FNR 305 Exam 2
1/95
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
96 Terms
gene pool
all alleles in a population
genetic diversity
the number and kind of genes (alleles) in a population; usually refers to a single species
genetic markers include:
microsatellites, mitochondrial DNA, allozymes, SNPs, etc.
genetic markers
most are codominant; serve as proxies for studying individual genes; allow us to estimate genetic diversity across entire genome
DNA typing
to genotype individuals by means of DNA extracted from tissue samples
population genetic structure
alleles at polymorphic loci may differ in frequency among subpopulations
genetic variation
allelic variation and heterozygosity are often assessed at microsattelite loci (genetic markers used for DNA fingerprinting)
genotype frequency
refers to the proportion of individuals in a population with a specific genotype
allele frequencies
count the # of times an allele occurs and divide by the number of chromosomes (2n in diploids); frequencies will sum to 1; basis for determining HE
average heterozygosity
the percentage of individuals that are heterozygous at a particular locus
Heterozygosity (H)
observed (H0), expected (HE)
allelic diversity (A)
mean number of alleles per locus
microsatellite genotyping
1. extract DNA sample, 2. characterize microsatellites if none described, 3. amplify each locus using PCR, 4. electrophorese to sperate alleles based on size
mtDNA markers
mitochondrial DNA: haploid, maternally inherited DNA marker; endosymbiont; evolves rapidly
Heterozygote superiority
when the fitness (measurment of viability and fertility) of heterozygotes is greater than homozygotes
overdominance
in sickle cell anemia, overdominance of the heterozygote carriers is observed because they are more resistant to malaria
Hardy-Weinberg Principle (HW)
analyzes the factors which may affect the frequencies of alleles in a population; uses a simple equation to calculate allelic frequencies: p+q=1
HWE assumptions/conditions:
1. Mating is random, 2. allelic frequencies are the same in males and females, 3. all genotypes have equivalent viability and fertility (i.e. no selections), 4. Mutation does not occur, 5. Migration into the population is absent, 6. Population is large so that allelic variations do not occur by chance (i.e. no genetic drift)
HWE calculating expected genotype frequencies:
p^2 +2pq+q^2=1
pp=homozygous dominant
qq=homozygous recessive
2pq=heterozygous
HWE calculating multiple allele frequencies:
p^2+2pq+2pr+q^2+2qr+r^2=1
Deviations from HWE
mutation, migration, selection, drift (or sampling error)
genetic drift
greater effects seen on smaller populations; sampling error that occurs at low population sizes; rare alleles may never get to next generation
microevolution
allele frequency change over time
macroevolution
refers to changes in the gene pool which produce phenotypic changes subject to the forces of natural selection
Causes of evolution:
mutation, migration, natural selection, random genetic drift
Under HWE, evolution...
would not occur and allele frequencies would remain unchanged over time
Principles of Evolution
1) more young are born that can survive, 2) heritable variation occurs within populations, 3) genetic variation may be beneficial, detrimental, or neutral, 4) organisms with genotypes most compatible with their environment are overrepresented in the next generation 5) populations change in genetic composition over time 6) over time, genetic differentiation of populations can give rise to new species
What can lead to speciation?
A reduction in gene flow between populations, accompanied by divergent selection or genetic drift.
Lack of gene flow and divergence
evolution of genetic reproductive barriers between discrete populations; may be caused by topography, habitat or climate then populations evolve separately
What is the evolutionary fate of gene pools in independent bison herds?
divergence
Divergence
can arise because of geographic barriers; can restrict gene flow; ultimately can lead to reproductive incompatibility
Rates of speciation are variable
1) instantaneous (polyploidy) 2)rapid (Drosophilia, cichlids) 3) 3 million years (estimated average time to full reproductive isolation) 4) > 20 million years for full reproductive isolation
Factors that promote rapid speciation
1) low dispersal rates 2) abundant opportunities for isolation of populations 3) strong sexual selection 4) opportunity for ecological specialization 5) bottlenecks in population size
Genetic diversity increases with
increasing population size
genetic drift
causes random changes in allele frequency in small populations; one allele may become fixed; loss of allelic diversity and lower H
Loss of genetic diversity
is related to the average increase in inbreeding, leading to reduction in fitness called inbreeding depression
Bottlenecks
longer the bottleneck, more diversity lost; rate of loss depends on population size
Mechanisms that decrease genetic diversity
Extinction of populations, fixation of favorable alleles by selection, genetic drift, inbreeding
Inbreeding
the mating of individuals related by ancestry; increases homozygosity, reduces heterozygosity; exposes rare deleterious recessive alleles; measured as the probability that 2 alleles at a locus are identical-by-descent (IBD)
inbred populations
will have a lowered mean fitness called inbreeding depression (i.e. Isle Royale wolves)
genetic load
the cumulative burden of deleterious recessive alleles found in a genome
coefficient of inbreeding, F
the probability that 2 alleles of a given gene in an individual are identical because they are descended from the same single copy of the allele in an ancestor
F
the probability that an individual carries alleles that are identical by descent (IBD); F=0 is completely outbred, F=1 is completely inbred
Inbreeding vs. Relatedness
inbreeding (F) is calculated for 1 individual; relatedness (r) between pairs and refers to the portion of genome shared btw. individuals
Inbreeding is cumulative
large populations accumulate inbreeding very slowly; smaller the population, more rapidly inbreeding accumulates
Inbreeding Depression (ID)
a reduction in fitness associated with inbreeding; Increases probability of extinction via the extinction vortex
extinction vortex
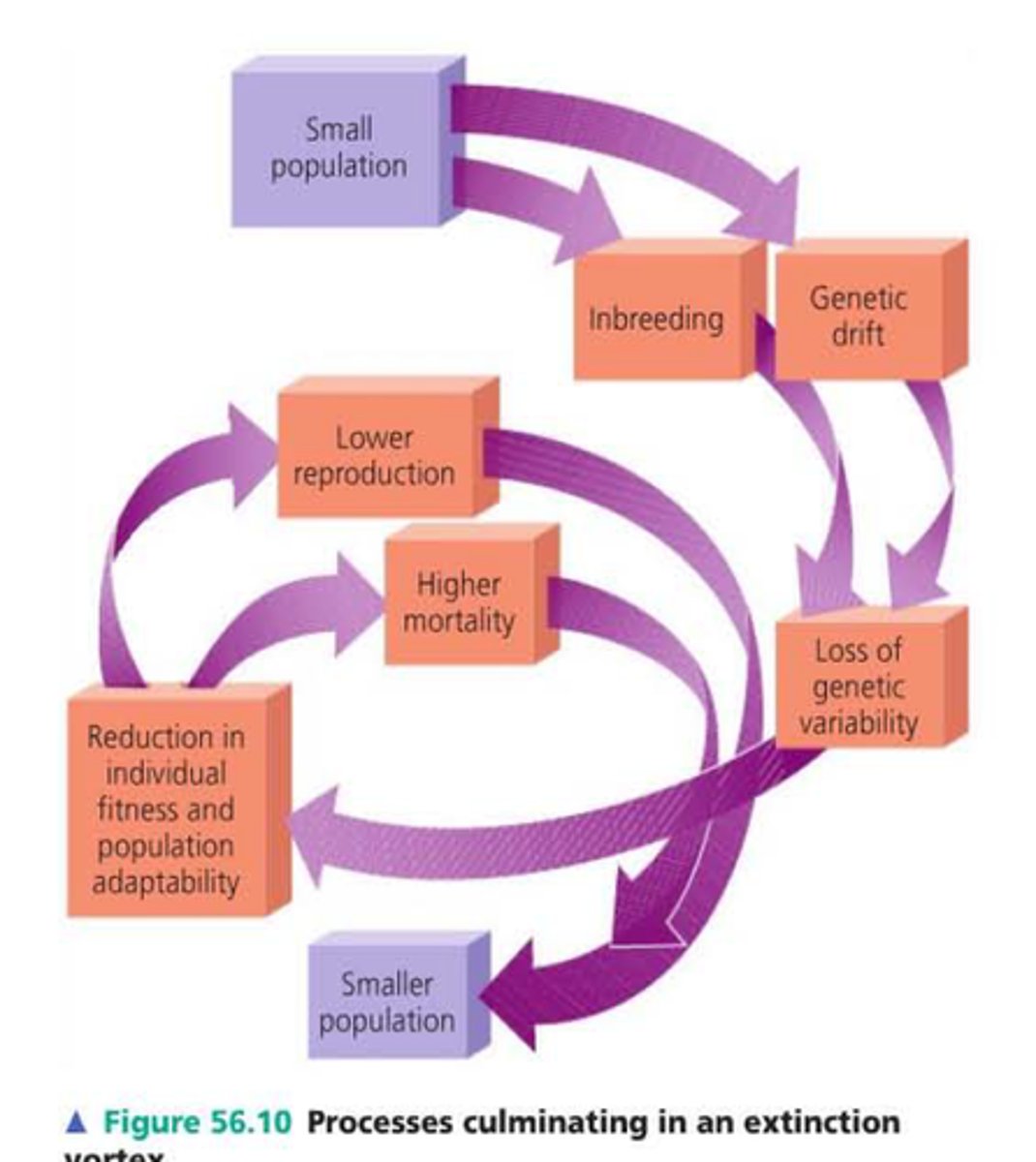
Measuring ID
= 1- (fitness inbred/fitness outbred)
10% increase in inbreeding leads to
10% reduction in fitness
Effective population size (Ne)
the size of an idealized population that would lose genetic diversity at the same rate as the actual population
Factors that reduce Ne/N
1)fluctuation in population size 2) variation in family size (few indv. monopolize mating, this skews the ratio) 3) variation in operation sex ratio
Other factors influencing Ne/N
overlapping generations and nonrandom mating
average ratio of Ne/N
0.10
Effective population size predict the effects on the population of:
random genetic drift, natural selection, mutation-selection equilibrium, inbreeding, genetic diversity
habitat fragmentation
leads to decreased Ne and reduced migration, may lead to genetic differentiation
small Ne
more drift, fewer immigrants
isolation-by-distance
the farther apart two populations are, the more likely they will differ; factor in allopatric speciation
F IT
is inbreeding of individuals in the total population
F IS
the probability that two alleles in the same individual are identical by descent
Fst
the probability that two alleles drawn at random from a subpopulation are identical by descent; widely used measure of divergence
If gene flow is high, Fst is generally:
low
Wind-pollinated trees have greater dispersal capacity than arboreal rodents such as squirrels. Given two woodlots separated by a very large field of row crops, would you expect Fst to be higher in the tree or the squirrel?
squirrel
m and Nm
m=migration rate Nm= number of migrants per generation
Consequences of fragmentation
1) fragmented populations genetically differ by chance 2)Cumulative diversification through inbreeding and drift within population fragments
How is Fst affected by dispersal rate?
Fst decreases as dispersal rate increases
How is Fst affected by habitat fragmentation?
Fst increases as habitat becomes more fragmented; Fst increases the more distant the habitat subdivisions
How is Fst affected by divergence time?
Fst increases with a longer divergence time
How is Fst affected by population size?
Fst increases when populations are smaller
How is FST generally affected by adaptive differences?
Fst increases with adaptive differences between populations
Self-fertilization
a form of inbreeding common in plants; increases homozygosity rapidly
Florida panther
isolated population that experienced a long-term bottleneck; 60-70 animals; Inbreeding depression: high incidence of abnormalities (heart defects, poor semen quality)
Minimal viable population size
minimum size required to retain reproductive fitness and evolutionary potential over thousands of years
According to Bailey, the more we intervene to manage wild populations, the more ______________________ will occur.
artificial selection
Bailey relates how a Russian geneticist selectively bred foxes for coat color. In so doing, he inadvertently selected for animals with mottled fur, floppy ears, and shorter tails. This is an example of:
linked traits
The International Union for Conservation of Nature lists what as a major obstacle for bison restoration?
regulatory status (most states recognize bison as livestock)
Population Viability Analysis (PVA)
a species specific risk assessment based on ecology and statistics; used as management tool to compare different options to recover a species
Deterministic factors
habitat loss, over-exploitation, pollution, introduced species
stochastic factors
demographics (sex ratio, birth rates), environmental variation (temperature, rainfall, competitors, predators), genetic (ID, loss of GD, divergences, etc.), catastrophes (hurricanes, floods, etc.)
Limitations of PVAs
-PVAs do not encompass the full genetic impacts on population viability
-Insufficient life-history data for most species
-Poisson variation in family sizes generally assumed (biased Ne estimates)
-May assume environmental conditions remain unchanged
-The process of conducting a PVA may be more important to conservation than the PVA output!
-mostly, species biology!
Goals for managing wild populations
-managing population size
-alleviate the effects of fragmentation
-alleviate genetic swamping due to hybridization between species
-minimize the impacts of harvesting
Hardy-Weinberg Equilibrium
a model that can be used to predict stable genotype frequencies from allele frequencies if certain assumptions are true: -no mutation, -no migration, -no selection, -no drift, -random mating
Which evolutionary force is most likely to cause genome-wide convergence in two different geographic bison populations?
migration
Biological Species Concept
group of populations that exchange (or have the potential to exchange) genes by interbreeding AND do not interbreed with other populations because of biological differences
Evolutionary "Options"
1) stasis
2) undergo gradual phyletic evolution (anagensis) to become a new species
3) undergo cladogensis to give rise to two distinct and independent daughter species
How does speciation occur?
mostly due to allopatry, but also sympatry, and some due to instant speciation (polyploid)
According to the biological species concept, the major criterion for determining whether two population are part of the same species is:
reproductive compatibility
Neutral Theory of molecular evolution
mutations leading to amino acid substitutions are rare, because they are usually detrimental; mutations that are neutral will be effected by other mutations and by genetic drift; most variation is the result of neutral mutations
Three primary questions when diagnosing genetic problems:
-Has a threatened species lost its genetic diversity?
-Is it suffering from inbreeding depression?
-Is it genetically fragmented?
Diagnosing Genetic Problems
often involves collecting tissue samples, then DNA is extracted and used to estimate allelic diversity (A), heterozygostiy (H), and the similarity of gene pools.
Recover small inbred populations
introduce individuals from other populations (outbreeding not always possible)
Genetic management of fragmented populations
-increase population size
-establish populations in several locations
-maximize reproductive rate
-insulate from environmental change
-translocation
-alternatives (aritificial insemination)
-re-establish gene flow (corridors)
Impacts of harvesting
directional selection occurs due to harvesting based on quality of individuals
Effective Population Size Equation
Ne = t/ (∑1/Nei)
Mammoths on Wrangel Island (at the tail end of mammoth existence)
Due to evolution occurring at low effective population sizes, detrimental mutations occurred such as deletions and premature stop codons. Also evolved to have a satin coat?
FOXQ1 locus
pleiotropic: satin coat and mucin secretion in GI tract, leading to gastic irritation
Nearly neutral theory
under small effective population sizes, detrimental mutations can accumulate in genomes.