ACS BIOCHEMISTRY EXAM
0.0(0)
Studied by 0 peopleCard Sorting
1/251
Earn XP
Description and Tags
Last updated 1:13 AM on 12/2/22
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
252 Terms
1
New cards
Henderson-Hasselbach Equation
pH = pKa + log ([A-] / [HA])
2
New cards
FMOC Chemical Synthesis
Used in synthesis of a growing amino acid chain to a polystyrene bead. FMOC is used as a protecting group on the N-terminus.
3
New cards
Salting Out (Purification)
Changes soluble protein to solid precipitate. Protein precipitates when the charges on the protein match the charges in the solution.
4
New cards
Size-Exclusion Chromatography
Separates sample based on size with smaller molecules eluting later.
5
New cards
Ion-Exchange Chromatography
Separates sample based on charge. CM attracts +, DEAE attracts -. May have repulsion effect on like charges. Salt or acid used to remove stuck proteins.
6
New cards
Hydrophobic/Reverse Phase Chromatography
Beads are coated with a carbon chain. Hydrophobic proteins stick better. Elute with non-H-bonding solvent (acetonitrile).
7
New cards
Affinity Chromatography
Attach a ligand that binds a protein to a bead. Elute with harsh chemicals or similar ligand.
8
New cards
SDS-PAGE
Uses SDS. Gel is made from cross-linked polyacrylamide. Separates based off of mass with smaller molecules moving faster. Visualized with Coomassie blue.
9
New cards
SDS
Sodium dodecyl sulfate. Unfolds proteins and gives them uniform negative charge.
10
New cards
Isoelectric Focusing
Variation of gel electrophoresis where protein charge matters. Involves electrodes and pH gradient. Protein stops at their pI when neutral.
11
New cards
FDNB (1-fluoro-2,3-dinitrobenzene)
FDNB reacts with the N-terminus of the protein to produce a 2,4-dinitrophenol derivative that labels the first residue. Can repeat hydrolysis to determine sequential amino acids.
12
New cards
DTT (dithiothreitol)
Reduces disulfide bonds.
13
New cards
Iodoacetate
Adds carboxymethyl group on free -SH groups. Blocks disulfide bonding.
14
New cards
Homologs
Shares 25% identity with another gene
15
New cards
Orthologs
Similar genes in different organisms
16
New cards
Paralogs
Similar "paired" genes in the same organism
17
New cards
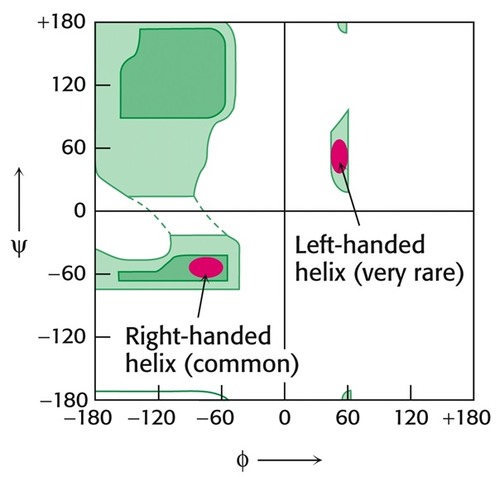
Ramachandran Plot
Shows favorable phi-psi angle combinations. 3 main "wells" for α-helices, ß-sheets, and left-handed α-helices.
18
New cards
Glycine Ramachandran Plot
Glycine can adopt more angles. (H's for R-group).
19
New cards
Proline Ramachandran Plot
Proline adopts fewer angles. Amino group is incorporated into a ring.
20
New cards
α-helices
Ala is common, Gly & Pro are not very common. Side-chain interactions every 3 or 4 residues. Turns once every 3.6 residues. Distance between backbones is 5.4Å.
21
New cards
Helix Dipole
Formed from added dipole moments of all hydrogen bonds in an α-helix. N-terminus is δ+ and C-terminus is δ-.
22
New cards
ß-sheet
Either parallel or anti-parallel. Often twisted to increase strength.
23
New cards
Anti-parallel ß-sheet
Alternating sheet directions (C & N-termini don't line-up). Has straight H-bonds.
24
New cards
Parallel ß-sheet
Same sheet directions (C & N-termini line up). Has angled H-bonds.
25
New cards
ß-turns
Tight u-turns with specific phi-psi angles. Must have gly at position 3. Proline may also be at ß-turn because it can have a cis-omega angle.
26
New cards
Loops
Not highly structured. Not necessary highly flexible, but can occasionally move. Very variable in sequence.
27
New cards
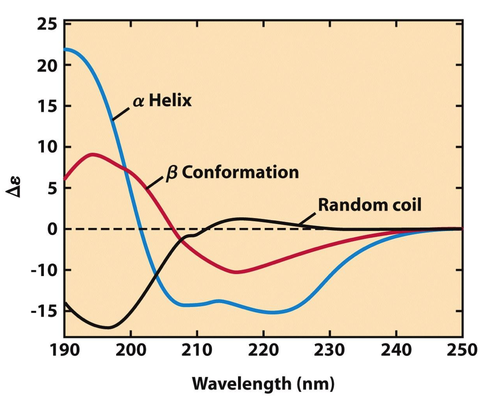
Circular Dichroism
Uses UV light to measure 2° structure. Can be used to measure destabilization.
28
New cards
Disulfide-bonds
Bonds between two -SH groups that form between 2° and 3° structure.
29
New cards
ß-mercaptoethanol
Breaks disulfide bonds.
30
New cards
α-keratin
formed from 2 α-helices twisted around each other. "Coiled coil". Cross-linked by disulfide bonds.
31
New cards
Collagen
Repeating sequence of Gly-X-Pro. 3 stranded "coiled coil". Contains gly core.
32
New cards
Myoglobin 4° Structure
Symmetric homodimer,
33
New cards
Hemoglobin 4° Structure
Tetramer. Dimer of dimers. α2ß2 tetramer.
34
New cards
α/ß Protein Folding
Less distinct areas of α and ß folding.
35
New cards
α+ß Protein Folding
Two distinct areas of α and ß folding.
36
New cards
Mechanism of Denaturants
Highly soluble, H-binding molecules. Stabilize protein backbone in water. Allows denatured state to be stabilized.
37
New cards
Temperature Denaturation of Protein
Midpoint of reaction is Tm.
38
New cards
Cooperative Protein Folding
Folding transition is sharp. More reversible.
39
New cards
Folding Funnel
Shows 3D version of 2D energy states. Lowest energy is stable protein. Rough funnel is less cooperative.
40
New cards
Protein-Protein Interfaces
"Core" and "fringe" of the interfaces. Core is more hydrophobic and is on the inside when interfaced. Fringe is more hydrophilic.
41
New cards
π-π Ring Stacking
Weird interaction where aromatic rings stack on each other in positive interaction.
42
New cards
σ-hole
Methyl group has area of diminished electron density in center; attracts electronegative groups
43
New cards
Fe Binding of O2
Fe2+ binds to O2 reversible. Fe3+ has an additional + charge and binds to O2 irreversibly. Fe3+ rusts in O2 rich environments.
44
New cards
Ka for Binding
Ka = [PL] / [P][L]
45
New cards
ϴ-value in Binding
ϴ = (bound / total)x100%
ϴ = [L] / ([L] + 1/Ka)
ϴ = [L] / ([L] + 1/Ka)
46
New cards
Kd for binding
Kd = [L] when 50% bound to protein.
Kd = 1/Ka
Kd = 1/Ka
47
New cards
High-Spin Fe
Electrons are "spread out" and result in larger atom.
48
New cards
Low-Spin Fe
Electrons are less "spread out" and are compacted by electron rich porphyrin ring.
49
New cards
T-State
Heme is in high-spin state. H2O is bound to heme.
50
New cards
R-State
Heme is in low-spin state. O2 is bound to heme.
51
New cards
O2 Binding Event
O2 binds to T-state and changes the heme to R-state. Causes a 0.4Å movement of the iron.
52
New cards
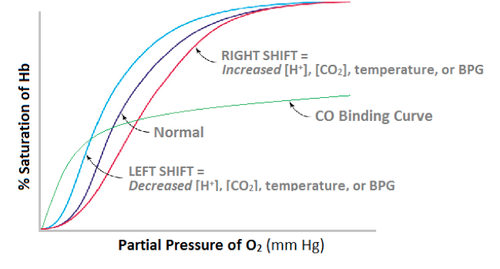
Hemoglobin Binding Curve
4 subunits present in hemoglobin that can be either T or R -state. Cooperative binding leads to a sigmoidal curve.
53
New cards
Binding Cooperativity
When one subunit of hemoglobin changes from T to R-state the other sites are more likely to change to R-state as well. Leads to sigmoidal graph.
54
New cards
Homotropic Regulation of Binding
Where a regulatory molecule is also the enzyme's substrate.
55
New cards
Heterotropic Regulation of Binding
Where an allosteric regulator is present that is not the enzyme's substrate.
56
New cards
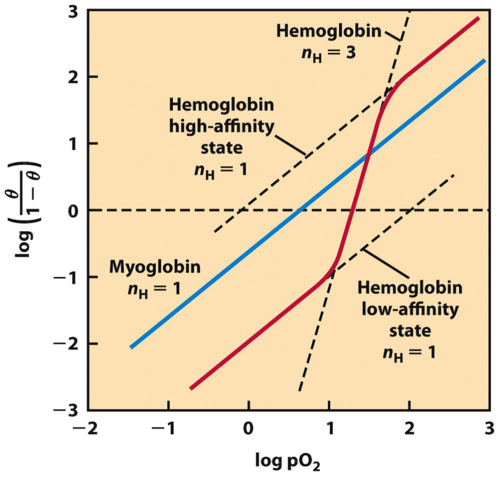
Hill Plot
Turns sigmoid into straight lines. Slope = n (# of binding sites). Allows measurement of binding sites that are cooperative.
57
New cards
pH and Binding Affinity (Bohr Affect)
As [H+] increases, Histidine group in hemoglobin becomes more protonated and protein shifts to T-state. O2 binding affinity decreases.
58
New cards
CO2 binding in Hemoglobin
Forms carbonic acid that shifts hemoglobin to T-state. O2 binding affinity decreases. Used in the peripheral tissues.
59
New cards
BPG (2,3-bisphosphoglycerate)
Greatly reduces hemoglobin's affinity for O2 by binding allosterically. Stabilizes T-state. Transfer of O2 can improve because increased delivery in tissues can outweigh decreased binding in the lungs.
60
New cards
Michaelis-Menton Equation
V0 = (Vmax[S]) / (Km + [S])
![V0 = (Vmax[S]) / (Km + [S])](https://knowt-user-attachments.s3.amazonaws.com/d1a21aa097074e93971e10044521eb15.png)
61
New cards
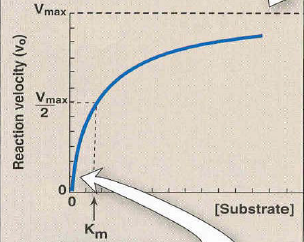
Km in Michaelis-Menton
Km = [S] when V0 = 0.5(Vmax)
62
New cards
Michaelis-Menton Graph
63
New cards
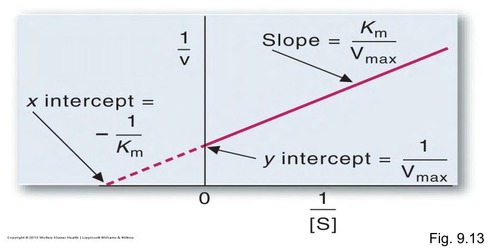
Lineweaver-Burke Graph
Slope = Km/Vmax
Y-intercept = 1/Vmax
X-intercept = - 1/Km
Y-intercept = 1/Vmax
X-intercept = - 1/Km
64
New cards
Lineweaver-Burke Equation
Found by taking the reciprocal of the Michaelis-Menton Equation.
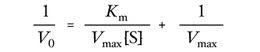
65
New cards
Kcat
Rate-limiting step in any enzyme-catalyzed reaction at saturation. Known as the "turn-over number". Kcat = Vmax/Et
66
New cards
Chymotripsin
Cleaves proteins on C-terminal endof Phe, Trp, and Tyr
67
New cards
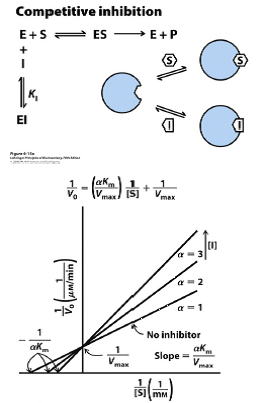
Competitive Inhibition Graph
Slope changes by factor of α. Slope becomes αKm/Vmax.
X-intercept becomes 1/αKm
Y-intercept does not change.
Vmax does not change.
X-intercept becomes 1/αKm
Y-intercept does not change.
Vmax does not change.
68
New cards
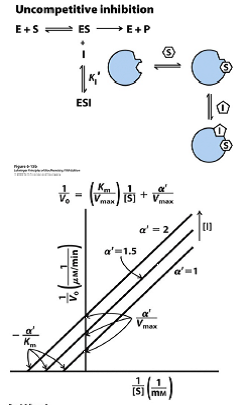
Uncompetitive Inhibition Graph
Does not change slope.
Changes Km and Vmax.
Results in vertical shift up and down.
Y-intercept becomes α'/Vmax
X-intercept becomes -α'/Km
Changes Km and Vmax.
Results in vertical shift up and down.
Y-intercept becomes α'/Vmax
X-intercept becomes -α'/Km
69
New cards
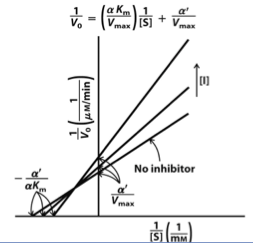
Mixed Inhibition Graph
Allosteric inhibitor that binds either E or ES.
Pivot point is between X-intercept and Y-intercept.
Pivot point is between X-intercept and Y-intercept.
70
New cards
Non-Competitive Inhibition Graph
Form of mixed inhibition where the pivot point is on the x-axis. Only happens when K1 is equal to K1'.
71
New cards
Ionophore
Hydrophobic molecule that binds to ions and carries them through cell membranes. Disrupts concentration gradients.
72
New cards
ΔGtransport Equation
ΔGtransport = RTln([S]out / [S]in) + ZFΔΨ
73
New cards
Pyranose vs. Furanose
Pyranose is a 6-membered ring.
Furanose is a 5-membered ring.
Furanose is a 5-membered ring.
74
New cards
Mutarotation
Conversion from α to ß forms of the sugar at the anomeric carbon.
75
New cards
Anomeric Carbon
Carbon that is cyclized. Always the same as the aldo or keto carbon in the linear form.
76
New cards
α vs. ß sugars
α form has -OR/OH group opposite from the -CH2OH group.
ß form has -OR/OH group on the same side as the -CH2OH group.
ß form has -OR/OH group on the same side as the -CH2OH group.
77
New cards
Starch
Found in plants. D-glucose polysaccharide. "Amylose chain". Unbranched. Has reducing and non-reducing end.
78
New cards
Amylose Chain
Has α-1,4-linkages that produce a coiled helix similar to an α-helix. Has a reducing and non-reducing end.
79
New cards
Amylopectin
Has α-1,4-linkages. Has periodic α-1,6-linkages that cause branching. Branched every 24-30 residues. Has reducing and non-reducing end.
80
New cards
Reducing Sugar
Free aldehydes can reduce FeIII or CuIII. Aldehyde end is the "reducing" end.
81
New cards
Glycogen
Found in animals. Branched every 8-12 residues and compact. Used as storage of saccharides in animals.
82
New cards
Cellulose
Comes from plants. Poly D-glucose. Formed from ß-1,4-linkage. Form sheets due to equatorial -OH groups that H-bond with other chains.
83
New cards
Chitin
Homopolymer of N-acetyl-ß-D-glucosamine. Have ß-1,4-linkages. Found in lobsters, squid beaks, beetle shells, etc.
84
New cards
Glycoproteins
Carbohydrates attached to a protein. Common outside of the cell. Attached at Ser, Thr, or Asn residues.
85
New cards
Membrane Translayer Flip-Flop
Typically slow, but can be sped up with Flippase, Floppase, or Scramblase.
86
New cards
Membrance Fluidity
Membrane must be fluid. Cis fats increase fluidity, trans fats decrease fluidity.
87
New cards
Type I Integral Membrane Protein
Membrane protein with C-terminus inside and N-terminus outside
88
New cards
Type II Integral Membrane Protein
Membrane protein with N-terminus inside and C-terminus outside
89
New cards
Type III Integral Membrane Protein
Membrane protein that contains connected protein helices
90
New cards
Type IV Integral Membrane Protein
Membrane protein that contains unconnected protein helices
91
New cards
Bacteriorhodopsin
Type III integral membrane protein with 7 connected helices.
92
New cards
ß-Barrel Membrane Protein
Can act as a large door. Whole proteins can fit inside.
93
New cards
α-hemolysin
Secreted as a monomer. Assembles to punch holes in membranes.
94
New cards
Cardiolipin
"Lipid staple" that ties two proteins (or complexes) together in a membrane. Formed from two phosphoglycerols.
95
New cards
Hydrolysis of Nucleotides
Base hydrolyzes RNA, but not DNA. DNA is stable in base because of 2' deoxy position.
96
New cards
Chargaff's Rule
Ratio of A:T and G:C are always equal or close to 1
97
New cards
DNA Double-Helix
Opposite strand direction. 3.4Å distance between complementary bases. 36Å for one complete turn.
98
New cards
A-form DNA
Condensed form of DNA. Deeper major groove and shallower minor groove.
99
New cards
B-form DNA
Watson-Crick model DNA. Deep, wide major groove.
100
New cards
Z-form DNA
Left-handed helical form of DNA