Lecture I&J
1/37
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
38 Terms
Sec 12
ER integral membrane protein – functions as guanosine-exchange factor (GEF) that catalyzes the exchange of GDP for GTP on Sar1
enriched at ER exit sites
soluble in its GDP bound form
functions as Sar1 receptor
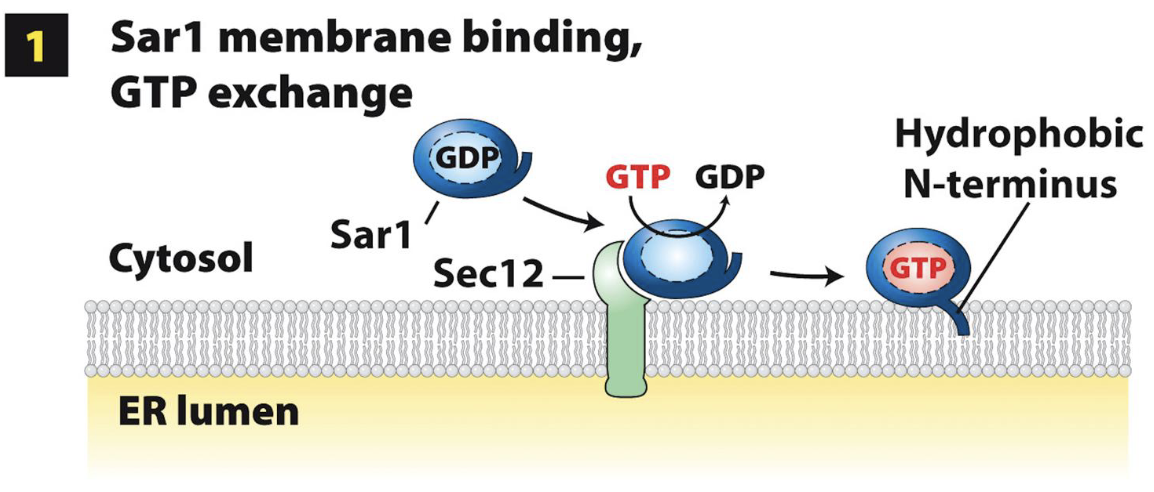
COPII-Coated Vesicle Assembly at ERES: Step 1
The soluble COPII component Sar1 (GTPase) is recruited from the cytoplasm to the ERES membrane (future site of vesicle budding) via binding to Sec12
Sar1 binding to GTP (Sar1-GTP) causes a conformational change – exposes the Sar1 hydrophobic N-terminus (serves as ERES membrane anchor)
Sar1-GTP integrated into cytoplasmic-facing leaflet of ERES membrane bilayer
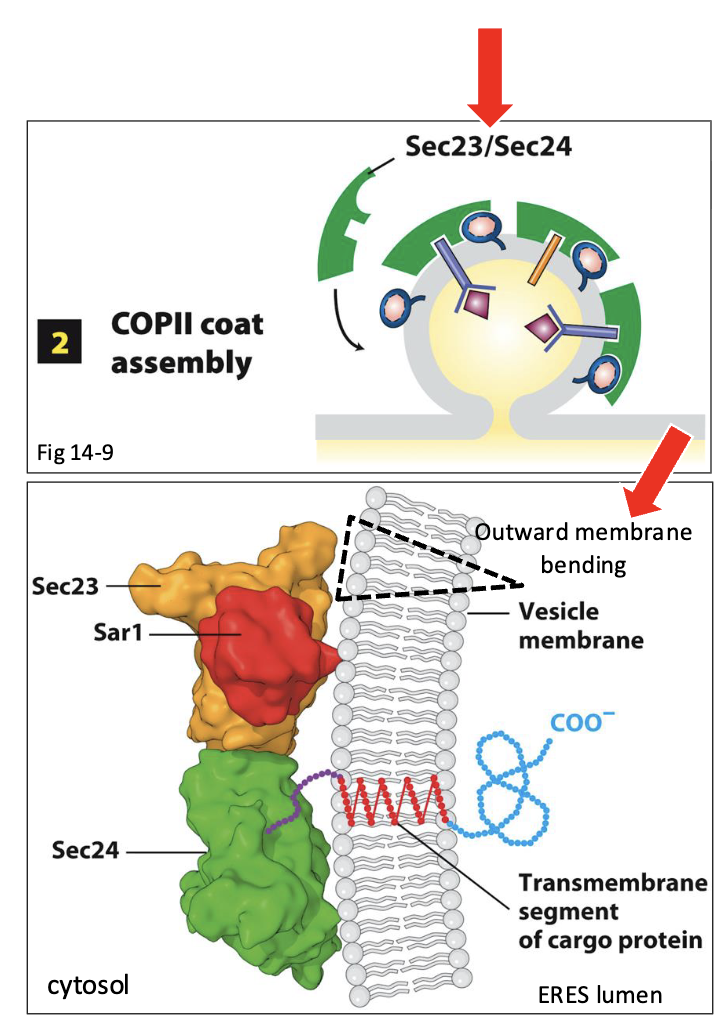
COPII-Coated Vesicle Assembly at ERES: Step 2 (till beginning of COPII ‘bud’ formation)
Sar1-GTP recruits several other COPII proteins from the cytosol to the ERES membrane surface
Sar1 initially recruits Sec23 and Sec24, soluble proteins that form ternary complex with Sar1 at the ERES membrane surface
Sec 23 binds to Sar1 but Sec24 is that mediator that gets material into vesicle
Sec23/24 act as structural scaffolding and promote the initial outward bending (towards cytosol) of the ERES membrane
This serves as the beginning of COPII vesicle ‘bud’ formation
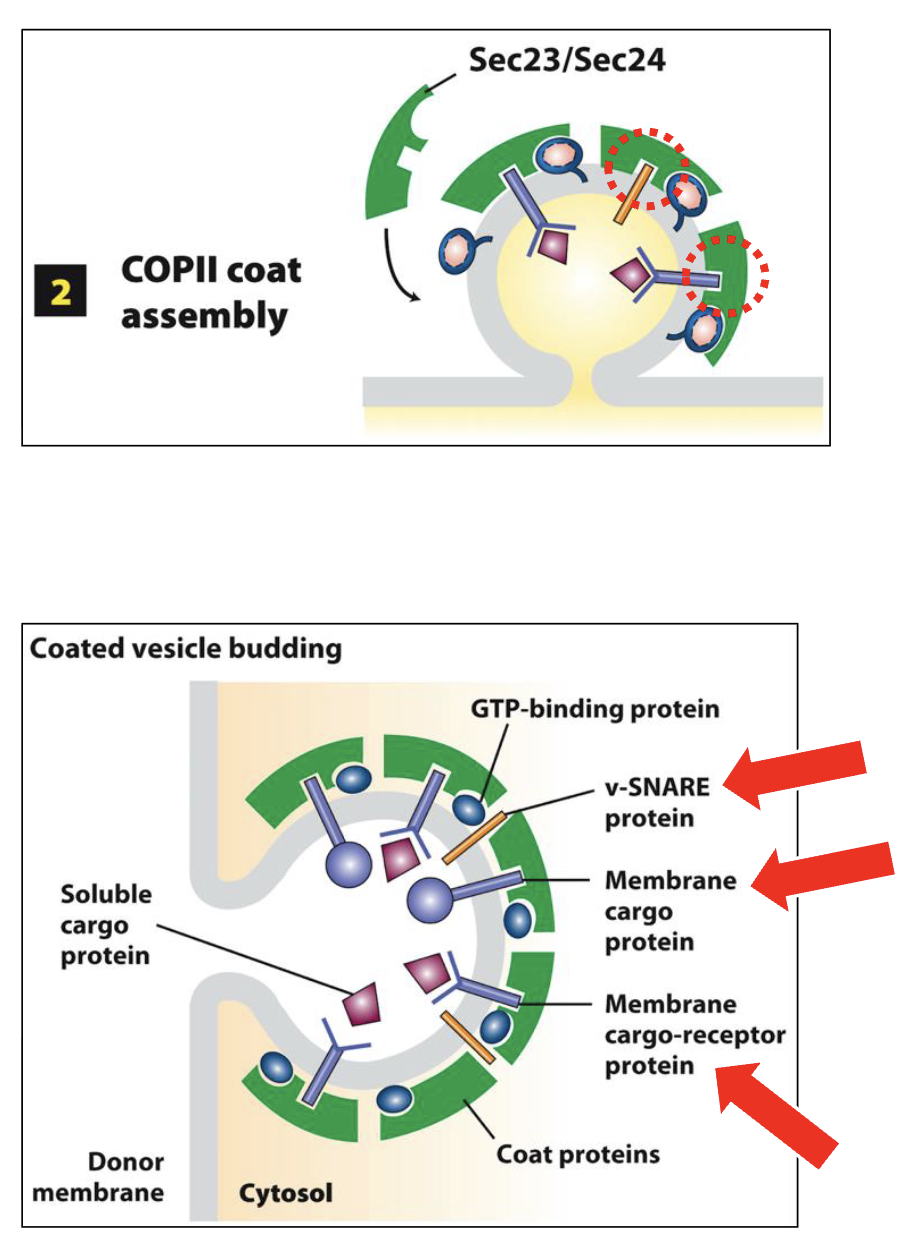
COPII-Coated Vesicle Assembly at ERES: Step 2 (vesicle protein selection)
Sec24 binds to cytoplasmic-facing domains to select several types of ER integral membrane proteins:
membrane cargo proteins - destined to exit ERES for the Golgi
membrane cargo-receptor proteins – bind via lumenal-facing domains to soluble lumenal ‘cargo’ proteins destined to exit ERES for Golgi
membrane trafficking proteins - required for the subsequent trafficking and docking of the vesicle with the proper acceptor membrane (Golgi) e.g., v-SNARES
ER export signal characteristics
di-acidic ER export signal (-Asp-X-Glu-)
located in the cytoplasmic- facing domains of membrane proteins in order to be detected and bound by Sec24
NOT found on ER resident protein
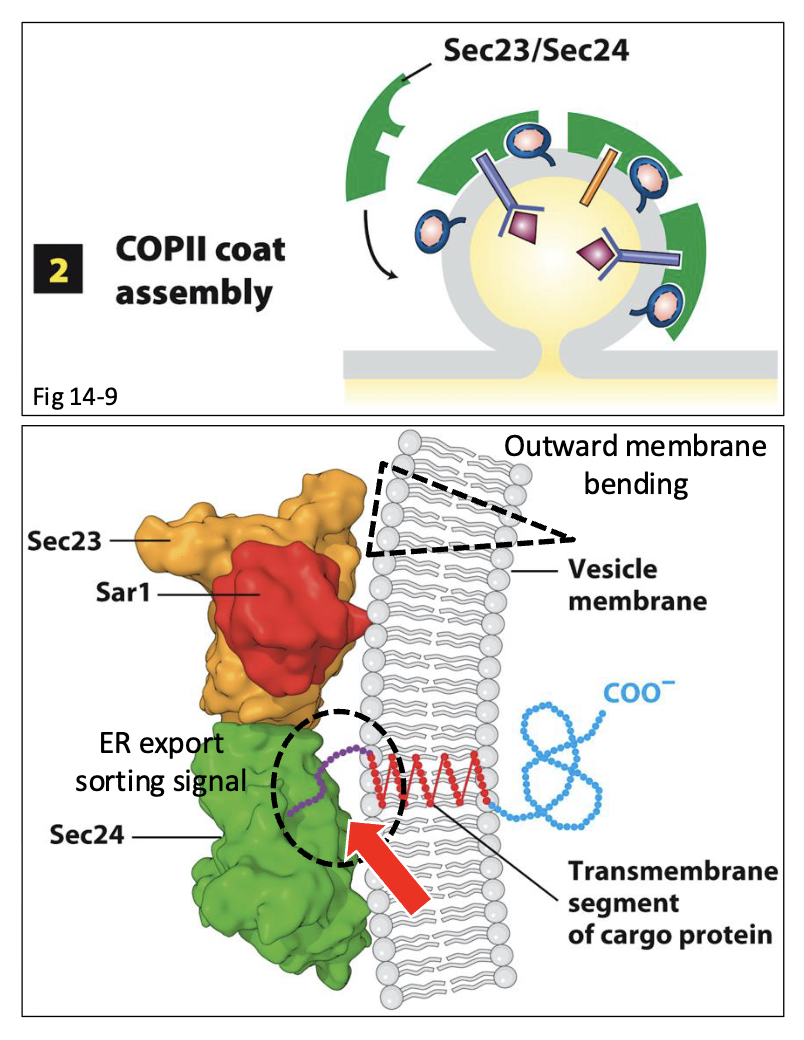
Purpose of ER export signal
Recognition and selection of vesicle membrane proteins by Sec24 is mediated by the ER export signal
Where are Sec24-bound proteins found
all Sec24-bound proteins (and soluble proteins bound by the membrane cargo-receptors) are concentrated $$within growing, COPII protein-coated vesicles
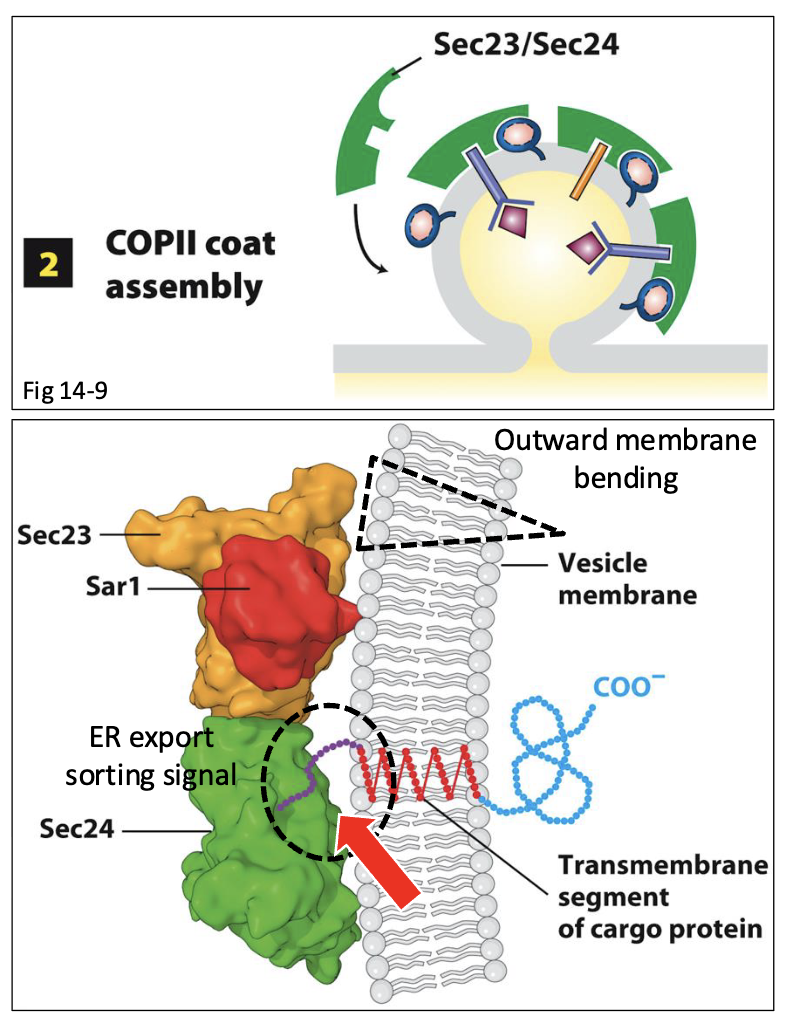
COPII-Coated Vesicle Assembly at ERES: Step 2 (till release of vesicle)
Sec23 and Sec24 recruit additional soluble COPII components from the cytoplasm to the surface of the growing vesicle
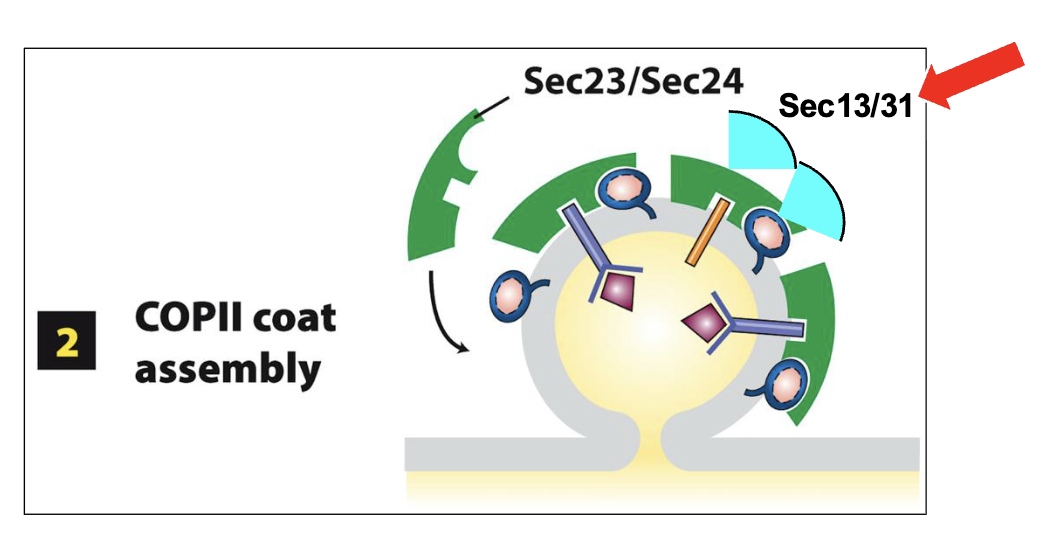
Sec13 and Sec31 assemble an outer cage-like lattice and act as structural scaffolding for the growing COPII vesicle bud
promotes additional outward bending of the ERES membrane
eventually... the nascent COPII vesicle is released (scission) from the ERES membrane into the cytosol
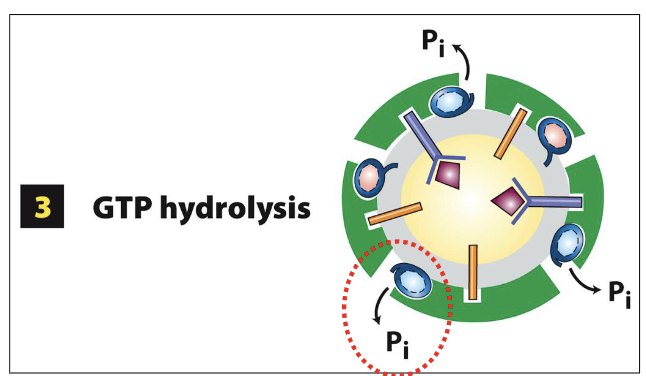
COPII-Coated Vesicle Assembly at ERES: Step 3
after the release of the COPII vesicle from the ERES membrane, Sec23 (GAP) promotes the hydrolysis of GTP in Sar1-GTP
Sar1-GTP converted to Sar1-GDP
COPII-Coated Vesicle Assembly at ERES: Step 4
GTP-hydrolysis in Sar1 results in the disassembly of the COPII protein coat
Sar1-GDP and all other soluble COPII proteins are released into the cytoplasm for additional rounds of COPII-coat assembly at ERES
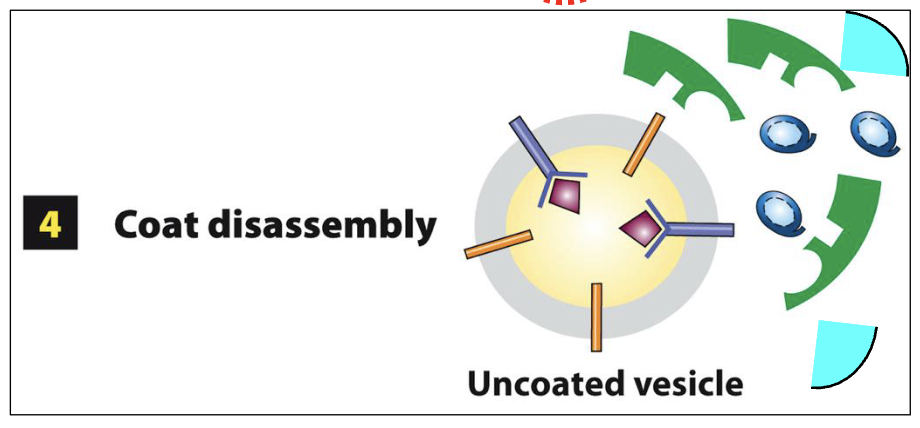
What does an ERES-derived vesicle contain
the soluble and membrane protein cargo AND the molecular machinery (e.g., v-SNAREs & Rabs) required for trafficking to and docking/fusion with the proper acceptor membrane
Where does EREs-derived vesicle go
vesicle traffics from the ERES to the cis-Golgi network (CGN)
incoming nascent vesicles fuse with one another to form the CGN
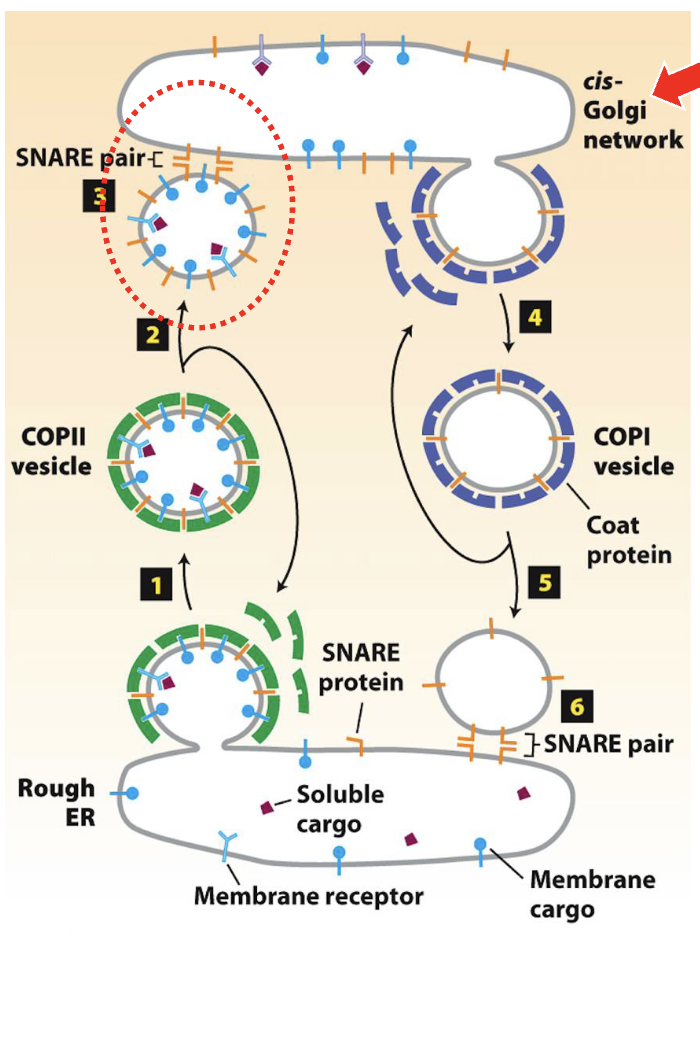
What does the cis-Golgi network consists of and where is it located
vesicle traffics from the ERES to the cis-Golgi network (CGN)
incoming nascent vesicles fuse with one another to form the CGN
Rab proteins and function
a large family of lipid-membrane anchored, GTP-binding proteins associated with all transport vesicle
key regulators of vesicle trafficking and fusion
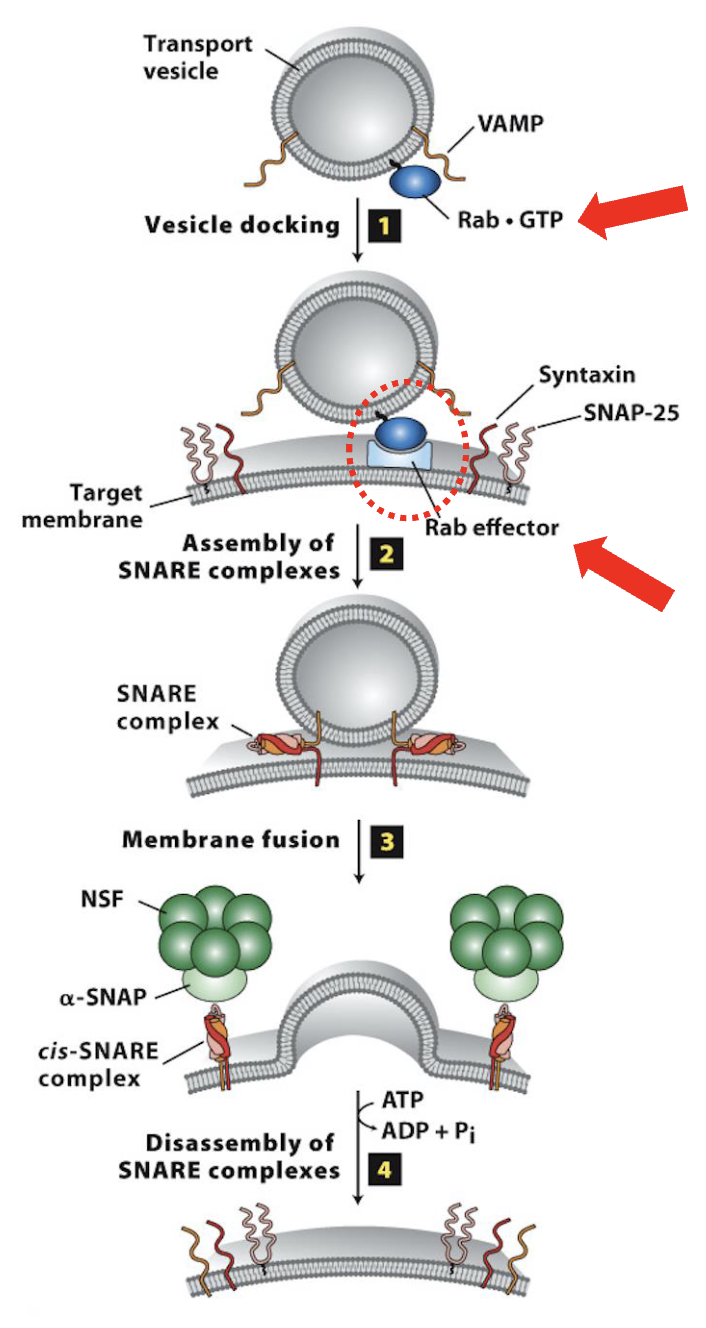
Vesicle Docking and Fusion at the CGN: Step 1
Vesicle Docking with target membrane (mediated by Rab)
specific activated vesicle Rabs (Rab-GTP) bind to specific Rab effector(s) proteins on the target membrane (e.g., CGN)
Rab effectors (Rab-binding proteins) also include cytoskeletal molecular motor proteins that enable the vesicle to associate with and move on cytoskeleton (kinesin and dynein)
Rabs and Rab effector binding conveys vesicle targeting specificity and docking
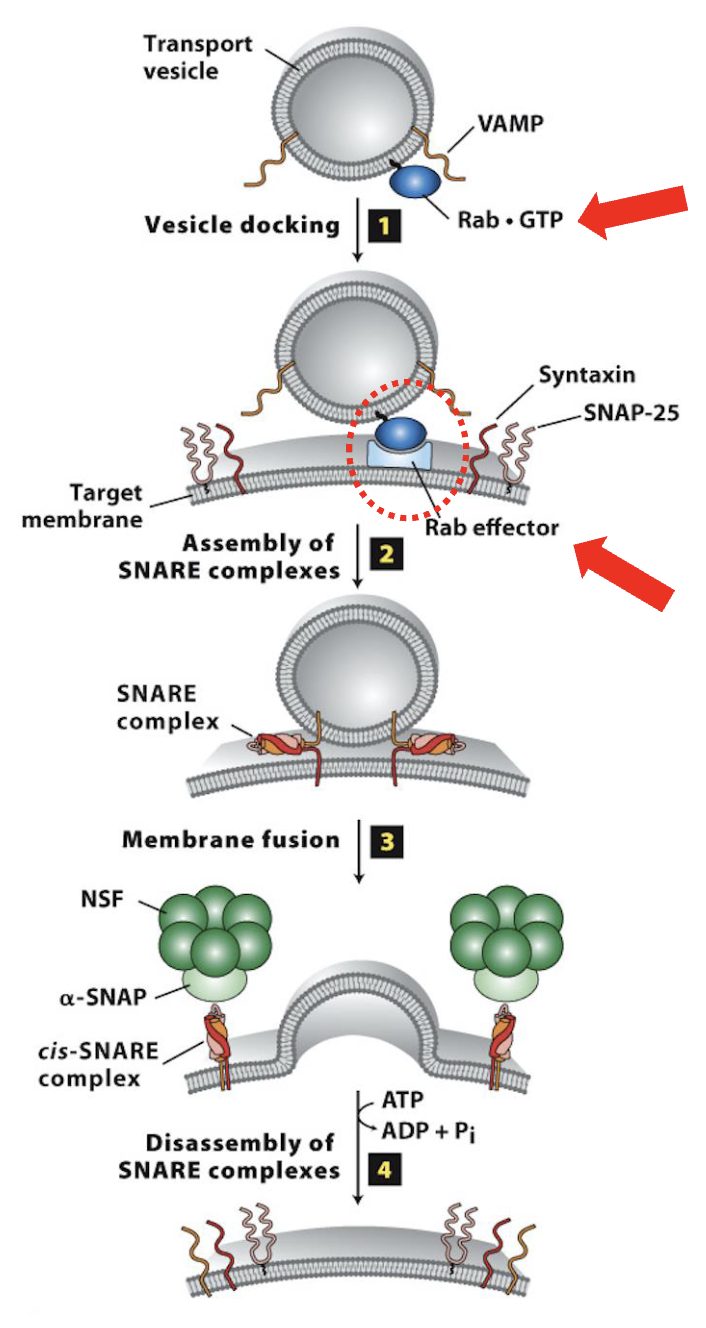
SNARE proteins and purpose
large family of integral (most) membrane proteins located on all transport vesicles AND all target membranes
unique SNAREs are associated with different membranes and provide additional vesicle targeting specificity (in addition to the Rab & Rab effectors)
What do all SNARE proteins possess
all SNARE proteins possess SNARE motif, a cytoplasmic- facing, coiled-coil domain in both v- & t-SNAREs that extend from the vesicle/target membrane surface
Two main classes of SNARE proteins and where they’re found:
v-SNAREs: found on transport vesicle (v) membranes and are incorporated into vesicle membrane at the site of budding
e.g., at the ERESs, v-SNAREs are specifically incorporated via Sec24 binding to a di-acidic ER export signal in cytoplasmic-facing domain of the v-SNARE into Golgi-destined vesicle membranes
t-SNARES – found on target (t) ‘acceptor’ membranes
e.g., at the cis-Golgi network
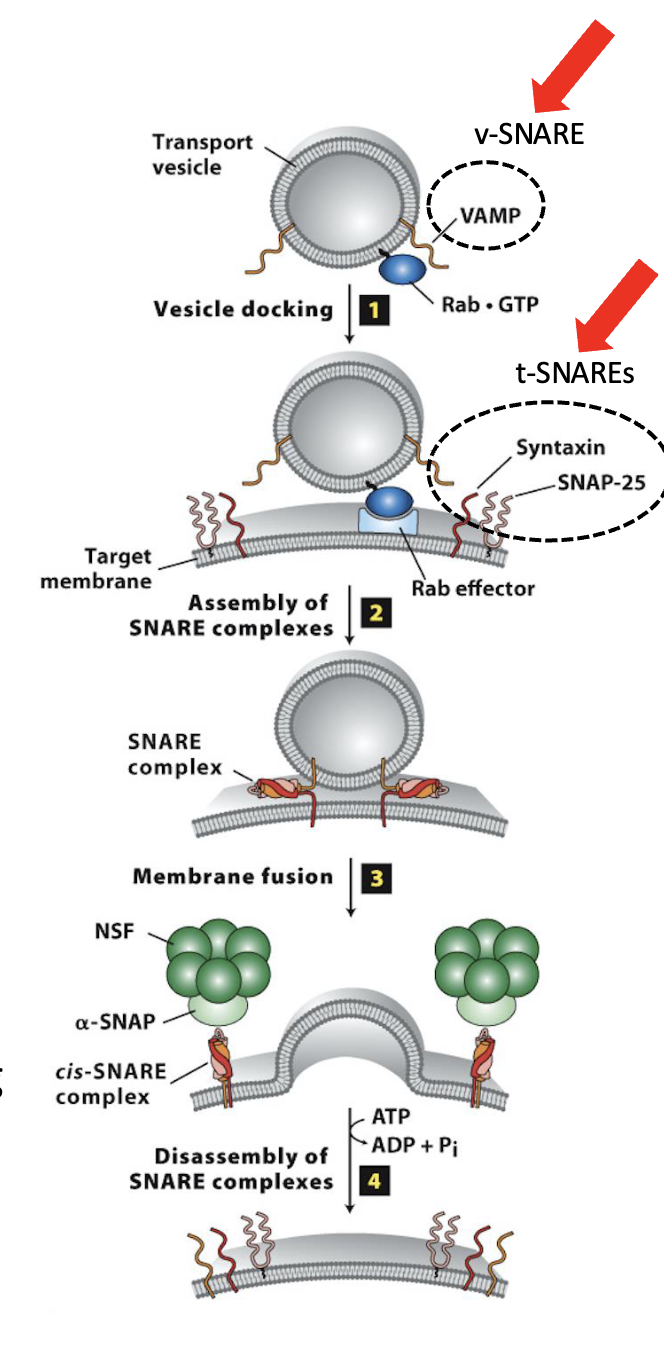
Vesicle Docking and Fusion at the CGN: Step 2
Assembly of the SNARE complexes
After docking of the vesicle, SNARE protein interactions brings membranes close together for fusion
SNARE motifs in both v-SNAREs and t-SNAREs interact to form stable SNARE complex
pulls vesicle and target membranes close together
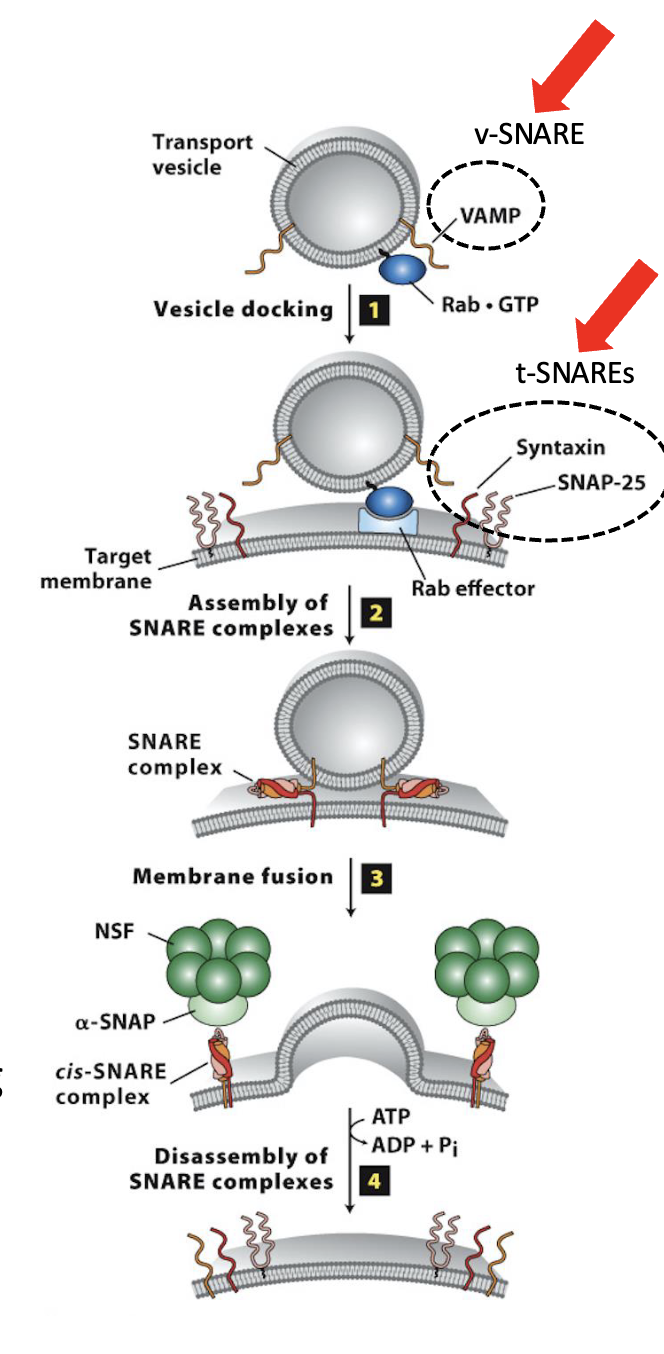
Vesicle Docking and Fusion at the CGN: Step 3
Membrane fusion
SNARE complex formation leads to membrane fusion (biophysical mechanism for membrane fusion is not well understood)
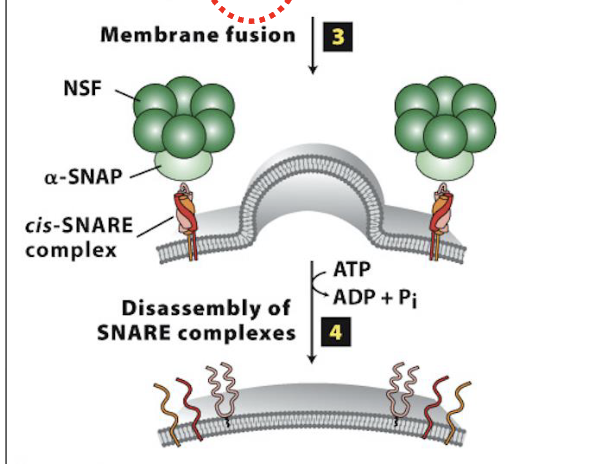
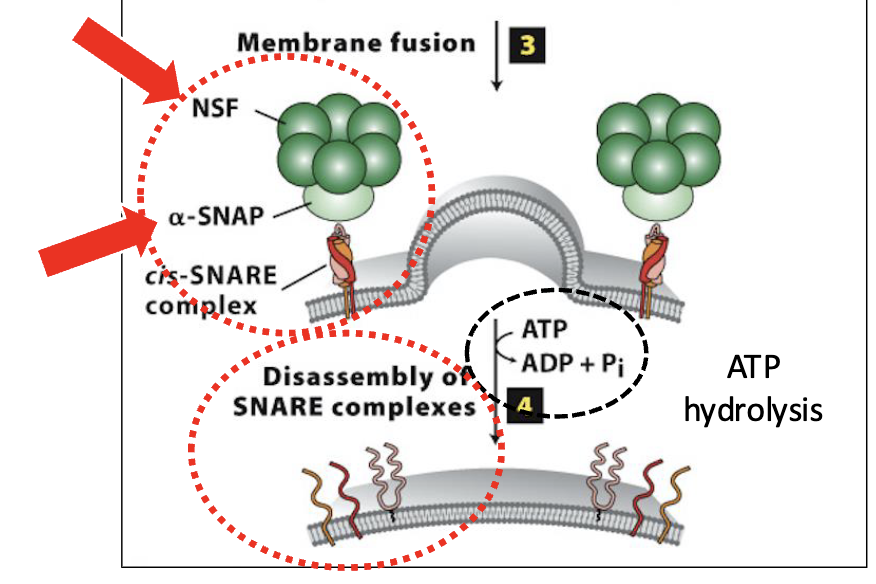
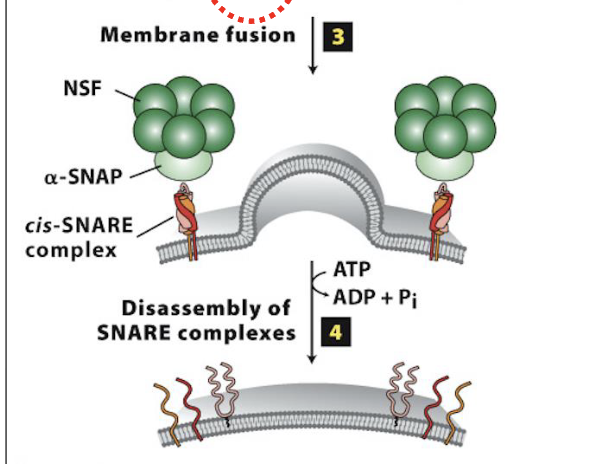
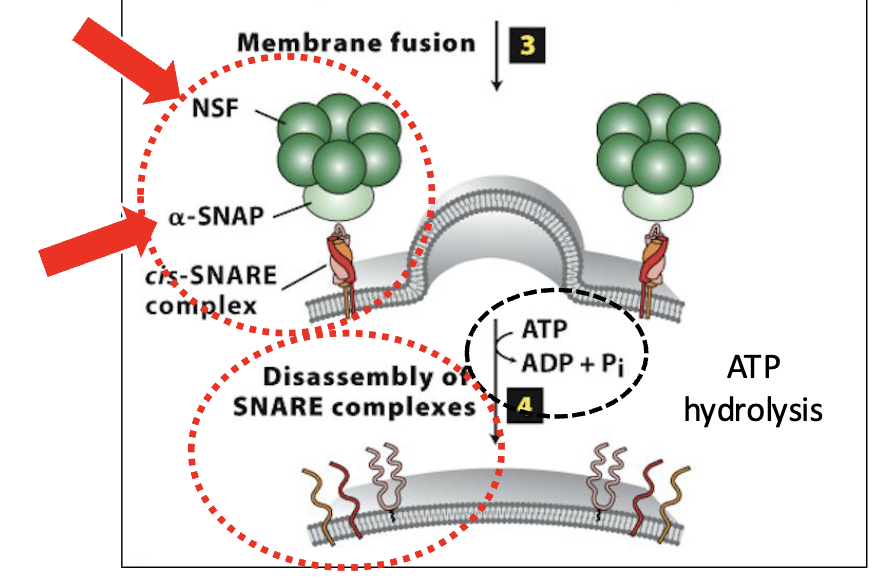
Vesicle Docking and Fusion at the CGN: Step 4
Disassembly of the SNARE complexes
after vesicle/target membrane fusion, SNARE complexes (and Rab/Rab-effector complexes) dissociate and are recycled for additional fusion events
disassembly of the SNARE complex is mediated by NSF & SNAP, soluble cytosolic proteins
they bind to SNARE complexes and unwind (via ATP hydrolysis) the SNARE domains linking v/t-SNAREs
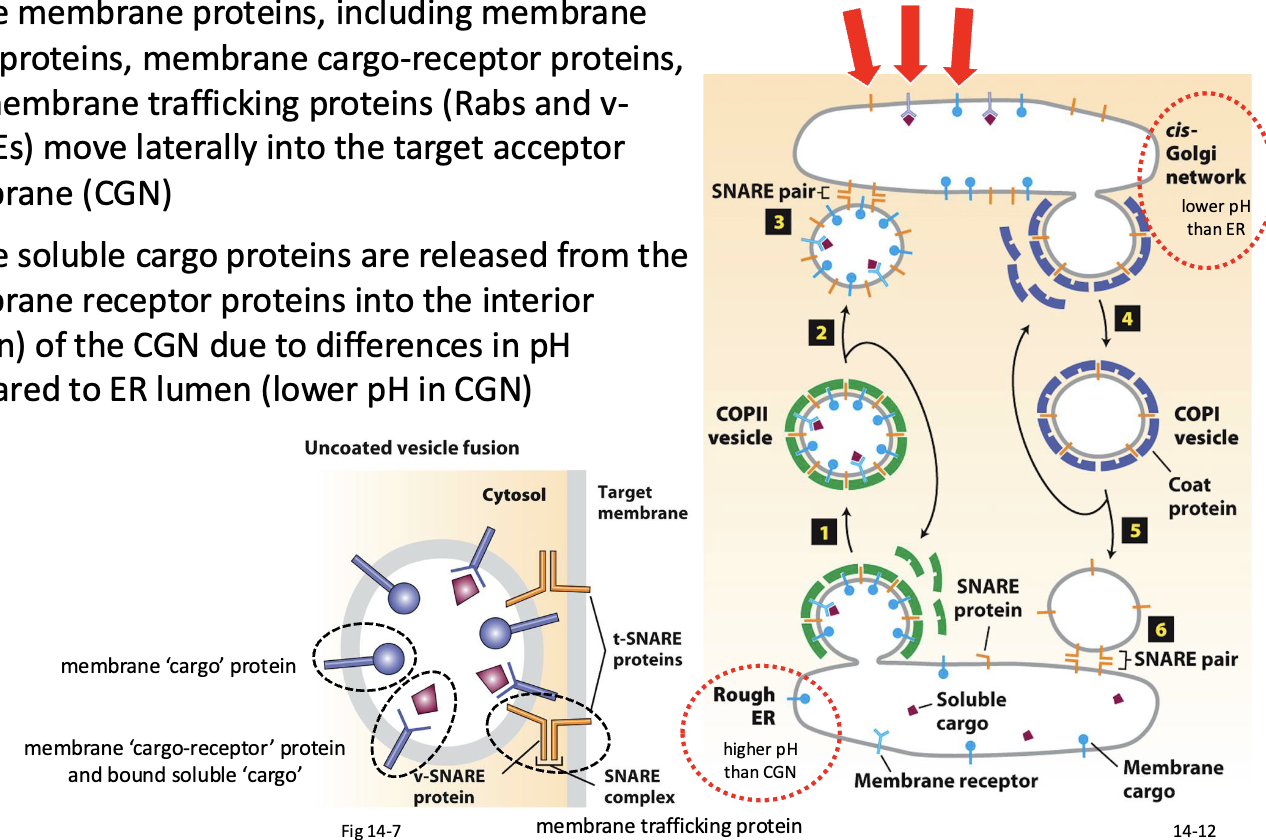
Vesicle-target membrane fusion results in…
vesicle membrane proteins, including membrane cargo proteins, membrane cargo-receptor proteins, and membrane trafficking proteins (Rabs and v- SNAREs) move laterally into the target acceptor membrane (CGN)
vesicle soluble cargo proteins are released from the membrane receptor proteins into the interior (lumen) of the CGN due to differences in pH compared to ER lumen (lower pH in CGN)
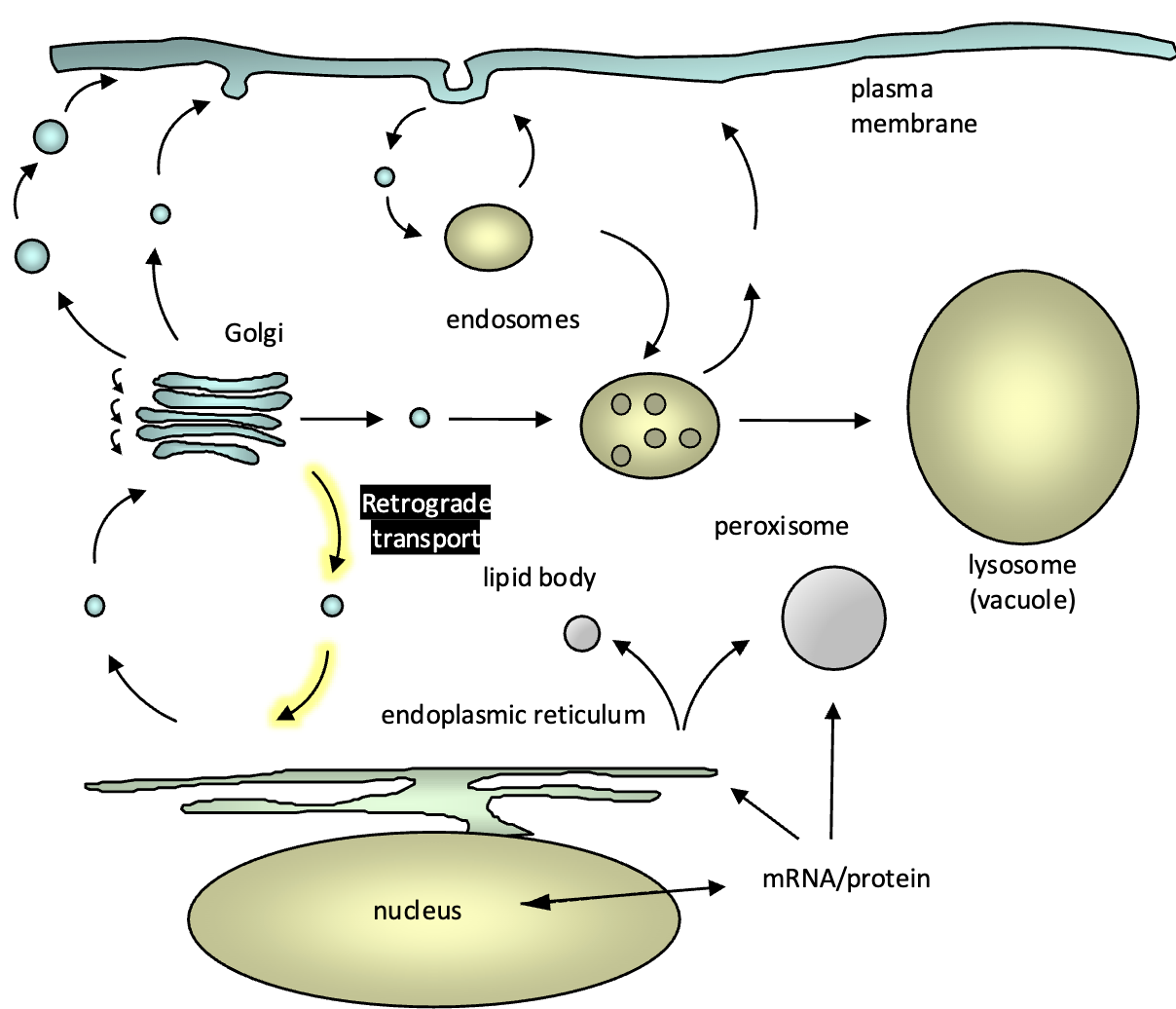
What is the fate of vesicle-specific proteins or proteins that ‘escape’ from the ER (ex. v-SNAREs, BiP, calnexin, cargo protein receptors, Rab)
most ER resident proteins are retained in the ER by exclusion from budding COPII vesicles at ERESs (they do not posses the di- acidic ER export sorting signal)
escaped ER resident proteins are returned from the CGN back to the ER (retrograde transport) by specific ER retrieval sorting signals
What sequence do most resident soluble ER proteins possess
most resident soluble ER proteins (e.g, BiP) possess a C-terminal KDEL sequence which serves as an ER retrieval sorting signal
What does the KDEL receptor have that helps incorporate it into a COPII vesicle
KDEL receptor also has the di-acidic ER export sorting signal (-Asp-X-Glu-) that is recognized by Sec24 for incorporation into a COPII vesicle
Steps of escape soluble ER protein in CGN-to-ER retrograde transport
escaped soluble ER proteins in the CGN lumen are recognized by the KDEL receptor, an integral transmembrane protein with a lumenal-facing domain that binds to the KDEL sequence of escaped soluble ER proteins in the CGN lumen
cytoplasmic-facing domain of the KDEL receptor contains a KKxx sequence and is recognized by COPI coat proteins
COPI coat proteins (like COPII at ERES) mediate the formation of transport vesicle at the CGN (and elsewhere in the Golgi complex [see later])
after COPI-coat disassembly, vesicles targets, docks, and fuses with the ER
soluble ER protein-KDEL receptor complexes traffic back (retrograde transport) to ER
KDEL receptor releases the resident soluble ER protein into the ER lumen
KDEL receptor binding is sensitive to the higher pH inthe ER lumen compared to the CGN lumen – results in a conformation change and release of the soluble ER protein back into the lumen of the ER
Empty KDEL receptor in the ER membrane returns to the CGN via COPII-coated ERES vesicles
![<ol><li><p>escaped soluble ER proteins in the CGN lumen are recognized by the KDEL receptor, an integral transmembrane protein with a lumenal-facing domain that binds to the KDEL sequence of escaped soluble ER proteins in the CGN lumen</p></li><li><p>cytoplasmic-facing domain of the KDEL receptor contains a KKxx sequence and is recognized by COPI coat proteins</p></li><li><p>COPI coat proteins (like COPII at ERES) mediate the formation of transport vesicle at the CGN (and elsewhere in the Golgi complex [see later])</p></li><li><p>after COPI-coat disassembly, vesicles targets, docks, and fuses with the ER</p></li><li><p>soluble ER protein-KDEL receptor complexes traffic back (retrograde transport) to ER</p></li><li><p>KDEL receptor releases the resident soluble ER protein into the ER lumen </p></li><li><p>KDEL receptor binding is sensitive to the higher pH inthe ER lumen compared to the CGN lumen – results in a conformation change and release of the soluble ER protein back into the lumen of the ER</p></li><li><p>Empty KDEL receptor in the ER membrane returns to the CGN via COPII-coated ERES vesicles</p></li></ol><p></p>](https://assets.knowt.com/user-attachments/b4bda6f7-c521-445d-9ef9-d5cc413421e4.png)
What sequence do most resident ER membrane proteins possess
most resident ER membrane proteins (e.g., calnexin) possess a cytoplasmic-facing, C-terminal sequence of KKxx which serves as a ER retrieval sorting signal
What recognizes KKxx sequence on escaped ER membrane proteins?
COPI proteins at the CGN
COPI-coated vesicles bud off the CGN and target/dock/fuse with the ER membrane
What proteins that have both KKxx and KDEL signal sequence
proteins that cycle between the ER and the CGN, including KDEL receptor, membrane cargo-receptor proteins, ER membrane trafficking proteins (e.g, v-SNAREs) also contain the KKxx ER retrieval signal
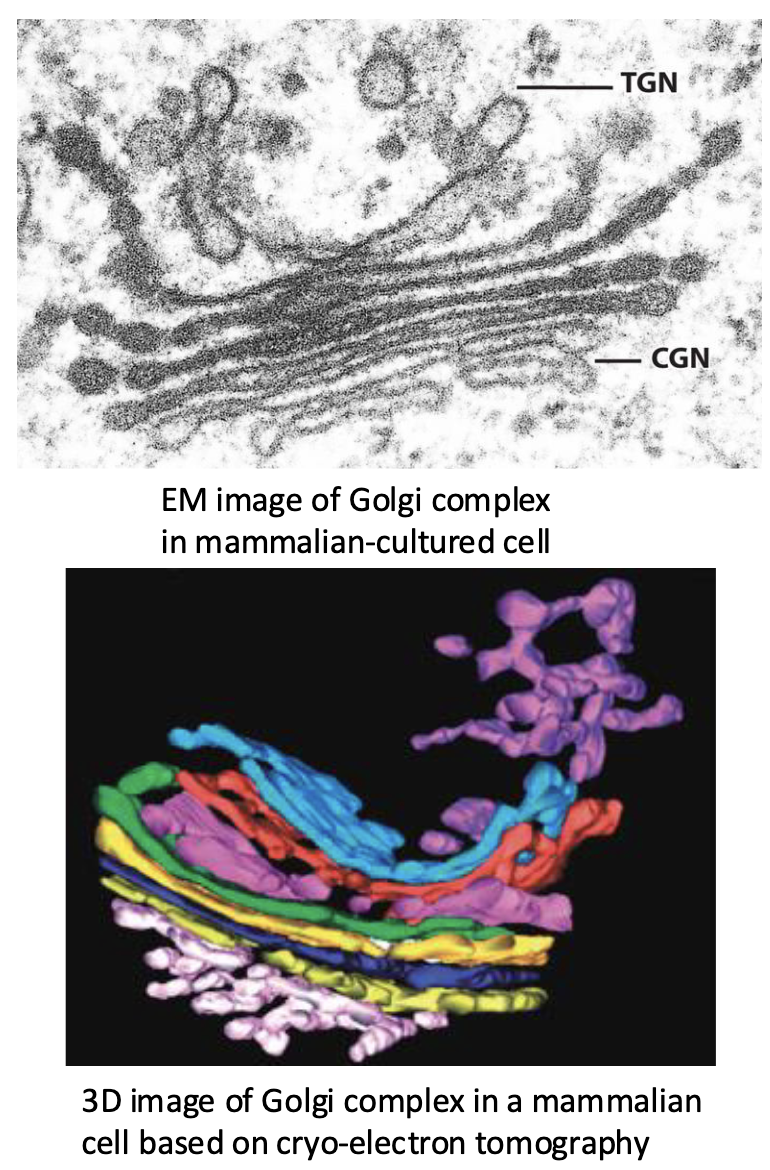
Characteristics of Golgi Complex
unique morphology – ‘complex’ or ‘stack’ of flattened, membrane-bound cisternae (sacs) with dilated edges and numerous associated tubules and vesicles
Possesses several subcompartments that results in the complex (stack) having distinct polarity: both structurally and functionally
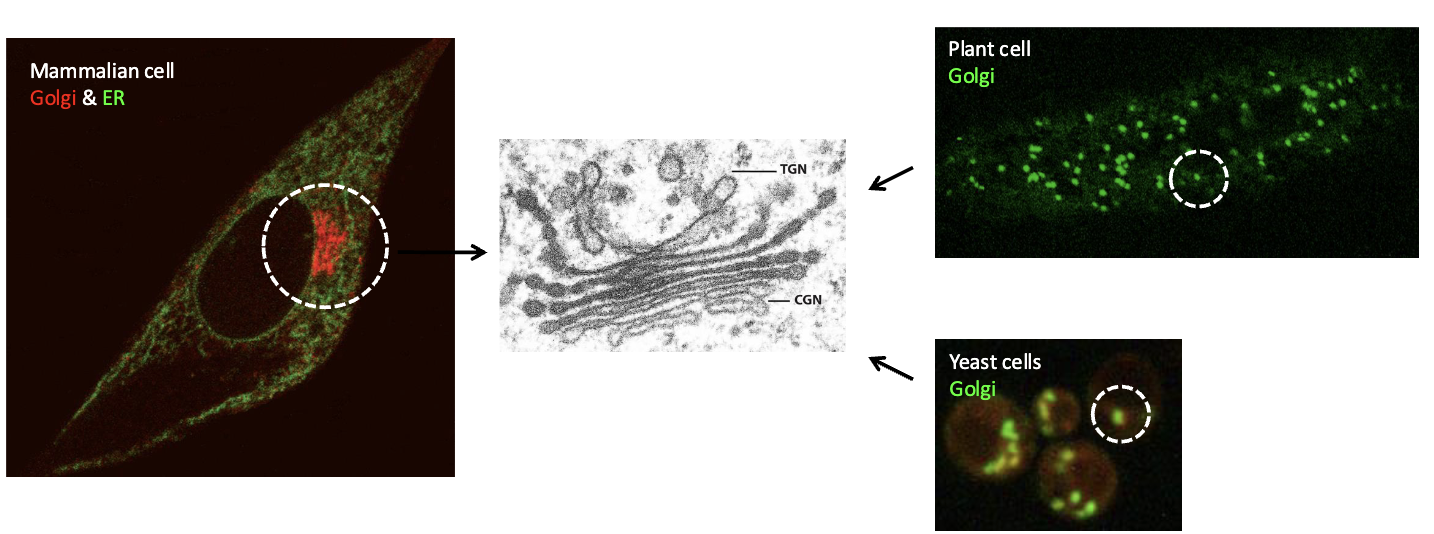
Number of Golgi complexes in cells
The number and distribution of Golgi complexes vary between different cell types
mammalian cell: typically contains one (large) Golgi complex located near the center of the cell
Plant/yeast cells: typically contain several Golgi complexes located throughout the cell
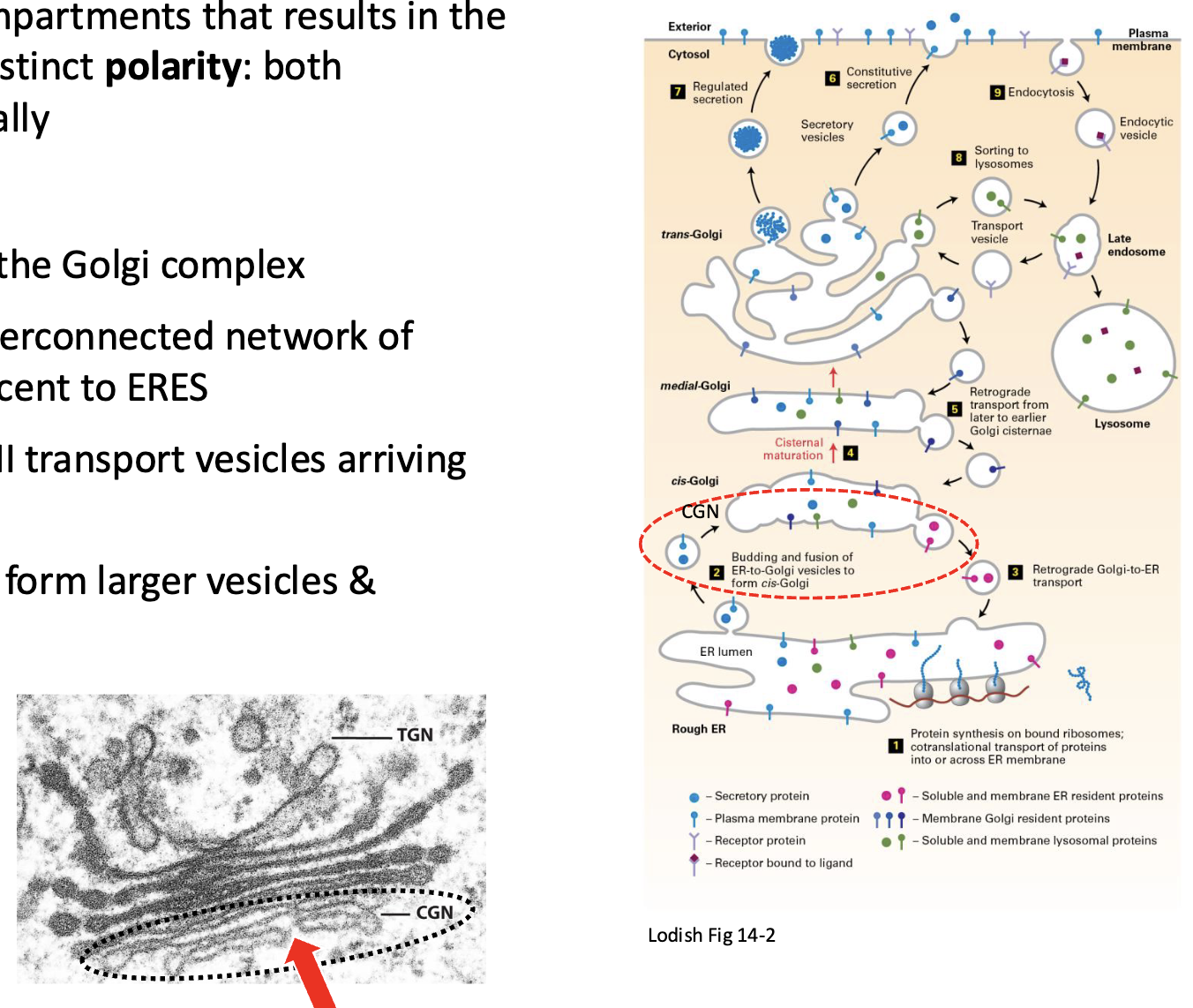
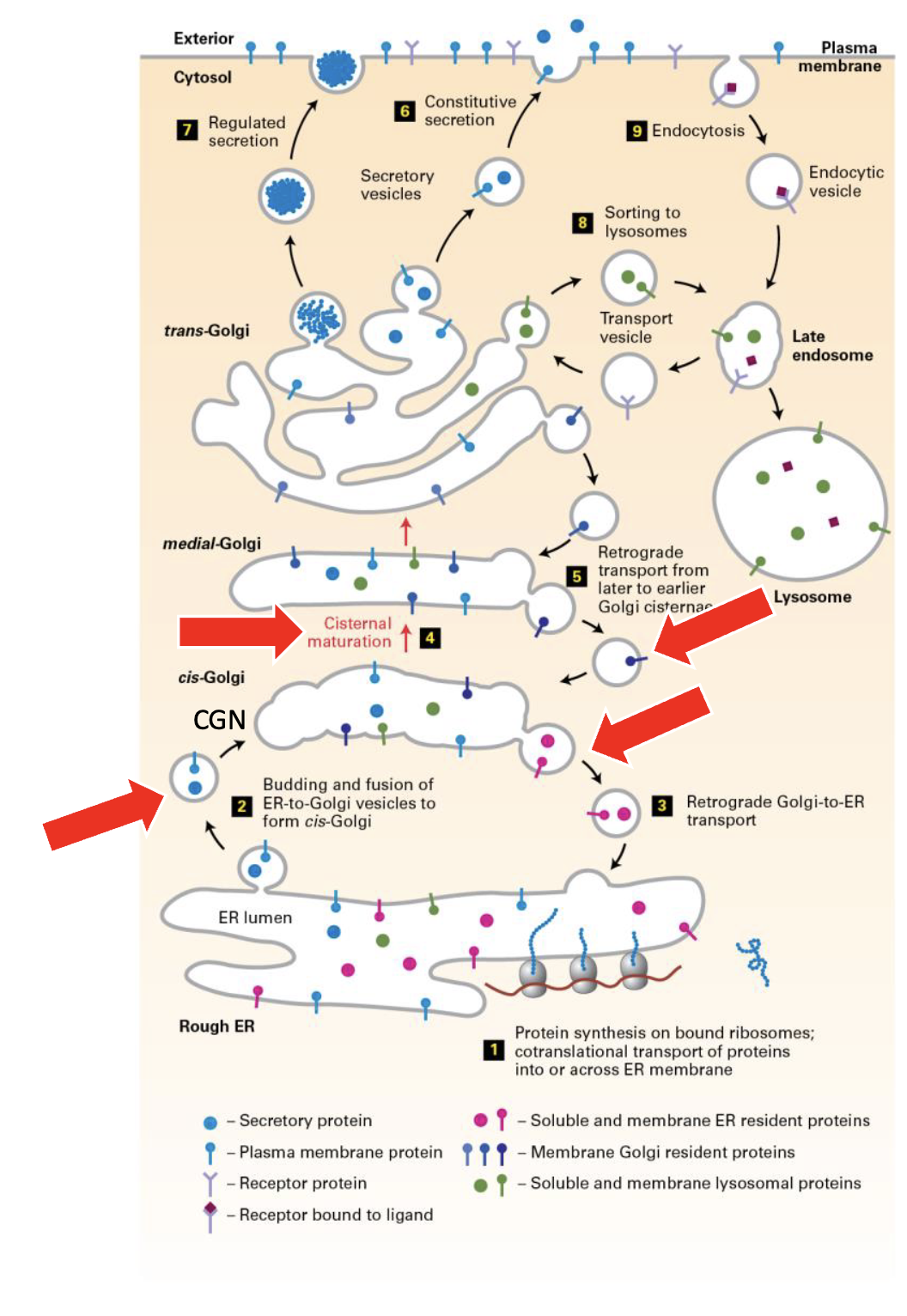
Formation of the whole Golgi network
These complexes are constantly in flux and forming in the order:
ER→cis-Golgi→Golgi Cisternae→Medial→trans→vesicles
cis, medial, and trans have different enzymes which move along when transitioning via COPI vesicles
cis-Golgi network (CGN) characteristics
located at the cis face of the Golgi complex
consists of a complex, interconnected network of tubules and vesicles adjacent to ERES
incoming vesicles fuse to form larger vesicles & interconnected tubules
Purpose of CGN
serves as a destination and sorting station of COPII vesicles coming from ERESs to the CGN (anterograde transport)
the site of COPI vesicle assembly for transport back from the CGN to ER (retrograde transport)
forward (anterograde) transport as the CGN maturesinto the next subcompartment of the Golgi complex (CGN → cis cisternae)
destination of COPI vesicles moving back (retrogradetransport) from the upstream subcompartment of the Golgi complex (cis cisternae → CGN)
Golgi Cisternae characteristics
series of three or more large, flattened cisternae that comprise the majority of the Golgi structure
3 main sections: cis, medial and trans cisternae
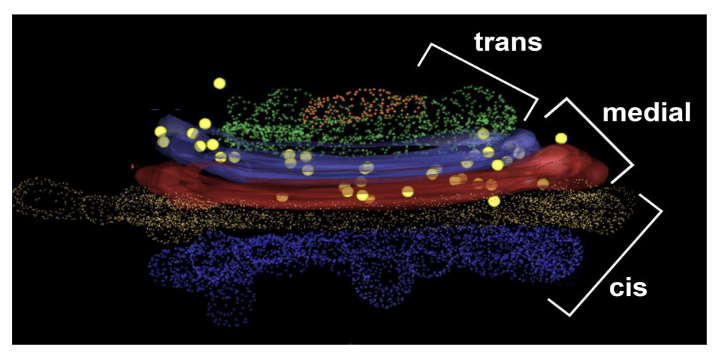
Golgi cisternae function
Cisternae are the sites of Golgi metabolism
synthesis of complex polysaccharides used for plant cell walls
modification (glycosylation) of proteins/lipids
phosphorylation of mannose units in lysosomal-destined proteins
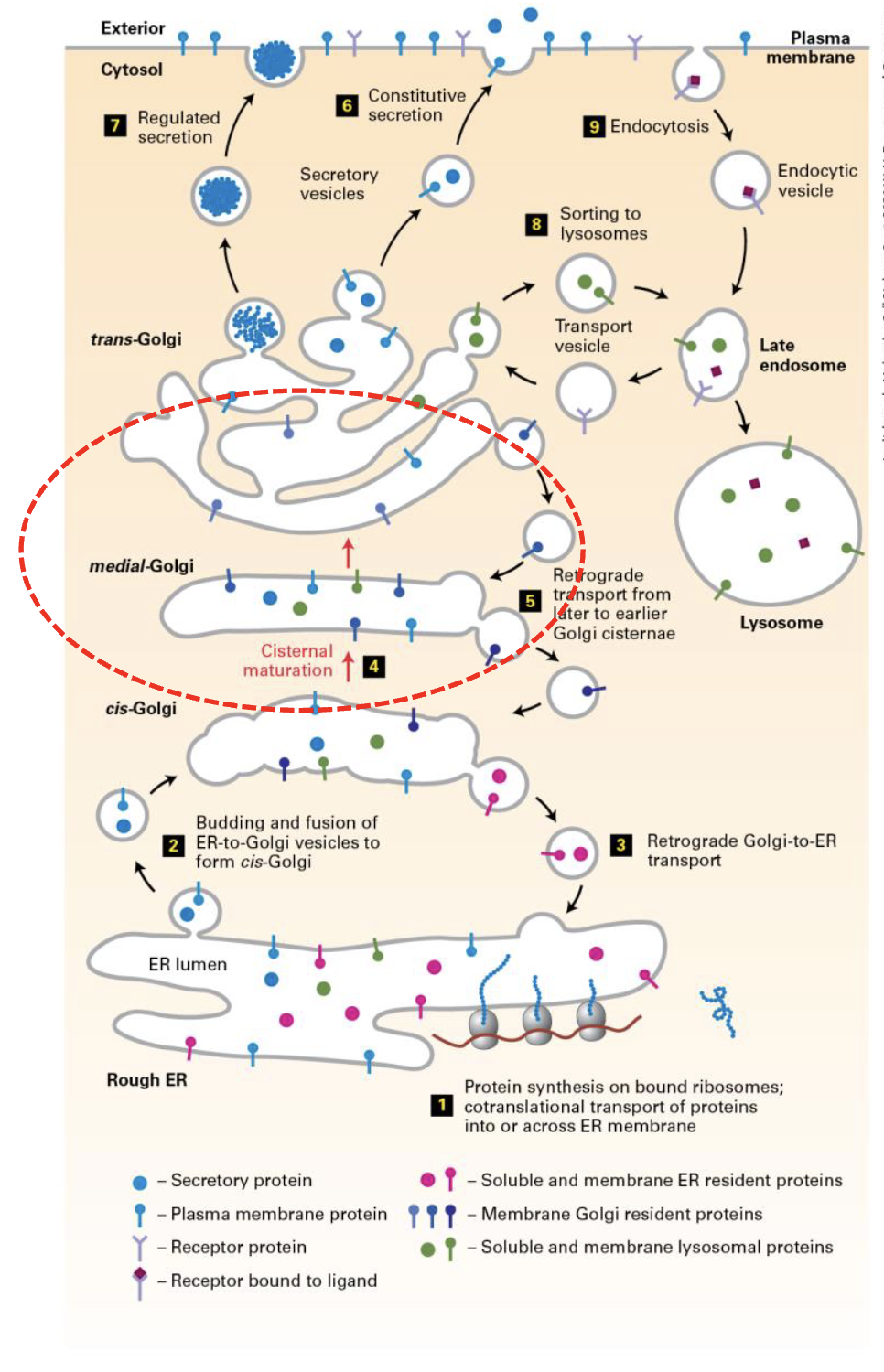
trans-Golgi network (TGN) characteristics
interconnected network of tubules and vesicles (similar to CGN)
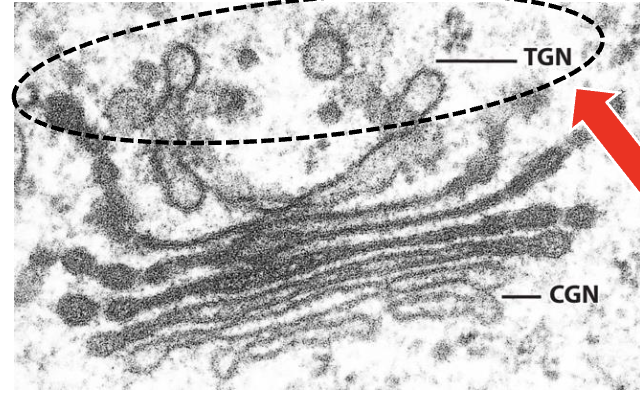
Purpose of trans-Golgi network (TGN)
serves as a sorting station ( CGN)
Mediates the forward transport once the previous subcompartment of the Golgi complex matures into the TGN (trans cisternae → TGN)
site of clathrin coat vesicle assembly for transport from the TGN to endosomes
site of secretory vesicle assembly for transport forward
(anterograde) to the plasma membrane and eventual
secretion into the extracellular space
site of COPI vesicle assembly for transport back
(retrograde) to the Golgi trans cisternae