Transcription and Translation - Bio 102
1/65
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
66 Terms
tRNA
transfer – transfers the amino acid that connects to mRNA
mRNA
messenger – template to make amino acid code
rRNA
ribosomal - makes up ribosomes
transcription
synthesis of mRNA from a single gene
template strand
the strand that mRNA is made off of
coding strand
the strand that is the same as the RNA strand (with exception to the thymine/uracil)
RNA Polymerase
major enzyme in transcription
polymerase activity
puts the building blocks of mRNA together
helicase activity
unwinds the double helix
unzips the two DNA strands
3 types of RNA polymerase in eukaryotes
I, II, III
RNA polymerase I
transcribes most rRNA genes
RNA polymerase II
the one we care about most
transcribes all protein-coding genes
miRNA genes
genes for other noncoding RNAs
RNA polymerase III
transcribes most tRNAs
5S rRNA gene
other small RNAs
promoter region
helps direct transcription
two components of promoter region
TATA box
transcription factors
TATA box
always upstream of the gene
define the direction of transcription and indicate where RNA polymerase should start reading the DNA
transcription factors
These lead to transcription
They bind to promoter and tell polymerase where to start
Help determine which is the coding strand and which is the template strand – tells them which direction the RNA polymerase should go
direction RNA polymerase reads DNA
3’ to 5’
direction RNA polymerase makes RNA
5’ to 3’
terminator sequence
blocks the RNA polymerase which causes it to fall off and release the DNA
how transcription is affected by drugs
helicase activity is halted
polymerase activity is halted
difference between prokaryotic and eukaryotic transcription
prokaryotes
In cytosol
1 type of RNA polymerase
Sigma factor (subunit of RNA polymerase) recognizes promoter region
Termination signal in DNA
Transcription and translation kinda happen at the same time, so splicing doesn’t happen
They just have coding regions, eukaryotes also have introns (noncoding regions)
Eukaryotes
In nucleus... TRANSLATION occurs in the cytosol
3 types of RNA polymerase
Requires transcription factors
Termination signal in mRNA (AAUAA)
3 processes to get RNA ready for translation in eukaryotes
capping
splicing
poly-A tail
splicing
removing introns, keeping exons
splicesome
takes two exon ends, puts them together, and takes out introns
capping
put on the 5’ end (first part made)
protects from degradation and determines how long the RNA will stay in your cells
poly-A tail
on the 3’ side
150-250 A’s
helps protect from degradation and determines how long the RNA will stay in your cells
alternative splicing
can produce different proteins by splicing out different introns
anything between the introns can be taken out, but the ends need to stay
exporting mRNA from the nucleus
mRNA is exported from the nucleus through pores and channels into the cytosol
cap-binding protein takes it out of the nucleus (this is a reason why we need capping)
poly-A binding protein hooks on to the tail
once the mRNA exits, initiation factors replace the other proteins
nuclear pore complexes
form channels in nuclear membrane and can regulate exit of mRNA
codon
3 mRNA letters used to create an amino acid
tRNA
contains anticodons that are complementary to the codons of the mRNA
start codon
AUG - methionine
keeps the mRNA in the correct reading frame
wobble base pairing
allows multiple codons to code for the same amino acid
allows for some mutations and mistakes
large subunit
holds the tRNA
small subunit
holds the mRNA
initiation
MRNA binds to the small subunit, the start codon is read (5’ to 3’), and this triggers the tRNA to come next
The tRNA is gonna come first, and then the large subunit comes on after
Every amino acid enters at the A site, EXCEPT the initiator tRNA which enters at the P (peptide) site
elongation
Another tRNA is added and this pushes the first tRNA to the P site where a peptide bond combines the amino acids
Termination
Stop codon is reached
Instead of a tRNA with an amino acid, a release factor comes in with no amino acid
Since there is nothing for the amino acid chain to bond with, it is released
Everything breaks apart and can get used again
A site
aminoacyl
where the tRNA enters
P site
peptide site
where a peptide bond combines the amino acids
E Site
exit
where the tRNA exits the ribosomes
chargaff’s rules
A goes with T
C goes with G
purines
A G
pyrimidines
C and T
purpose of genetic information
serves as “instructions” for building and maintaining cells
what genetic information is made of
DNA
how genetic information is stored
genes
base that is not common to DNA and RNA
thymine is replaced with uracil in RNA
5’ end
5th carbon on a sugar that connects to the phosphate group at the end of the strand
3’ end
3rd carbon on the sugar
number of bonds between G-C
3 hydrogen bonds
number of bonds between T-A
2 hydrogen bonds
differences between DNA and RNA
DNA
double stranded
deoxyribose (less oxygen)
contains T
RNA
single stranded
ribose (more oxygen)
contains U
genes
segments of DNA that code for RNA which is then translated into a protein
amino acid structure
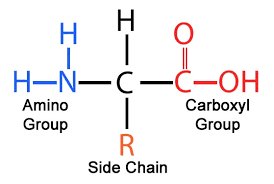
number of different side chains in amino acids
20
how peptide bonds are formed
The –OH of the carboxyl group of the first
amino acid and the H from the amino
group of the next one are removed as a
water molecule – a condensation
(dehydration) reaction
bond between the C and the N
N terminus
beginning of amino acid
C terminus
end of the amino acid
primary protein structure
linear sequence of a chain of amino acids
linked by peptide bonds
secondary protein structure
formed by hydrogen bonds between protein backbone
alpha-helix
beta-pleated sheet
tertiary structure
overall 3-D structure
involved interactions between side chains
interactions that contribute to tertiary structure
hydrophobic interactions
hydrogen bonds
disulfide bridge
ionic bond
quaternary structure
multiple subunits assembled into a larger complex
chaperone proteins
help ensure that proteins fold properly
form isolation chambers to provide the idea environment for protein folding