Biochemistry Methods
1/31
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
32 Terms
SDS PAGE
polyacrylamide gel electrophoresis- molecular separations of proteins based on gel filtration (size/shape) and electorphoretic mobility
SDS detergent denatures proteins, creating a rod-like shape in a consistent ratio, and masking charges
SDS-treated proteins have similar shape and charge-mass ratios
relative mobilities vary linearly with the log of molecular masses=
Because SDS disrupts noncovalent interactions between polypeptides, SDS-PAGE yields the molecular masses of the subunits of multisubunit proteins.
xray crystallography
images molecules in crystals- allows determination of 3d structure of proteins
crystal of the molecule is exposed to colimated beam of x-rays
radiation detector gives diffraction pattern
intensities of peaks create mathematical image of 3d crystal structure
—>electron density image of molecule
Knowing primary sturcture allows fitting of amino acid residue to to the electron density map
refinement reduces uncertainty to .1 angstroms!
cryo-electron microscopy
directly images macromolecular structures
wave particle duality- electrons have wavelike properties
beam of electrons of sufficiently high energy will have a wavelength small enought o image molecules at atomic resolution
hydrated sample is cooled to low temps rapidly - becomes a vitreous glasslike state.
sample retains native structure —> high resolution
setbacks: requires higher dosage of electrons to resolve atoms, damaged samples through chemical bonds, —> imaging with lower electron doses allows higher atomic resolution
measuring free energy
determine by
primer tricks: site directed mutagenesis- individual changes to amino acids and quantify the effects of various forces to stabilization
ex: test for a cavity- alanine for valine—> measure free energy of unfolding to see the heat for a protein to denature? is that different from this amino acid change
centrifugation
ELISA
Pulsed H/D exchange
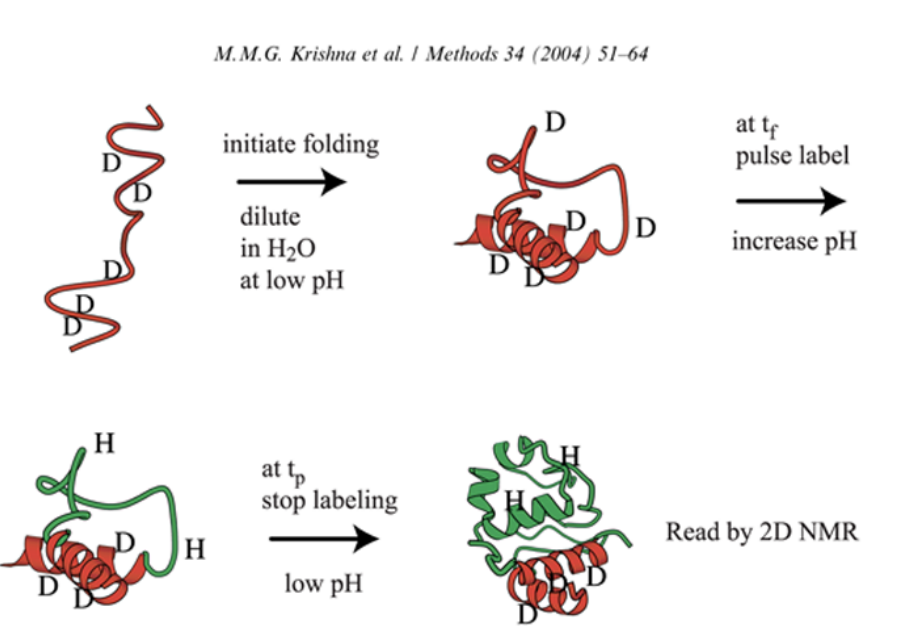
How do we know how proteins fold?
generally done with backbone amines protonated—> let sit in deuterium until we get an exchange to occur
These results look different in NMR- we can see the odd nucleus on the hydrogen but not on the deuterium
rapid exchange of hydrogen for deuterium if we put a small molecule in -
if involved in a hydrogen bond - exchange rate will slow because the h-bond is stabilizing in an energetic minimum
start letting the protein fold
dilute it into water (with syringe on rapid automated timescale) with lower pH to control the exchange- no exchange with the solvent of deuteriums
increase pH that is amenable to eltting the amine exchange with the protons on the water- will exchange more rapidly than in low pH - won’t exchange to the same degree
hit with final time with low pH to stop the exchange, let protein continue to fold
—>throw inot nMR
pulse it, hit with stock at low pH, this will tell us what happened at the beginning
by doing this in different time scales, can see the processes at different points in folding process
sanger dideoxy sequencing
restriction enzyme
bacterial proteins that act as "molecular scissors," cutting double-stranded DNA at specific palindromic sequences.
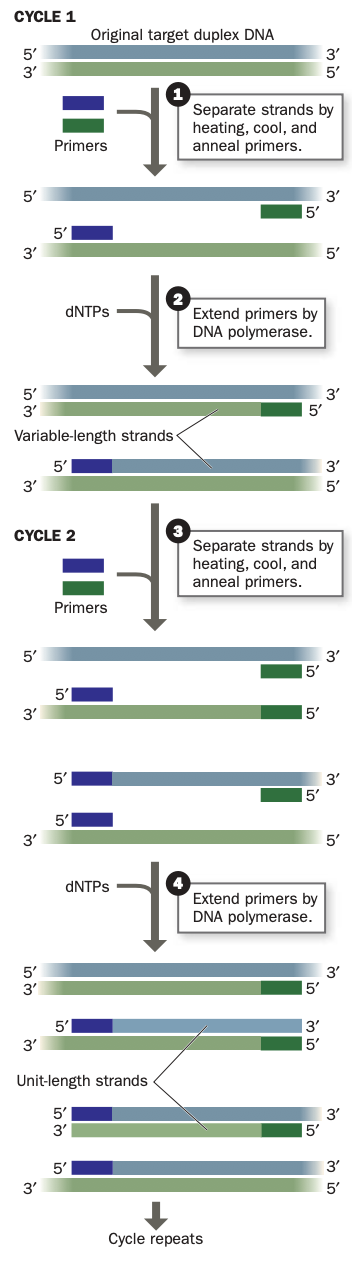
PCR
a DNA sample is separated into single strands and incubated with DNA polymerase, dNTPs, and two oligonucleotide primers with sequences that flank the DNA segment of interest. The primers direct the DNA polymerase to synthe- size complementary strands of the target DNA.
n each cycle, the two strands of the duplex DNA are separated by heating, then the reaction mixture is cooled to allow the primers to anneal to their complementary segments on the DNA.
Heat-stable DNA polymerase directs the synthesis of the complementary strands. The use of a heat-stable DNA polymerase, such as Taq polymerase- allows the cyclic variation of temperature
obtain/amplify dna procedure
1. A fragment of DNA of the appropriate size is generated by a restriction enzyme, by PCR (below), or by chemical synthesis.
2. The fragment is incorporated into another DNA mole- cule known as a vector, which contains the sequences necessary to direct DNA replication.
3. The vector—with the DNA of interest—is introduced into cells, in which it is replicated.
4. Cells containing the desired DNA are identified, or selected.
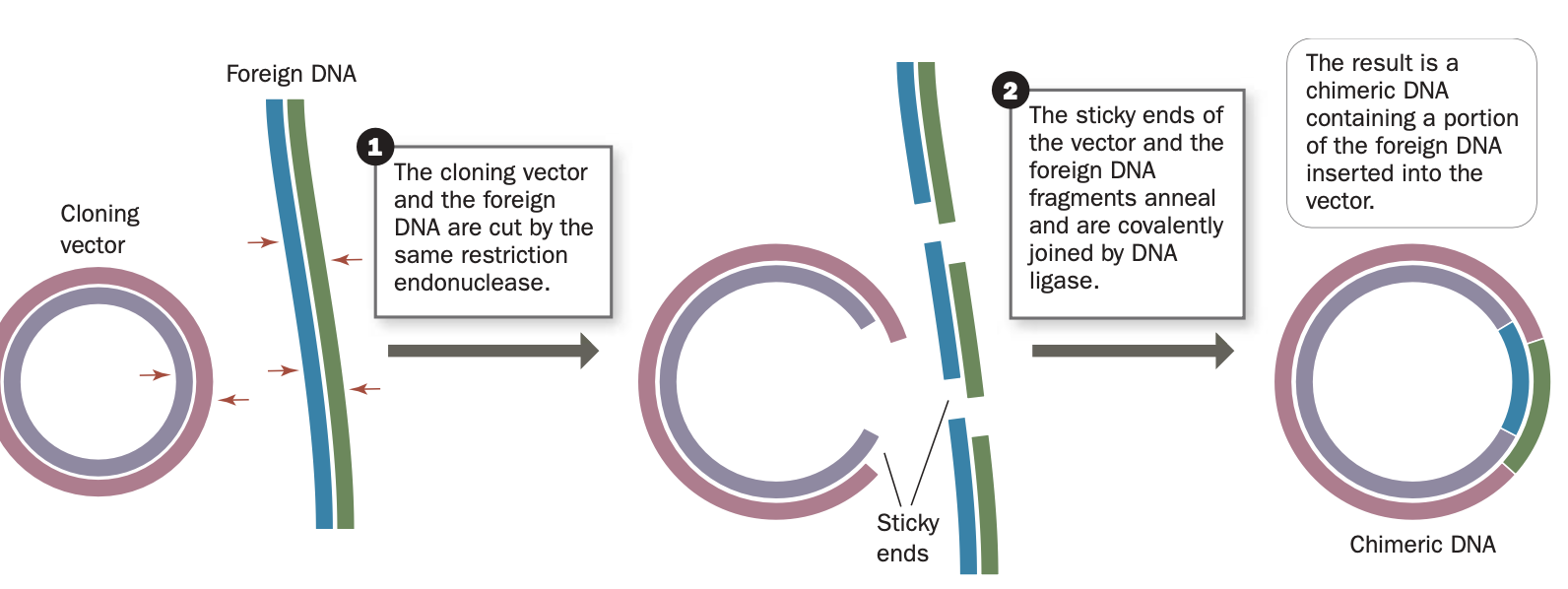
ligase
restriction fragment can be insert into a cut made in a cloning vector by the sam restriciton enzyme
the complemtary ends are spliced together
Construct recombinant dna molecules
selection
detects the presence of cloned dna
essential to select onlyhost organizations that have been transformed and contain a properly constructed vector
transformation
vector becomes permanently established in its bacterial host
vector
a carrier that transfers genetic material or infectious agents into another cell or organism.
small, autonomously replicating dna molecule
ex: plasmid
plasmid
small, circular, double-stranded DNA molecule found in bacteria, archaea, and some eukaryotes
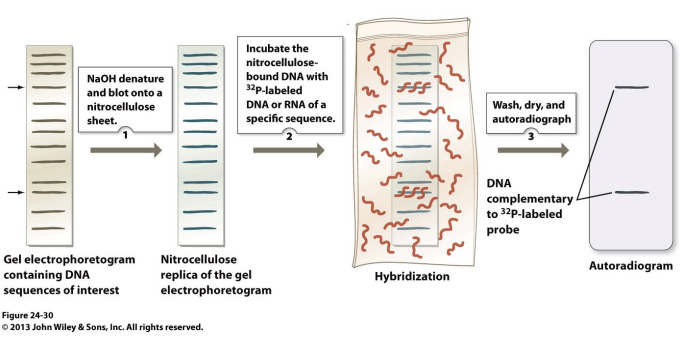
southern blot
in cases that molecules size/charges aren’t well-resolved by the gel:
Technique identifies different sequence compositions of fragments
gel electrophoresis
soak in .5 NaOH to convert to single stranded form
overlaid with nitrocellulose- molecules forced through with a absorbant towel
single stranded dna binds to nitrocellulose at same position in gel
dried with 80 degrees
moistened with solution containing a complementary single stranded probe tagged
hybridizes to the target sequence
pcr steps
Heat to separate strands.
Add synthetic oligonucleotide primers; cool (to allow annealing)
Add thermostable DNA polymerase to catalyze 5'3'DNA synthesis.
keep repeatimg
After 25 cycles, the target sequence has been amplified about 106-fold.
pcr+restriction enzymes
Combining PCR and restriction enzymes enables targeted DNA amplification followed by precise cutting for cloning, mapping, or analysis. Primers are designed to introduce restriction sites, allowing the PCR product to be digested and inserted into vectors.
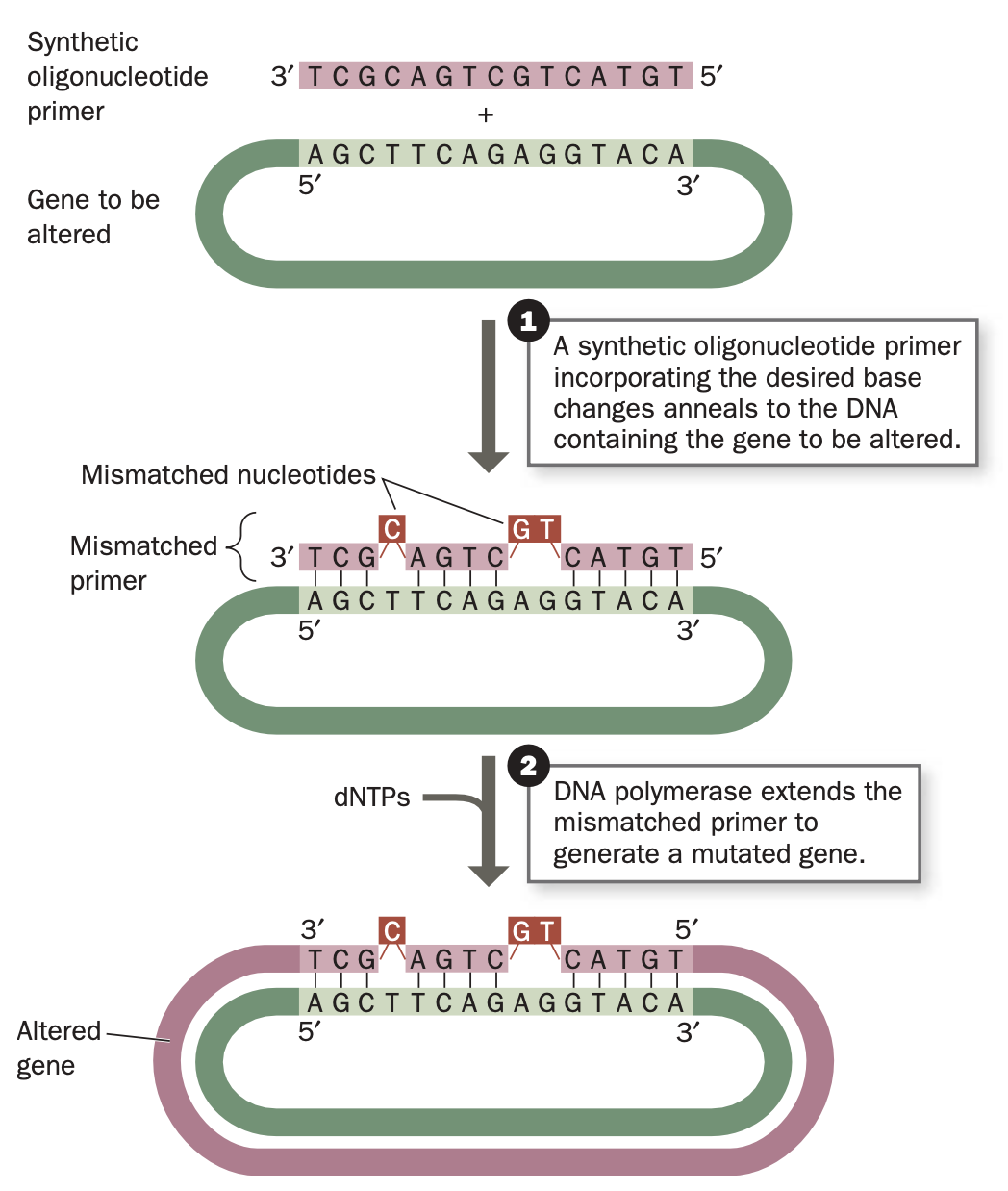
site-directed mutagenesis
modify the nucleotide sequence to alter the amino acid sequence of the encoded protein
an oligonucleotide with an identical sequence, except for the desired base changes directs replication of the gene.
Hybridizes to wild-type sequence
DNA polymerase extends the mismatched primer to generate the gene
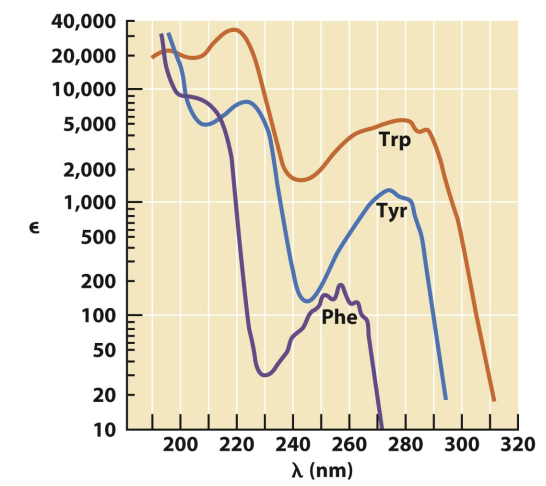
aromatic amino acids
absorb at different wavelengths
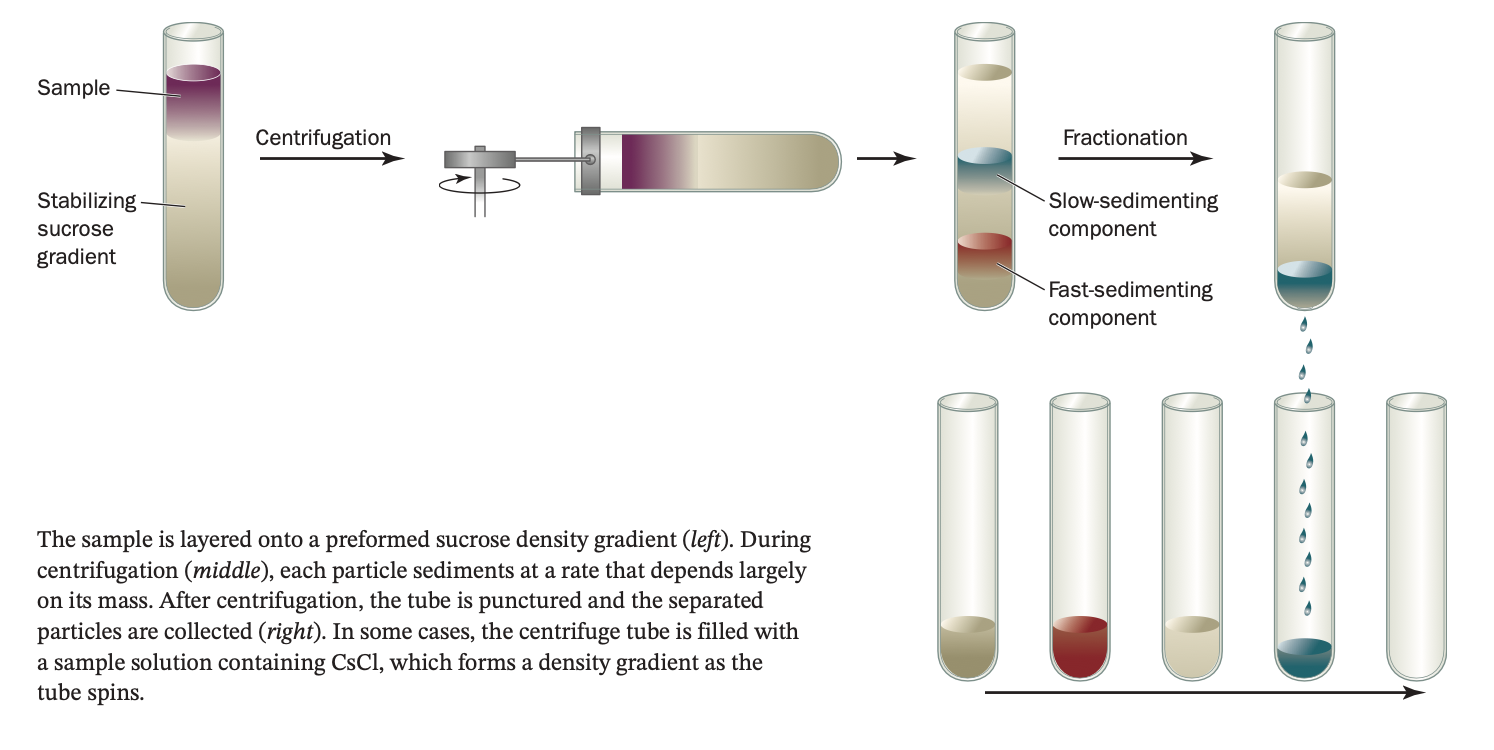
ultracentrifugation
uses immense centrifugal fields (up to 1,000,000 × g or higher) to force macromolecular sedimentation.
The rate at which a particle sediments in the ultracentrifuge is related to its mass (the den- sity of the solution and the shape of the particle also affect the sedimentation rate).
western blot
secondary antibody
Western blotting bind to the primary antibody to detect specific proteins, usually acting as a signal amplifier.
primary antibody
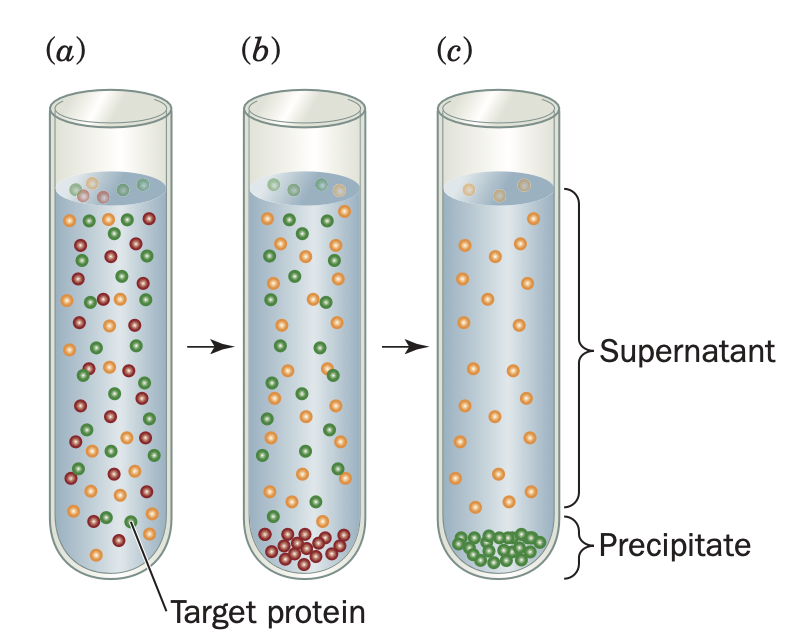
salting out
This salting out effect is primarily a result of the competition between the added salt ions and the other dissolved solutes for molecules of solvent. At very high salt concentrations, so many of the added ions are solvated that there is significantly less bulk solvent available to dissolve other substances, including proteins.
Adjusting the salt concentration in a solution containing a mixture of proteins to just below the precipitation point of the protein to be purified eliminates many unwanted proteins from the solution. Then, after removing the precipitated proteins by filtration or centrifugation, the salt concentration of the remaining solution is increased to precipitate the desired protein. This procedure results in a significant purification and concentration of large quantities of protein. Ammonium sulfate, (NH4)2SO4, is the most commonly used reagent for salting out proteins because its high solubility (3.9 M in water at 0 °C) allows the prepa- ration of solutions with high ionic strength. The pH may be adjusted to the approximate pH where the desired protein possesses a net charge of zero (the isoelectric point, pI) because a protein is least soluble under these conditions.
isoelectric focusing
2-mercaptoethanol
reducing agent breaks disulfide bonds
can be added to SDS-Page if subunits are linked
autoradiograph
2d electrophoresis resolves complex mixtures of proteins