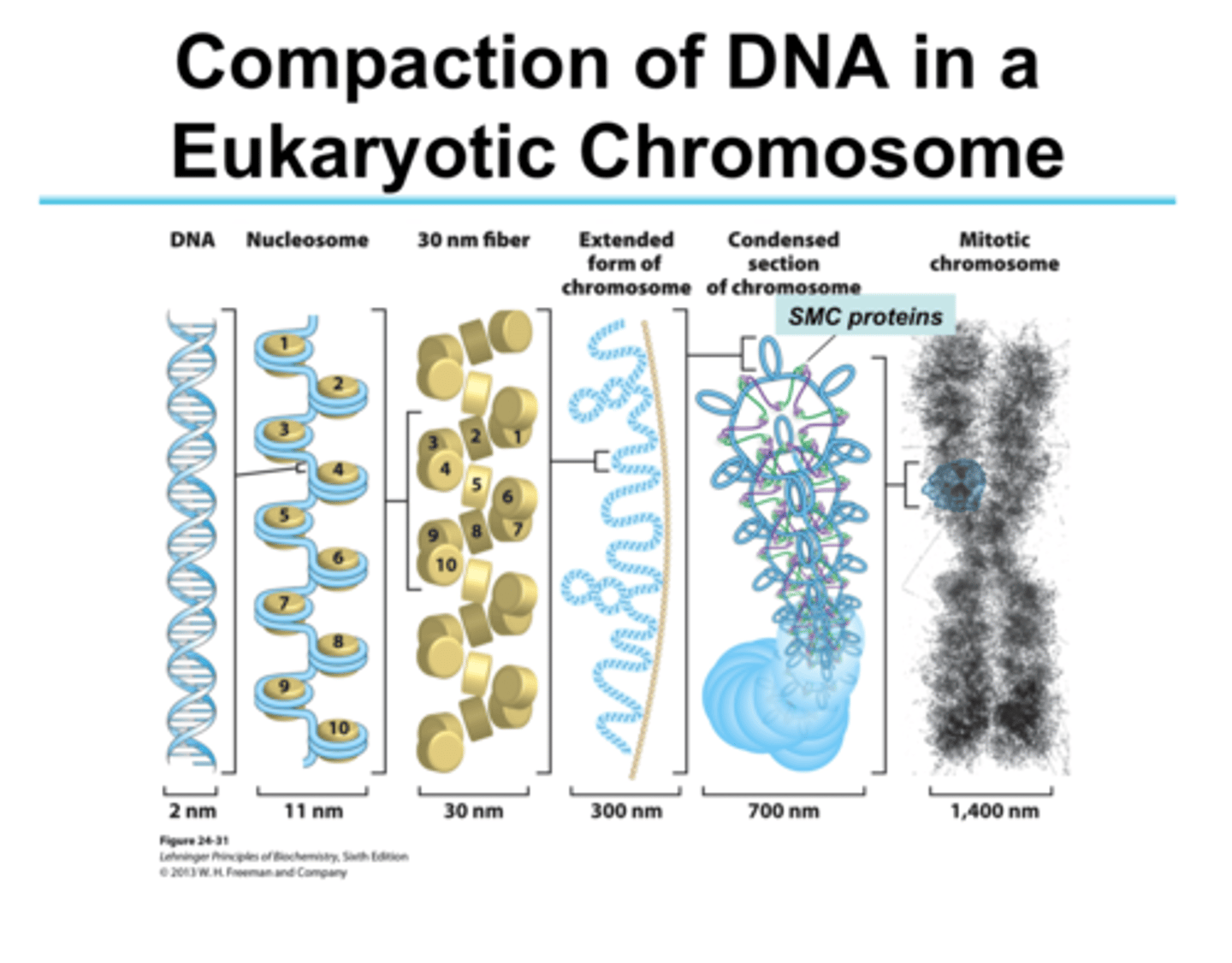
Alternate model of super coiling in DNA: the Zig Zag model
the DNA backbone (and solenoid) is not flexible enough to bend between nucleosomes; straight linker DNA connects opposite nucleosomes
The zig zag model takes the bends out, stacks a few solenoids on top of each other, allows more compaction because it puts DNA in the middle
Is still a 30 nm model
experimental results show that both the solenoid and zig zag topologies may both be present in the chromatin fiber
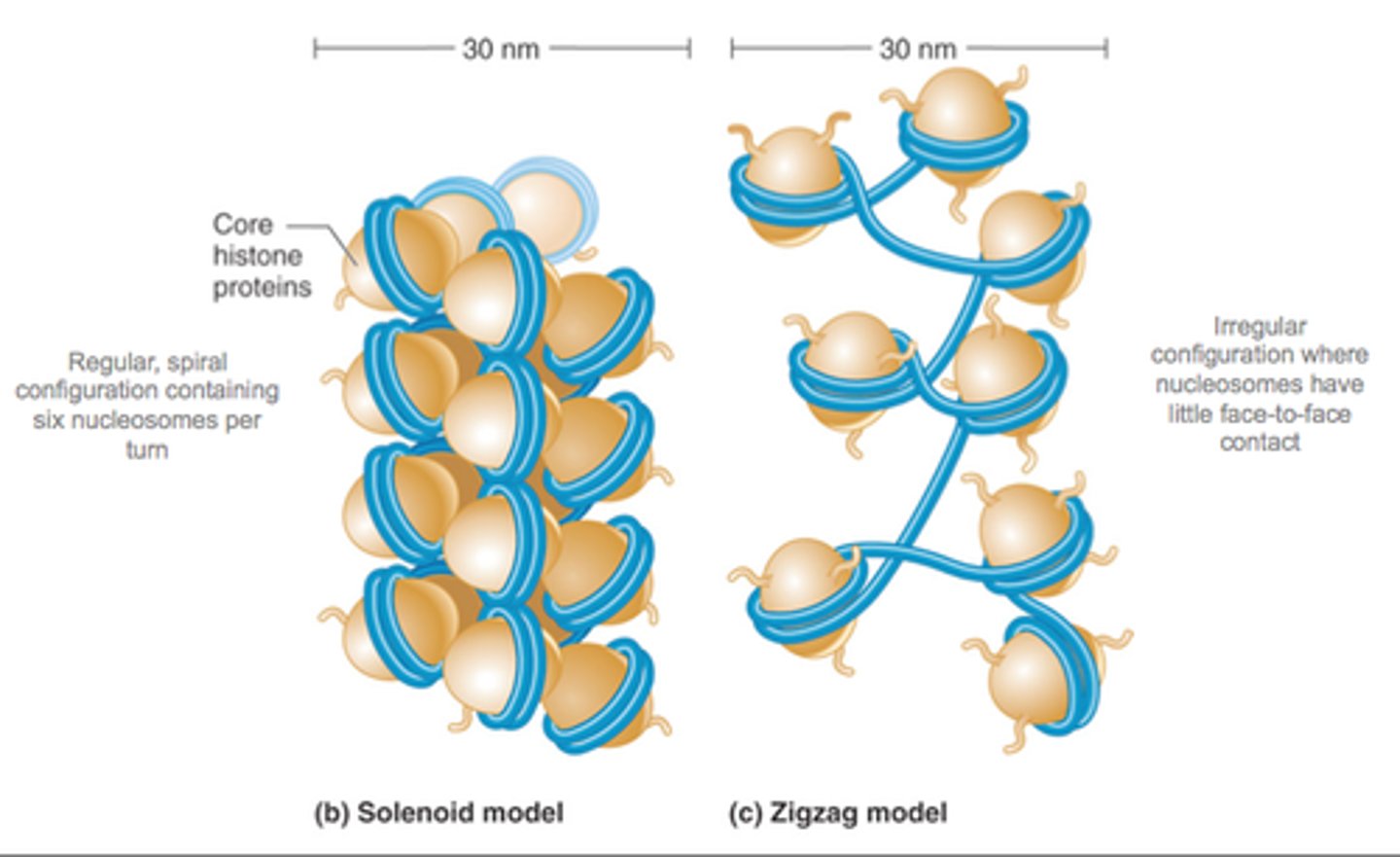
higher order coiling in the 300 nm fiber: chromatin loops
rosettes of chromatin loops are built around a scaffolding of topoisomerase 2
the loops have solenoids
have a compaction level of EUCHROMATIN (loose, genes are accessible, transcription can happen)
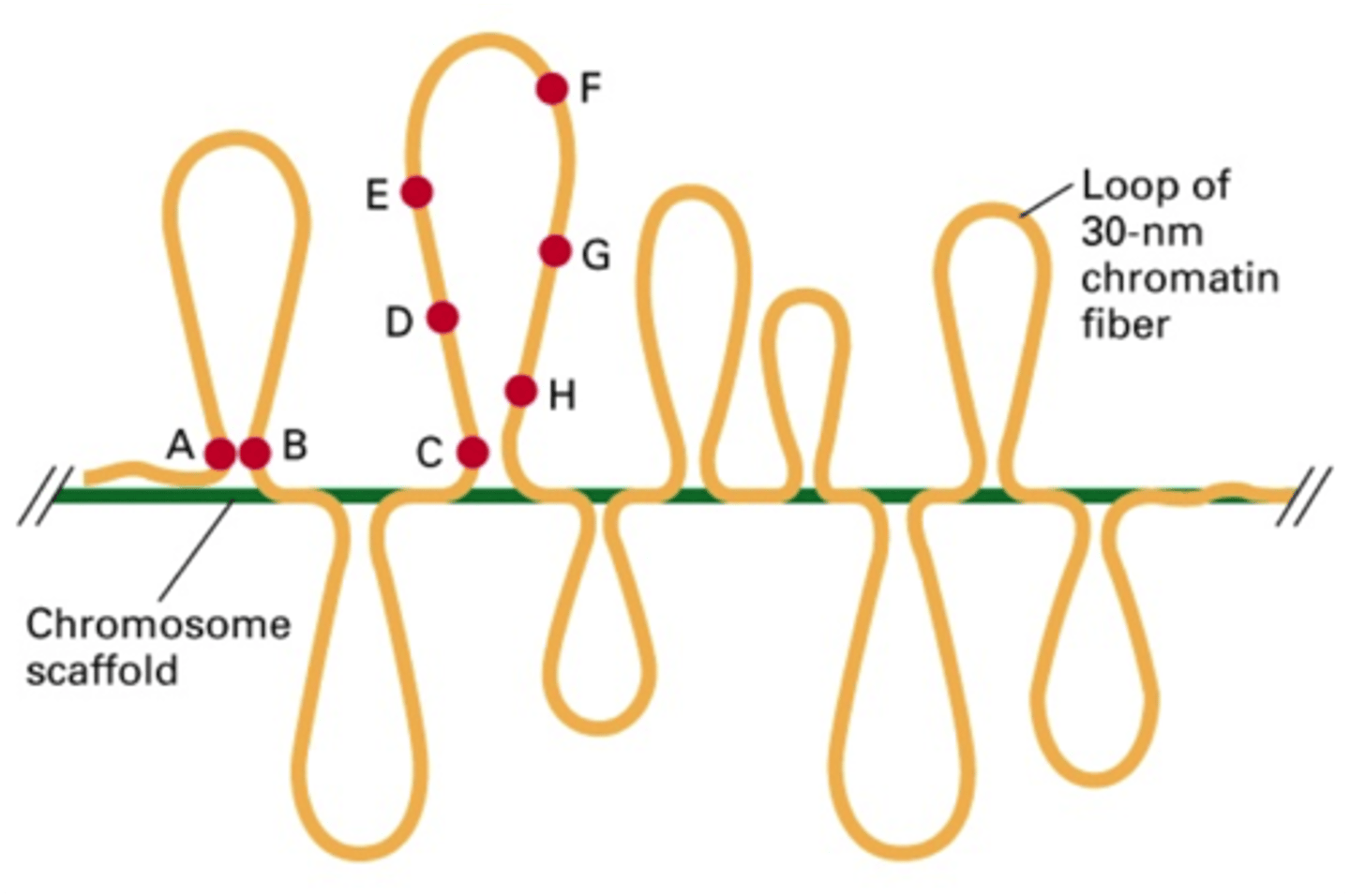
final condensation: the metaphase chromosome
DNA diameter about 700 nm
spiral scaffolding composed of topoisomerase 2 and about 15 non-histone proteins
compaction level of HETEROCHROMATIN
Chromatin Formation and the Cell Cycle
...
interphase
DNA is replicated in the S phase, DNA is accessible and lightly packed (G1, S, and G2 phase)
Early prophase
chromosome condenses, you cant access the genes
Late prophase
nuclear envelope breaks down, mitotic spindle assembles
Metaphase
mitotic spindle arranges chromosomes at cell's equator
anaphase
sister chromatids separate/torn apart
telophase
cell plate forms, cell division in cytokinesis; then we go back into G1 of interphase and it starts over
where the chromatin structures are during cell division
2 nm fiber-10 nm: not compacted, G1 phase
30 nm fiber- end of the S
300nm fiber- G2 phase
700 nm chromatin- early prophase
chromosome: in metaphase of mitosis
karyotype
picture of metaphase chromosomes in the nuclues of eukaryotic cells
let us looks at: chromosome size, position, and hetero vs euchromatin
we have 46
apes have 48
Down's Syndrome (Trisomy 21)
primary risk factor: mothers age
parents usually genetically normal
failure of chromosome 21 to separate in sperm or egg
Klinefelter's syndrome (XXY)
parents usually genetically normal
failure to separate the X chromosome during meiosis
true of false: large DNA molecules must be highly condensed to fit within the nucleus of a cell
true
stages of condensation
nucleosomes
solenoids
chromatin loops
condensed chromatin
chromatin folded around a protein scaffold
chromosome condensation is intimately associated with the
cell cycle
Pre-Darwian "great chain of being" thinking
Where human genome considered the most complex, but does the amount of DNA actually lead to complexity?
biased data set based on already sequenced "small" genomes
Junk DNA
discovery of repetitive DNA sequences
What defines complexity? (2)
1. number and types of cells
2. degree of cellular organization
C-value paradox
genome size does not correlate with organismal complexity
-excess (junk) DNA is present in the genome that does not seem to be essential for the development/evolutionary divergence of an organism
Gregory's Onion test
an onion
-diploid 2n=16
-haploid n=17 pg
-so, 17x 10^6 bp (5X MORE NON-CODING DNA)
humans
-diploid 2n=48
-haploid n=3.5 pg
-so, 3.5x 10^6 (LESS THAN AN ONION)
human vs onion: which one is more advanced?
amount of DNA: onion
number of genes: both the same
nutrition: onion
alternative splicing: humans
transcription factors: humans
genome size and cell volume
We are more complex because we have higher levels of control of our genome
Genome size and cell volume: plant cells typically larger than animal cells, DNA has a structural element in defining nucleus shape
Coding sequences in the human genome
2-5% of the human genome
about 20k protein coding genes
G-value paradox
the number of genes does not correlate with organism's complexity
C vs G value paradox
genome size vs number of genes
Types of DNA in the genome: 3 classes of nucleotide sequences
1. highly repetitive DNA sequence (HR)
2. moderately repetitive DNA sequence (MR)
3. single copy DNA sequence (unique)
highly repetitive DNA sequence (HR)
10 % of the human genome
mostly in heterochromatin regions around centromere/telomere
"non coding DNA regions"
function range: structural and organizational role to junk
example: alpha satellite DNA
moderately repetitive DNA sequence (MR)
30% of human genome
mostly in euchromatin
300 bp in size
10-10^6 copies per genome
includes "redundant" genes for histones, rRNA, and proteins
example: micro satellite DNA (repetitive sequences 2-5 bp) and interspersed repetitive DNA (transposable elements)
Microsatellites
variable number of tandem repeats typically occurring in non-coding regions of the genome
useful genetic markers as they tend to by highly polymorphic
used to sequence the human genome, markers for certain diseases, primary markers for DNA testing in forensics
can be di, tri, or tetra
occurs through a mutation process knows as "slippage recognition"
-repetitive sequence can cause slip forwards (skip info) or slip backwards (replication of old info)
single copy DNA sequence (unique)
1-5% human genome
most thoughout euchromatin
genes= "coding DNA regions"
about 20k genes
present at single or low copy number
HR+ MR+ unique
10 + 30 + 5= 45%
not 100%, the rest is junk DNA
where do we find the different types of DNA in the genome?
1. HR: in the heterochromatin, tightly coiled
-telomeres and centromeres
2. MR: scattered throughout euchromatin
-facultative heterochromatin
3. unique: euchromatin
-GENES
What is a gene?
the basic physical and functional unit of heredity
gene
sequence of unique nucleotides (genotype) that carry the genetic information which is to be expressed (phenotype)
the instruction manual for our bodies
every person has 2 copies of each gene
molecular level "gene"
DNA sequence that gives rise to an RNA molecule
the transcriptional unit
extends from the promoter to the terminator
On the DNA:
*regulatory sequence: site for the binding of regulatory proteins
1. promoter: signals beginning of transcription
2. transcribed region: part of this region contains the information that specifies an amino acid sequence
3. terminator: signals the end of transcription
is transcribed to mRNA
exon
coding sequences= phenotype
introns
intervening sequences= areas of genes that do not generally code for phenotype
flanking regions
5' untranslated region and 3' untranslated region
5' untranslated region (5' UTR)
mRNA that is directly upstream from the initiation code
-5' region untranslated forms a hairpin
3' untranslated region (3' UTR)
section of mRNA that immediately follows the translation termination codon
promoter
is a DNA sequence onto which the transcription machinery binds and initiates transcription
TATA box
highly conserved sequence in DNA serving as the binding site for transcription factor binding
core DNA sequence: 5'- TATAAA- 3'
on/off switch for transcription, where transcription factors bind
basal level of transcription (to produce one gene product)
Regulatory sequences
CAAT box
GC box (enhancer)
CAAT box
5'- GGCCAATCT- 3'
consensus sequence that occurs upstream by 60-100 bases to the initial transcription site; typically required for inducible genes to be produced in sufficient amounts
GC box
ENHANCER region of the DNA that can be bound with proteins (activators) to activate transcription of a gene or genes
"ramped up" level of transcription
termination/terminator
region of DNA where transcription ends
termination: the end of the genes
terminator: section of nucleic acid sequence that marks the end of a gene or operon in genomic DNA during transcription
helps identify the addition of the poly-A tail to the transcript
start site of transcription
+1
AUG codon codes for start
5 types of genes
1. solitary genes (unique)
2. duplicated genes
3. multigene families
4. psuedogenes
5. repeated genes
solitary gene (unique)
-a single copy of a gene (haploid situation); 2 copies in diploid
-comprises the bulk of euchromatin
duplicated genes
process by which a portion of a chromosome is duplicated in an additional copy of a gene
-results in a copy of the original gene called a paralog gene
-either of the 2 genes may mutate and change the original function of the gene
-usually occurs due to an error during meiosis
multigene families
-set of several similar genes, formed by duplication of a single original gene, and generally with similar biochemical functions
-most often located in similar regions of the chromosome
-most often used or synthesized at different times
psuedogenes
-dysfunctional relatives of genes that have lost their protein-coding ability
repeated genes
multiple copies of small genes clustered throughout the genome at specific chromosomal sites; present in a high copy number and in a tandem (head to tail) configuration
Central Dogma of Molecular Biology
DNA-transcription-RNA-translation-protein
1 gene (genotype) to 1 mRNA (intermediate) to 1 "protein" (phenotype)
but this isnt really accurate, more than one mRNA and protein being made
Transcription
DNA to mRNA
Where: in eukaryotes, cell nucleus
When: either in G1 or G2 (period when genes coding for cellular organelles are synthesized)
A. Not happening in mitosis because the genetic material it tight here (heterochromatin)
B. Not in S phase (replication) because different machinery
How: see basic rules below
basic rules of transcription
-results in an RNA compliment
A. Antiparallel
B. Unidirectional
-only one of the two DNA strands serves as a template strand
A. The strand being transcribed is the template strand because both strands cant be transcribed at the same time
-selective process
A. Replication does the entire DNA strand, transcription is controlled by transcription factors (which ones turn a gene on or off), where on= you are transcribing the gene
B. Histones: house keeping genes, actin- proteins we need all the time; always on
C. Heat shock proteins, kinases are tuned on/off based on cellular environment
-always in the 5' to 3' direction
A. Complimentary RNA will be read 3-5' (opposite direction)
steps of transcription
0. Pre-recognition
1. Recognition
a. Pre-initiation complex formation
2. Initiation
a. Binding of the RNA complex
3. Elongation
a. Movement of RNA pol 2 and formation of mRNA
4. Termination
a. Cleavage of new transcript
step 0. Pre-Recognition (DNA access)
1. DNA is packaged in the nucleus as Heterochromatin
2. DNA must convert to Euchromatin prior to Transcription so that the machinery can access the DNA
3. Part of the gene expression regulation
Transcription machinery accesses the DNA strand in the "active" euchromatin region
Transcription machinery is not able to access DNA strand in the "silent" heterochromatin region
Nucleosome (10 nm fiber) and acetylation
The core is very positive (basic) and DNA is negative, thus they attract
a. Decrease the positive charge of the histone core loosens the DNA strand; Its easier to loosen the core
b. Lys (K) acetylation
i. Histone acetyl transferase (HAT)
ii. Histone deacetylase (HDAC)
Histone acetyl transferase (HAT)
Transfers acetyl group to histone; this decreases the positive charge of the histone core, making it more negative (but is still positive)
Histone deacetylase (HDAC)
1) Takes away an acetyl group from a histone, more positive direction, DNA gets tighter
step 1. recognition: pre initiation complex formation
1. Recognition process: pre initiation complex formation
a. Formation of a large complex of proteins (PIC_ required for RNA pol 2 to bind
i. First step: TATA binding protein (TBP) binds promotor region (TATA box)
ii. General transcription factors (TF 2D) recruited by TBP
a) Form the pre-initiation complex
step 2. initiation
Initiation: binding of RNA pol complex
-other transcription factors and RNA pol 2 recruited to complex
-mediator complex: about 20 proteins (ATPase and helicase unwind DNA, about one turn of DNA unwinds and forms transcription bubble)
-template strand: binds to RNA pol 2 active site
-RNA pol 2 is unphsphorylated at its carboxyl end (CTD): is the on/off switch (phosphorylation turns it on)
step 3. elongation
Elongation: movement of the RNA pol 2 and formation of the mRNA
-RNA pol 2 is phosporylated at its carboxyl end (CTD)
-RNA pol2 traverses the template strand (3' to 5') and creates an RNA copy, but transcription occurs 5'- 3'
-exact copy of the coding strand (except the T and U, and there is ribose 5c sugar for RNA)
coding and template strand
coding 5-3'
template 3-5'
step 4. termination
termination: cleavage of new transcript
-two protein complexes carried by CTD recognize pol-A signal (AAUAAA)
1. CPSF (cleavage and polyadenylation specialty factor)
2. CSTF (cleavage stimulation factor)
-other proteins are recruited to carry out cleavage
-poly-A polymerase adds the poly-A tail, IS NOT TEMPLATE DEPENDENT
-final product is an mRNA
why do we need post-transcriptional regulations?
Transcription occurs in the nucleus. To get the mRNA to be translated, we must move it to the cytoplasm, which is a harmful environment on single stranded nucleotides. We have to protect the ends.