Biology H - Molecular Genetics unit
1/42
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
43 Terms
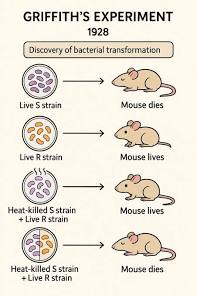
Frederick Griffith’s experiment
- In 1928, Griffith worked with two strains of a bacteria:
A pathogenic (disease-causing) “S” strain
A harmless “R” strain
- When he mixed heat-killed remains of the pathogenic strain with living cells of the harmless bacteria, some living cells became pathogenic. He named this phenomenon Transformation
Genotype
An organism’s underlying genetic makeup, such as DNA or alleles
Phenotype
An organism’s visible traits, such as physical appearance or behavior
Transformation
- Phenomena where a change in genotype and phenotype occurs due to assimilation (absorption) of foreign DNA
DNA is organized into…
genes & chromosomes
Oswald Avery, Maclyn McCarty, and Colin MacLeod’s experiment
- In 1944, Avery, McCarty and MacLeod announced that the “transforming substance” (one that transforms hereditary info) was DNA
- As the first ones to ever claim such thing, their conclusion was based on experimental evidence that only DNA worked in transforming harmless bacteria into pathogenic bacteria
- Many biologists remained skeptical about DNA’s "transforming” properties even after their study, as so little was known about it at the time
Bacteriophages (or phages)
- Virus that infects organisms by putting its DNA in their cells to affect them
- Often used in molecular genetics research
Alfred Hershey’s and Martha Chase’s experiment
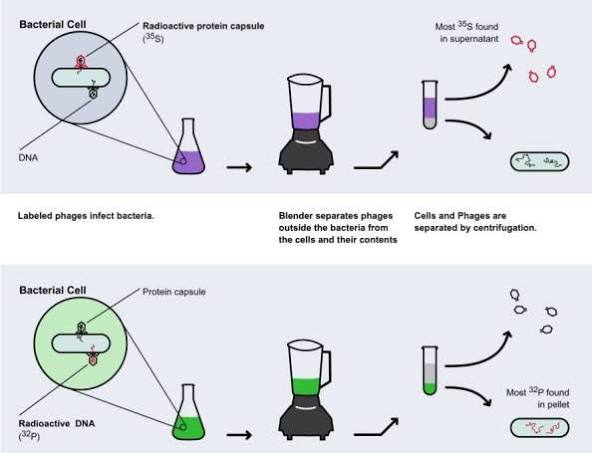
- In 1952, to determine the genetic material in the phage T2, Hershey and Chase designed an experiment to show that only one of the two components (DNA or proteins) enter an E. Coli cell during infection
- Hershey and Chase labeled the DNA of phages with a radioactive P isotope and their protein coats with a radioactive S isotope (to track which component entered bacteria during infection).
- After infecting E. coli and using a blender to remove empty phage coats (made out of proteins), they found that only the P-labeled DNA entered the cells, providing conclusive evidence that DNA was the genetic material
Erwin Chargaff’s experiment
- In 1947, Chargaff reported that DNA composition varies from one species to another, making DNA a more credible candidate for genetic material (due to diversity)
- He also found out that the molar ratio of specific bases to other specific bases was always 1:1, despite differences in base compositions across species. Soon, he created Chargaff’s rule:
% of Adenine = % of Thymine
% of Cytosine = % of Guanine
Maurice Wilkins and Rosalind Franklin’s experiment
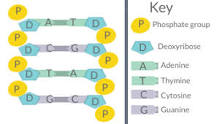
- In 1953, Franklin used a technique called X-ray crystallography to study molecular structure. Through it, they concluded that DNA was composed of two anti-parallel sugar-P backbones; with the nitrogenous bases in the middle of it
- As they continued to research DNA, Franklin made the photo that Crick and Watson would later use to determine the structure of it, as it proved that the molecule was helical and made up of two strands
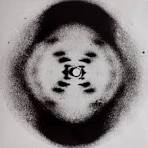
James Watson and Francis Crick’s experiment
- In 1953, Watson and Crick introduced a double-helical model for the structure of DNA, conforming to previous X-rays and chemical properties of the molecule
- They did this primarily through looking at the photo Franklin made, as its width determined that a purine (A or G) always pairs with a pyrimidine (C or T), explaining Chargaff’s findings
- With this discovery, it soon became widely accepted among scientists that:
Hereditary info is encoded in DNA and reproduced in all cells of the body
DNA directs the development of biochemical, anatomical, physiological and (to some extent) behavioral traits
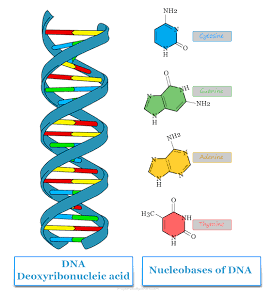
While DNA has Adenine, Thymine, Cytosine and Guanine as its nitrogenous bases, RNA has…
Adenine, Uracil, Cytosine and Guanine
DNA is … stranded, while RNA is … stranded
double; single
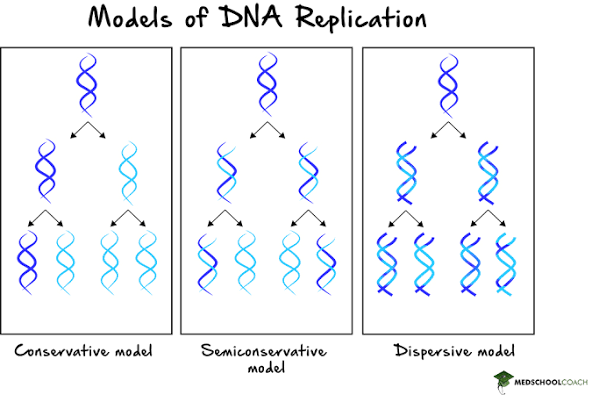
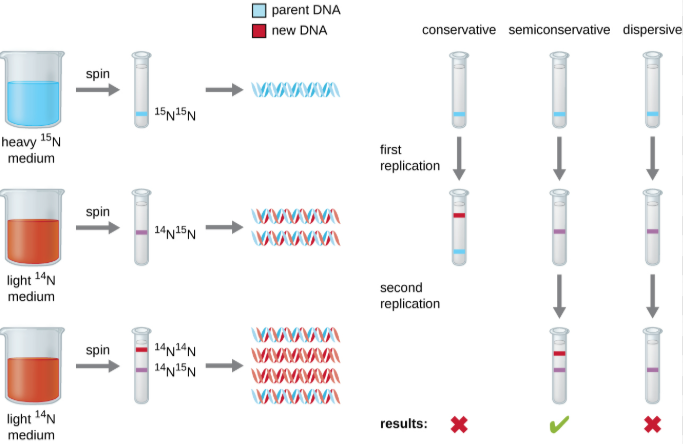
After the structure and general functions of DNA were discovered, scientists now needed to discover how exactly hereditary information was passed down. Soon, 3 models of DNA replication were pitched:
- Conservative model: Predicted that after the original double-helix acts as a template to create a completely new DNA molecule, the two original strands come back together and reassociate, restoring the original heavy DNA molecule and producing 3 identical light DNA daughter molecules (after two rounds of replication)
- Semi-conservative model: Predicted that the original parental strands separate and pair with newly synthesized strands, resulting in half of the products being hybrid DNA molecules and the other half being light DNA molecules (after two rounds of replication)
This model was supported by Watson & Crick from the very beginning, and would later be proven to be the correct one through the Meselson and Stahl experiment
- Dispersive model: Predicted that all the products would be hybrids, as the original parental strands “randomly” mixed with new strands to produce daughter DNA molecules
Only … DNA is available after … round(s) of replication
newly-synthesized (light); two
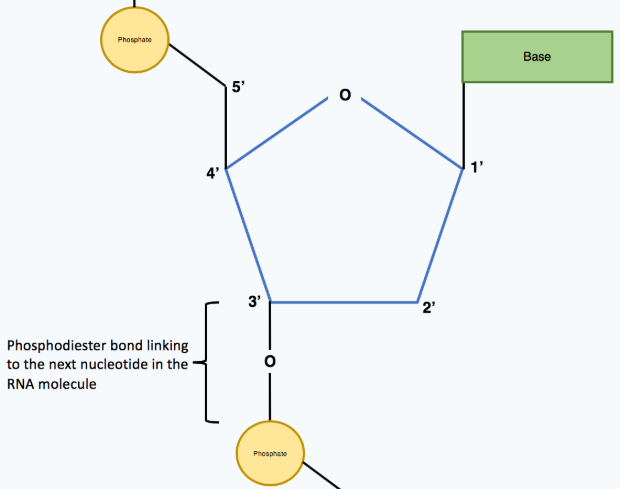
Phosphodiester bonds in DNA/RNA link…
Phosphate groups to adjacent pento-sugars
During Meselson and Stahl’s experiment, you would expect to see the light-labeled DNA at the … of the medium, as it is less dense
top
During Meselson and Stahl’s experiment, you would expect to see the heavy-labeled DNA at the … of the medium, as it is more dense
bottom
Meselson and Stahl’s experiment
- In 1958, Meselson and Stahl labeled old DNA strands with a heavy Nitrogen isotope (¹⁵N), while any new strands were labeled with a lighter isotope (¹⁴N)
- The first replication produced a group of hybrid DNA, then eliminating the conservative model (as you would expect one fully heavy and one fully light molecule)
- The second replication produced two light and two hybrid DNA, then eliminated the dispersive model (as you would expect all hybrid molecules)
- Since the phenomena perfectly fit with what the semi-conservative model predicted, the experiment ultimately proved that that was the way DNA replicated
Origin of replication
- Origins of replication are specific sequences within the parental (original) DNA molecule where the process of replication begins, seen as DNA helicase begins to unwind the double helix
- A typical eukaryote usually has hundreds/thousands of origins of replication
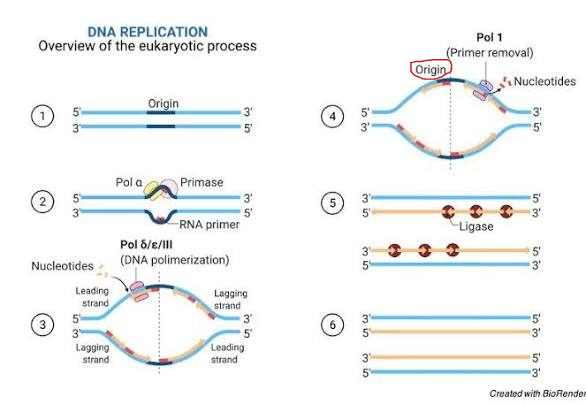
Replication “bubble"
- Replication bubbles are open regions of DNA where replication occurs, formed as DNA helicase separates parental strands at an origin of replication
- A typical eukaryote usually has multiple replication bubbles per chromosome
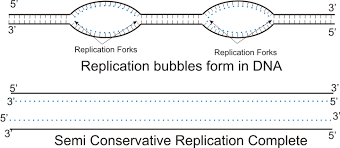

Replication fork
- Replication forks are Y-shaped structures formed when helicase unwinds DNA’s double helix, separating the strands to act as templates for replication
- Site of DNA elongation (catalyzed by DNA polymerase)
- Two forks move bidirectionally from each replication origin, with one leading and one lagging strand forming at each
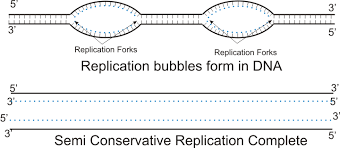

Since DNA replication is bidirectional (proceeds in both directions), … move opposite ways until the entire molecule is copied
replication forks
Because DNA polymerase only adds nucleotides to the … end of a parental strand, new DNA strands can only elongate in the … direction. This is why both leading and lagging strands towards the same ‘
3’; 5’-3’
Leading DNA strands
- Leading strands are complementary strands (ones that follow base-pairing rules) synthesized by DNA polymerase, continuously (in 5’ to 3’ direction)
- Always move towards replication fork
Lagging DNA strands
- Lagging strands are complementary strands (ones that follow base-pairing rules) synthesized by DNA polymerase, discontinuously (in 5’ to 3’ direction)
- Always move away from replication fork
Okazaki fragments
Fragments of DNA that make up lagging strands, and join together after ligase connects with them
DNA polymerase
- Enzyme that catalyze the synthesis of new DNA strands by adding complementary deoxyribonucleotides to a pre-existing DNA template during DNA replication and repair (where the nucleotides are supplied by primers produced through primase), also producing leading & lagging DNA strands
- Also proofreads/repairs DNA by overseeing newly made DNA and replacing incorrect nucleotides
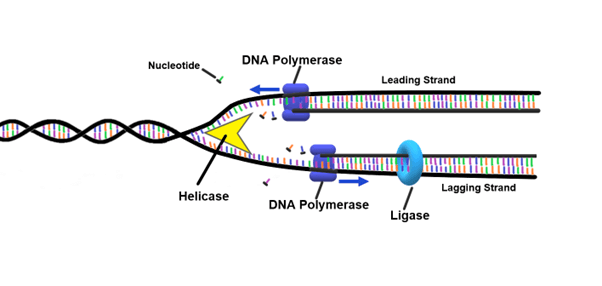
Primase
Enzyme that “kick-starts” replication by synthesizing RNA primers which DNA polymerase needs
Primers
- Groups of RNA nucleotides that initiate replication by supplying 3'-hydroxyl groups that DNA polymerase needs to begin adding nucleotides to the 3’ end of an elongating DNA strand
- Available in 3’ end only, as they can only be added from there
- While only one primer is needed for leading strands, lagging strands require multiple
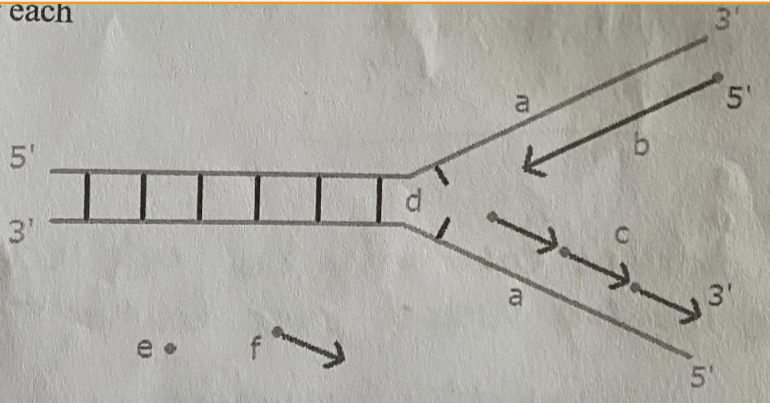
Label A-F:
A - Parental strand (old DNA)
B - Leading strand (new DNA)
C - Lagging strand (new DNA)
D - Replication fork
E - RNA primer
F - Okazaki fragments
While in leading strands DNA polymerase adds to the 3’ end of …, in lagging strands it adds to the 3’ end of …
new DNA strands; Okazaki fragments
DNA helicase
Enzyme that untwists DNA’s double helix and separates template DNA strands at the replication fork by breaking H bonds between nitrogenous bases
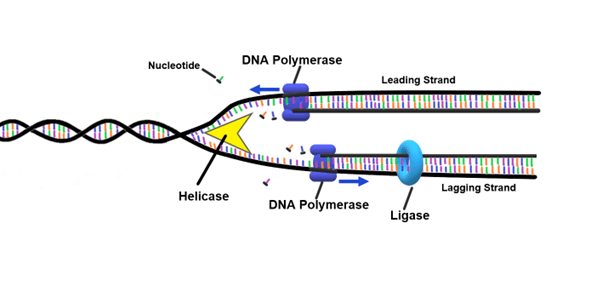
DNA ligase
Enzyme that joins Okazaki fragments together by sealing the “nicks” between the strands with phosphodiester bonds
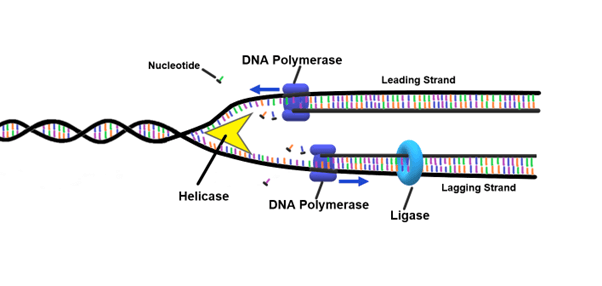
DNA topoisomerase
Enzyme that corrects “overwinding” ahead of the replication fork by breaking, swiveling, and rejoining DNA strands

There are two things that can happen when mistakes are found in nucleotides. Describe them:
- Mismatch repair (if the nitrogenous bases aren’t paired up correctly): Repair enzymes correct errors in base pairing
- Nucleotide excision repair (if nucleotides added to new DNA strand have some sort of mistake): Repair enzymes cut out and replace stretches of DNA affected by nucleotide(s)
Describe how a limitation of DNA polymerase affects the linear DNA of eukaryotic chromosomes:
Because the DNA replication “machinery” provides no way to complete the 5’ ends of strands, repeated rounds of replication produce shorter strands, leading to erosion of DNA (aging)
Telomeres
- Repetitive DNA at the end of eukaryotic chromosomes that act as a protective measure against the production of shorter DNA strands with time (due to DNA polymerase’s limitation)
- While they do not completely prevent the shortening of DNA strands, they postpone the erosion of genes near the ends of DNA molecules
Because telomeres delay DNA erosion, shorter telomeres in eukaryotes are associated with…
shorter life expectancies
Telomerase
- Enzyme that produces telomeres
- Aging is strongly related to the shortening of these
Scientists researching have tried to “block” telomerase in cancerous cells because:
The sudden stop in the production of telomeres should (supposedly) prompt the cancer cells to die, as DNA erosion would happen quicker than normal with no protective layers (telomeres)
List major difficulties that come with blocking telomerase in cancer cells:
- Impairment of fertility
- Deceleration of wound-healing
- Reduction in production of red/white blood cells
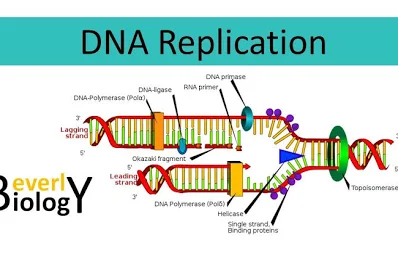
Describe the process of DNA replication, step-by-step:
- Helicase breaks H bonds between nitrogenous base pairs from the origins, creating a replication “bubbles” (and replication forks inside of them)
- Concurrently, DNA topoisomerase is repairing “overwinding” DNA strands ahead of the replication fork by breaking, rearranging or rejoining them
- Primase synthesizes RNA primers to supply DNA polymerase with nucleotides
- DNA polymerase then adds those new nucleotides in a 5′→ 3′ direction, forming the leading strand (continuously)
- On the lagging strand, DNA polymerase synthesizes discontinuously, creating short fragments called Okazaki fragments
- DNA ligase then seals the gaps between Okazaki fragments, forming a continuous strand
- As DNA polymerase produces strands solely towards the 3’ direction, telomeres (repeated DNA) at the furthermost 5’ point protect valuable genetic info by delaying its erosion
- As proofreading, DNA polymerase molecules (and other repair enzymes) proofread the pairing of nitrogenous bases/arrangement of nucleotides added to new DNA strands
- By the end, two identical DNA molecules are produced, each containing one old strand and one newly synthesized strand (hybrids)