L18 Oxidative Phosphorylation
1/73
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
74 Terms
what is glucose catabolized into
pyruvate via glycolysis and then acetyl-CoA
what is fatty acid catabolized into
acetyl-CoA via beta-oxidation
what are amino acids catabolized into
acetyl-CoA via deamination an doxidation
what is acetyl-CoA put into
citric acid cycle which moves electrons into the electron transport chain/oxidative phosphorylation
how are electrons transported in the ETC
specialized electron carried NADH and FADH2 which transfer electrons to components of the ETC in the inner mitochondrial membrane
what ultimately drives ATP
free energy derived from oxidation of NADH and FADH2
what happens in the ETC
electrons are passed to the ultimate electron acceptor, O2, and protons (H+) are pumped from the mitochondrial matric to the inner mitochondrial space
what does ATP synthase use to provide energy for making ATP
the proton electrochemical gradient
what is cellular respiration
the process of reoxidizing NADH and ubiquinol by oxygen - break down glucose and other organic molecules with oxygen to produce energy in the form of ATP, releasing carbon dioxide and water as byproducts
what is the outer membrane of the mitochondria
freely permeable to small molecules and ions
what is the inner membrane of the mitochondria
impermeable to most molecules and ions - contains complex I-IV and transporters and electrons transport across to create a proton gradient which is used to fuel ATP synthase
what is the inter mitochondrial space of the mitochondria
H+ pumped here via ETC actions - when NADH gives up an electron in complex I
what is the matrix in the mitochondria
cristae convolutions provide extensive surface area - enzymes, cofactors and intermediates are found here
what generates most of the cell’s NADH and QH2
the citric acid cycle inside the mitochondria as well as fatty acid oxidation inside the mitochondria
what do the reduced cofactors NADH and QH2 do with their electrons
transfer them to the protein complexes of the respiratory ETC which are tightly associated with the inner mitochondrial membrane
why do NADH produced by glycolysis have to use reducing equivalents
these and other oxidative processes in the cytosol cannot directly reach the respiratory chain
why do we need electron carriers
electrons cannot diffuse freely throughout the cell so they must be carried on molecular electron carriers which function as coenzymes that cycle between their oxidized and reduced states to support oxidation-reduction reactions in the cell
what is the malate-aspartate shuttle system an example of
reducing equivalents because NADH cannot enter the mitochondria via diffusion, so the enzyme cytosolic malate dehydrogenase uses electrons from NADH to reduce OAA into malate - transfers reducing potential from NADH to malate which can cross the membrane via a transporter - malate converted back to OAA which reduced mitochondrial NAD+ back into NADH
what does ATP translocase protein do
imports ADP and exports ATP and a symport protein pump then permits simultaneous movement of Pi and H+ - this helps us to regulate when ATP needs to be synthesized and the appropriate movement of Pi and H+
what is oxidative phosphorylation
the process of the electron transport chain and ATP synthase and is the final step of cellular respiration
what are thermodynamics of electron flow in the ETC
electrons move from the more negative reduction potential to the more positive reduction potential
what is Coenzyme Q/ubiquinol
a mobile liquid molecule that carries 2 electrons from comple I/II to III that is anchored to the mitochondrial matrix by 10 isoprenyl units
what is cytochrome C
a water-soluble protein that has iron-containing, heme prosthetic groups and moves on the outer surface of the membrane- the heme group carries only 1 electron from complex III to complex IV and there are three classes (A,B, and C)
what is the function of complex I
(e- → NADH → Q) NADH-ubiquinone oxireductase accepts 2 electrons from NADH, produces ubiquinone, transports 4 H+ from matrix to intermembrane space - releases NAD+, QH2, and the 4 H+ (transports H+)
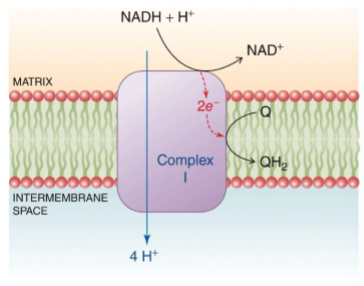
what does electron movement require
flavin mononucleotide and iron-sulfur centers - flavin picks up electron in complex I and noncovalently binds it as a prosthetic group similar to FAD and transfers the electrons one at a time to a second type of redox center (iron) - electrons ultimately are trasnferred to CoQ which is transferred to uniquinol (+2e-)
can the movement of electrons through complex I generate ROS
yes because the process can be reversible - electrons can prematurely escape the normal ETC and react directly with molecule O2 and flavin (FMN) site, and Q binding site - ROS produces as a signaling byproduct of cell stress can be leaked into matric and be converted to hydrogen peroxide
what is complex II
succinate dehydrogenase and is the electron acceptor from FADH2
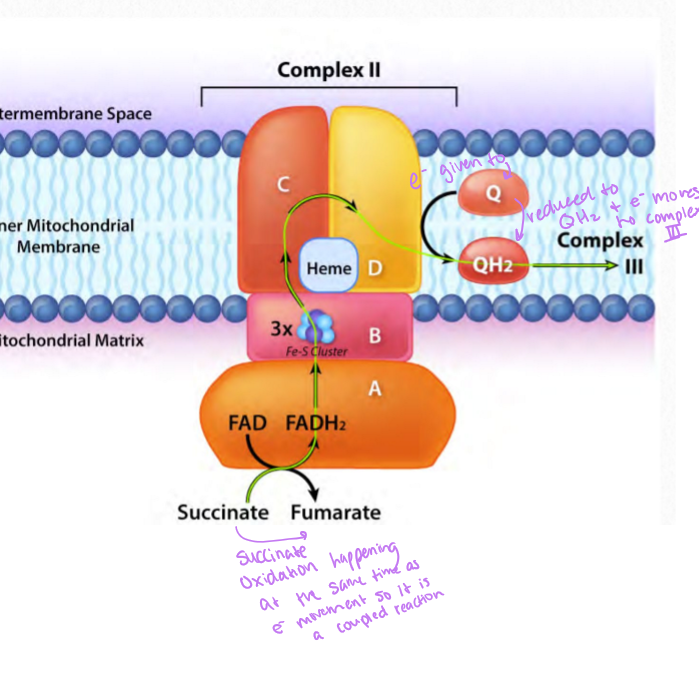
what does complex II do
succinate-ubiquinone reductase accepts electrons from FADH2, produces ubiquinone also know as succinate dehydrogenase from the TCA cycle
what is complex II coupled with
electron movement and succinate oxidation to fumarate and because there is no proton movement, there is no ROS generation
what does complex III do
2 electrons from QH2 reduce 2 molecules of cytochrome c - 4 protons are translocated to the intermembrane space - reoxidizes ubiquinol via cytochrome c oxireductase and transfers electrons from ubiquinol to the peripheral membrane protein cytochrome c (transports H+)
what are cytochromes
proteins with heme prosthetic groups
why is complicated about the flow of electrons through complex III
the two electrons donated by ubiquinol must split in order to travel through a series of one-electron carriers - 2 FE-2S cluster of iron-sulfur protein, cytochrome c1, and cytochrome b - the Fe-S protein must change its conformation by rotating and moving over in order to pick up and deliver an electron
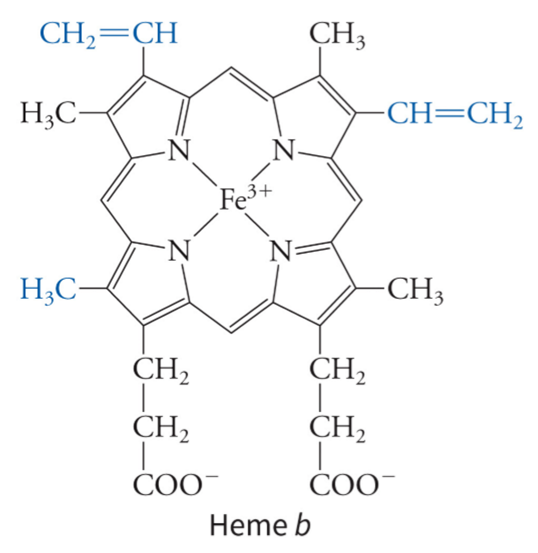
what is the heme group of a b cytochrome
undergoes reversible one-electron reduction, with the central Fe atom cycling between the Fe3+ (oxidized) and Fe2+ (reduced) states - this is why the complex 3 has a complicated path for electron movement - they can accept electrons due to the heme iron group
what is the Q cycle
the route of electrons from ubiquinol to cytochrome c and there are 2 rounds of it
what happens in the first round of the Q cycle
QH2 donates one electron to iron-sulfur protein - electron travels to cytochrome c1 and then to cytochrome c - QH2 donates the second electron to cytochrome b - the two protons from QH2 are released into intermembrane space - oxidized ubiquinone diffuses to another quinone-binding site - accepts the second electrons from cytochrome b becoming a half-reduced semiquinone
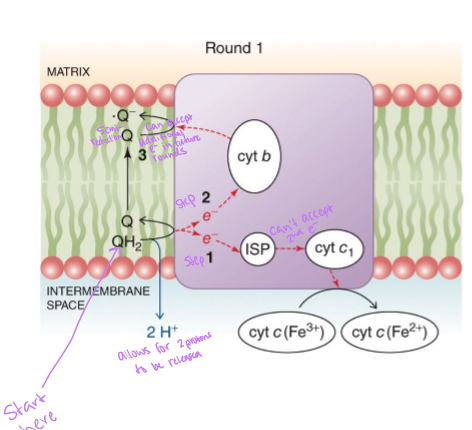
what happens in the second round of the Q cycle
a second QH2 surrenders 2 electrons to complex III and its two protons to intermembrane space - one electron goes to reduce cytochrome c - other electron goes to cytochrome b and then to the waiting semiquinone from the first round - this regenerates QH2 using the protons from the matrix
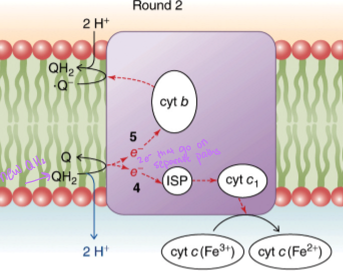
what is the net result of the Q cycle
2 electrons from QH2 reduce 2 molecules of cytochrome c and 4 protons, translocated to the intermembrane space (2 from QH2 in the first round and 2 from the QH2 in the second round)
what is the overall formula of the Q cycle
QH2 + 2 cytochrome c (Fe3+) = Q + 2 cytochrome c (Fe2+) + 2 H+
how is cytochrome c reduced
It shuttles a single electron between Complex III and Complex IV. To reduce one molecule of to in Complex IV, four individual molecules of cytochrome c must successively deliver their electrons to the complex which are consumed in the reduction of molecular oxygen to water and four additional protons are relayed from the matrix to the intermembrane space
what does complex IV (cytochrome c oxidase) do
4 cytochrome c dock here and donate electrons to copper center that is passed through the complex - the 4 electrons pass through which forms water - this process requires 4 protons from the matrix that are also pumped into the intermembrane space - no ROS formation (transports H+)
what complexes are there ROS formation
complexes I and III
what does the constant fluid interaction do to TCA, ETC, and Ox Phos
about 10 H+ are transported to the intermembrane space per NADH oxidized which creates n electrochemical gradient of protons and of charge - this is a form of energy storage that can be used to drive ATP synthesis - Q and cyt C are utilized for electron movement, but the final acceptor is O2
what is the proton motive force
the energy of the proton gradient and is derived from the concentration gradient of protons across the inner mitochondrial membrane, and the electrochemical potential gradient derived from the separation of charged molecules
how does the concentration gradient of protons help energize the proton motive force
the charge difference across the inner mitochondrial membrane produces a membrane potential gradient that is more negative inside delta trident looking thing
what is the membrane potential gradient of the ETC max operations
about -180 mV = energized → the higher the number, the less energized
what are metabolic inhibitors used for with proton motive force
to affect various stages of the ETC and are frequently used as diagnostics in assessing ETC functionality
how is the proton gradient generated
the proton-translocation activity of the electron transport complexes in the inner mitochondrial membrane - this is a source of energy that can drive the activity of an ATP synthase
what are protonmotive forces
the imbalance created by the protons that cannot diffuse back into the matrix because the membrane is impermeable to ion
how does the proton motive force drive the phosphorylation of ADP into ATP
the free energy from passage of the proton back into the matrix (delta Gdegree’ = -20kJ) would not be enough to drive the phosphorylation of ADP (needs 30.5) so the 10 protons translocated for each pair of electrons transferred from NADH to O2 have an associated protonmotive for of 218.2 which is enough the drive synthesis of several molecules of ATP
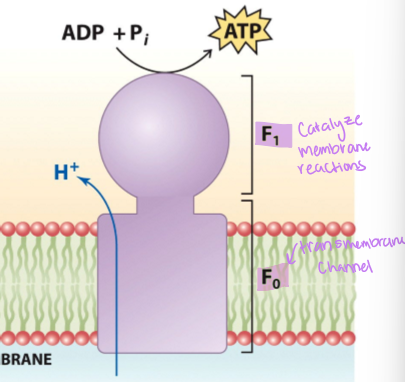
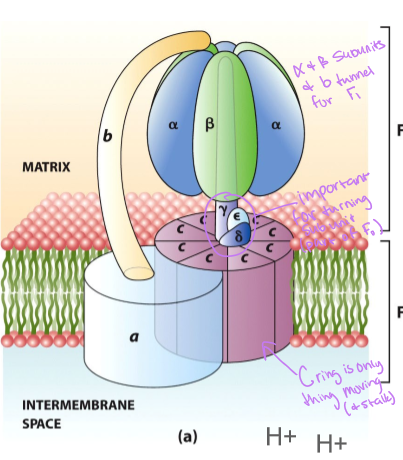
what are the components of ATP synthase
F0 , which has a transmembrane channel, and F1 which catalyzes the reaction
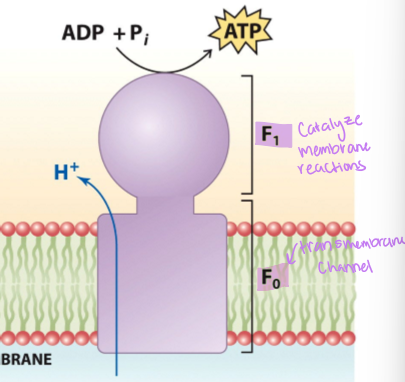
what does ATP synthase do
couples exergonic H+ transport of the accumulated protons in the intermembrane space to endergonic ATP synthesis - proton motive force drive ATP synthesis
what is required for proton transport in ATP synthase
rotation of the c ring past the stationary a subunit - the carboxylase side chain of a highly conserved aspartate or glutamate residue on each c subunit serves as a proton binding site
what are some of the features of the F0 component of ATP synthase
integral membrane protein complex - there are three subunits (a1b2c10) that rotates
what does the c subunit of F0 do
forms a rotor that passes H+ from intermembrane space to matrix through the adjacent a subunit - the overall c complex rotates relative to the a subunit
what does the b subunit of F0 do
dimer that connects alpha to sigma subunit on F1
what does the binding of protons to the c subunit of F0 do
takes up a proton from the intermembrane space and the binding neutralizes the carboxylate group which frees it from the electrostatic attraction of a positively charged arginine residue on the a subunit - the protonated c subunit then moves away, and this slight rotation of the c ring brings another c subunit into position so that it can release its bound proton into the matrix - pick up, bind, neutralize, shift, repeat
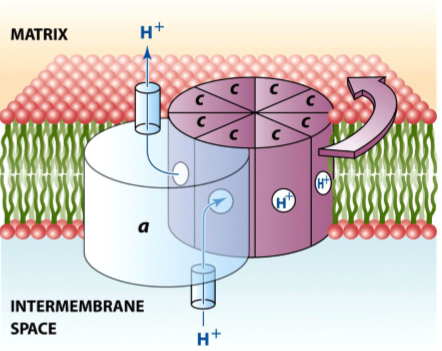
what is the structure of the F1 component of ATP
the three alpha-beta pairs change their conformations as the gamma subunit rotates - as each proton moves across the membrane, the c ring and gamma subunit rotate together relative to the alpha-beta complex and causes the conformational changes in beta
does the alpha-beta hexamer rotate in F1
no since it is held in place by the peripheral stalk b that is anchored to the alpha subunit
how many ATPs are synthesized by one full cycle of ATP synthase turning
3 ATPs
how is ATP produced by ATP synthase
uses mechanical energy (rotation) to form a chemical bond which is the attachment of a Pi group to ADP - the rotation driven conformational changes alter the affinity of each catalytic beta subunit for an adenine nucleotide
what are the different conformations of ATP synthase
at any moment, each catalytic site has a different conformation and binding affinity - open, loose, or tight state - the F1 region of ATP synthase has 3 alpha-beta pairs, each of which has a binding site, and the rotation of the F0 stalk causes them to switch between conformations
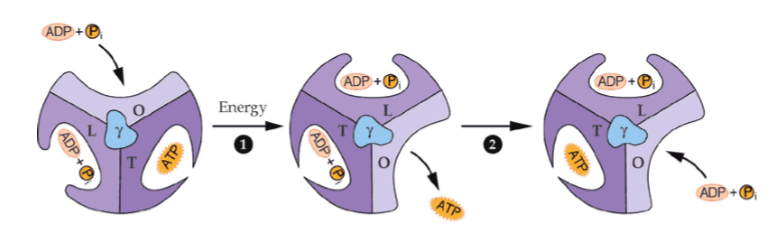
what is the open conformation of an alpha-beta pair in the F1 complex
nothing is bound (accept and release conformation)
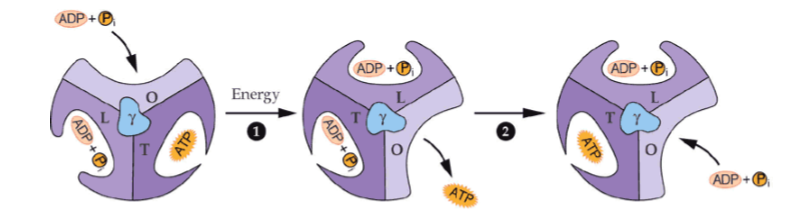
what is the loose conformation of an alpha-beta pair in the F1 complex
had bound ADP and Pi
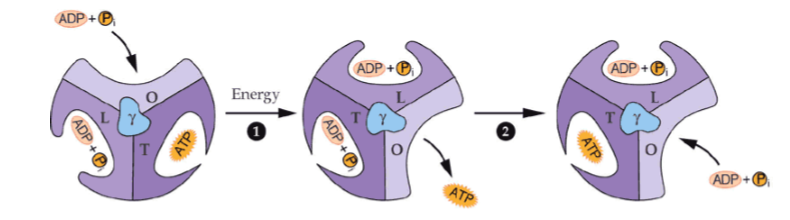
what is the tight conformation of an alpha-beta pair in the F1 complex
ADP and Pi combine to form ATP
how does PMF change the ETC flow and ATP synthesis
since electron transport is directly coupled to proton translocation, the flow of electrons through the electron transport system is regulated by the magnitude of the PMG - the high the PMF, the lower the rate of electron transport and vice versa
what does a high PMF do to the ETC flow rate
slows ETC electron flow by increasing the energy required to pump protons while providing the necessary power for ATP synthase to turn and generate ATP efficiently coupling these processes
what happens to the ETC flow rate under resting conditions
the demand for new synthesis of ATP is limited and although the PMF can be high, flow of protons back into the matric through ATP synthase is minimal - under these conditions, NADH accumulates and reduces the activity of malate dehydrogenase
what happens to ETC flow rate when energy demands are increased
cytosolic ADP rises and is exchanged with intramitochondrial ATP via the transporter activity - increased intramitochondrial concentrations of ADP cause the PMF to become discharged as protons pour through ATP synthase, regenerating the ATP pool
what do the uncoupling proteins do
proton transporters - facilitate H+ transport in the mitochondrial inner membrane which dissipates the proton motor force and reduces the amount of ATP that is produced - the energy from the proton gradient is released as heat thereby contributing to body temp
what is rotenone
an ETC inhibitor in complex I
what is antimycin
an ETC inhibitor in complex III
what is oligomycin
an ETC inhibitor in ATP synthase
what is dinitrophenol
an ETC inhibitor in the membrane for integrity
what is ubiquinone
a hydrophobic mobile electron carrier that stays within the lipid bilayer of the inner mitochondrial membrane