Cells to Organisms
1/194
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
195 Terms
What are the 3 main domains of life
Eukarya, Archea, and Bacteria
What are the differences between prokaryotes and eukaryotes
Eukaryotes
Larger
Compartmentalisation
Nucleus
Prokaryotes
Smaller
No compartmentalisation
No nucleus
What are the 3 main forms of prokaryote
Coccus
Rod/Bacillus
Sprillum/Spirochete
Describe prokaryote cell structure
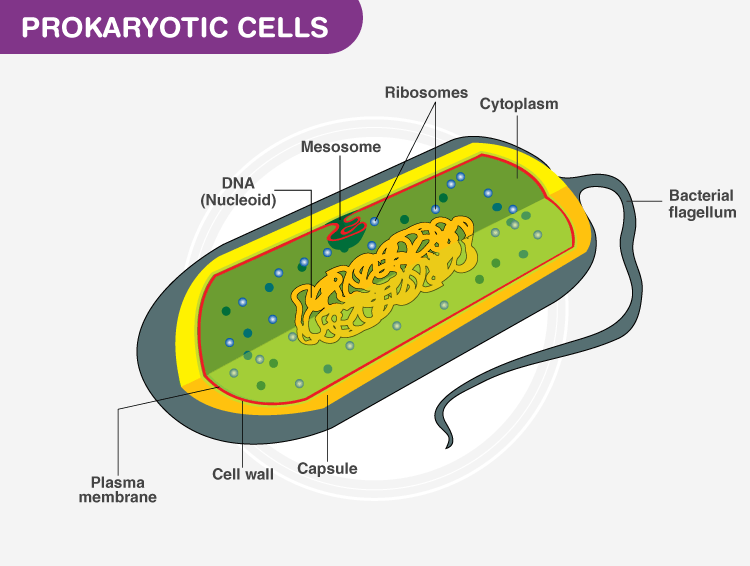
What is the difference between gram positive and gram negative
Named after the ability to take up a gram stain
Gram positive- turns purple
Gram negative- turns pink
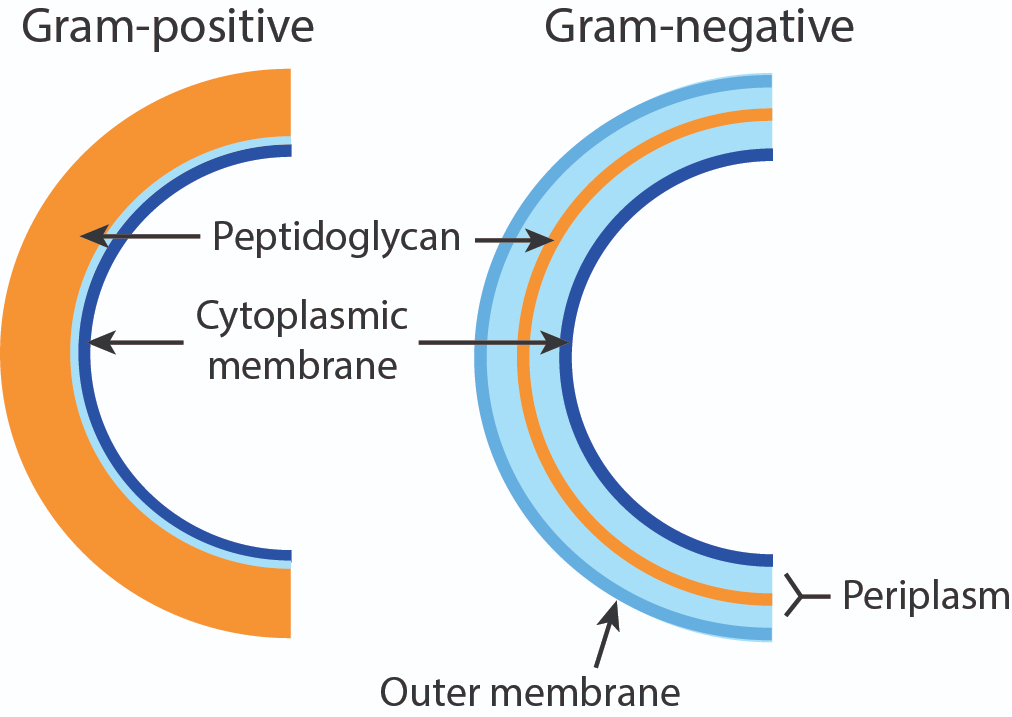
How does the structure of the outer membrane differ in gram negative and gram positive bacteria.
The gram stain wont stick to the gram -ve due to the outer membrane. Peptidoglycan is made out of alternating sugars and amino acids.
What are some functions of the prokaryotic membrane
Permeability barrier
Protein anchor
Energy conservation
Pumps control permeability
Membrane uses proton motive force for ATP production
How is genetic material organised in prokaryotes
Prokaryote chromasomes are circular, supercoiled and there is only one copy. Plasmids carry accessory genetic material used in horizontal gene transfer
Describe prokaryotic ribosomes
70s size (smaller than eukaryotes)
Due to lack of compartmentalisation, transcription and translation are coupled
mRNA : RNAP : Ribosome complex - multiple ribosomes get loaded onto a single mRNA (polysome)
What is the glycocalyx / EPS
A sticky/slimy coat made up of polysaccharides, proteins and glycoproteins / glycolipids
EPS stands for extracellular polymeric substance
Subdivided into capsules and slime layers
What are the differences between capsules and slime layers
Capsules - layer organized into a tight matrix with excludes small particles
Slime layers- Easily deformed and loosly attached
What is the role of capsules/slime layers
Adherence of bacteria to surfaces
Formation of biofilms
Capsule acts as a virulence factor
Helps bacteria avoid destruction by the hosts immune system
What are fimbrae
Plays a role in adherence of bacteria to host cells in symbiotic or pathogenic situations
What are Pilli
There are only a few on a cell and they are longer than fimbrae. Sex pilli are responsible for donor + recipient cells in bacterial conjugation
Type 4 pilli support adhesion and twitching mobility
What is the function of flagellum
Used for locomotion. Hollow and made out of the protein flagellin
What are the 4 types of flagellum and how do they differ
Monotrichous - A singular flagellum on one polar end of cell
Lophotrichous- Multiple flagella on one polar end of cell
Amptrichious - A singular flagellum on each polar end of cell
Petritrichous- Flagella surrounding the cell
Why is gene expression controlled in prokaryotes
Resource preservation- transcribing and translating all the time is resource intensive
Regulating bacterial behaviour- infection, number of flagella, behaviour, biofilm and spore formation
Response to the environment- Circadian rhythms, Quorum (being able to sense when other bacteria are around), immune evasion in pathogenic bacteria, gravitaxis
How are spores formed
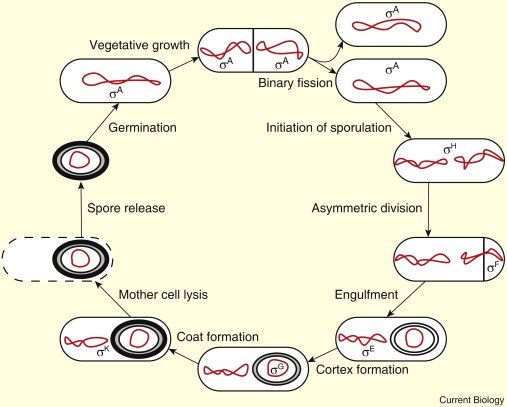
What are sigma factors
A subunit of RNAP holoenzyme
Acts as a landing pad for the rest of RNAP
Initiates binding and then falls off
The binding is initiated in promotor sequences
What are 3 methods of controlling gene expression
Only activate some genes at once (use different promotor regions for different genes, requiring different sigma factors, however this requires a lot of different sigma factors)
Inhibit genes until we need them (use repressor/regulator genes which prevent sigma binding, or use anti sigma factors which sequester the sigma factor)
Degrade sigma factors (Use degraders to degrade the sigma factor before binding)
Example of controlling gene expression by degrading sigma factors
Uses the gene RPOH and the sigma factor H
Chaperone proteins (proteins which handle sigma factors) bind to the sigma factor and degrade it at 30 degrees, so RPOH isnt translated
However if heat shock occurs some of the proteins denature and unfurl, causing the chaperone proteins to be sequestered to refold the proteins
This means the sigma factors can activate RPOH
What are regulator proteins
Proteins which bind to the operator region
What is the operator region
Usually near the transcribed region
What is an operon
A cluster of genes in the same functioning unit
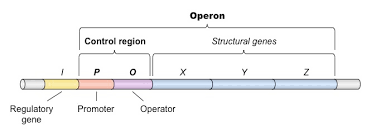
What is polycistronic
Described how multiple proteins can be produced from 1 mRNA
What is the function of operator regions
repressor protein binds to operator
Hinders DNAP (steric hinderance)
Genes are not transcribed
What are the 2 types of negative gene regulation
Repression - uses a repressor binding to operator region
Induction - inducer binds to repressor, removing it (eg LAC operon)
What is positive gene regulation
Induction - Inducer binds to activator protein, which binds to operator this changes the DNA structure to induce transcription
What is EMSA (gel shift assay)
Proteins (eg. sigma factors) in the sample will bind to DNA fragments, meaning the fragments will migrate slower in gel electrophoresis. The labelling of nucleotides enables the detection of the protein - nucleotide complexes
What are promotor and consensus sequences
Promotors are found at the 5 prime end and contain conserved consensus sequences which allow sigma factor binding. The consensus sequence is the most common order of nucleotides in a promotor sequence
How does the genome of prokaryotes and eukaryotes differ
Prokaryotes- Circular DNA, 1 chromosome, lacks introns
Eukaryotes - Linear DNA, multiple chromosomes in pairs, introns
How does the DNA location of prokaryotes and eukaryotes differ
Prokaryotes- Nucleoid region in cytoplasm
Eukaryotes- Within membrane enclosed nucleus
How does the cell wall of prokaryotes and eukaryotes differ
Prokaryotes- usually present and made of peptidoglycan
Eukaryotes- when present made of cellulose or chitin
How does the internal membrane of prokaryotes and eukaryotes differ
Prokaryotes- May have energy inducing lamella
Eukaryotes- Extensive membranous organelles
How does the gene structure of prokaryotes and eukaryotes differ
prokaryotes- Contains operator region and no introns/exons
Eukaryotes- No operator region but includes introns and exons
How is RNA processed in eukrayotes
Introns are spliced out, exons are ligated
Forms mRNA
Called splicing
Novel proteins can be created by ligating exons in different ways
What is compartmentalisation
It enables specialization of cells, allowing different biological processes to happen simultaneously and separately. Allows different reactions to have different conditions (eg. pH)
What is the golgi apparatus
Made out of membrane bound stacks of organelles called cisternae. Post transcriptional modifications occur here.
What is the endosomal system
Acts as a “way station” for proteins coming in/out of the cell
How does the nuclear pore complex control import/export
It is a multi unit holoenzyme, proteins and nucleic acids can pass if they have the NIS (proteins) or are attached to a chaperone protein (nucleic acids).
What is the NIS
Nuclear localisation signal, a sequence of amino acids which is required for proteins to enter the nuclear pore complex.
What is the difference between heterochromatin and euchromatin
Heterochromatin- tightly wound, inactive chromasomes tethered to the nucleus by lamina
Euchromatin- active DNA, less tightly wound
What does the mitochondrial genome contain
Circular DNA, genes for the ETC subunits and ribosome function (tRNA). Proteins are imported from nucleus by chaperone proteins.
How does the endomembrane system transport proteins
Endomembrane system consists of
Endoplasmic reticulum
Nuclear membrane
Golgi apparatus
Vesicles
Lysosome
Proteins are shuttled from the ER to the golgi to a vesicle
What do the free ribosomes in cytosol produce
Proteins which are shuttled to the nucleus, chloroplasts, mitochondria and peroxisomes
What do the ER membrane bound ribosomes produce
Proteins which are shuttled to the golgi apparatus which are then transferred to secretory vesicles to the plasma/nuclear membrane, endosomes or lysosomes
What are the 3 types of immunostaining
Immunofluorescence- Uses florescent dye
Immunohistochemistry- Uses antibodies in a tissue
Immunocytochemistry- Uses antibodies in a cell
What are some advantages of multicellularity
Cells are bigger and therefore have greater protection from predation and better buffering from environment
Allows specialised cells to develop
Explain the flagellar synthesis constraint hypothesis
Flagella allow a simple multicellular organism to move.
However the microtubule organising machinery needed for flagella formation is also needed for spindle fibre apparatus in cell division
Therefore there is competition for the microtubule machinery
The presence of both flagellated and non-flagellated cells allows movement and development in one simple colony
Explain weissmans nuclear determinants theory
Unequal division of determinants leads to cell specialisation and differences.
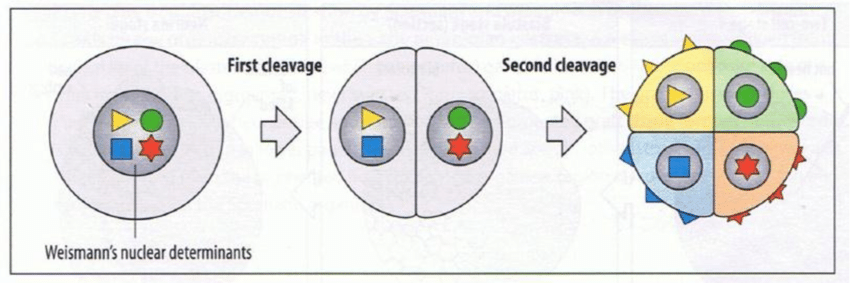
Explain Hans Dreish’s experiments and theory
He used sea urchin blastomeres (2 cell embryos) to observe development
Saw that each blastomere developed into a complete larva
Shows early embryonic cells retain ability to develop into a full organism
Disproves weissmans nuclear determinants theory
Why is cell-cell signalling important
Balance between differentiated cell types (flagellar constraint hypothesis)
Homeostasis and maintenance of internal conditions
Regulating development
Recognising self and non self
Cell adherance
What is cell lineage
The cells which are switched on/off in a cell. Important as all cells have the same genetic material so there must be a key process regulating gene action in development.
Eg. Dolly the sheep
What are the different types of cellular response
Movement (eg. slime mould)
Contraction (eg. lactation)
Metabolism alteration (eg. Diabetes response)
What is gene expression
The process by which a genes coded information is converted into the structures which are present and operating in the cell
How is gene expression altered
A gene may/may not be translated into RNA
Splicing
At export from nucleus
Proteins encoded by genes may be regulated by post-transcriptional modifications that alter activity and stability of the protein
How is a eukaryotic protein coding gene organized
Enhancers/silencers - Appear throughout genome
Promotor- Composed of core and proximal regions
Open reading frame- Contains exons and introns
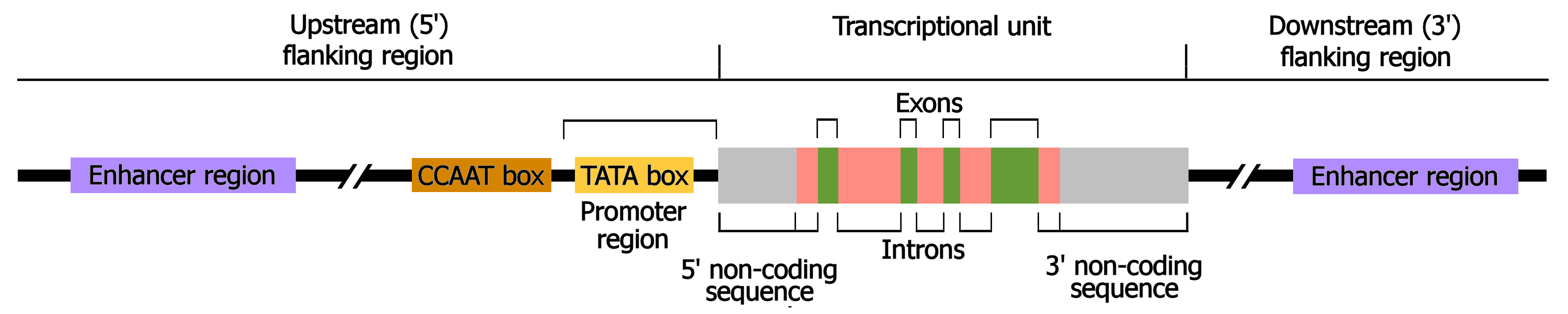
What is the difference between cis and trans regulatory elements
Cis - Regions of DNA involved in gene regulation (enhancers, silencers, promotors)
Trans- binding factors which bind to cis regulatory elements (eg. transcription factors)
How do transcription factors operate
They alter the activity of cis regulatory elements by binding to the major groove of DNA (does not melt it)
What are the functions of general transcription factors
They form the RNAP complex at the TATA box
What is the function of TBP (transcription binding protein)
Recognises TATA box
Bends the DNA 80 degrees to separate strands
As there are fewer H bonds between TA and GC, it is easier to melt at the TATA box
What is the full mechanism of RNAP complex binding in eukaryotes
TF11D (made up of TBP and TAF) recognises TATA box and separates strands
TF11B recognises binding recognition sequence in promotor region and positions RNAP at the start site
TF11F attracts and stabilises RNAP interaction with TF11H and E
TF11H and E unwinds at start point and phosphorylates. Releases RNAP from promotor
RNAP complex is formed, transcription factors are released and transcription begins.
How do specific transcription factors function
They recognise regions of 6-12 bases long (motifs) and influence the binding of transcription initiation complex by binding to silencer/enhancer regions. They can also recruit other proteins to these regions.
How do specific transcription factors influence silencers / enhancers
They form loops in the DNA to bring the regions back into proximity with the promotor
What is the wnt signalling pathway used for
Embryonic development and tissue regeneration
What is the process of the wnt signalling pathway
wnt messenger molecule activates the frazzled receptor in the cell membrane
This activates dishevelled which inhibits the b catenin destruction complex
This leads to the stabilisation of b catenin which activates transcription factors
However if the pathway is mutated, the transcription factors wont be activated as b catenin is still degraded.
How does the hypoxia pathway function in normal oxygen conditions
HIF1A is marked by proline hydroxylase which requires oxygen
Marked HIF1A is recognized by pVHL which targets it for degredation
No genes are activated
How does the hypoxia pathway function in hypoxic conditions
HIF1A cannot be marked due to lack of oxygen
It is not degraded so will bind to genes and activate them
Eg. VEGF which is the gene for blood vessel developement
How is binding of a transcription factor measured
CH1P - qPCR
DNA is crosslinked to transcription factors using formaldehyde
DNA is sheered using ultrasound and purified using antibodies for the transcription factor
The region in which TFs are bound is amplified
If no TFs are present then the region is not amplified.
What is extracellular signalling
The process by which cells communicate with each other via signalling molecules
What is the function of receptors
Enable the cell to respond to a signal
The carriers of these signals will bind to specific receptor proteins.
What are the 4 types of signalling
Contact
Signalling requires cells to be in direct contact between signaling and receiving molecules
Paracrine
Signalling secreted in the extracellular space
Synaptic
Specialized paracrine using neurotransmitters released in synapses between neurons
Endocrine
(hormones) produced in a local group of cells and secreted
What are some examples of signalling molecules
Peptides, e.g. Insulin/VEGF. These are the products of genes and encoded like any other protein.
Small molecules, e.g. nitrous oxide – short lived and breaks down fast.
Metabolic products, e.g. Steroids. These are not encoded in genes but are the product of a series of enzymes.
Lipids, e.g. phospholipids. These remain bound to membranes.
What are the different types of signal termination
Receptor internalisation/sequestration
the receptor is brought into the cell so it can no longer respond to the signal.
Degradation/Down-regulation of the signalling molecule/receptor:
The molecule may break down on its own or enzymes may break down the signalling molecule receptor.
Feedback inhibition:
Activation of the receptor leads to feedback loops that make the receptor less responsive to activation.
How do hydrophilic molecules cross the plasma membrane
As the receptor is bound to the surface the transduction is usually more complex, as the signal has to pass through the cell surface and cytoplasmic and onto the intracellular target. These can be very complex.
How do hydrophobic molecules cross the plasma membrane
They can cross the membrane so they use intracellular receptors
Often transported by carrier proteins, they can diffuse across the membrane; the receptor can be found in either the cytoplasm or even the nucleus. The consequence is the signal transduction pathway is less complex.
What are steroid hormones
Synthesized from cholesterol
Small organic molecules
How do steroid hormone receptors operate
Nuclear Hormone Receptors are both receptors and transcription factors. This feature of the receptor means it directly connects the binding of the steroid as a ligand to the transcription of a gene.
In the absence of a hormone, the signaling molecule and ligand for the receptor. The receptor is held in an inactive complex. Typically this is a monomer in bound to heat shock proteins.
Binding of the ligand to the receptor alters the shape of the nuclear receptor, activating it. The active form then releases from inhibitory proteins and acts as a transcription factor.
What is amplification
Amplification in the pathway can be achieved by a single receptor activating an enzyme.
The enzyme may then activate many other proteins or generate many other secondary messenger molecules. The result is that one molecule (an enzyme) amplifies the signal of many other molecules to enable the cellular effect.
What is integration
Integration in a pathway is often caused by two signalling pathways by using a shared common component.
This means two signals can activate the same downstream effect. Alternatively, the second pathway may inhibit the output of the first to provide regulation of the second pathway.
How does regulation of signalling pathways operate
A single pathway can be regulated at multiple points, using the Wnt pathway as an example again.
Eg.
The Frizzled receptor expression
Regulation of β-catenin Stability
TCF/LEF Transcription Factors (and cofactors)
Crosstalk with other pathways, e.g. Notch
What is protein phosphorylation
The addition, or removal, of a phosphate group to a protein can alter the structure and activity of the protein. The phosphate group is typically added to a tyrosine residue in the protein, but serine, threonine are also common targets for phosphorylation. The phosphate group is highly charged, and therefore alters the protein confirmation. One third of eukaryotic proteins are phosphorylated.
What are protein phosphatases and kinases
Kinases
Specific to the residue of the phosphate
Specificity is determined by the amino acids which surround the target amino acid residue, these are recognized as the kinases
Protein phosphatases
Remove phosphate groups from proteins, they catalyze the reverse reaction to remove the phosphate group.
What is SDS page
SDS-PAGE stands for Sodium Dodecyl Sulphate Polyacrylamide Gel Electrophoresis.
The technique separates proteins based on their molecular weight by using an electric field to move them through a polyacrylamide gel.
How are the proteins treated before being analysed using SDS page
Proteins are treated with SDS, a detergent that denatures them and imparts a uniform negative charge, and mercaptoethanol to remove disulphide bonds.
This denaturation ensures that proteins unfold into linear structures and carry a consistent negative charge per unit mass, facilitating separation based on size alone.
What is the process of western blotting used for
Western blotting allows for the specific detection of a target protein within a complex mixture. This uses specific antibodies. It also generates quantitive data by measuring the intensity of bands on the blot . This is useful for comparing protein expression between different samples or conditions.
What steps are involved in the process of western blotting
Gel Electrophoresis
We initially run an SDS-PAGE gel, but don’t stain.
Transfer to Blotting Membrane
After separation, the proteins are transferred onto a membrane. This transfer allows the proteins to be immobilised and accessible for antibody binding.
Incubation with Antibodies
The blotted membrane is then incubated with primary antibodies, which specifically recognise and bind to the target protein of interest. Following this, secondary antibodies that are conjugated to enzymes or fluorescent markers are introduced.
Detection Methods
Depending on the type of secondary antibody used, the detection of the target protein can occur through various methods.
Why are secondary antibodies used
Amplification of Signal
Secondary antibodies amplify the signal generated by the primary antibody. This amplification is crucial for enhancing the detection sensitivity, as secondary antibodies can bind to multiple sites on the primary antibody, leading to an intensified signal.
Cost-Efficiency
Secondary antibodies are generally less expensive to produce than primary antibodies. Using a universal secondary antibody with various primary antibodies can be a cost-effective approach, especially when conducting experiments involving multiple target
Describe fluorescent detection as a secondary antibody detection method
Secondary antibodies are conjugated with fluorescent dyes.
Fluorescent signals are visualised using a fluorescence microscope or gel documentation system.
Pros- High sensitivity and multiplexing capabilities, minimal sample degradation
Cons - Potential for photobleaching
Describe chemiluminescent as a secondary antibody detection method
Enzyme-catalysed reaction produces light, often through luminol or other chemiluminescent substrates.
Emitted light is captured and visualized using specialized equipment.
Pros - High sensitivity and wide dynamic range, minimal background noise
Cons- Relatively short signal duration, limited options for multiplexing.
Describe colorimetric detection as a secondary antibody detection method
Enzymes (e.g., horseradish peroxidase) conjugated to secondary antibodies catalyse a reaction producing a visible product.
Colorimetric (colour change) or chemiluminescent (light emission) signals are generated.
Pros- Moderate sensitivity
Cons- Signal decay over time, limited dynamic range
Describe radioactive detection as a secondary antibody detection method
Utilizes radioactive isotopes or labelling probes.
Radioactive emissions are detected by autoradiography.
Pros - High sensitivity, quantitive measures
Cons- Safety concerns due to radioactivity, short half life of some isotopes.
Why are positive controls used in western blots
Importance: Confirming the efficacy of antibody binding, detection, and overall success of the western blot experiment.
Example: Known protein sample with expected band size under the chosen experimental conditions.
Why are loading markers/housekeeping proteins used in western blots
Importance: Normalizing for variations in sample loading, ensuring equal protein amounts, and validating consistent transfer efficiency.
Example: Housekeeping protein, like GAPDH, serving as an internal control for gel-based western blot analyses.
Why are negative controls used in western blots
Importance: Assessing antibody specificity, detecting potential contamination, non-specific binding and preventing false positives.
Example: Using a cell line that doesn’t express the protein.
Why are molecular weight markers used in western blots
Importance: Facilitating accurate size estimation of target proteins, confirming successful transfer, and aiding in result interpretation.
Example: Confirming band is appropriate mass for the known protein, and therefore specific.
What are RTKs
Receptor tyrosine kinases
They exist for many different signals and all function by a similar pathway for activation by dimerisation to a ligand.
They span the cell membrane
How does activation of an RTK occur
It is a 2 stage event
A signal protein will bind to an RTK
This causes dimerisation of the inactive RTK
Then trans-autophosphorylation occurs which activates kinase domains
The phosphorylation spreads across the RTK
This creates binding sites for signalling proteins
How does autophosphorylation of RTKs cause the next step of signalling
The ability for the RTK to phosphorylate itself enables the binding of adaptor proteins which cannot bind until the tyrosines on the receptor are phosphorylated.
The proteins that bind the RTK may also be enzymes to phosphorylate other proteins or other forms of messaging, e.g. membrane lipids.
What is the experimental data that supports the theory that dimerisation isn’t enough to fully activate signalling?
Data using chimeric insulin/EGF receptors
The insulin receptor is always dimerized, as its subunits are covalently linked by disulphide bonds.
A chimeric (pieced together) receptor was created using the insulin receptor external domains (containing the disulphide bonds) and the EGF receptor (the RTK being studied).
The new hybrid receptor was only able to transmit signals when occupied by insulin, demonstrating that dimerization alone is not sufficient for signal transduction.