Greenblatt Research
1/39
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
40 Terms
Combining genomics and model organism genetics to understand infertility and autism spectrum disorders
Human Primordial Follicles vs. Human Mature Oocytes
Primordial follicles: stable for 4-5 weeks
mature oocyte: stable for days - weeks
aneuploidy in 30-70% of human embryos
question: how do oocyte survive long periods of arrest (cell cycle is paused/transcriptionally inactive)
Transcriptionally Inactive Drosophila Oocyte
transcriptionally inactive Drosophila mature oocytes have a limited lifespan
uses genetic info differently than the central dogma, they do not produce new RNA
survive for a limited period by relying entirely on stored RNA transcripts
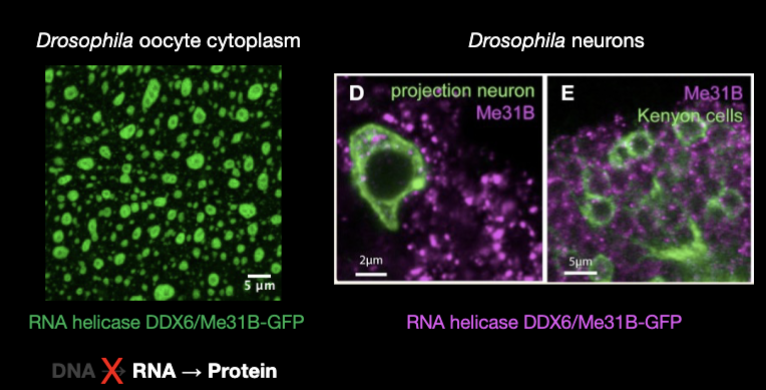
Drosophila oocyte cytoplasm:
RNA is spatially organized, storage and enzymes keep RNA stable for during transcriptionally silent phase
Drosophila neurons: store embryonic programs in RNA until synaptic activation triggers translation
Questions to ask based on previous slide
how stable is a cell whose gene regulation is based entirely on stored mRNAs
how do defects in this type of system contribute to reproductive/neural disorders
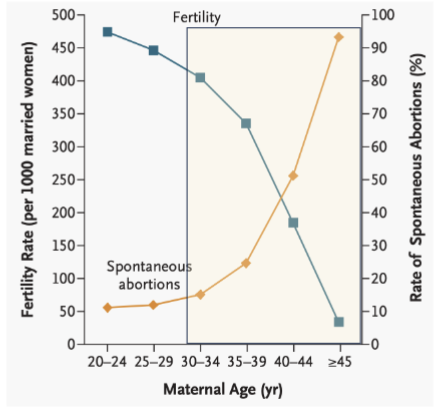
What is the largest risk factor for female infertility?
Age
the ability of oocytes to maintain proper chromosome segregation declines.
leads to meiotic errors, resulting in aneuploidy (eggs with the wrong number of chromosomes)
causes spontaneous abortion
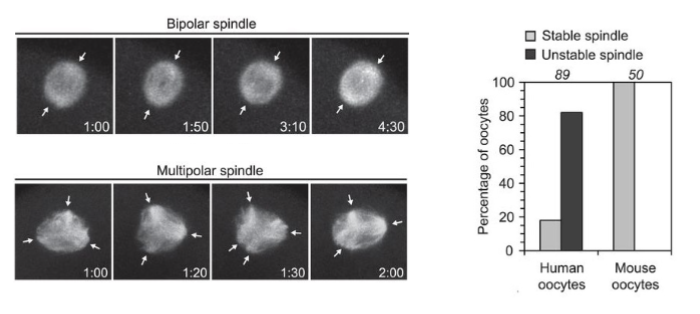
Is the meiotic spindle stable in human oocytes?
no the meiotic spindle is uniquely unstable in human oocytes
normally, the spindle ensures each daughter cell receives an equal amount of info
Bipolar spindle → normal chromosome segregation
Multipolar spindle → abnormal segregation → aneuploidy
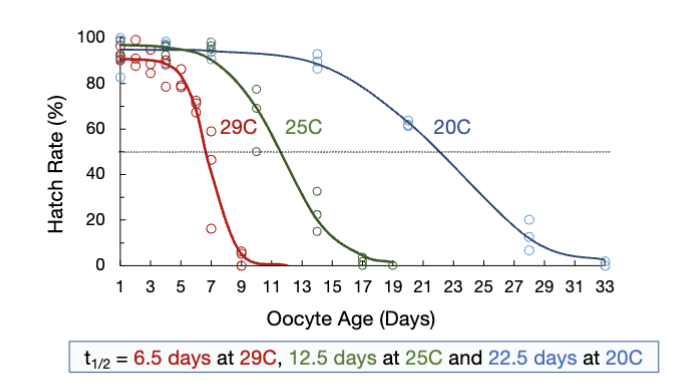
How to quantify the half-life of an oocyte?
tracking hatch rate over time at diff temperatures
half-life of oocytes decreases as temp increases
oocyte viability declines over time
Questions to ask based on previous flashcard
what changes in arrested oocytes?
what genes are essential during oocyte arrest?
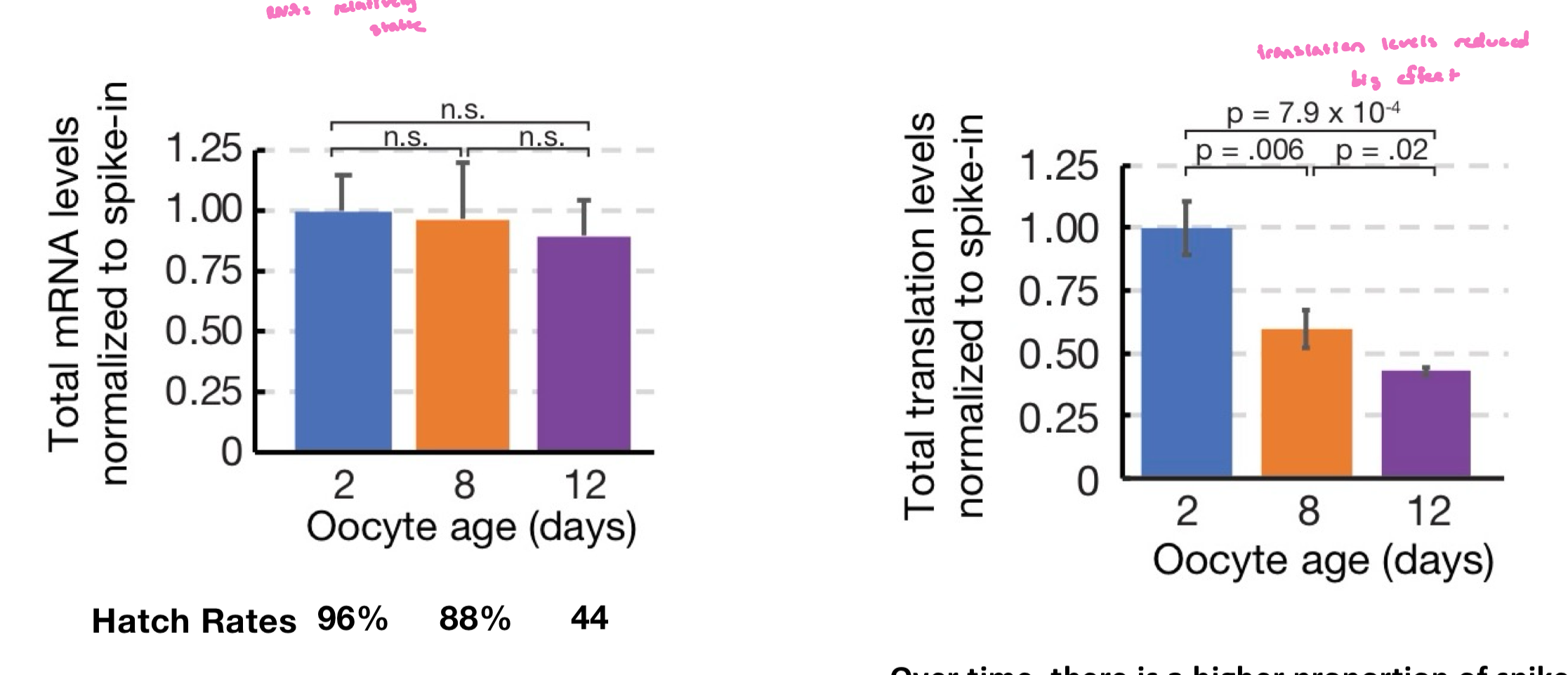
How stable is oocyte gene expression during aging? Two options
1) reduced mRNA levels
2) reduced translation (when hatched, already have all the ribosomes)
Measuring changes to transcript and translation levels
spike-in = Drosophila pseudoobscura ovary extract
allows us to measure changes globally and in individual genes (normalizes the data so you can accurately compare transcript or translation levels between diff oocyte ages)
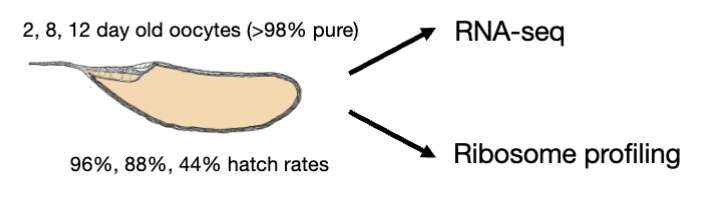
Spike ins are…
spike-ins are loading controls for sequencing experiments
the total number of reads from a sequencing experiment does not necessarily direct related to the amount of starting material, many steps in the process of producing a library may vary in their efficiency from prep to prep
spike-in controls help us compare the relative amounts of starting materials between samples
Spike ins: Jelly bean analogy

Is translation from stable mRNA consistent over time?
no, translation from stable mRNAs decline over time
over time, there is a higher proportion of spike-in reads due to decreased translation
Questions to ask from previous slide
Are the genes translated in developmentally arrested oocytes important?
What genes are actively translated in oocytes?
most mRNAs are stored in the oocyte and used for events in early embryogenesis
are there some mRNAs that are there specifically to help preserve the egg?
experiment: compare translation in oocytes vs early embryos → if an mRNA is translated specifically in stored oocytes, then it may be there to help oocytes survive
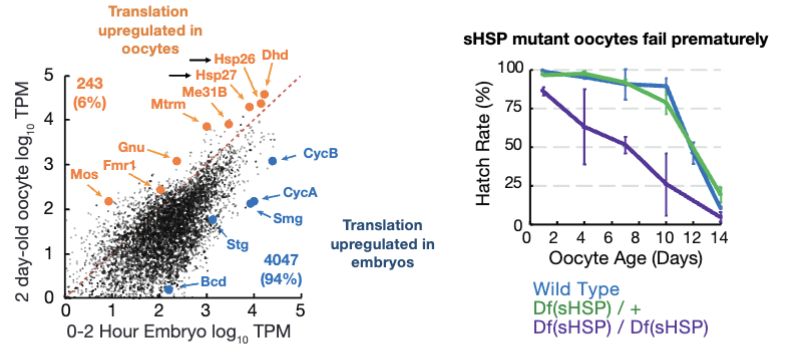
What percent of genes are translationally up-regulated during arrest?
~6% of genes are translationally upregulated during arrested
Some mRNAs are actively translated during the arrest period to maintain oocyte health/survival
small heat shock proteins help preserve oocytes, w/o these proteins, oocytes fail prematurely
oocyte-regulated genes represent a potential pilot light for oocyte survival
Questions to ask on previous slide
what genes are translationally reduced with aging?
Does oocyte translational decline affect key pathways?
yes, oocyte translational decline affects key pathways (no protection for genes)
As oocytes age, their ability to translate stored mRNAs declines, even though the mRNAs are still present
Translation is critical for producing proteins that maintain cellular structures and prepare the oocyte for fertilization
e.g. Proteins involved in meiotic spindle assembly and cell cycle regulation are particularly impacted
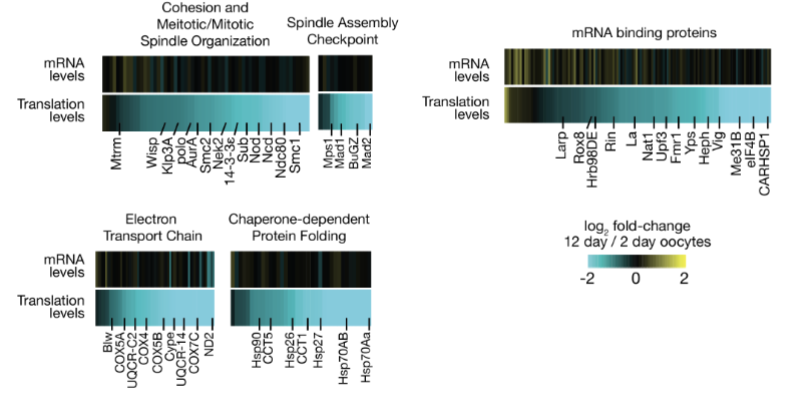
Could reduced translation of spindle proteins contribute to instability?
yes!
aged Drosophila oocytes and majority of human oocytes exhibit meiotic spindle defects
Hypothesis
the inability of oocytes to maintain their translation may be driving their failure during storage
whether the oocyte can use its stored mRNA effectively.
If oocytes lose the ability to translate these stored mRNAs efficiently, then essential proteins are not produced in sufficient amounts.
This translational decline could explain why older oocytes fail, e.g., lose spindle integrity, fail to divide properly, or cannot support embryo development
Approach
ask the fly “what pathways are bottlenecks for egg aging”
hypothesis: if a pathway is a bottleneck, putting more pressure on this pathway should have a big effect
Chromosomal deficiencies remove (“knock out”) sections of chromosomes, causing a partial loss of dozens of genes simultaneously
By observing which deficiencies cause premature oocyte failure, researchers can pinpoint pathways that act as bottlenecks
Ribosomal Subunit Haploinsufficiency
ribosomal subunit haploinsufficiency causes premature oocyte failure
screen performed to find genes whose partial loss reduces oocyte survival
9 genes (0.3%) caused premature oocyte when reduced
8 out of these 9 genes encode ribosomal subunits → suggests the ribosome is a critical bottleneck for oocyte survival.
Control: 50% survive 14 days; Df(RpL29)/+oocytes [haploinsufficient for a ribosomal subunit]: 50% survive 5.5 days
Ribosomal Subunit Haploinsufficiency Figure
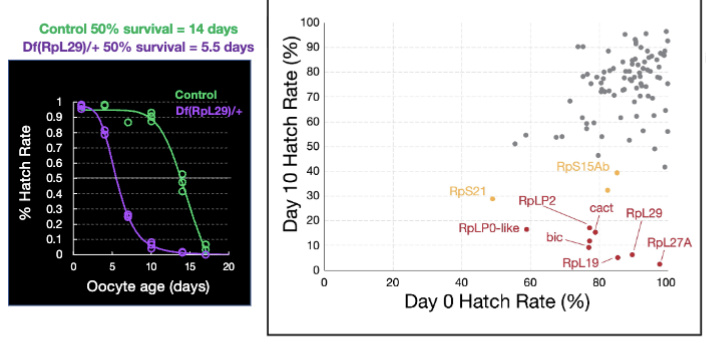
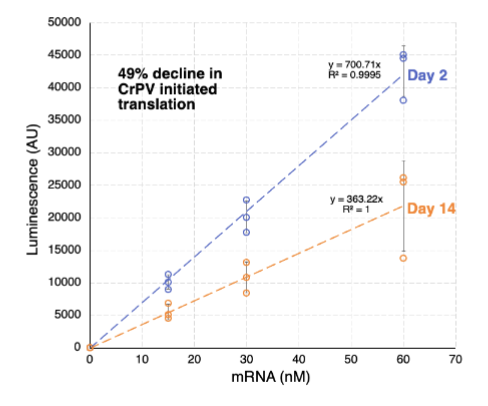
Are aged oocyte ribosomes less active?
yes
aged oocyte ribosomes are ~50% less active
Is translational control in oocytes/neurons linked to RNA granules?
yes
RNA granules are specialized structures that store and regulate mRNAs
In Drosophila oocytes, RNA granules help keep maternal mRNAs stable during developmental arrest
In Drosophila neurons, RNA granules store mRNAs for later use, e.g., until a synapse is activated
Questions based on previous slide
are RNA granule genes important for the survival of arrested oocytes?
Result of loss of RNA granule genes
loss of RNA granule genes leads to premature oocyte failure
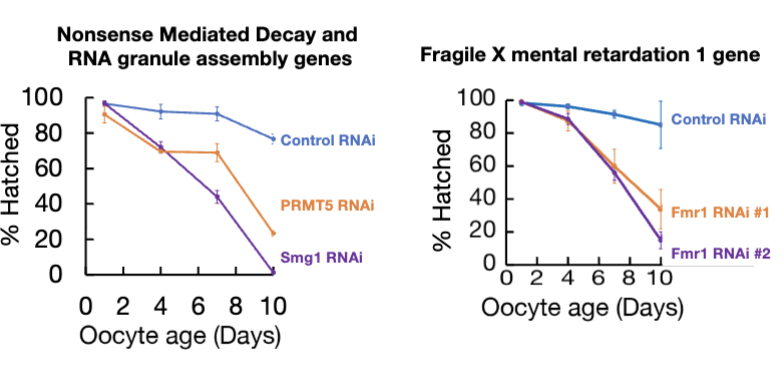
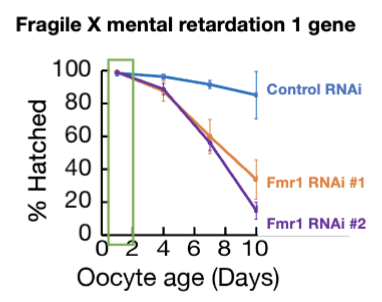
Fmr1 loss
Fmr1 loss specifically impairs arrested follicles
Fmr1’s WT function is critical for maintaining the viability and proper translation of mRNAs in oocyte during arrest, but less important in new oocytes (rescue in figure)
Fmr1 mutations is the single largest cause of two major human conditions: fragile X-associated primary ovarian insuffiency and fragile X syndrome (heritable cause to autism)
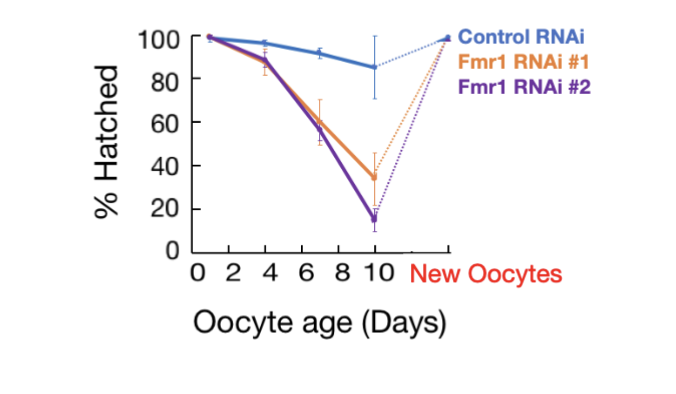
Are Fmr1 mutations pleiotropic?
yes, highly pleiotropic (single gene or genetic variant influences multiple phenotypic traits)
Fmr1 is an RNA binding, represses translation in vitro but unclear in vivo
Fmr1 binds transcripts from dozens of autism genes
cellular defects in Fmr1 KO animals:
enhanced long term depression
mRNA transport defects
compromised homeostatic plasticity
PI3K/mTOR overactivation
dendritic spine pruning/maturation defects
Hypothesis
Finding Fmr1’s primary function could aid drug development targeting all defects
Could studying Fmr1 RNAi oocytes before any development reveal Fmr1’s (singular) primary function?
experiment: perform RNA-seq and ribosome profiling comparing gene expression in WT vs Fmr1 RNAi oocytes
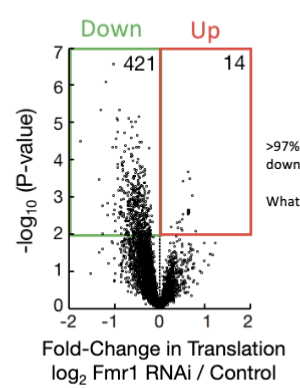
Loss of Fmr1
loss of Fmr1 leads to decreased ribosome abundance
As a result, >97% of mRNAs that depend on Fmr1 are translated less (downregulated at the translational level)
Interpretation: Fmr1 is particularly important for efficient translation of long mRNAs, which are otherwise hard to translate. Without Fmr1, these essential mRNAs are not properly translated, impairing oocyte or neuron function
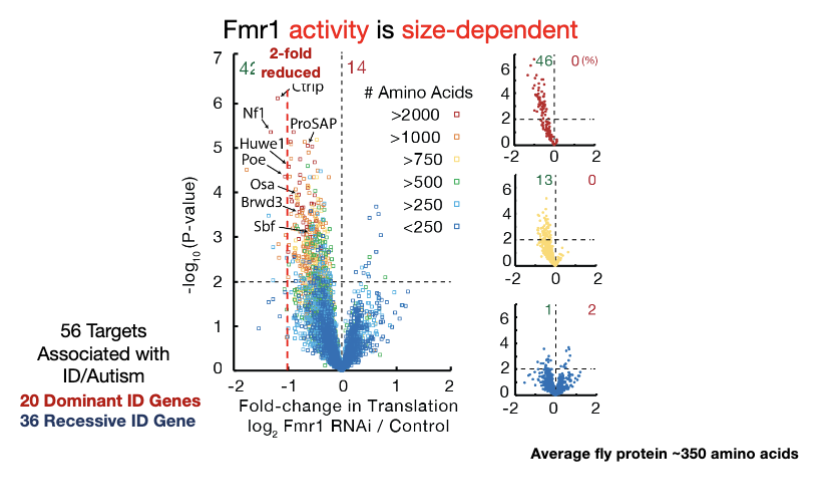
Is Fmr1 activity size dependent?
yes! extreme selectivity to help cell make large proteins
Targets: Fmr1 regulates 56 mRNAs that are linked to intellectual disability (ID) or autism.
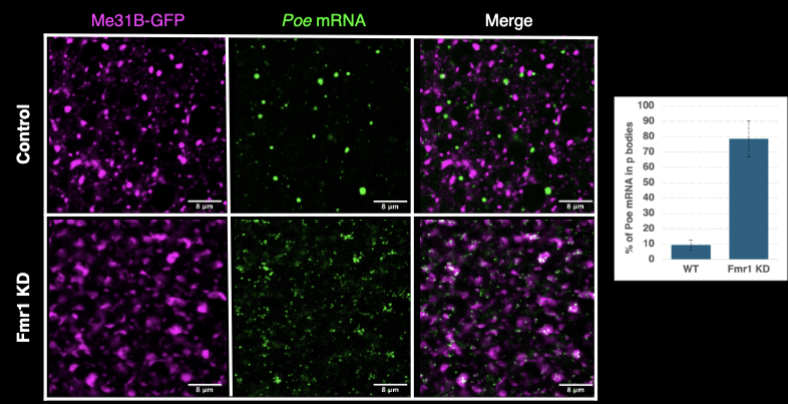
Poe
Fmr1 particles are potential sites of translation of the Fmr1 target Poe
Poe mRNA localizes to granules in Fmr1-dependent manner
without Fmr1, Poe mRNA fails to localize correctly
co-translational targeting: Poe mRNA is actively translated while being localized to the granules, suggesting these granules are sites of regulated translation
Puromycin blocks translation, poe mRNA may no longer localize properly to granules
because poe mRNA localization to Fmr1 granules is translation dependent, puromycin disrupts this co-translational targeting
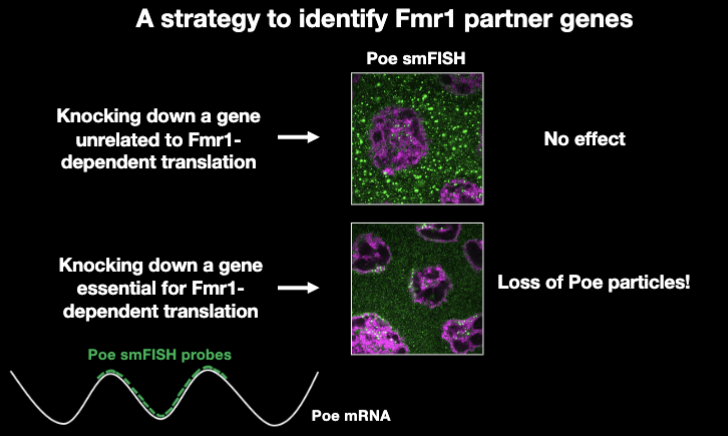
Binary Assay for Fmr1 Function
a strategy to identify Fmr1 partner genes
Core stress granules
Core stress granule proteins are the top putative FMRP-cofactors
When core stress granule proteins were disrupted, the amount of Fmr1 target mRNA in granules changed significantl
single-molecule FISH: allows visualization of individual mRNA molecules inside cellsMeasured the fraction of mRNA molecules located in granules/particles
Proper localization of Fmr1 target mRNAs depends on stress granule components.
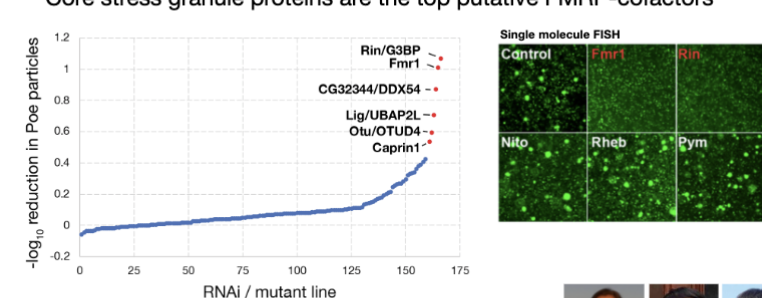
Genes required for Fmr1-dependent translation in flies are linked to human intellectual and neurodevelopmental disorders.
Genes required for Fmr1-dependent translation in flies are linked to human intellectual and neurodevelopmental disorders.
Fmr1 regulates translation through specific cofactors
Suggests that defects in: RNA granules, Translational control, Ribosome regulation are central mechanisms underlying intellectual disability and autism-related disorders.
What does FMRP block
FMRP blocks Poe mRNA from engaging repressive P bodies (from being silenced)
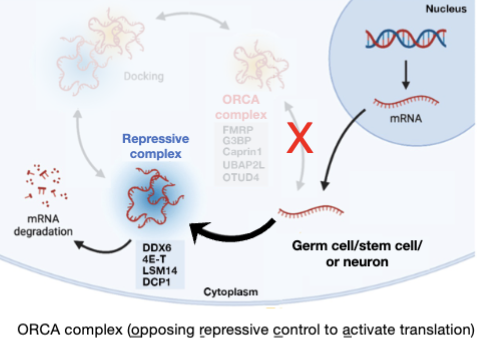
P-bodies
P bodies and stress granules are distinct RNA granules formed under stress
Under normal conditions in vivo proteins associated with P bodies and stress granules may promote translation repression or activation
FMRP’s de-repression activity enables the genome to encode very long transcripts
analogous to transcription control by mutually antagnozing DNA-associated (e.g. polycomb vs. thrithorax)
Ribosome profiling data show that oocyte translation declines during aging leading to a loss of production of key proteins
Ribosome profiling experiments show that Fmr1 is an activator rather than a repressor – the loss of translation likely underlies fragile X disorders
Using single molecule FISH to visualize Fmr1’s activity, we are teasing apart mechanisms of autism-relevant gene translation