Midterm 3 Bio A
1/66
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
67 Terms
Non-Mendelian Genetics
1) X-linked
2) Sex-linked
3) Multiple Alleles
4) Codominance
5) Incomplete dominance
6) lethality
Mendel’s Experiment
Used pea plants
Focused on flower color, seed shape, and pod color
Law of segregation
Law of independent assortment
Traits are inherited as discrete genes
Dominant traits can mask recessive ones
Law of segregation (Mendel)
During the formation of gametes, the two alleles for a trait segregate from each other
each gamete only carries one allele for each gene
Principle explains how offspring inherit one allele from each parent
Results in genetic combinations
Law of independent assortment (Mendel)
Alleles for different traits segregate independently of one another during gamete formation
Inheritance of one trait will not affect the inheritance of another trait
Only applies to genes located on different chromosomes or those far apart on the same chromosome
Leads to genetic variation
Crossover
occurs during meiosis
Homologous chromosomes exchange segments of genetic material
Exchange happens during prophase I
Increases genetic diversity by allowing variations of traits to be passed on
Leads to evolution and adaption
Relationship between Independent Assortment & Crossover
Contribute to genetic variation during meiosis
Independent = split without influencing other traits
Crossover = exchange of genetic material
Linked Genes
Located together on same chromosome
Tend to be inherited together
Do not assort independently
Crossing over can break linkage
The further the genes are from each other, the more likely a crossover event will happen
Can have multiple crossover events
0-50% (Independent assortment max is 50%)
Explain how crossover happens
Occurred during meiosis I (specifically during Prophase I)
Synapsis
Homologous chromosomes must find each other (one from each parent) before crossing over can happen
They line up side-by-side, gene-by-gene
Tight pairing is called synapsis
Held together by protein structure (synaptonemal complex)
Chiasmata Formation
Once paired, the non-sister chromatids (one from each parent) overlap at specific points
Contact points called chiasmata
This is where actual break and swap happens
DNA swap
specialized enzymes break the sugar-phosphate backbone of the DNA at the same location on both non-sister chromatids
Broken segments re-sealed to the opposite strand
Genetic Recombination
As meiosis continues, the homologous chromosomes are pulled apart
Recombinant chromosomes
Allele
Specific variation of gene
Gene —> general characteristic —> alleles → determine specific expression of trait
Found on locus
Homozygous
Heterozygous
Dominant
Recessive
Polygenic Traits
Polygenic inheritance controlled by 2+ genes
Creates a continuous gradient (bell curve)
Each dominant allele “adds” a small amount of phenotype
skin
height
eyes
Pleiotropy
One single gene influences multiple traits
Usually happens because the gene codes for a protein that is used by many different types of cells throughout the body
Synergistic genes
Synergistic interactions results in more intense phenotype (polygenic)
happens because the genes are part of the same metabolic or signaling pathway
Often, a certain trait won't appear at all until a specific combination of synergistic genes is present
Epistasis
When the effect of one gene is hidden or masked by the presence of a different gene
one gene overriding
gene that does the masking is called epistatic
the gene being masked is hypostatic
usually happens because the epistatic gene controls a step that occurs earlier in a biological pathway
Recessive Epistasis
two copies of the "masking" allele to hide the other gene
Dominant Epistasis
one copy of the "masking" allele to hide the effects of the other gene
Monohybrid Cross
Tracks the inheritance of one single trait
3:1 (phenotype)
1:2:1 (genotype)
Dihybrid Cross
tracks the inheritance of two independent traits simultaneously
9:3:3:1 (Phenotype)
Test Cross
discover the unknown genotype of an individual showing a dominant phenotype
Self Cross
occurs when an individual is crossed with itself or
with an individual of an identical genotype
Pedigree Analysis
visual chart (a "family tree") that tracks the inheritance of a specific trait through multiple generations
can determine if the gene is dominant or recessive and if it is located on an autosome or a sex chromosome
Autosomal dominant: every generation rule; males and females with similar frequency
Autosomal recessive: Unaffected parents can have affected children (meaning the parents are "carriers")
X-linked
X-Linked Dominant
If a father is affected, all of his daughters will be affected, but none of his sons
An affected mother has a 50% chance of passing it to any child
Hemizygous
describes an individual who has only one copy of a particular gene or chromosome segment, instead of the usual two
we inherit two alleles for every autosomal gene, if one copy is missing—either naturally or due to a deletion
Sex Chromosomes
Females: XX
Males: XY (hemizygous) - fewer genes (shorter)
Y has a region similar to x for meiosis
Multiple alleles
Wild type (+) → Dominance; refers to phenotype/genotype most commonly observed
Variants (-) → differs from wild type
If variants becomes common (>1%) then it becomes a polymorphism
Dominant hierarchies
forms when multiple alleles exist in a population
the alleles are ranked by their ability to mask one another
Codominance + Blood type
equal expression of alleles
Blood types:
Type A: people have A antigens → anti-B antibodies
Type B: people have B antigens → anti-A antibodies
Type AB: people have both equal parts A and B antigens → no antibodies → Universal acceptor
Type O: people have no antigens → have both A and B antibodies → universal donor
Rh Factor (follows normal mendelian)
+ → have protein (dominant)
- → no protein (recessive) - produce protein to attach Rh protein
Incomplete dominance
intermediate blend
dominant allele isn't strong enough to fully mask the recessive one
the heterozygous offspring ends up with a "diluted" version of the trait
Most times the recessive allele is a loss of function allele
Genotypes: 1:2:1
Phenotypes: 1:2:1
Snapdragons
the plant produces the full amount of red pigment (dominant)
recessive produces no red pigment, flowers are white
Pink flower
Lethality
refers to a situation where a specific combination of alleles is so disruptive to an organism's development or survival that it results in death
usually happens because the gene involved is essential
organism only dies if it inherits two copies of the lethal allele (homozygous recessive)
Carriers: may show a distinct phenotype that looks like a dominant trait
Only one copy of the allele is needed to cause death, allele isn’t passed on
lethal effect doesn't kick in until after reproductive age (like Huntington’s Disease in humans), the allele can persist in a population
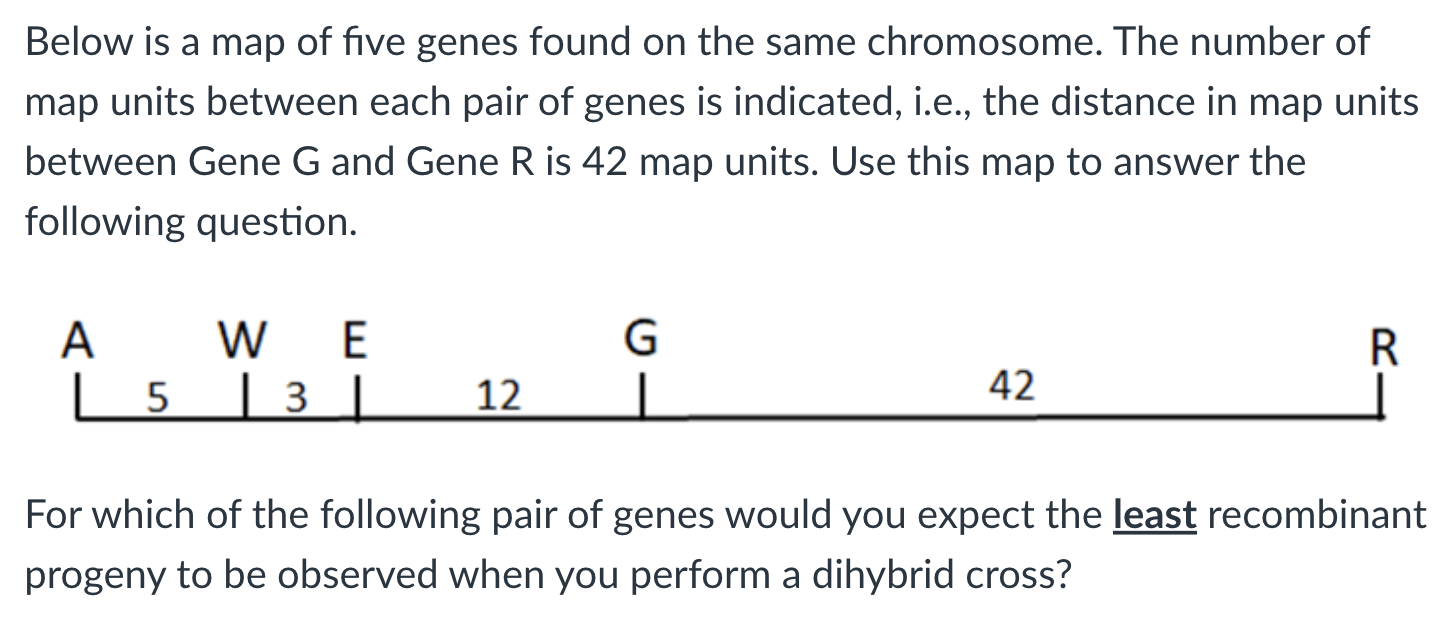
a) A and W
b) G and R
c) W and E
d) E and G
C
A series of dihybrid crosses are done between Labrador dogs with black coats. The ratio of phenotypes of the offspring is found to be 9 black to 3 brown to 4 yellow coat color. This ratio of phenotypes results from _____.
a) gene linkage
b) codominance
c) incomplete dominance
d) epistasis
D
A homozygous plant with red flowers is crossed to a homozygous plant with white flowers and three phenotypes are observed among the offspring in the following ratios: 1 red to 2 pink to 1 white. The ratio of the phenotypes of the offpsring of this cross is likley caused by _____.
a) incomplete dominance of the red allele.
b) gene linkage between the red and white alleles.
c) Mendelian dominance of the white allele.
d) epistasis of the white allele
A
If you cross two heterozygous organisms and observe that the ratio of three offspring phenotypes is 1 to 2 to 1 then the ratio can only be the result of two codominant alleles. (T or F)
F
Based on Figure 12.8 and the paragraph immediately preceding the figure, you crossed two rabbits with Himalayan fur to each other and observed that the ratio of the offspring was 3 Himalayan fur to 1 albino fur. What must be the genotypes of the two parents?
a) chch and cc
b) chc and chc
c) cc and cc
B
Linkage
Less than 50% new recombinants
Reduced crossover
Use test cross to see linkage by using phenotypic ratio and summing the recombinants, and dividing over total DNA
Gives distance and order of genes (ex 17% crossover = 17 map units)
Unlinked
Mendelian 1:1:1
Expect 50% novel allele combos
Half parental, half recombinant
Cell cycle
Interphase: cell grows and DNA is replicated - DNA semi-condensed
G1
S
G2
Mitotic: replicated DNA and cytoplasmic contents are seperated
PMAT
Interphase
G1: Cell accumulates building blocks of chromosomal DNA, proteins, and energy
S: Formation of identical pairs of sister chromatids attached at centromere; centrosome duplicated → mitotic spindle
Centrioles (rod-like) + centrosomes
G2: Cell replenishes its energy stores and synthesizes proteins necessary for chromosome movement
Cell organelles duplicated
Cytoskeleton dismantled
Final prep
G0: Cells not actively dividing
Environment
Will stay until improved or triggered
Mitotic
PMAT
Prophase
Nuclear envelop dissociates
Membranous organelles
Nucleolus disperses
Centrosomes move to opp ends
Sister chromatids coil more tightly with aid of cohesin proteins
Prometaphase
Nuclear envelope fragments
Mitotic spindle continues to develop as more microtubules assemble and stretch
Each sister chromatids develops kinetochore in centrometric region
Chromosome oriented until the kinetochores of sister chromatides face opposite ends
Spindles that do not engage are called polar microtubules
Microtubules overlap midway b/w 2 poles and contribute to cell elongation
Metaphase
All chromosomes align along metaphase plate
Sister chromatids are sitll tightly attached by cohesin proteins
Chromosomes maximally condense
Anaphase
Cohesin proteins degrade
Sister chromatids separate at the centromere
Chromatid = single chromosome
chromosomes pulled rapidly toward the centromere
cell elongates
Telophase
Chromosomes reach opposite poles and begin to decondense
Relaxed in stretched-out chromatin configuration
Mitotic spindle depolymerized into tubulin monomers
Will be used to develop cytoskeletal components for each daughter cell
Cytokinesis
optional - if doesn’t happen then daughter cells are multi-nucleated
Physical seperation of daughter
Animal cell:
starts in late anaphase
contractile ring composed of actin filaments forms inside the membrane
Actin filaments pulls the equator of the cell inward forming a fissure (cleavage furrow)
Furrow depends a the actin ring contracts
Plant cell:
new cell wall forms
during interphase, the golgi apparatus accumulates enzymes, structural proteins, and glucose propor to breaking into vesicles and dispersing throughout the dividing cell
telophase: golgi vesicles fuse and coalesce from the center to walls
as more fuse, gets longer
Internal checkpoints
Near end of G1: DNA damage, cell size, adequate reserves; cell fix, G0 until triggered or improved
G2/M: All chromosomes replicated/DNA damaged; complete DNA replication OR try to fix damage
Metaphase: Determines whether all sister chromatids are anchored to at least 2 spindle fibers from opposite sides
External regulators
apoptosis
human growth hormone (HGH)
Lack of HGH: inhibit proliferation (dwarfism)
Too much of HGH: excessive proliferation (gigantism)
Crowding of cells could inhibit proliferation
If a cell becomes too big, and starts to lack in nutrients, will divide
Regulator molecules
For the cell cycle to continue:
All positive regulators on
All negative regulators off
Regulator molecules may act independently, influence activity or production of other regulatory proteins
Positive Regulation
Promotes progress
cyclins
Cdks
Cyclins & Cdks
Cdks are enzymes (kinases) that phosphorylate other proteins (shape change)
Cyclins regulate cycle when tightly bound to Cdks
4 cyclin proteins fluctuate based on timing of cycle
Cdk concentration is constant, what changes is whether it’s active or not
1) Specific cyclin binds to Cdks, promoting Cdk shape change
2) Cdk → target protein - takes p from ATP and attaches to protein
3) Target protein shape change
4) Cyclin degrades
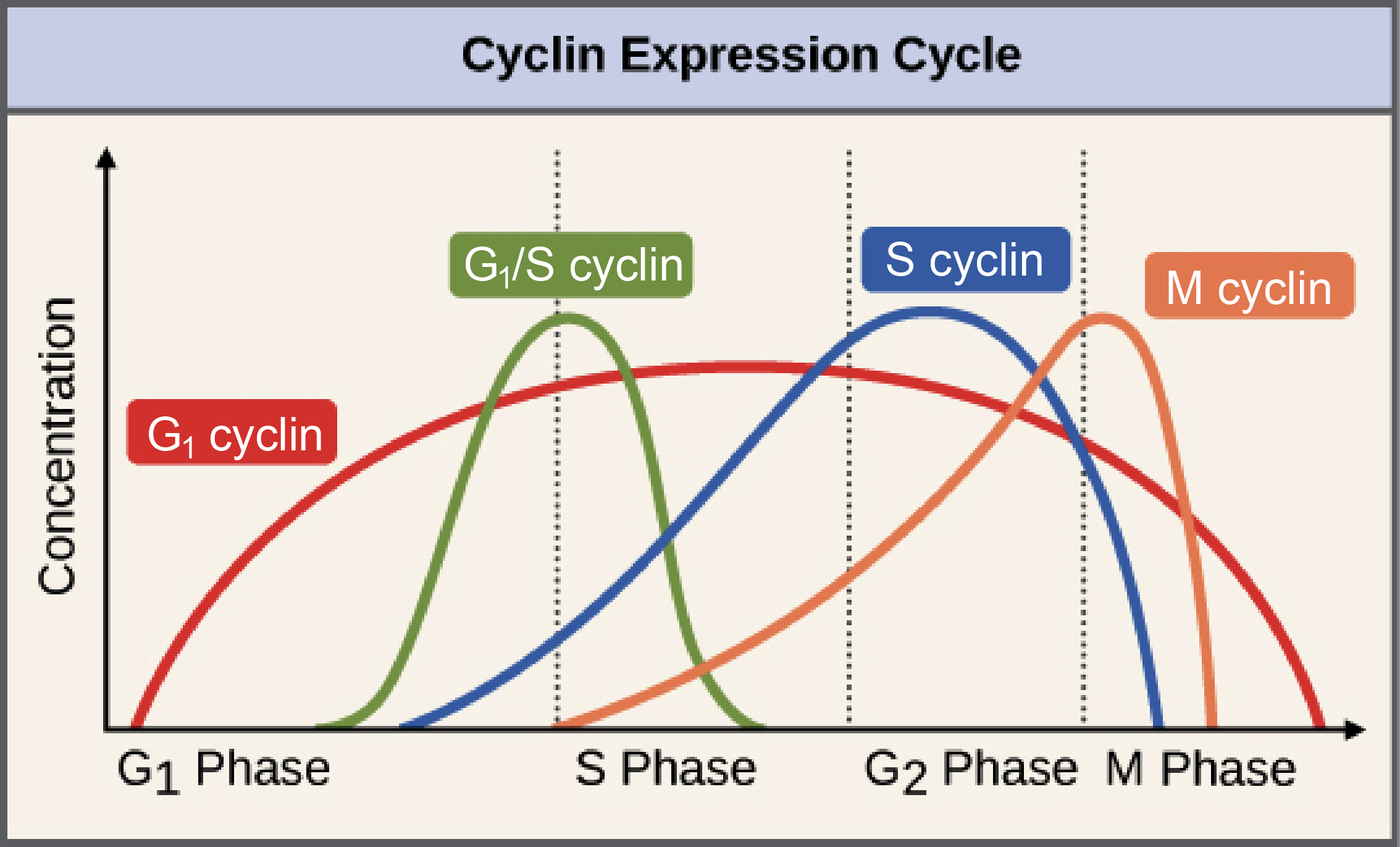
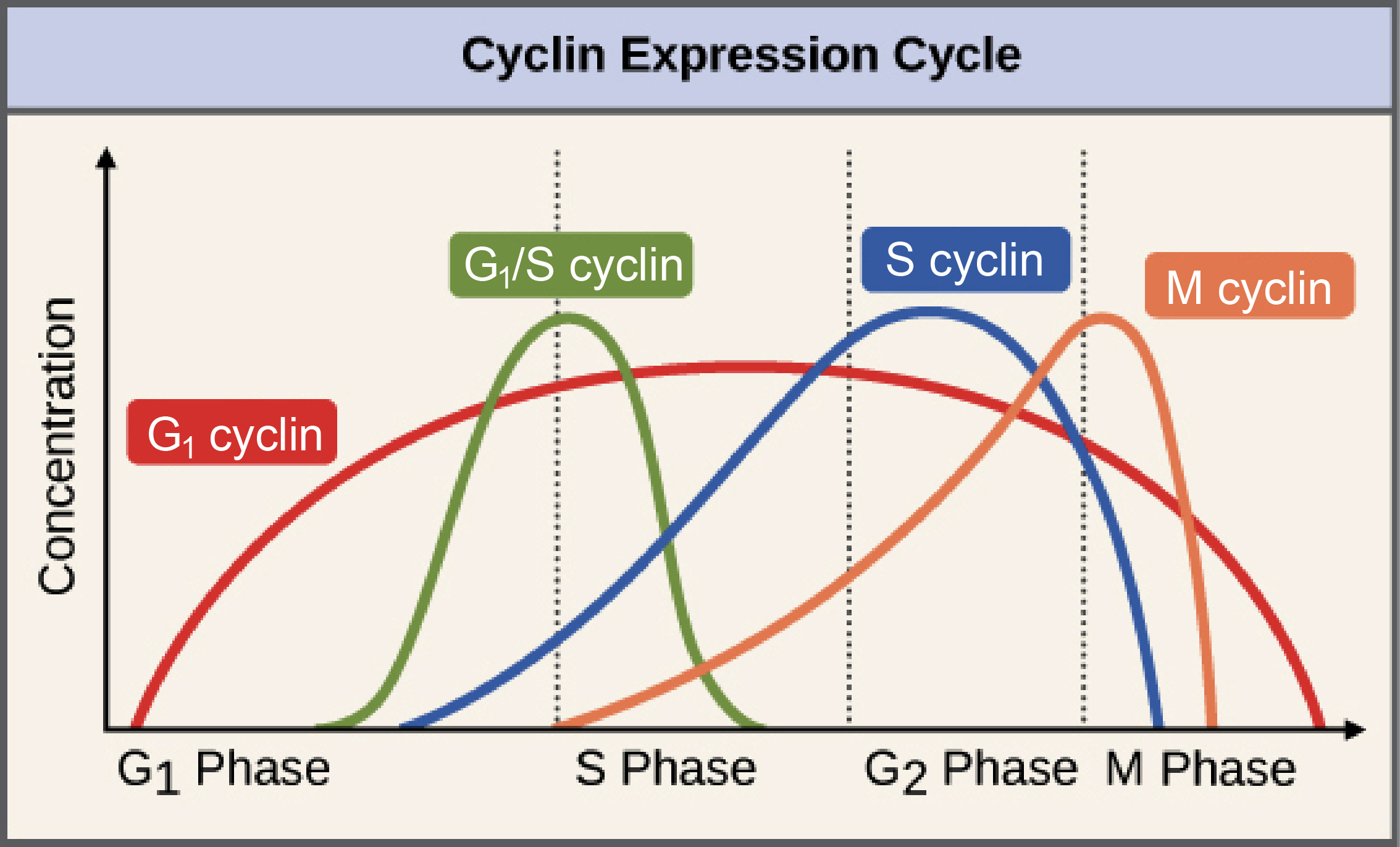
Cyclin Fluctuation Throughout Cell Cycle
1) Once G1 initiated, cyclin D synthesized and drives the G1/S phase transition
Rb phosphorylates, bound to E2F
2) Rb releases E2F which starts transcription of cyclin E which fully phosphorylates Rb
3) Cylclin E produces p27 which inhibits cyclin D
Phosphorylation of p27 tags it for degradation
4) Degradation promotes production of cyclin A by remaining CCRE from promotor (negative feedback) allowing it to enter S phase
E2F on cyclin A is part negative feedback loop
Cyclin A + CDK2 complex phosphorylates E2F, preventing from removing suppressor
5) Cyclin B → Maturation-Promoting Factor (MPF) → cells exit M phase
Negative Regulation
Cdk inhibitors: can trigger apoptosis
Rb (retinoblastoma protein): tumor-suppressor proteins
p53: if damaged, halts cell cycle and recruits specific enzymes to repair DNA, tumor-suppressor
p21: enforces the halt in the cycle by binding to and inhibiting cyclin/Cdk complexes
increase of p53 triggers production of p21
These all act primarily at the G1 checkpoint
retinoblastoma protein (Rb)
largely monitors size
inactive, dephosphorylate Rb binds to TF (E2F)
Rb → E2F, production of proteins for G1/S block
Increase in size, Rb becomes inactive and releases E2F
p53
1) DNA damage activates kinases that phosphorylate p53
2) Phosphorylated p53 turns on genes that inhibit cell cycle
Helps repair DNA after G1 phase (before it replicates DNA in S-phase)
3) Inhibiting the cell cycle gives the cycle time to repair the damaged DNA
2 Hit Hypothesis
Both copies of a tumor suppressor gene must lose function for cancerous phenotype to develop
2 mutations = 2 hits for cancer to develop
If have one copy, the second copy could later develop
Hereditary: passed two copies down
Nonherediatry (sporadic): Normal → develops one mutated tumor suppressor → the other one develops
Cell Signaling Pathways
MAPK/ERK
Wnt
Both increase cyclin D
MAPK/ERK
MAPK (Mitogen-activated protein kinase)
Activated by TGF(alpha) → EGFR regulates (growth factor receptor) through sos/Grb2 → activates Ras
Cascade: Ras → B Raf → MEK ½ → Erk ½
Erk once activated translocates to nucleus, activating several genes and TF
c-myc, Ets, c-Jun, c-fos → all relate to cell proliferation, survival, and metastasis
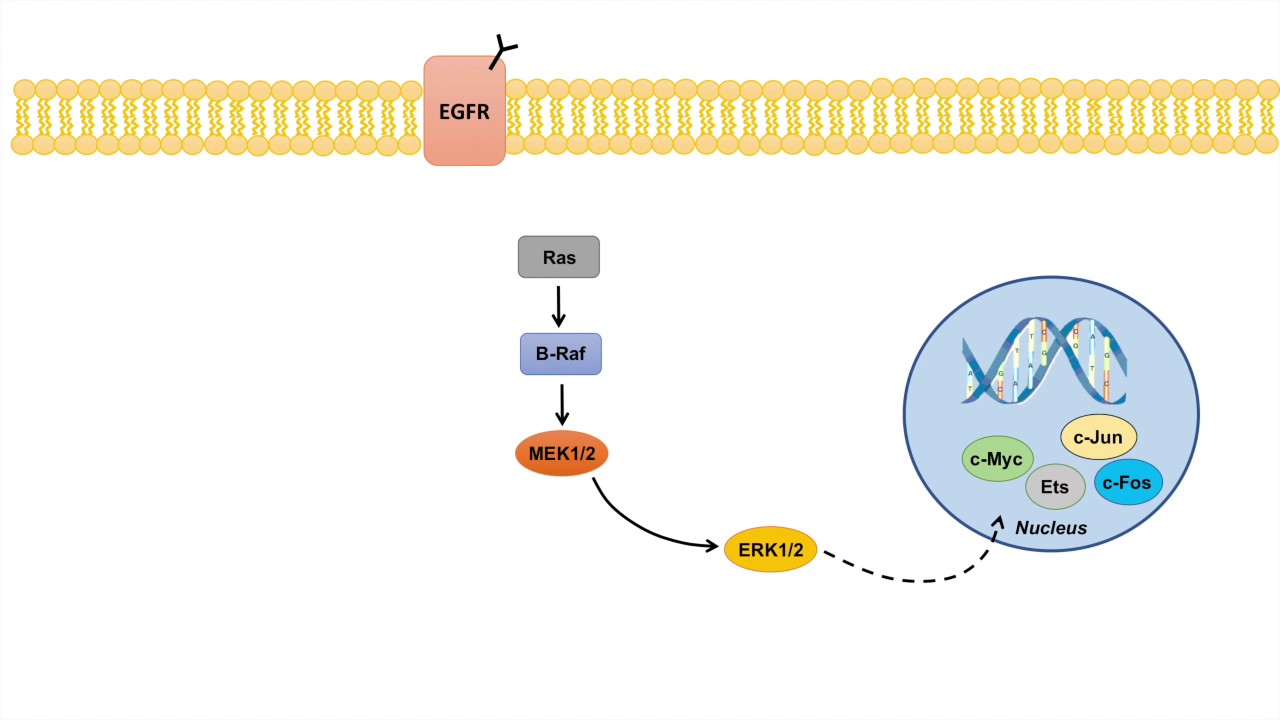
c-myc
proto-oncogene, may become an oncogene
only need 1 copy mutated for mutation
excess amounts: codes for proteins that increase proliferation
3 oncogenic mutations
1) Mutation/Deletion - proteins with increased function
2) Gene duplication - more copies
3) Translocated enhancer/promoter increases transcription - more copies
Immediate Early Genes (IEG)
c-fos, Elk-1
ERK immediately phosphorylates c-Fos and Elk-1 because they don’t require new protein synthesis to be activated
Normal protein
Regulatory: allow to make protein when needed
Proto-oncogene: need to replicate
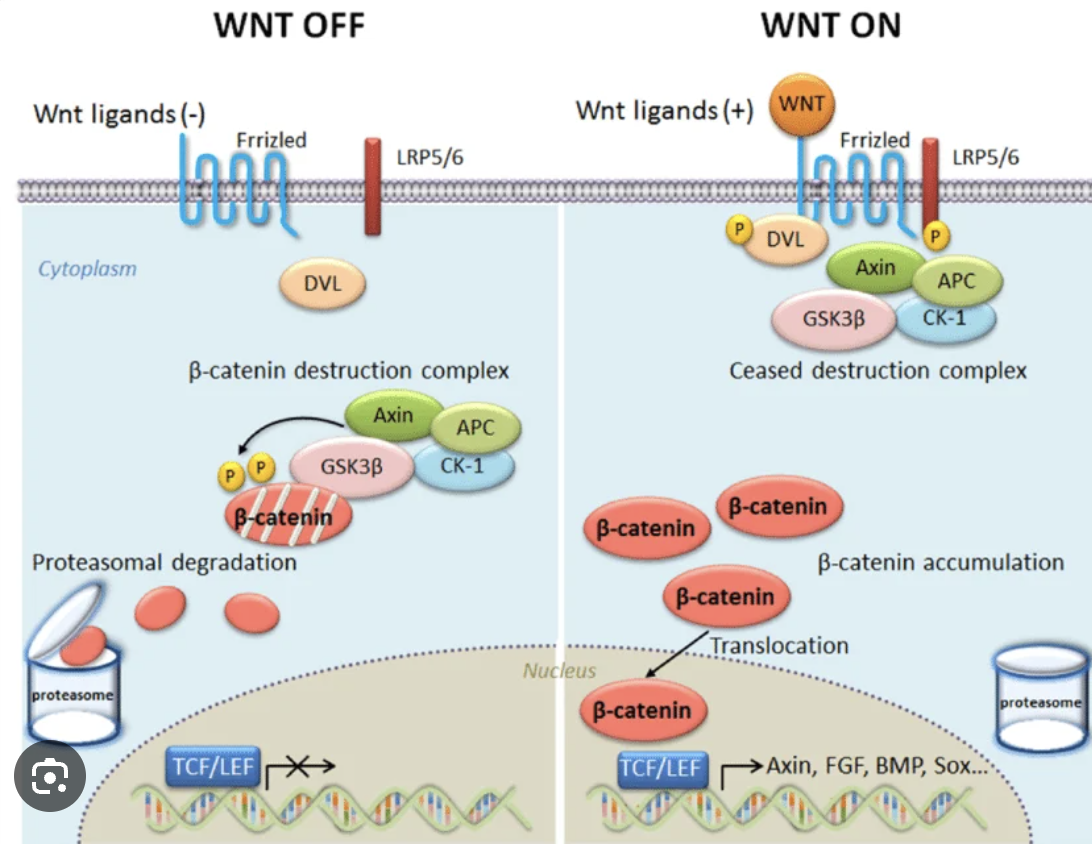
Wnt Pathway
On: beta-Centenin goes inside nucleus to tell it to divide
Off: Proteosome, no gene transcription for division
During which phase of the cell cycle does DNA replication occur?
a) G1
b) S
c) G2
d) G0
b
Sister chromatids are pulled apart by mitotic spindle fibers during which phase of mitosis?
a) Prometaphase
b) Metaphase
c) Prophase
d) Anaphase
d
The phase of mitosis in which the nuclear envelope re-forms in preparation for cytokinesis is called _____.
a) prophase
b) metaphase
c) telophase
d) prometaphase
c
Which statement about cyclins is FALSE?
a) Cyclins are positive regulators of the cell cycle
b) Cyclin concentrations rise and fall throughout the cell cycle
c) Cyclins bind to CDKs
d) Cyclins phosphorylate target proteins
d
What is the normal function of Rb?
a) Rb is a positive regulator of the cell cycle
b) Rb phosphorylates target proteins
c) Rb activates transcription of multiple genes
d) Rb inhibits progression through the cell cycle
d