CH29: rna function, biosynthesis, & processing (2)
1/30
Earn XP
Description and Tags
29.4-29.5
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
31 Terms
True or False: virtually all transcription products are further processed in eukaryotes, what about in prokaryotes?
True for euk, but some prok transcripts are modified
what is pre-mRNA and what 2 things happne to pre-mRNA molecules?
pre-mRNA: immediate product of RNA polymerase II
pre-mRNA molecules:
are spliced to remove introns
undergo modifications to both 5’ and 3’ ends
what is the 5’ cap? what 2 things do they do?
5’ cap: distinct terminus at 5’ containing an unusual 5’ - 5’ triphosphate linkage
protects 5’ ends from phosphatases & nucleases, contributing to mRNA stability
enhances mRNA translation by eukaryotic protein-synthesizing systems
what is the poly(A) tail and how is it added? what can it do?
poly(A) tail: a string of ~250 adenine nucleotides at the 3’ end
added by poly(A) polymerase after cleavage of pre-mRNA by endonuclease
may enhance translation efficiency & mRNA stability
what are the 3 steps to form cap 0?
trimming: one phosphate is clipped off the front of new RNA (hydrolysis)
attaching: the RNA 5’ end atks α-phosphorous atom of GTP to form 5’ cap
decorating (methylation): methyl group is added to nitrogen at the 7th position of new guanine (N-7) to form cap 0
adjacent riboses can be methylated to form cap 1 or cap 2 to help cell distinguish its own RNA from viral RNA
what does it mean when primary transcripts in eukaryotes are polyadenylated?
it means a long tail of adenine nucleotides (poly A tail) is added to the 3’ end of RNA mc after it has been transcribed from DNA
what is RNA splicing?
process by which introns are excised and exons are linked to form the final mRNA
what are the common structural motifs of introns that mark the beginning and end?
begin with GU (5’ splice site) and end with AG (3’ splice site)
describe the consensus sequence found at the 3’ end of a vertebrate intron
3’ end has polypyrimidine tract (~10 pyrimidines), followed by sequence NCAG, where AG marks actual end of intron
where is the “branch site” located and why is it significant?
branch site: 20-50 nucleotides upstream of 3’ splice site
important bc it is the point where the 5’ end of the intron attaches during the splicing process to form a loop structure
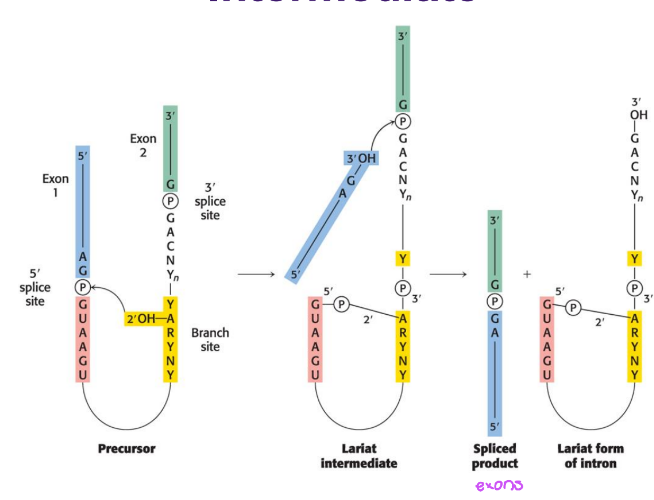
describe the 2 sequential steps of the splicing reaction (transesterification), what intermediate does it generate?
lasso: branch site adenine atks 5’ splice site, freeing exon 1 and forming lariat (2)
stitch: newly freed 3’-OH of exon 1 atks 3’ splice site, joining 2 exons and releasing intron
generates a branch and forms a lariat (loop) intermediate
Consider the sequence 5–AGGUAAGU(N)245(Py)10NCAGG–3′.
How long is the intron in this sequence?
start at GU, GUAAGU = 6, N245= 245 nucleotides, Py10= 10 pyrimidines, NCAG end at AG = 4
6+245+10+4 = 265
what are snRNPs, and which specific ones are considered essential for the splicing of mRNA precursors? what about them is highly conserved?
small nuclear ribonucleoprotein particles: complexes formed by small nuclear RNAs (snRNAs) and specific proteins
essential ones: U1, U2, U4, U5, U6
snRNAs’ secondary structures are highly conserved bc their specific folded shapes are critical for recognizing splice sites
how is the 60S spliceosome formed, and what is its role?
formed by association of essential snRNPs, hundreds of splicing factors, & pre-mRNA
role: catalyze precise excision of introns and ligation of exons
describe the roles of snRNPs for U1, U2, U5, U4, U6
snRNP Size of snRNA (nucleotides) Role
U1 165 Binds the 5' splice site
U2 185 Binds the branch site
U5 116 Binds the 5' splice site and then the 3' splice site
U4 145 Masks the catalytic activity of U6
U6 106 Catalyzes splicing
what are the 6 steps of the splicing process?
The Assembly
U1 binds to the pre-mRNA 5’ splice site
U2 binds intron branch site (req energy from ATP)
U4-U5-U6 tri-snRNP binds complex of U1, U2, and mRNA precursor to form spliceosome
The Activation
spliceosome rearrange, kicking out U1 & U4, allowing U6 to twist (intramc rearr) and pair with U2 to make catalytic center that will perform cut (interact w/ 5’ end of intron)
The Execution
First cut: 5’ exon is snipped off, intron forms lariat loop
Final stitch: U5 lines up 2 exons (5’ w 3’) and they will go transesterification to glue tgt
mature mRNA released, leaving spent snRNP, U2, U5, U6 still attached to lariat intron
what is the difference between a “cis-acting” and a “trans-acting”’ mutation in the context of splicing disease?
cis-acting mutation happens within the pre-mRNA sequence (eg. at splice site), affecting only that specific transcript
trans-acting mutation occurs in splicing machinery (eg. snRNP protein), which could affect splicing of many diff transcripts
what are some thalassemias (diseases from defective hemoglobin synthesis) caused by? what about retinitis pigmentosa?
thalassemias: caused by mutations at the 5’ or 3’ splice sites in either of the two introns of hemoglobin beta chain or in its exons
retinitis pigmentosa: disease of acquired blindness is due to mutation in pre-mRNA splicing factor that is part of U4-U5-U6 tri-snRNP
what is alternative splicing and what could it lead to?
alternative splicing: mechanism by which diff combo of exons from same gene may be spliced into a mature RNA
generates protein diversity and leads to changes in coding sequence
what is splice site selection determined by?
determined by the binding of trans-acting splicing factors to pre-mRNA cis-acting sequences
what are some human disorders attribuuted to defects in alternative splicing?
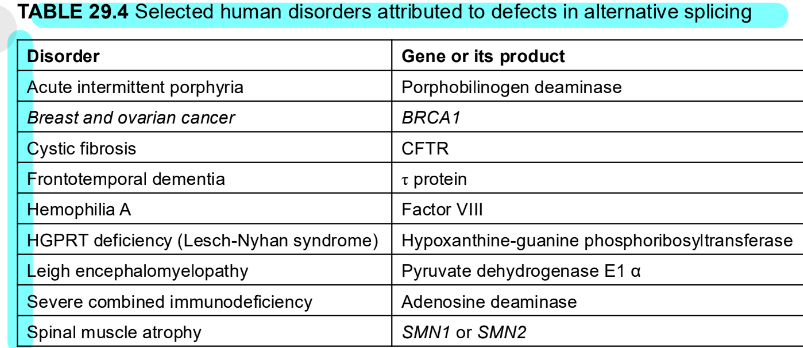
what are transcription and mRNA processing coordinated by?
the CTD (carboxyl terminal domain) of RNA polymerase II
CTD has repeated seq YSPTSPS, where S2, S5 or both may be phosphorylated
CTD caps enzymes, are components of splicing machinery and are endonuclease that cleaves transcript at poly(A) addition site
describe the structural transition a microRNA undergoes from its precursor form to its active regulatory form
it starts as a large transcript that folds into a hairpin structure, which is then cleaved by specific nucleases into short, single-stranded form that binds to regulatory proteins
what are microRNAs?
small (20-23 nucleotides) RNAs that regulate gene expression in eukaryotes
what are ribozymes?
RNAs that function as catalysts
what are group I self-splicing introns? what do they require as a cofactor?
introns that can excise themselves
initially identified in rRNA from tetrahymena
require guanosine as a cofactor
describe the transesterification reaction mechanism of group I self-splicing intron, how does its released form differ from spliceosome-mediated splicing?
it aligns splice sites using an internal guide sequence (IGS) that base-pairs with exons (base pair between IGS in intron and 5’ & 3’ exons)
then a phosphodiester bond is formed between 2 exons and intron is release
unlike spliceosome-mediated splicing which releases a lariat, group I introns are released as linear molecules
in self-splicing mechanisms, how is the catalytic site formed?
by the intron itself
in group II self-splicing introns, what is the attacking group?
atking group is a 2’-OH of an adenylate in the intron
intron released is released in lariat form
in vitro experiments, there’s the possibility of self-ligating ribozymes, what does this mean?
RNA molecules capable of joining other short RNAs to their own ends
How does group II self-splicing resemble spliceosome-catalyzed splicing of mRNA precursors? (choose all that apply )
a. A ribose hydroxyl group attacks the 3′ splice site and the
newly formed 5′-OH terminus of the upstream exon then
attacks the 5′ splice site, form a phosphodiester bond.
b. Both reactions are transesterifications with the phosphate
moieties at each splice site retained in the products.
c. The attack at the 5′ splice site is carried out by a part of the
intron itself.
d. The number of phosphodiester bonds increases.
e. The intron is released in the form of a lariat.
b, c, e