Bio 1002B cycles 1-7 with 100% correct answers
1/186
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
187 Terms
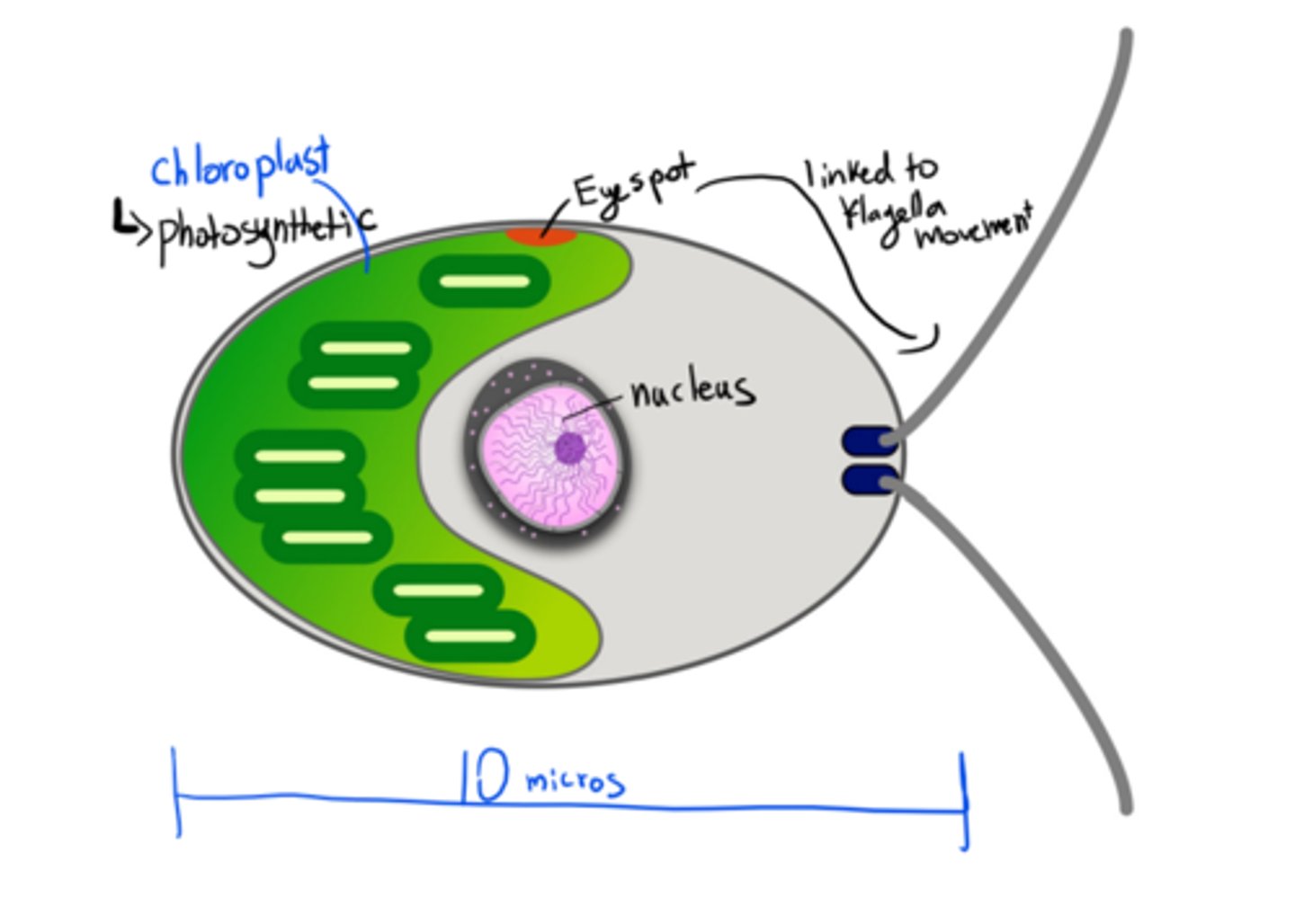
Major features of a Chlamydomonas cell
-eukaryote
-Single-celled green algae
-Has a chloroplast: photosynthetic, autotroph
-It's a model of study in bio (like lab rats) because it has a short generation time
-Has an eyespot
-Haploid dominant life style (haplontic)
-mitochondria
-has a flagella
-nucleus
-photosynthetic autotroph
-10 microns in length
-can grow in liquid culture and in agar plates, making them ideal to use in labs
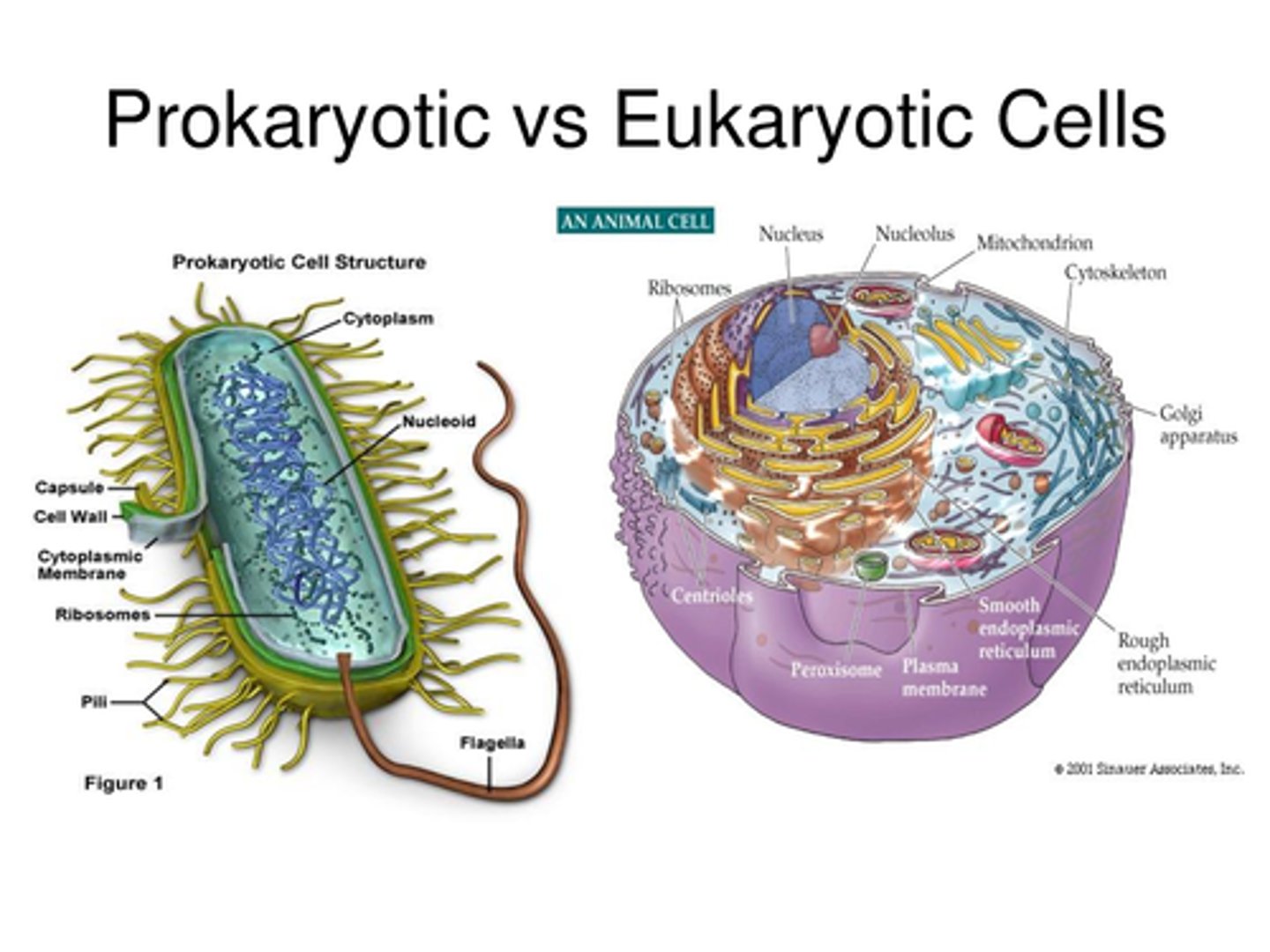
Prokaryotic vs. Eukaryotic Cells
Common:
-Plasma membrane
-Cytoplasm
-DNA
-Ribosomes
Prokaryote-specific:
-Unicellular
-No membrane-bound organelles (so no nucleus)
-DNA found in nucleoid region (part of the cytoplasm)
-Double-stranded circular DNA
-Includes: bacteria, archaea, cyanobacteria
-Asexual reproduction by binary fission
-no mitochondria or chloroplasts
-transcription occurs in the cytoplasm
Eukaryote-specific:
-Multicellular (can also be unicellular like chlamy)
-Has membrane-bound organelles (like nucleus, mitochondria and chloroplasts)
-Larger than prokaryotes
-Double-stranded linear DNA that are wound in a double helix
-Has introns and exons
-Could be haplontic (ex. Chlamy, Fungi, protists) or diplontic (animals, plants)
-transcription occurs in the nucleus
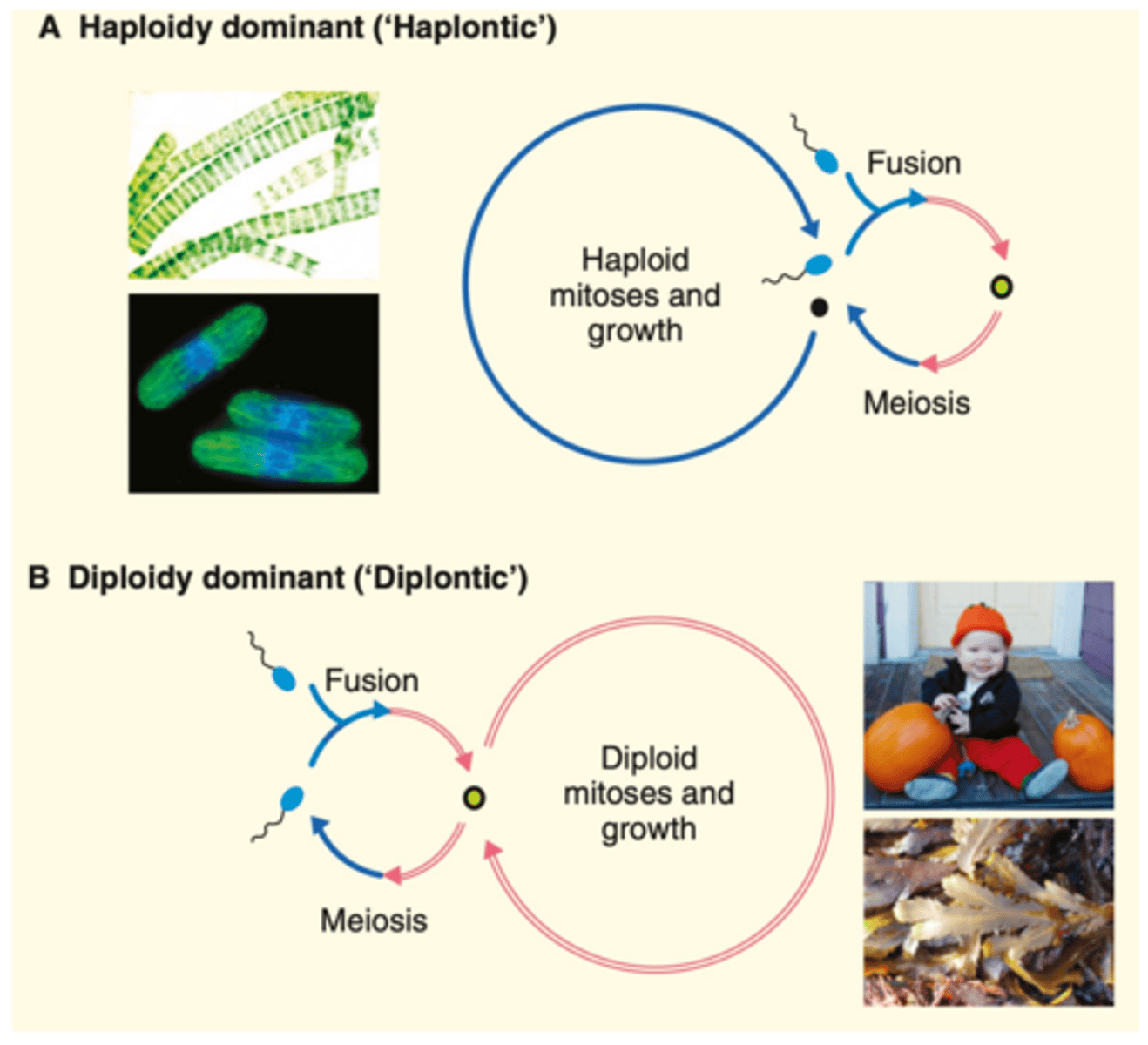
Two eukaryotic lifecycles
-Haplontic: haploid gametes fuse, diploid zygote created, the diploid zygote undergoes meiosis and becomes haploid, cells stay haploid and do mitosis to grow for most of their life
-Diplontic: haploid gametes fuse, diploid zygote created, zygote does mitosis and grows (spends most of its life diploid), some diploid cells undergo meiosis to generate gametes
-chlamy specific:
undergo sexual and asexual reproduction. Under normal conditions, chlamys undergoes asexual reproductions producing copies of itself. Under stressful conditions, chlamys undergoes sexual reproduction (sexual reproduction allows for recombination during meiosis, which results in variation and therefore increased adaptation in natural selecton).
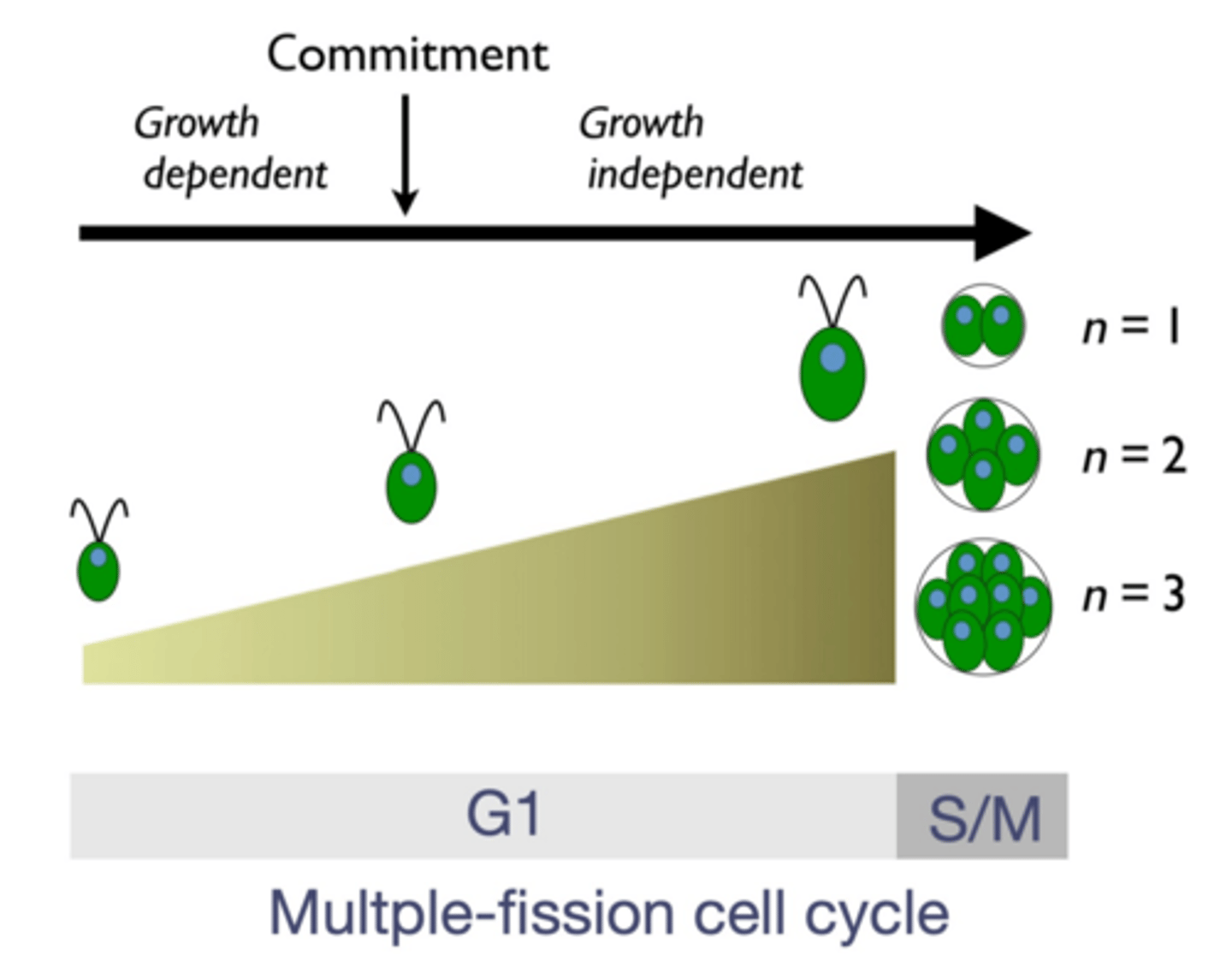
Features of Chlamydomonas growth
-During G1
-Asexual division of haploid cells (the main part of their life)
3 steps: growth-dependent, commitment, growth independent
~Growth dependent: they cannot divide yet, they need to get bigger
~Commitment: they don't NEED to get bigger, although they usually do so they can divide more later, in general this is a threshold point where it is now able to divide
~Growth independent: the phase where they're just getting bigger to get bigger, they can divide if they want to. They do this to be able to divide more (the bigger they are, the more times they will be able to divide)
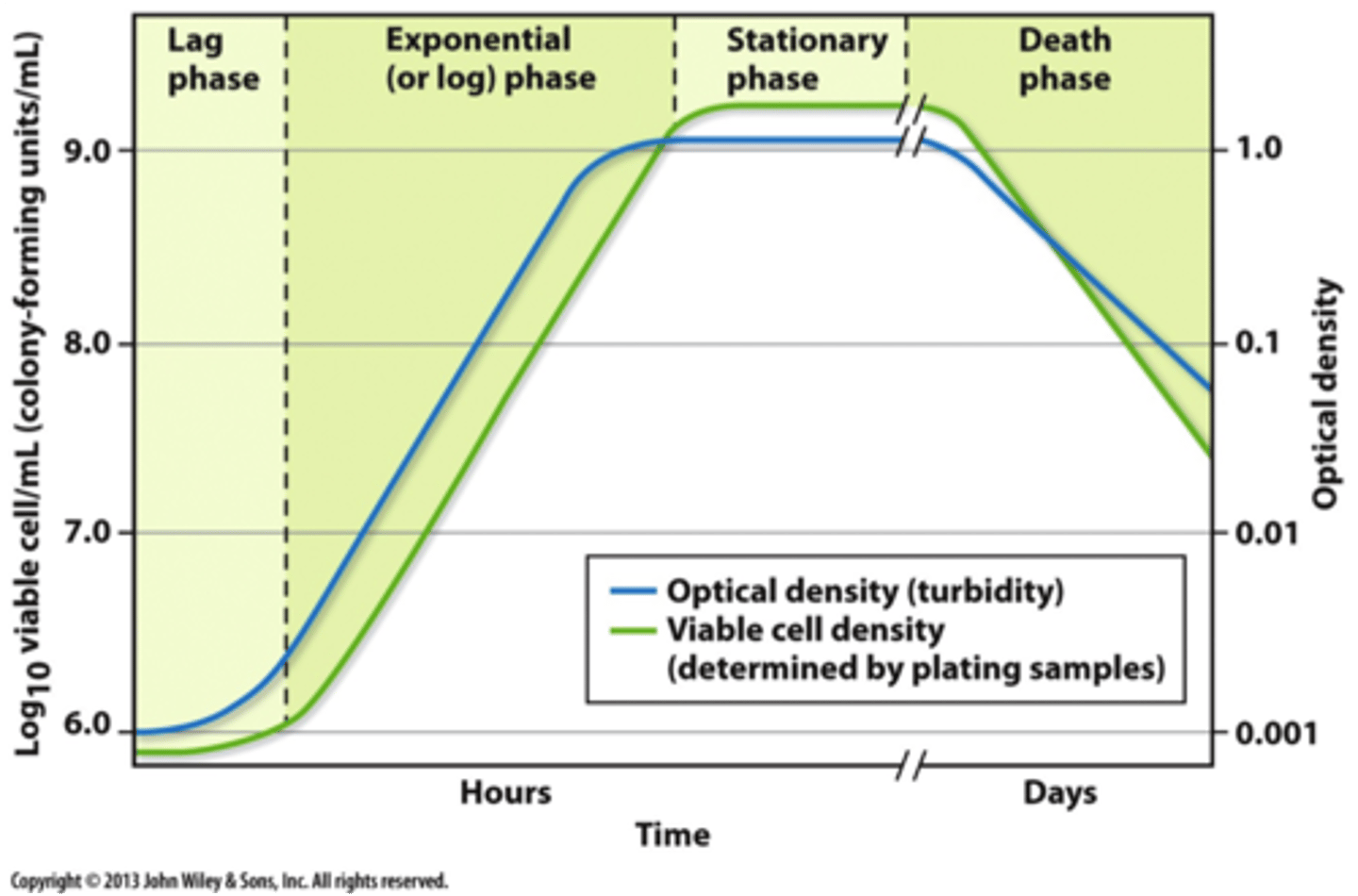
Features of Chlamys microbial growth curve
pic is missing: transitional phase between exponential and stationary + stationary phase is represented as a straight line but in reality is closer to a sin function/ wavy line
-All cells grown in the lab follows this pattern
Steeper slope= higher growth rate
Growth rate: how long it takes for one chlammy to become 2 (generation time)
-Lag phase: they are not dividing, they are simply growing and preparing to divide, they are metabolically active (not dead lol)
-Exponential growth phase: highest rate of growth, very high nutrient consumption from the growth media
-Stationary phase: almost complete stop of division due to the sample reaching its carrying capacity(K). There will be negative feedback that reduces population size every time it gets beyond carrying capacity (hence the sinusoidal function seen on the graph).
-Chlamy doesn't have enough food to stay alive, starts to die
-When doing experiments, we want the sample to be at its highest growth rate (exponential). So, when the cell is in the late exponential stage, 1ml of the sample is taken and put into a fresh flask of media. This is because the original flask has not run out of nutrients, so the sample would not reach its carrying capacity just yet and will not enter a stationary phase
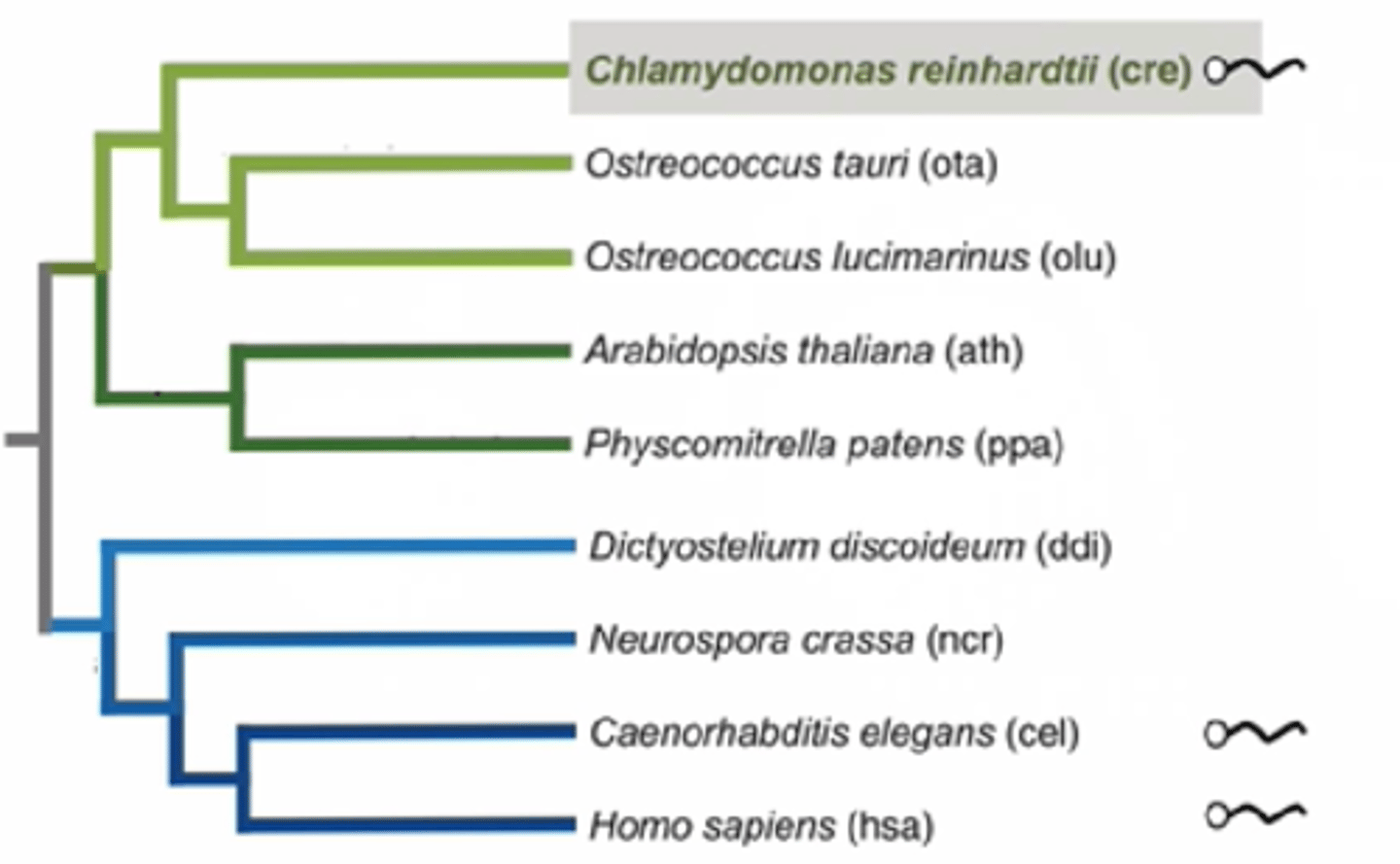
Chlamydomonas phylogeny
-Evolutionary relationship between chlammy and other species
-Chlammy (algae) is related to plants (diverged a billion years ago) since both have chloroplasts and are photosynthetic
-chlamy and humans both have flagella (cilia in humans), plants don't have flagellum, meaning that they evolved to lose flagellum, meaning that the common ancestor possessed flagellum
How can we explain that Chlamydomonas and humans have flagella but plants do not?
-Flagella exist in prokaryotes, but it's analogous to eukaryotic flagella (same function, different structure and origin)
-Flagella and cilia are practically identical (cilia is just shorter flagella and is non-motile)
-Flagella in chlammy and in humans are identical
The odds of the same structure evolving twice is very low
-Assumption: flagellum is an ancestral trait that evolved before humans, plants and algae, the common ancestor must've possessed them and lost it
-Plants lost them through evolution, chlammy and humans kept them
Why are chlamys a good model system to study human diseases related to flagella/cilia?
Mutations that alter cilia structure/function causes many human disease states (ciliopathies)
motile vs. non-motile cilia involved in sensory perception
-Motile cilia: they move lol
-Non-motile cilia (sensory cilia): cilia is a projection of the cell. Has sensory proteins to read the environment around, these can interact with light, sounds and can detect the change of flow of liquid in the urinary tracts
Analysis of 7,476 Chlamy proteins
-Proteome: total number of proteins found in chlammy
-Of 10% of the chlammy proteome, there is a related protein in humans
-Of 26% of the chlammy proteome, there is a related protein in plants
-Of the 33% of chlammy proteome, there is a related protein in both plants and humans
-The rest is unique to chlammy
-Chlamy would likely share its photosynthesis and phototactic associated proteins with only Arabidopsis, while it would likely share some flagella associated proteins with humans only. Proteins associated with functions such as DNA replication and cell respiration is shared with all three.
What characteristics does Chlamy have that makes it a "model experimental system"?
-it has a short generation (doubling time), easy to care for, can be generalized to humans so it is useful (ciliopathies)
In the lab it is useful that some systems (like Chlamy) are haploid...Why do scientists like working with haploid systems?
-It can grow and reproduce in liquid growth media and agar plates
-has a short generation time
-can undergo meiosis and adapt to changes in the environment, contains cilia that helps us study ciliopathies
-chlamy's haploid life cycle gives an experimental advantage as there is only one copy of the genome, no genetic masking that usually happens in diplontic organisms occurs, making it much easier to spot mutations
Why do you think cells stop growing in the Stationary Phase?
-it runs out of nutrients in the liquid growth culture
-for a cell to divide it needs nitrogen taken from macronutrients and trace minerals to create cofactors for enzymes
-the cells are not able to divide once they run out of these nutrients and food
Characteristics of the electromagnetic spectrum and relationship with photosynthesis
-Complete range of wavelengths
-Light: visible portion of the spectrum
-only visible light can be used in photosynthesis
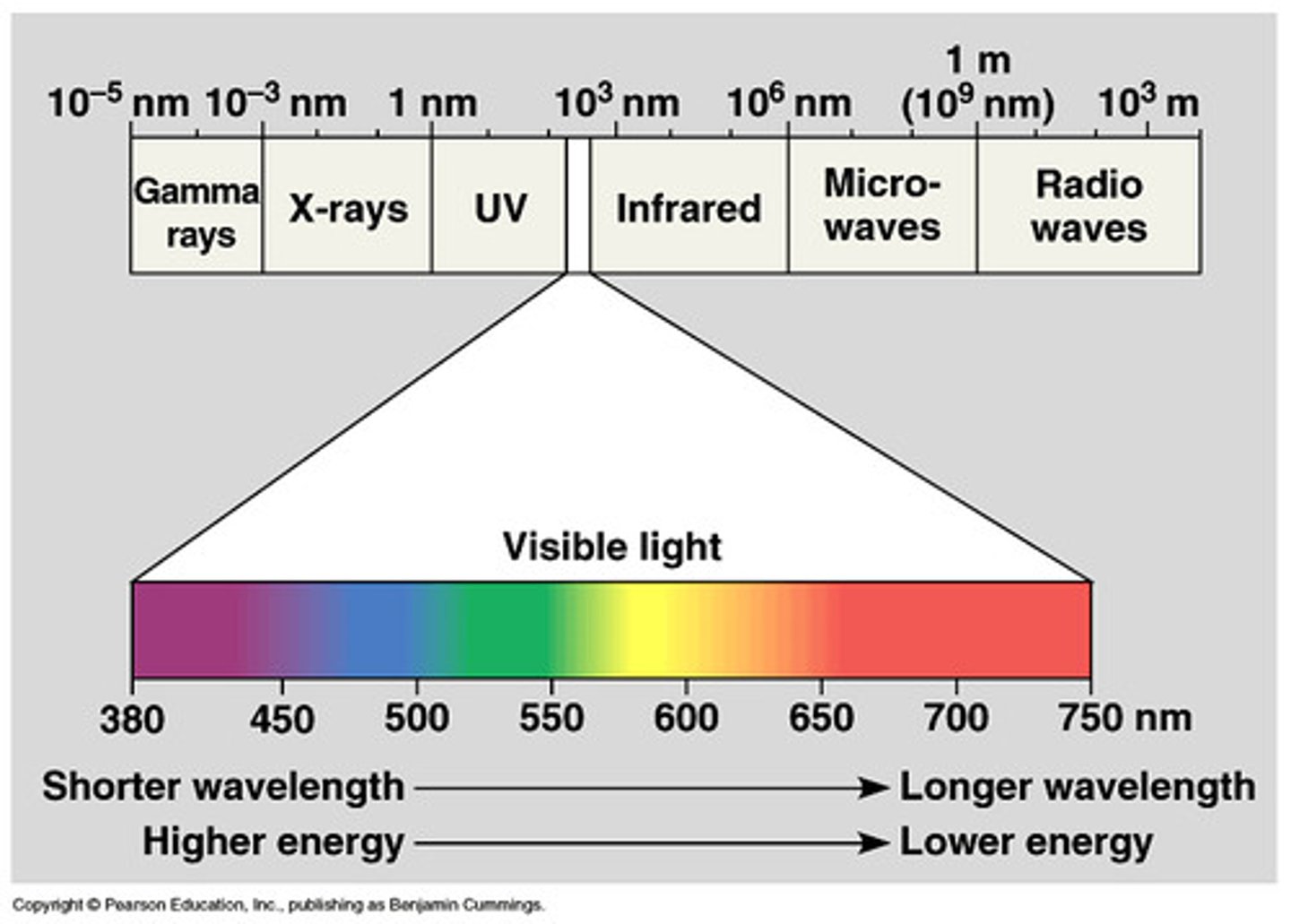
Relationship between wavelength and energy content of a photon
Wavelength is inversely proportional to energy (frequency)
Definition of photon absorption
Photon: particles of light that have wave/particle duality.
When light (photons) hit an object, either:
Reflected, Transmitted or Absorbed
-When light is absorbed, its energy is transmitted to an electron of the material. This electron will enter an excited state if the light is the appropriate wavelength that corresponds to its change between ground and excited state
Molecular characteristics of pigments that make them able to absorb light
-Pigments have a conjugated system
-Conjugated system: carbons attached covalently alternating single/double bonds
-The colour of the pigment is the colour of the wavelength that it does NOT absorb
-Ex. Chlorophyll absorbs all wavelengths of light except for the one associated with green (which is why we see it green)
-Example of pigments:
~chlorophyll a (photosynthesis)
~retinal (vision)
~indigo (jeans colours)
Understanding of why biological systems primarily absorb visible wavelengths of the electromagnetic spectrum and not x-rays or radio waves
-visible light is the dominant form of electromagnetic waves reaching Earth's surface
-visible light contains an ideal amount of energy to excite electrons of biological systems (not too strong not too weak; remember electrons can only be excited by a specific amount of energy)
-any stronger source of energy and rapid damage can occur to cells/organisms causing them do die from radiation
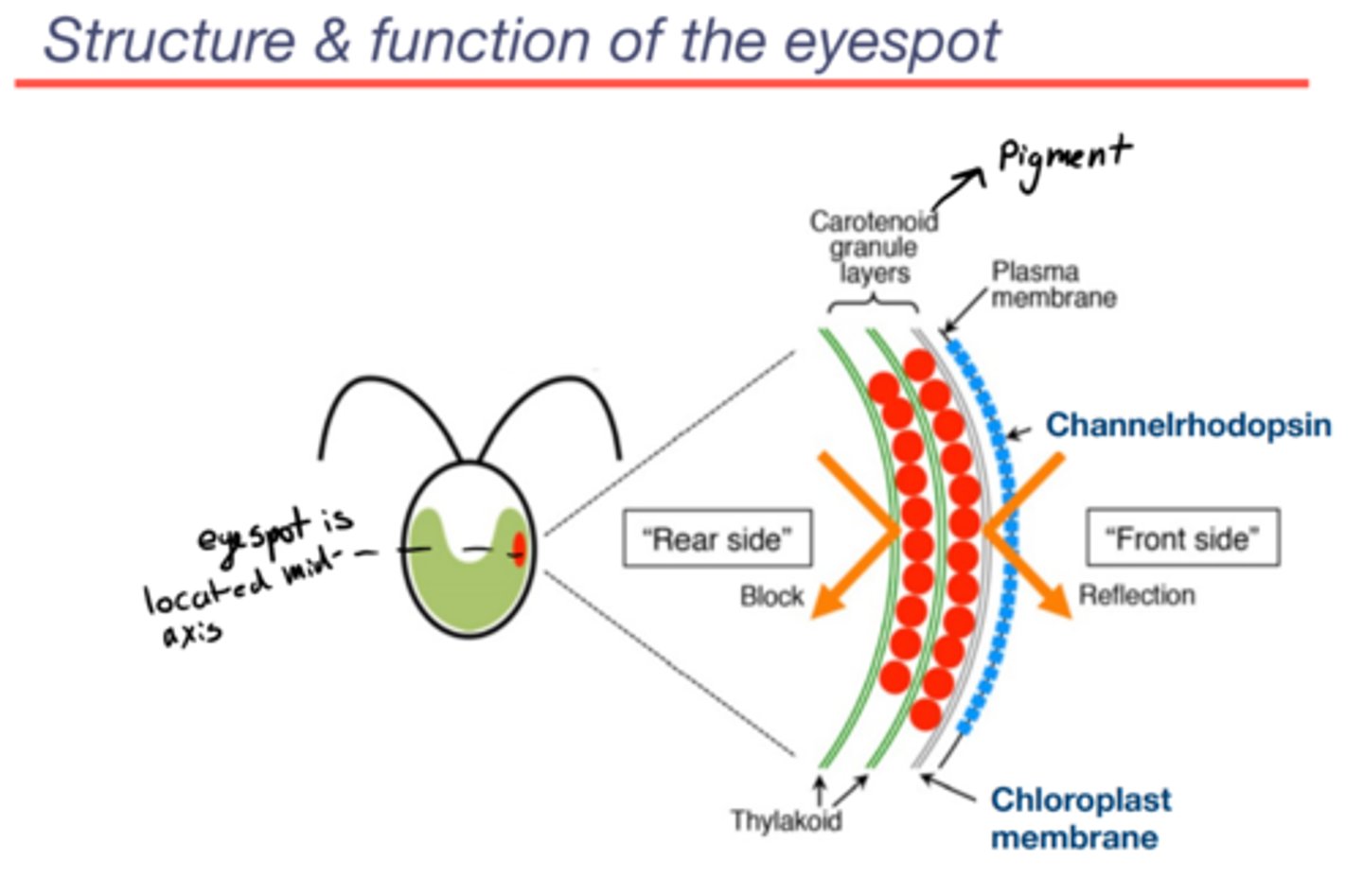
Basic organization and functional features of the chlammy eyespot
Channelrhodopsin:
-Protein complex
-Light-gated ion channel
-Sits on the plasma membrane
Carotenoid granule layer:
-component of eyespot found in the chloroplast
-the major visual component of eyespot
-when the light comes from the front side:
~either light hits channel
~the carotenoid layer is reflective: if light misses chan, it hits the granule layer, bounces off and gets absorbed by chan on the rebound... light only ever hits the back of channelrhodopsin
-when the light comes to form the rear side: it does not get to chan because the carotenoid granule layer blocks it (so light can only be sensed from the front side which gives chlammy a sense of direction)
phototaxis:
-the motion of chlammy as a response to light
-negative phototaxis: cells moving away from light
-positive phototaxis: cells moving towards the light
Basics of how an action potential is generated + Mechanism by which the signal is transduced from eyespot to flagella
light transduction: the transformation of light into an electrical message
-there is a voltage difference across the plasma membrane
(negative inside, positive outside)
1.the gate of chanrhod is closed
2.closed gate interacts with light
3.light gets absorbed
gate opens
4.cations (calcium and protons) go down the electrochemical gradient and enter the cell
5.depolarization of the membrane: inside becomes less negative
6.motion control: this inducted action potential runs down the plasma membrane of chammy and the information gets interpreted by flagella
~action potential: depolarizing→ repolarizing → hyperpolarizing → stabilizing
~action potential travels along the membrane and activates different components
~Gates opening near the eyespot causes adjacent gates to open in a chain reaction
7.flagella controls movement regarding where and if the light is present
8.gate closes
Is Channelrhodopsin is a photoreceptor?
YES
-chann = retinal + opsin
retinal: pigment (absorbs light)
-opsin: protein (a gated channel that opens or closes in response to light absorption)
-opsin alone cannot respond to light as it's a protein and does not have that ability. Which is why it is joined by retinal (retinal is the lock, light is the key and opsin is the door that is meant to open and close)
-retinal absorbs blue light best
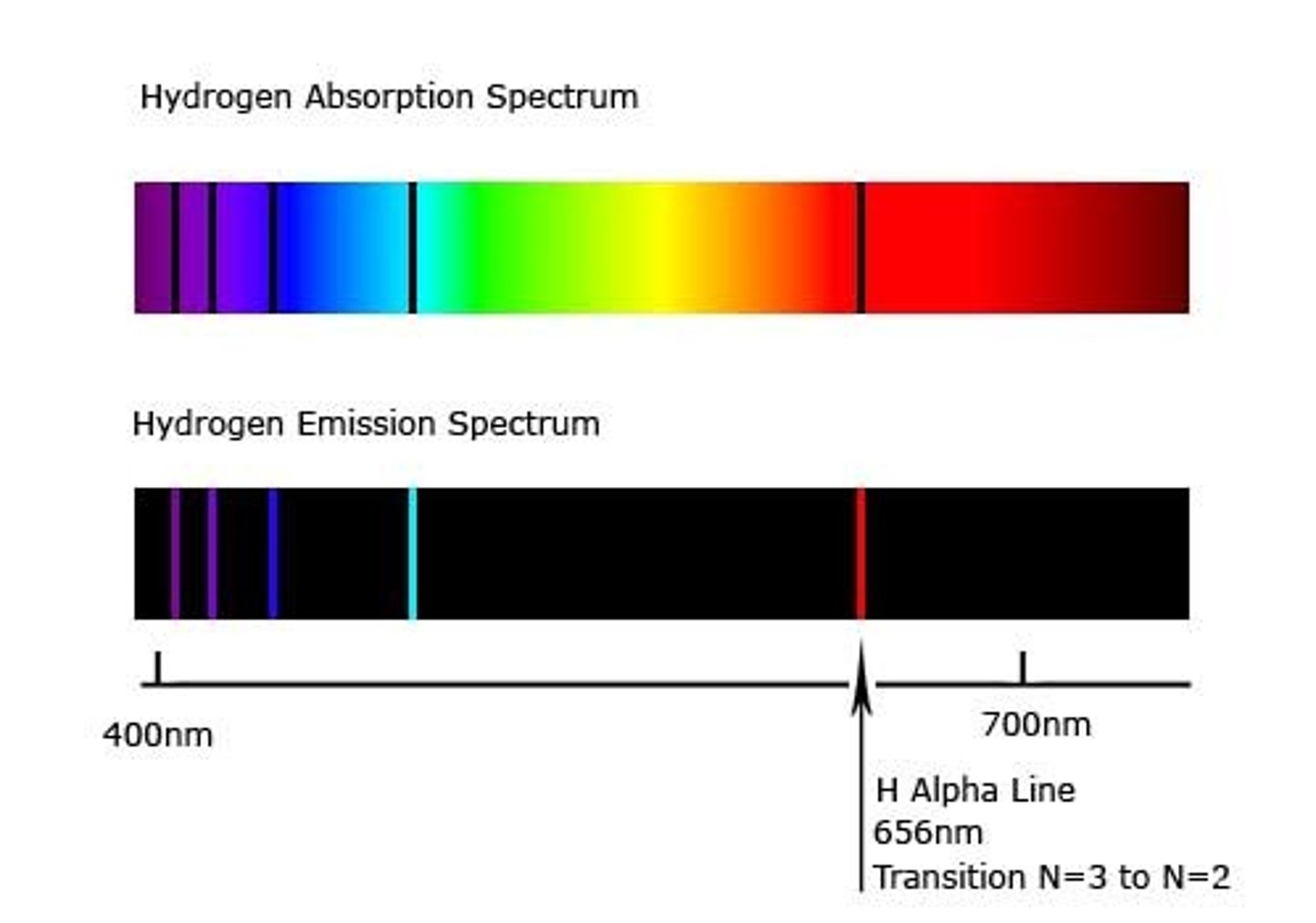
Define light absorption and what is an absorption spectrum
-light absorption: energy of the wavelength gets transferred to an electron
-the electron only gets excited if the wavelength matches the difference in energy between its ground and excited-state
-absorption spectrum: spectrum unique to each element with colourful background and some black bands. Those black bands correspond to the wavelengths of lights absorbed by the material from the entire range of light.
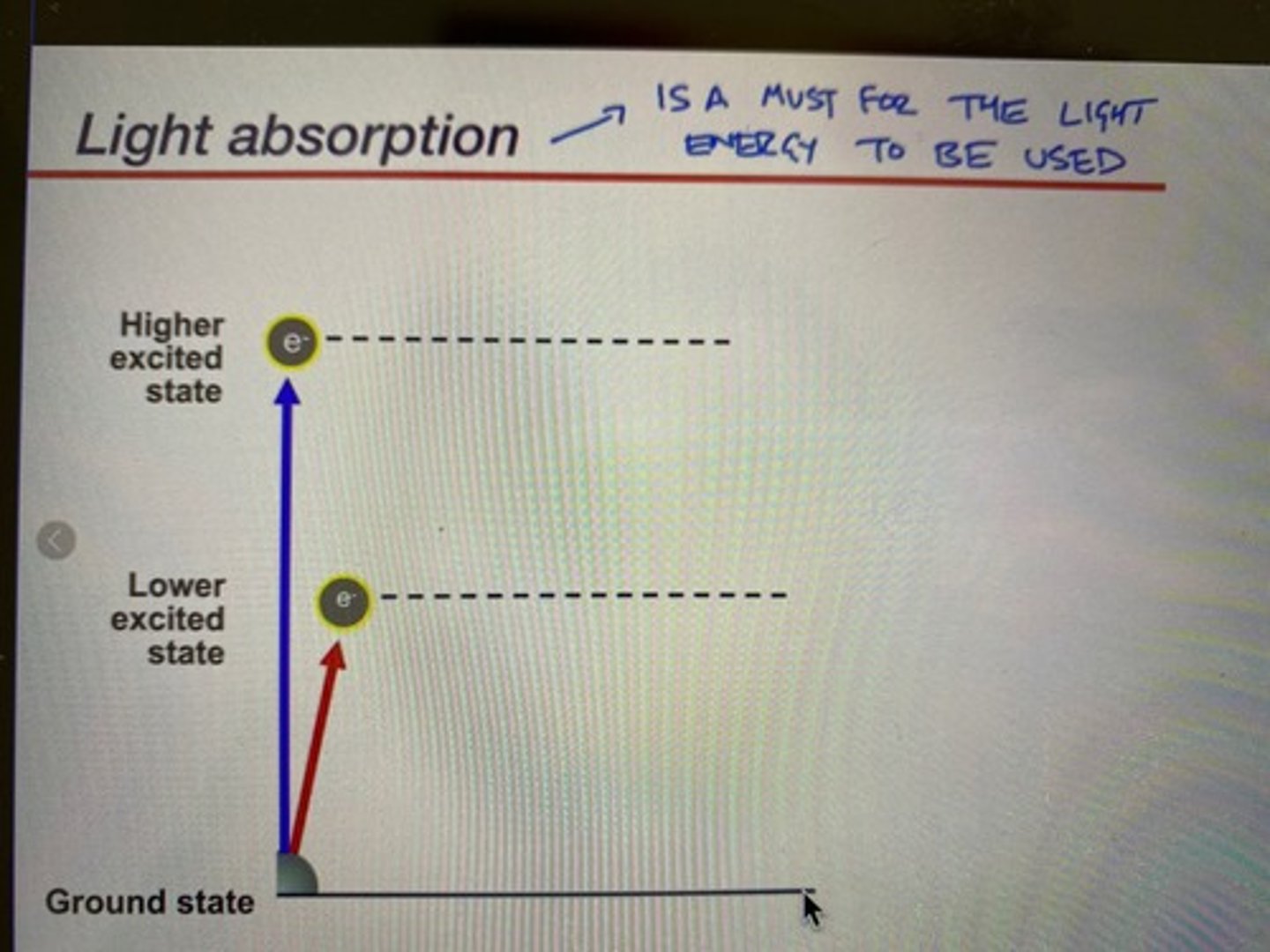
Fundamental principles of light absorption by a pigment (chlorophyll)
-Chlorophyll can absorb both red (lower excited state) and blue (higher excited state) wavelengths
-(remember that retinal only absorbs blue)
-blue light has more energy, so it excites the electron to a higher state: this state decays shortly as heat because it is very unstable
-red light has relatively less energy: more stable, an electron excited by a red wavelength will go down during the electron transport chain
-for light absorption to occur, the light hitting the photoreceptor must have the exact amount of energy to excite the electrons to a certain level
Why doesn't chlorophyll absorb green light ( added more for clarity)
-complimentary colours: since red is the one used by chlorophyll, and the complementary colour of red is green, we see chlorophyll as green
-chlorophyll won't absorb green light as it is green itself
-plants use light to make ATP etc. from the lower excited state electron (red) to power the Calvin cycle
~they don't use blue light as the blue excited state decays to red excited state anyway due to it being unstable
~since the the chlorophyll absorbs the colour red, it reflects green due to complementary pairings of colours in physics and chemistry
Fundamentals of photoisomerization of retinal and its link to a change in opsin shape (conformation)
-photoisomerization: changing from one isomer to another by the energy of light
-isomer change of retinal drives the conformational change of opsin "channels" , opens the channelrhodopsin pores
~the specific double bond of all-trans-retinal that can only be excited by blue light gets broken
~the formed double bond is a single bond for a few moments
~single bond can move around (unstable)
~the unstable single bond decays
~new cis double bond formed
~trans retinal=close gate chanrhod
~cis retinal=open gate chanrhod
~essentially breaking and swiveling of a specific bond to open an close the channel gate due to photon absorption
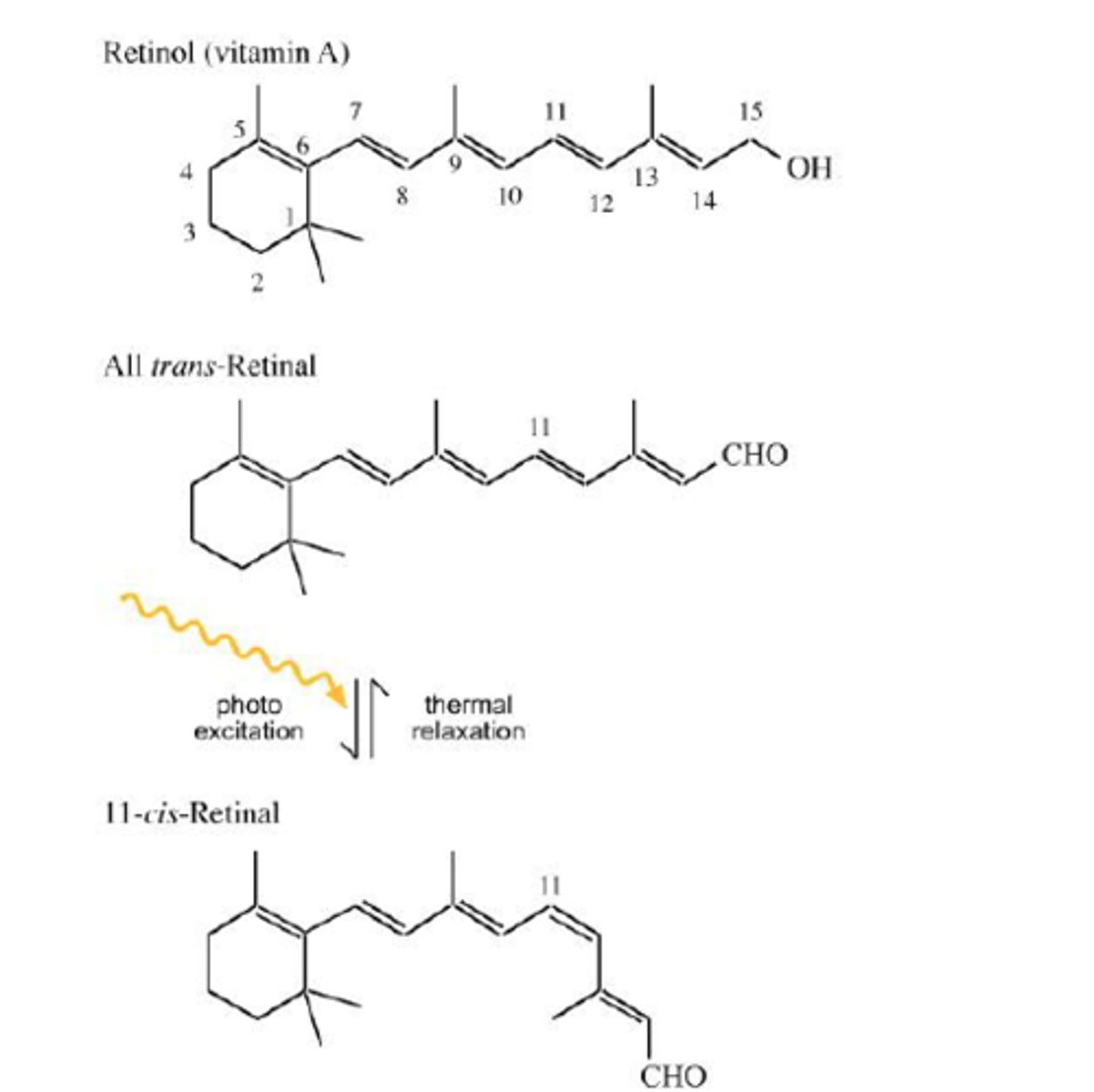
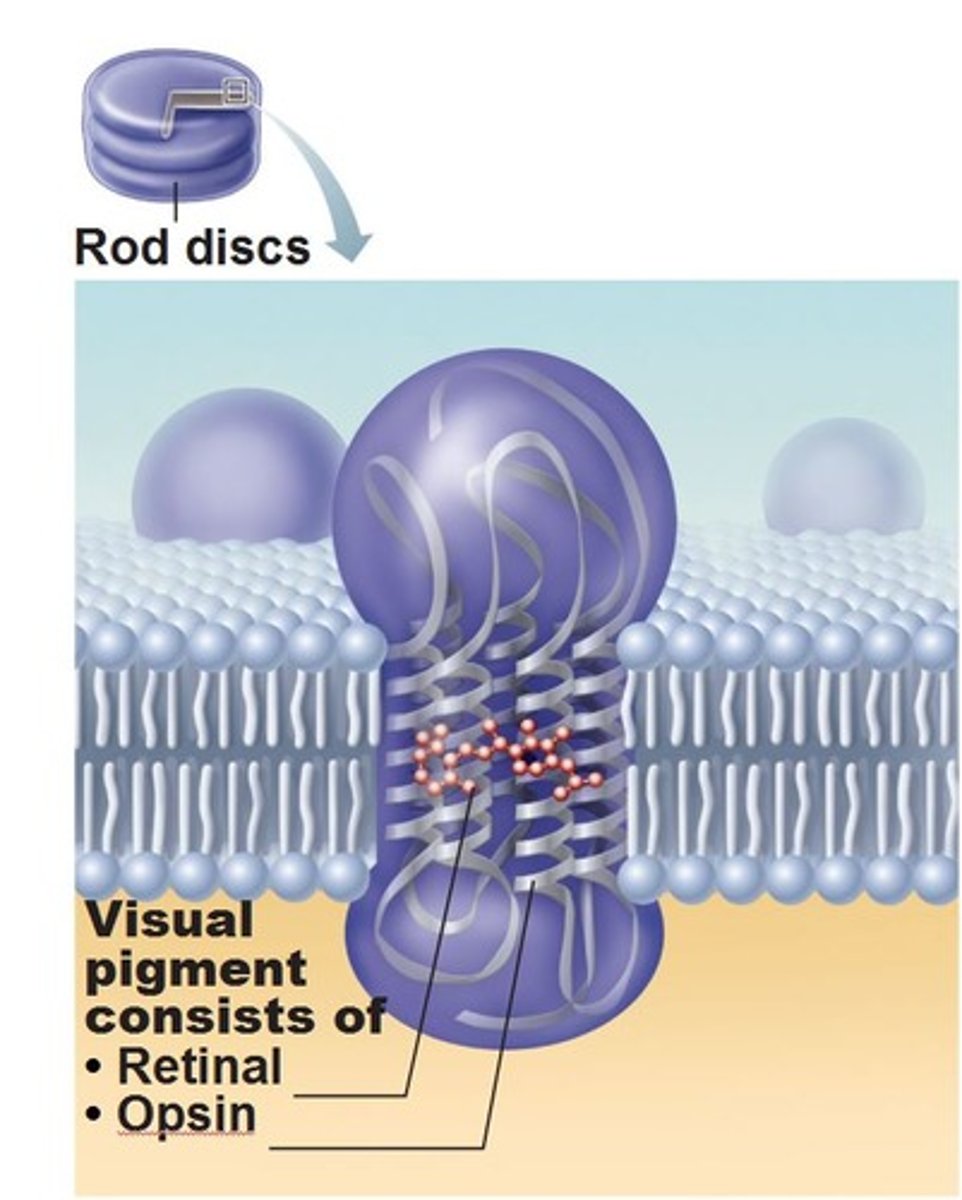
The basic structure and features of a ROD CELL (one of the two major photoreceptor cell types found in human eyes)
-rod cells in the human eye are modified neurons (and technically modified cillium)
-rhodopsin (photoreceptor in eye) = opsin (protein) + retinal (pigment) -> similar to chanrhod in chlammy
-responsible for vision at low light (does not mediate colour vision)
-function:
~pigment absorbs light
~rhodopsin changes shapes
~triggers downstream proteins (G proteins):
extracellular molecules that bind to receptors on the cell surface in order to cause an event inside of the cell
~indirect activation of change in ion current (important: rhodopsin in our eye is not a channel and cannot be used as a direct path for ions to diffuse in and out. However, it triggers the signal transduction pathway where external signals cause internal events to occur)
~ion flux generates an action potential
~the change in shape of opsin is caused by light
~the change in shape triggers a signal transduction pathway which triggers an indirect flow of ions which triggers an action potential
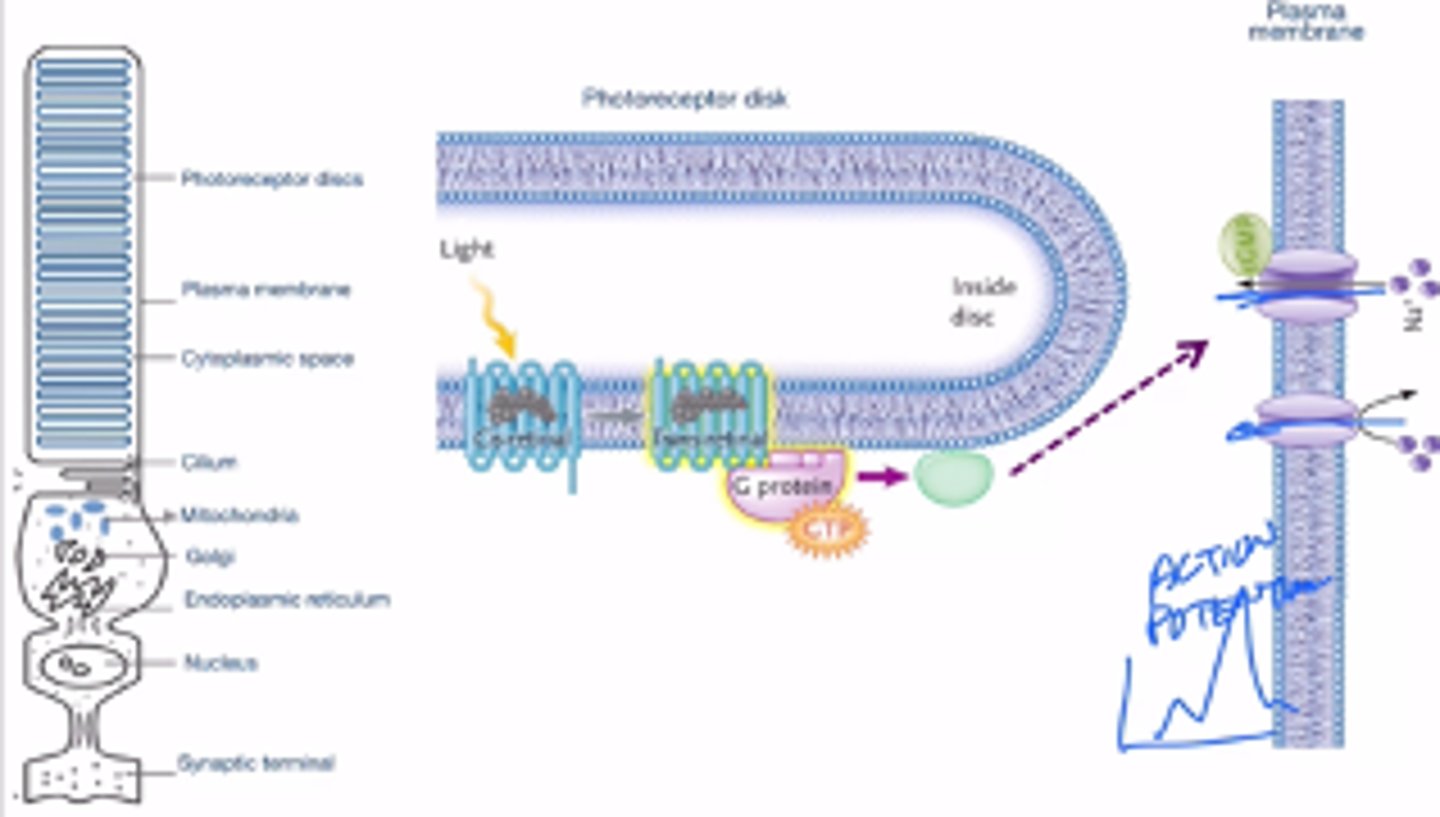
The distinction between photochemistry as it occurs in eyes and eyespots compared to how it occurs within a photosystem in photosynthesis
-photochemistry happens in photosystems and photoreceptors
-during photosynthesis, a photosystem is oxidized (loses an electron) this electron will go down electron transport chain
-In photoreceptors, photochemistry is used to do the isomerization of retinal, and in turn changes the shape of opsin
-loss of an electron occurs in a photosystem but NOT in photoreceptor
-the photochemical event in photoreceptors triggers shape change of a protein, but not in photosynthesis
-both events required excitation of an electron within a pigment (retinal for eye and chlorophyll in photosynthesis
-isomerization happens in the eye for retinal but not in photosynthesis
-phototransduction: photoisomerization occurs to open/close channels and relay info, oxidation doesn't occur in phototransduction
-photosynthesis: oxidation of chlorophyll occurs instead, excited state of oxidized chlorophyll is called Chl*, this occurs in a photosystem of chlorophyll
Key similarities and differences between the function of channelrhodopsin in the eyespot and rhodopsin in the human eye.
Channelrhodopsin Function (Chlamy)
-photons change the conformation from trans-cis
-directly activates change in ion movement
-ion channel
-protein and pigment
-embedded within the chloroplast membrane
Rhodopsin Function (humans)
-photons change the conformation from cis-trans
-not an ion channel
-indirectly activates change in ion movement through activation of a signaling pathway
-protein only
-more complex
-located inside the rods and cones
Similarities between Chlamy channelrhodopsin and human rhodopsin
-photon causes changes in conformation
-both use light for information
-7 transmembrane domains
-both bind retinal
-retinal changes shape upon light absorption
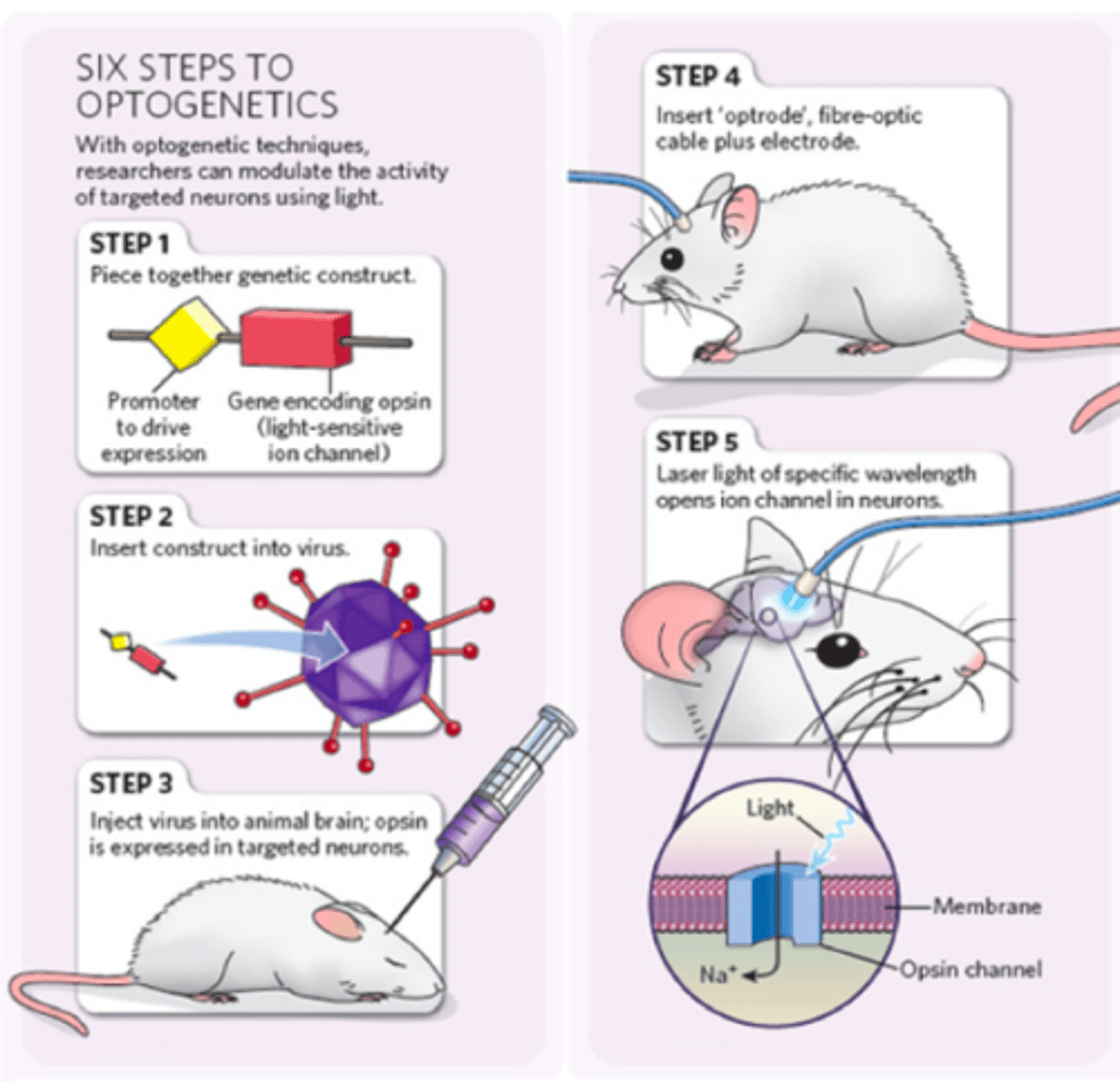
Basic understanding of optogenetics: what is it, why is it useful. What are the basic steps required for transfection and stimulation.
-express chann in the brain neurons
-using chann to activate neurons by photons using blue light
-chann can use the K+/Na+ ion channels involved in traditional action potentials by blocking or allowing ions to cross into the neuron, and hence do the same function in the neuron
-mapping the brain (designating what area if associated to what function)
-transfection: making a foreign gene system and inserting it into a new organism
~remove the native chlammy promoter from the coding region of the chlammy chan gene that makes the promotes
~replace the promoter with a mouse promoter
~gene expression will happen (chan will develop in mouse)
~put the new gene (with a new promoter) in a virus that specifically infects mouse brain cells, this is called transfection
~the virus will insert its genome (including the chann gene) into the mouse's brain aka neurons
~trigger chann by shining blue light on it
~track the position of the blue fiber optics and record changes in physiology and behaviour of the mouse
What is LUCA, why do we know it existed, relationship to first forms of life
-LUCA: last universal common ancestor
-hypothetical: there is no concrete evidence for its existence, but we have good proof to believe it
-characteristics shared by all life that make us believe LUCA existed:
~DNA
~RNA
~lipid membrane
~glycolysis (metabolic pathway that creates ATP)
~ATP
~ribosome
~proteins
-LUCA may not have been the first form of life that ever existed, but it is the most recent form of life that had all the characteristics shared by all forms of life today
3 stages of prebiotic evolution, how much is known about each
-What is needed to produce a living cell?
-DNA, RNA, proteins... must've been formed before the first ever cells
-abiotic synthesis
~when looking at cells today, many of their components were formed before the actual cell itself (like ATP)
~we are studying the synthesis of these biologically important molecules before life (living cells) actually existed
-metabolism
-formation of cells
-three stages of evolution:
~geophysical stage: a study of the composition of the earth and atmosphere (what are they made up of that sustain life?)
>the early atmosphere was a reducing atmosphere with many reducing species, this is important because all biologically important molecules are reduced
~chemical stage: how were the building blocks of life (aka molecules, atoms) ever synthesized when life (living cells) didn't exist, through abiotic synthesis
~biological stage: how did the building blocks derived from the chemical stage organize into becoming living cells (WE KNOW NOTHING ABOUT THIS!!)
Conditions in the primordial atmosphere, why were they conducive to the development of complex molecules
-the early atmosphere was reduced (electron-rich): contain a lot of carbon-rich molecules
-we say carbon-rich molecules are reduced because there is a lot of C-H bonds in a carbon-rich molecule
-the more C-H bonds are in a compound, the fewer other bonds with other atoms more electronegative than carbon will exist (such as C-O). Therefore, the C-H bond will keep all the electrons and won't lose them to more electronegative atoms
~since O_2 is a strong oxidizing molecule it likes to steal electrons, so the absence of oxygen was very important in regards to allowing C-H bonds to form since carbon combusts easily
-because of the absence of oxygen and abundance of C-H, the place was set perfectly for the formation of other relevant molecules (most biologically relevant molecules are reduced)
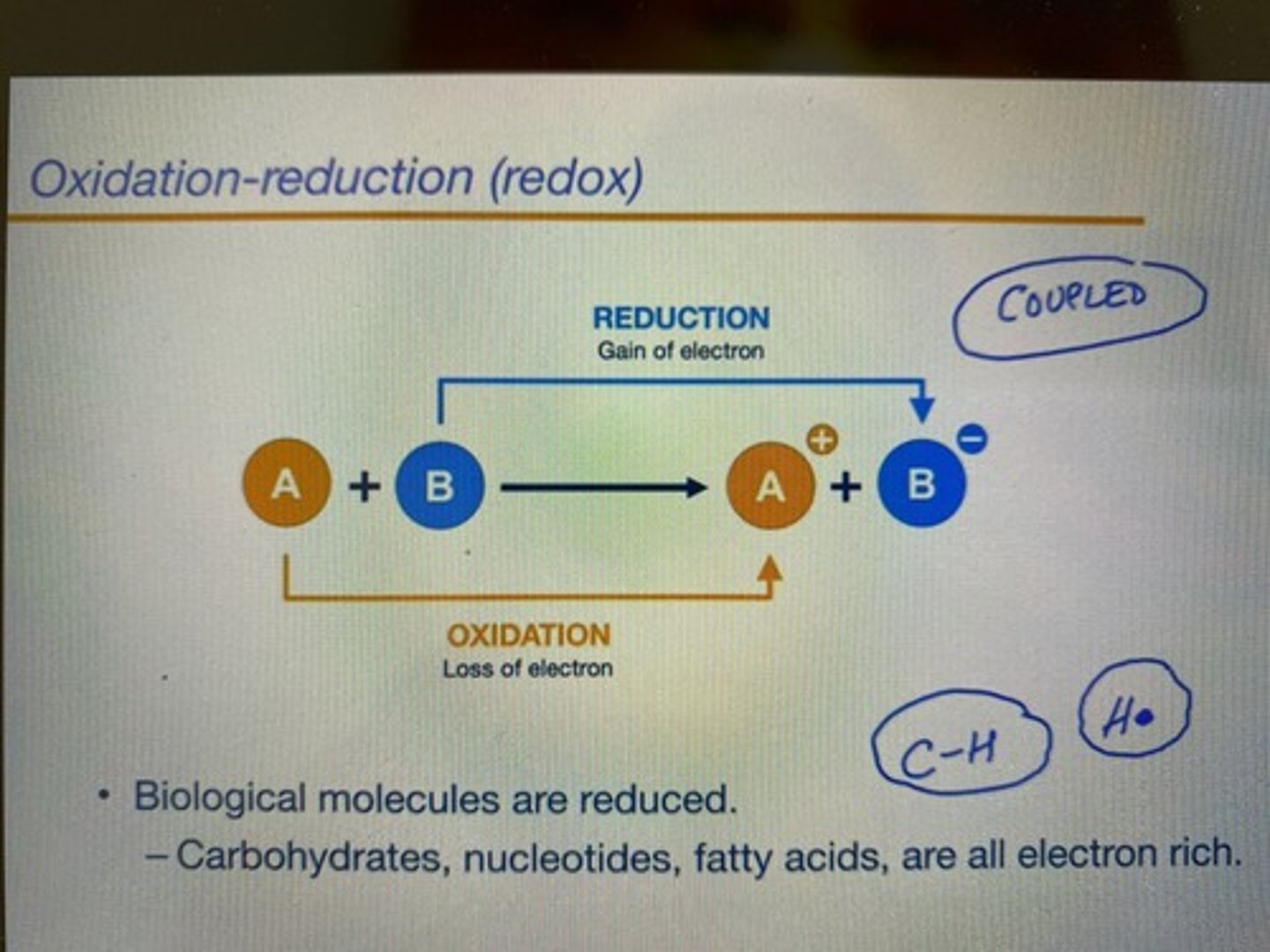
Basics of Oxidation - Reduction reactions
-coupled reactions (always happen together at the same time)
-oxidation: loss of electrons or hydrogen (increase in oxidation state)
-reduction: gain of electrons or hydrogen (decrease of oxidation state)
-tip: an oxidizing agent (molecule) oxidize OTHER molecules, so it becomes reduced itself because it wants more electrons.
-O2 is an oxidizing agent that gets reduced into H2O (H2O is the reduced form of O2)
What was the Miller-Urey experiment? What did it show, didn't show.
-the question that the experiment tried to answer: can we make organic molecules from inorganic molecules? (it tried to explain how life and biological molecules were formed in the absence of life and living cells)
-he tried to recreate the chemical conditions of early life (only CH4, NH3, H2O and H2 present in the early atmosphere)
-put the gases in a tube, and induced a spark discharge (spark was to recreate the source of energy or lightning that occurred in early life)
-included boiled water to recreate water vapour that was also present on Earth
-outcome: several of the protein precursors necessary for life were created just by these conditions alone, showed that an intact biological system isn't needed to create organic molecules, only a reducing atmosphere and a source of energy (ie. lightning)
-50% of either enantiomer of each molecule was created
-a limit of the experiment: life is homochiral (only one enantiomer of each molecule is produced) but the experiment produced half of each
-another limit of this experiment also was that RNA and DNA was never created either
The distinction between racemic and homochirality
-racemic mixture: 50-50 mixture of both enantiomers (like Miller experiment)
-racemic does not happen in real life biology
-homochirality: only one enantiomer is produced by living systems (almost never both together at the same time) this is the form that occurs in biology
Importance of homochirality to the evolution of life
-life is homochiral
-L (left-handed): amino acids
-D (right-handed): sugars
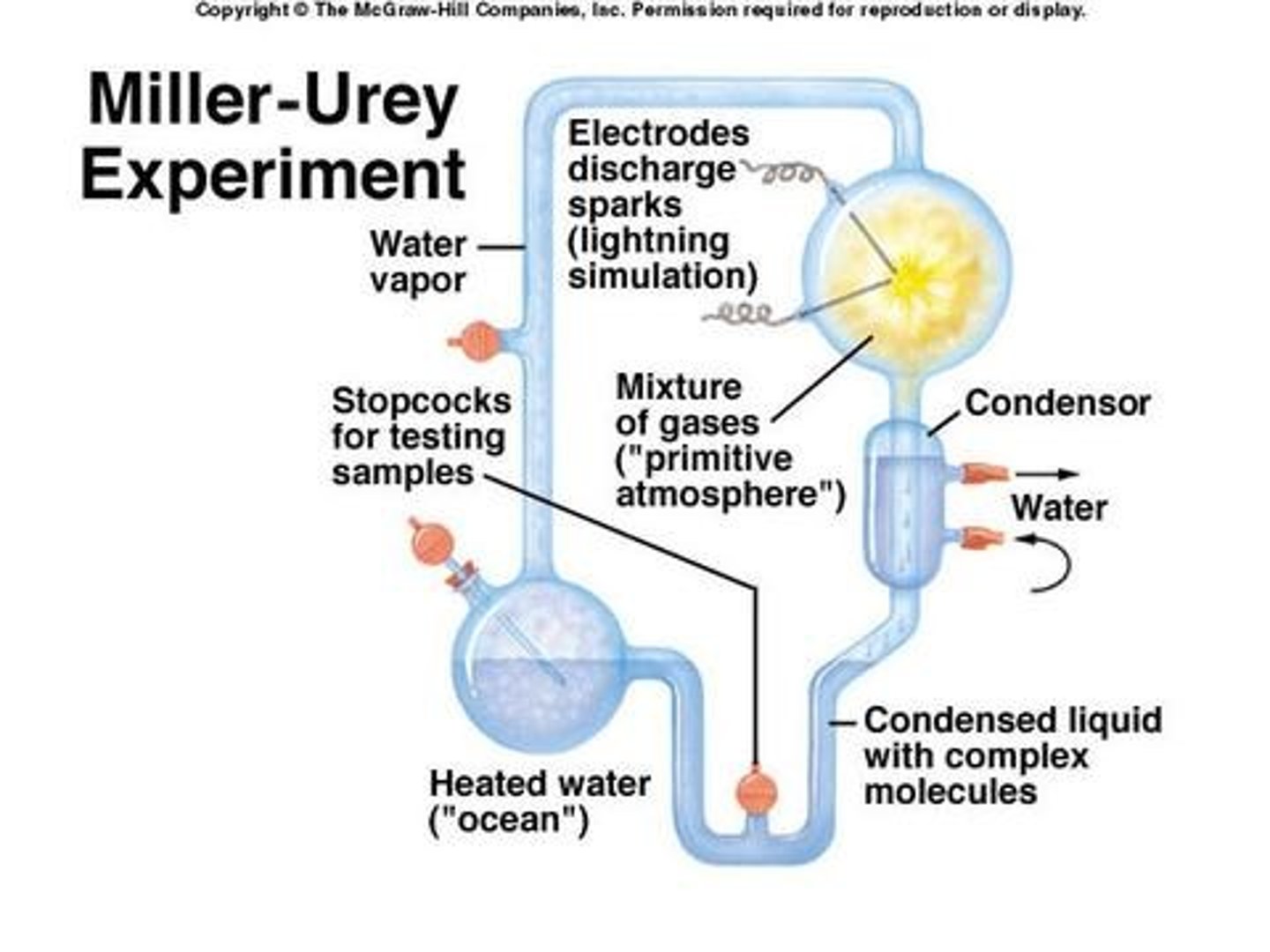
-Enzymes interact differently with substrates depending on which enantiomer they are (they are not programmed to work with both since enzymes work like a lock and key and can only use one type of chiral molecule)
-If an organism wants to have both enantiomers (substrates) of a substance, it needs enzymes that can interact with both of them
-having both enzymes is a complex process and not favourable, so most likely eliminated by evolution through natural selection
-exception: limonene in humans (we have two different receptors that can tell both enantiomers apart, so we can smell both oranges and limes)
The central dogma of molecular biology
-information processing only goes one way and not the other (DNA -> RNA -> protein)
-each arrow is the work of enzymes
-DNA, RNA and proteins are polymers
-the Miller experiment only produced the building blocks and not the actual polymers (suggesting that under initial conditions, only monomers could be made)
-question: how did these chained polymers form? how did they even develop in the absence of enzymes (enzymes came first or polymers it's like chicken or egg)
-hypothesis: polymerization happened slightly by mineral surfaces ( rocks were formed around them so they were pushed together and kinda forced to polymerize)
-you can also create small polymers using clay, small chains of RNA can be made this way
Structural similarities and differences of DNA and RNA
Both:
-phosphate group (5 carbon sugar)
-the linear sequence of amino acids (order of amino acids) codes for some specific information
-use H bonding (RNA h-bonds to itself to become circular, DNA h-bonds for the amino acids of each strand to interact)
-they are a polymer of nucleotide
DNA
-double-stranded
-amino acids: A, G, T, C
-has the genetic information necessary for the development
-stores genes much better as it's double stranded and has 2 copies of itself, giving it a high fidelity
-because it has no loose, unbonded ends, it is much more difficult to degrade via enzymes in the cell
RNA
-single-stranded
-amino acids: A, G, U, C
-can have a 3D (mainly circular) shape
-mainly involved in protein synthesis and sometimes gene regulation
-can bond with itself to create these 3D structures and provide structural support within the cell
-can catalyze as well but not very effectively
What is a nucleotide?
-building blocks of DNA and RNA
-composed of sugar base (either ribose or deoxyribose nucleic acid), a phosphate group and a nitrogen base that differentiates the nucleotides
What is a ribozyme, what roles do they serve in cells
-some enzymes are catalytic, as they can fold/act like a protein
-catalyzers that are not proteins (they act as enzymes)
-they are RNA under the 3D shape
-ex1: ribosome. the main function part of ribosomes used for translation of RNA into proteins is made of ribozymes (RNA), although the rest are proteins (⅓ protein and ⅔ RNA)
-ex2: spliceosome. protein/RNA complex (made from SnRNA) that slices introns and brings exons together
What are the advantages of Protein over RNA and DNA over RNA?
Protein over RNA:
-proteins are more diverse (RNA is built around 4 (AGCU) bases while proteins can use a combination between 20 amino acids). Therefore, they can adapt even more 3D structures(hence even more diversity because each 3D structure has different properties)
-enzymes (protein catalyzers) are faster and more efficient than ribozymes (RNA catalyzers)
-proteins are also a much more powerful structural molecule compared to RNA (more stable)
DNA over RNA:
-better energy storage in DNA because it's double-stranded
-DNA fidelity: if one strand has a mistake, the second one will be correct, the fact that the nucleotides have ot base pair with each other within the double helix also allows for proof reading to occur due to inconsistencies in the strands, allows for proofreading of mismatches, leading to greater genetic fidelity
-DNA is less likely to breakdown or gets degraded due to the lack of the oxygen molecule in its backbone
Chemically, why is RNA more susceptible to breakdown than DNA
-ribose has an H bonded to O (deoxyribose does not, hence its name)
-there is a lot of hydroxyl OH- groups in an aqueous environment (water is in dynamic equilibrium with H+ and OH-)
-OH- attacks the extra O on ribose to make water
-so, the RNA strand gets cut and the nucleotide gets free
-the same problem does not happen on DNA because it does not have that extra oxygen
What explains why Chlamy stops growing in the stationary phase
-at stationary phase, the chlammy sample reaches its carrying capacity (there is no more nutriments or resources to sustain further growth)
-cells need nutrients to grow (they need a lot of macronutrients and a small amount of trace minerals)
-cells need nitrogen taken from macronutrients and trace minerals to produce cofactors for enzymes to divide, without the ability to produce these cofactors, they won't be able to divide
What is in the Chlamy growth media and why (in general terms)
-a lot of macronutrients
-a small number of trace minerals
-a lot of inorganic salts in the clear media
-nitrogen
Purpose of RNA blot analysis
-all body cells have the same genes, the cells only differ in which of the genes that they express
-Blot analysis is used to see if a certain gene is transcribed (expressed) or not
-useful in cancer research
-since the transcribed version of a gene is a protein, we identify mRNA in a mixture to see if a gene gets expressed
Blot analysis: Basic understanding of all the steps
-take two samples: samples of the DNA of the same organism put in two different environments to see the gene transcriptions of both samples of a specific gene (how that gene expression would differ in different environments)
~the example used: bacteria grown in glucose vs galactose
-extract the mRNA from both samples
-purify mRNA (with different techniques)
-put the mRNA of both samples on an agarose gel that separates different mRNA
-different mRNA separate based on size
-transfer the mRNA onto a piece of paper (because the gel is thick and it's hard to interpret results)
-mix a probe of DNA (labelled single-stranded galactose DNA ) with the transfer mRNA on the paper
-over time, gene probe will complementary base pair with galactose mRNA
-because one of the samples will have more galactose mRNA than the other (because it had higher gene expression) there will be more DNA probes binding to it
-put an x-ray sheet on top of the new mix (mix with the bound DNA)
-one of the samples will show a black line
the line represents the bounded DNA probe to mRNA
-the bigger and darker the line, the greater is the level of transcription as there is literally more mRNA that is present
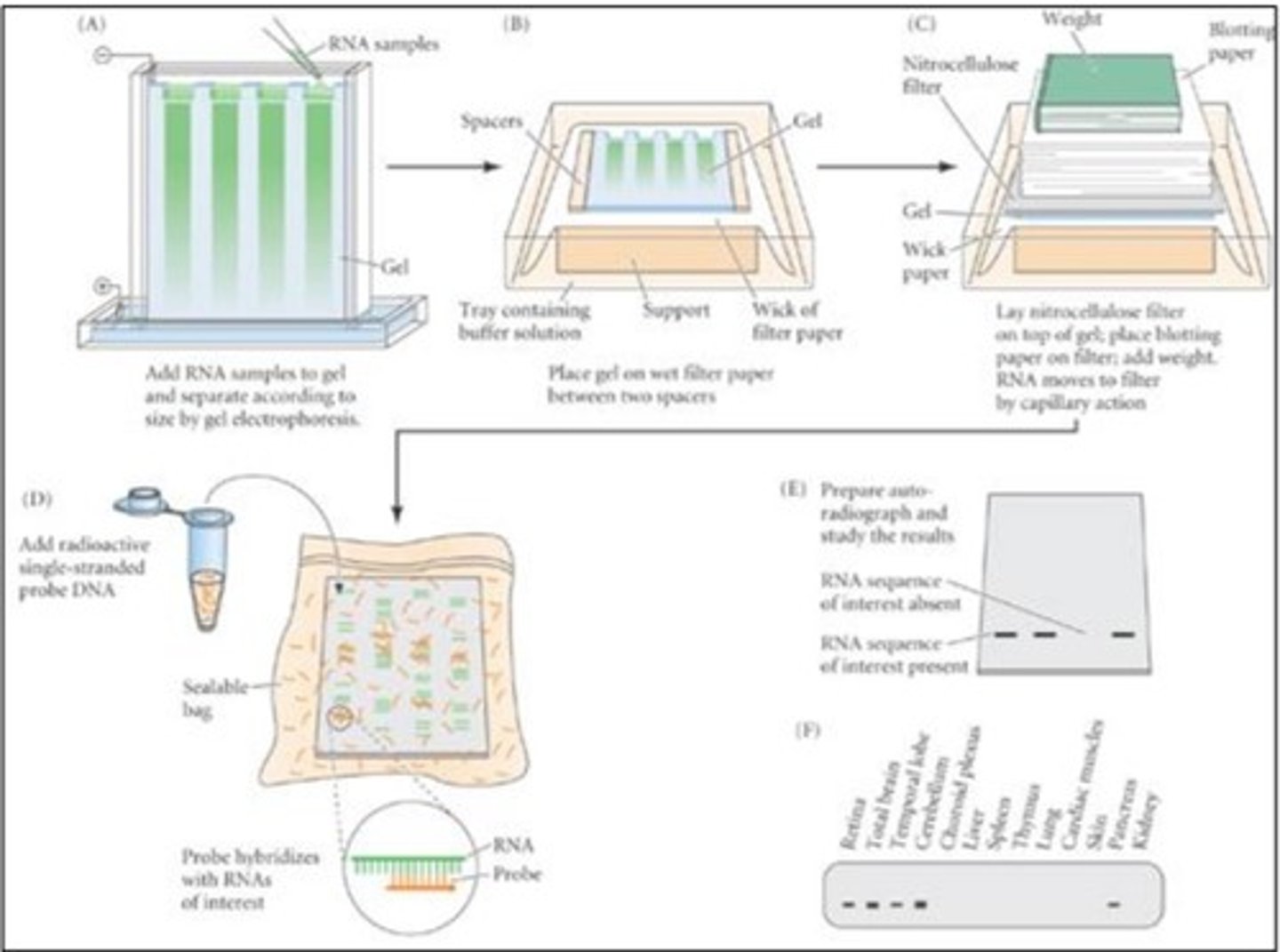
Blot analysis: Importance of the probe being "labelled" and single-stranded
-single-stranded: it should be single-stranded so it could complementary base pair to the corresponding mRNA sequence. If it's double-stranded it would just not bind to anything else.
-labelled: fluorescent labelling is needed because it allows the mRNA of interest to be distinguished among the many other mRNA's in the solution
Blot analysis: explain the concept of "complementary base pairing" that occurs between the single-stranded DNA probe and the mobilized RNA molecules (the GAL transcript)
-mRNA is made from complementary base pairing itself to one of the strands of a specific gene (this means that one of the DNA strands is actually going to be a copy of the mRNA)
-The DNA probe will complementary base pair to the mRNA (putting the complementary nitrogen base for each nucleotide)
-This allows for the probe to attach itself to the RNA and act as a marker to aid in identification
Definition of the term: GENE
A sequence of DNA (not the whole DNA!) that codes for an mRNA that will later be translated into a protein
Different types of RNA
-mRNA is messenger RNA, it can be transcripted to be able to translate for functional proteins that run our bodies and cells
-tRNA is transfer RNA it allows our bodies to build functional proteins based off of the genes encoded in the mRNA (brings nucleotides to ribosome)
-rRNA is ribosomal RNA, it helps build the ribosome which can facilitate tRNA to interact with mRNA and essentially translates the mRNA into functional proteinson
Genomics, transcriptomics, proteomics, metabolomics....what are these and what is the usefulness of each...(in broad terms)
genomics:
-all the genes of an individual, to look at an entire genome profile and identify the genetic risks that it carries within
transcriptomics:
-study of the transcriptome—the complete set of RNA transcripts that are produced by the genome, under specific circumstances or in a specific cell
-catalogue all species of the transcript, including mRNAs, non-coding RNAs and small RNAs; to determine the transcriptional structure of genes, in terms of their start sites, 5′ and 3′ ends, splicing patterns and other post-transcriptional modifications
-if a gene is expressed in a transcript—> a corresponding mRNA is made
proteomics:
-large scale study of proteins
-important because most diseases are manifested at the level of protein activity
-the mRNA is making the functional proteins
metabolomics:
-large-scale study of metabolites
-metabolites are intermediate of end results of catalyzation reactions (metabolism)
-metabolome: low weight (smaller) metabolites
-study of all metabolites like ATP, hormones, sugar that lead to an expression of a trait due to a gene
Aspects of gene expression: transcription, translation, mRNA and protein breakdown (decay)
-transcription: gene-> mRNA
-translation: mRNA -> protein
-protein breakdown: all molecules break down (have half-lives). Protein breakdown is the last step as at that protein the protein has fulfilled its function and can be digested by the enzymes in the cell
~in a human, RNA has a half life of about 8 hours and proteins have a half life of about 46 hours, note the discrepancy, this is important in regulating proteins and cellular function
~RNA is generally more unstable due to the HO- groups that attack the ribose, there are also specific enzymes that break down mRNA called ribonucleases or RNases
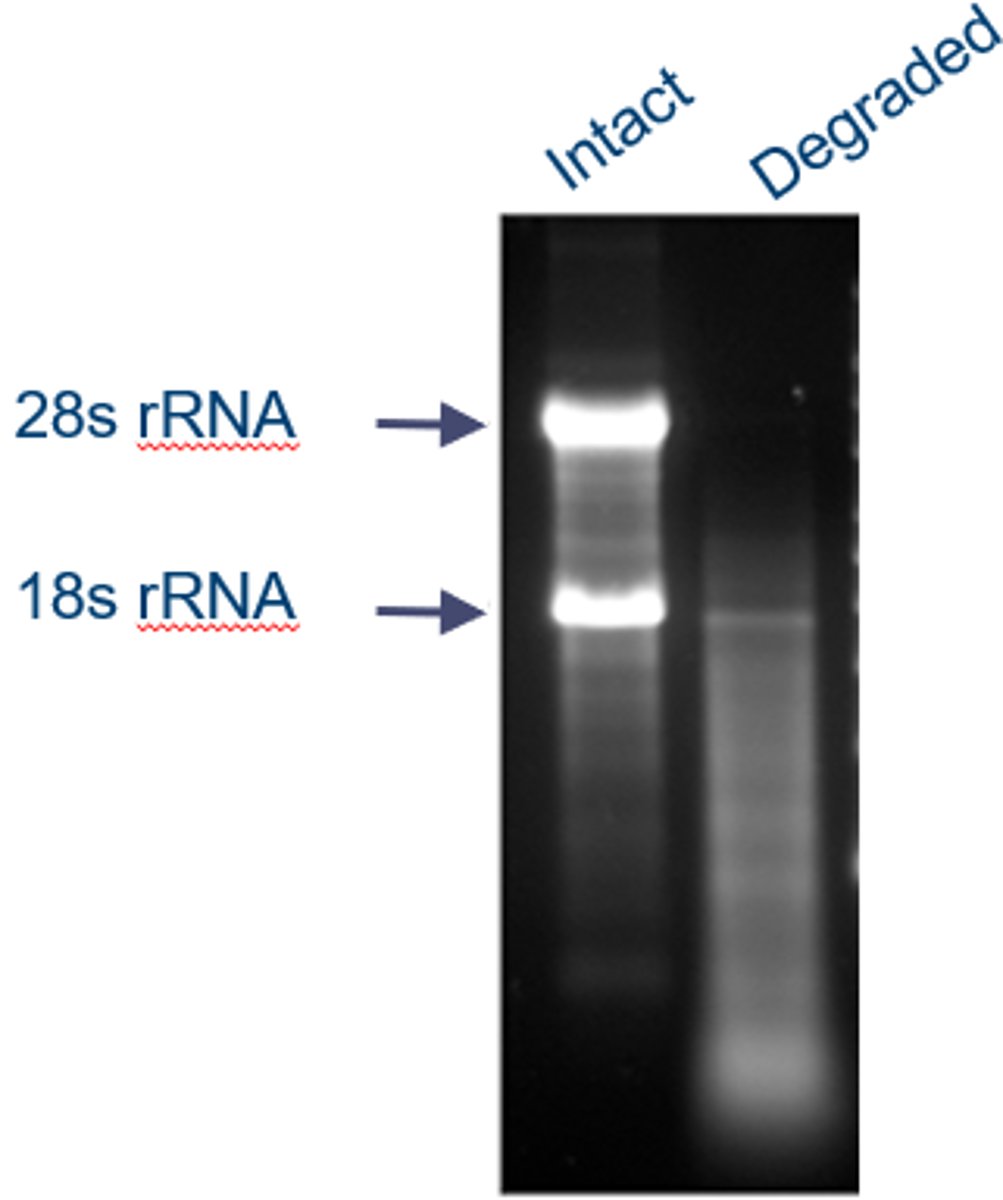
Identify intact versus degraded total RNA
-within cells, tRNA and rRNA are stable
-mRNA is unstable because:
~OH- attacks the extra O on ribose
ribonucleases enzymes attack mRNA and break it down
~as seen on northern blotting: (the degraded RNA is in the presence of RNAase)
-when an RNA is degraded, there is a gradient of the fluorescent probe, no strong bands of any sort and no clear readings off the Northern Blot can be made
-when an RNA is intact, very strong and clear bands of the probe can be seen
-RNA degradation is useful because it allows us to get rid of defective mRNA and not make defective proteins. It is not bad for the cell because there are a lot of mRNAs that get produced so degradation of some does not affect overall performance. Also, mRNA is constantly being made as genes are always being transcribed so it's degradation allows for mRNA not to be oversaturated.
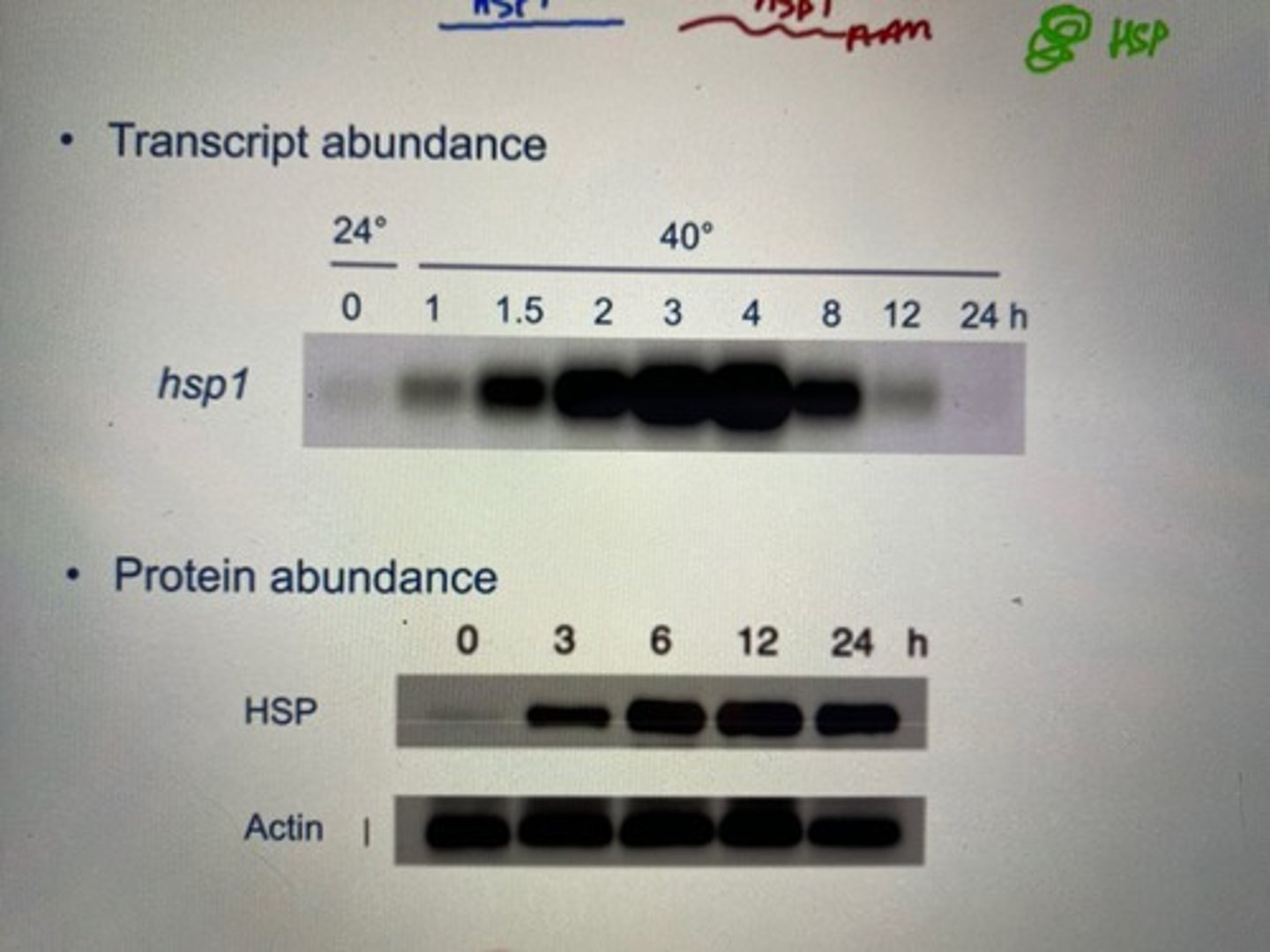
The distinction between constitutive, induced and repressed gene/protein expression. Think about how the rate of synthesis and decay of mRNA/protein can be altered to change expression levels (relate with HSP1 in chlamy)
-When put in stressful environments (ex. change in temperature), chlamy or other organisms can adapt by switching some genes on and off to make proteins or stop making proteins in order to survive the environment
Chlamy:
-when looking at transcript abundance with northern blotting:
HSP1 (heat shock protein) transcript abundance (rate of mRNA making) increased as the temperature approached 40 degrees.
-Looking at protein abundance of the same gene, we notice that the peak of protein abundance comes AFTER the peak of mRNA abundance.
-This is because mRNA is produced before protein. So mRNA synthesis reaches its max (we see it in northern blotting on transcript abundance), and then all this mRNA gets translated into protein (so protein abundance increases on northern blotting)
three types of change is possible for genes when put in a changing environment:
-constitutive: no change in protein abundance (for example, actin's abundance stays constant throughout change in temperature). This is due to house-keeping proteins that maintain function of that protein so that it does not change under stress.
-induced: expression of protein is higher (like HSP1 in chlammy) due to an expression promoter being added onto the promoter site to enable a stronger rate of transcription
-repressed: expression of the protein is less due to inhibitors being present at a promoter site due to stress, this inhibits RNA polymerases' transcription rate for protein production and thus expression levels of a trait
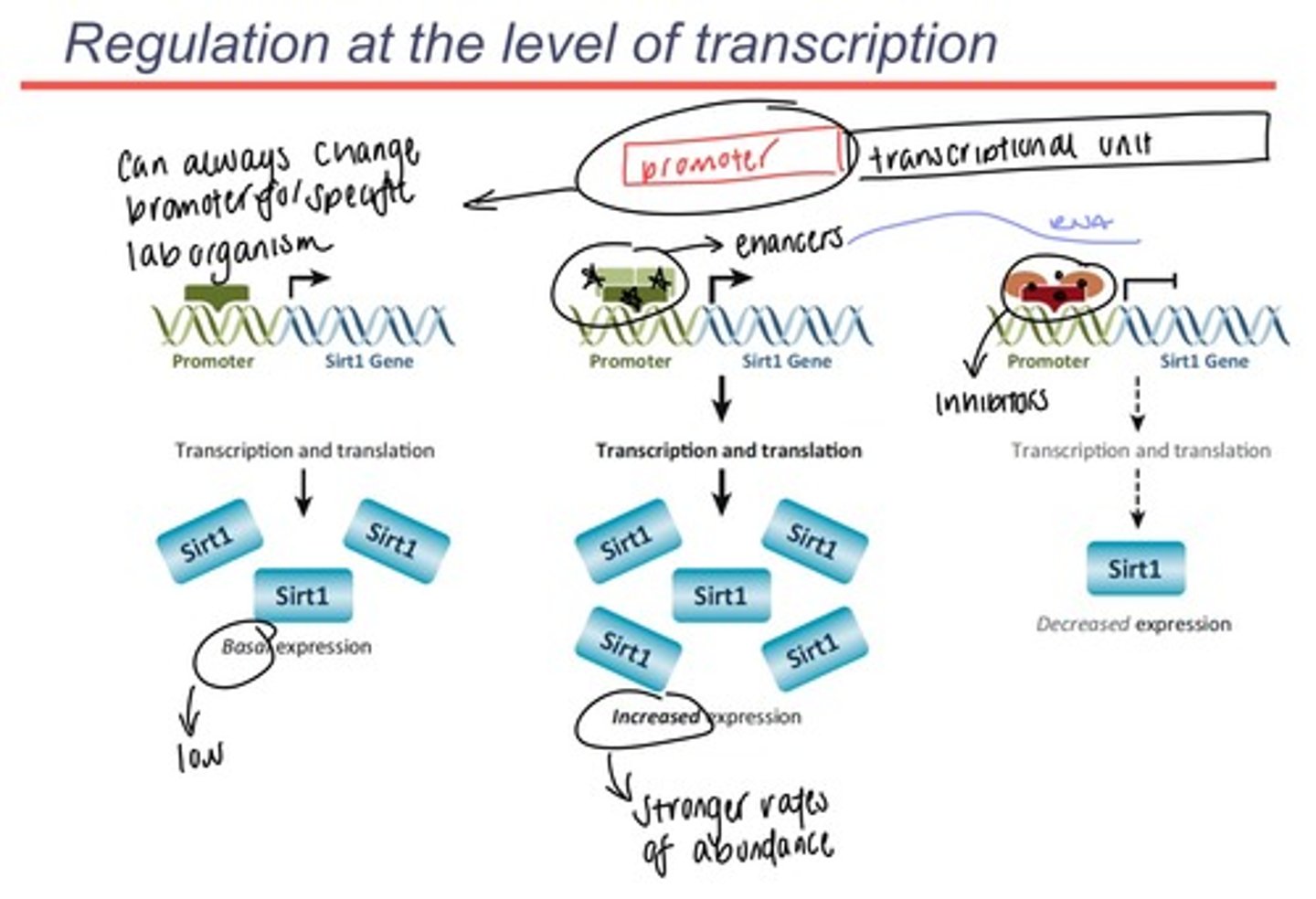
Promoter vs transcriptional unit and the factors that act at the level of transcription that alter transcript abundance.
-promoter sits before the beginning of the gene (it is what designates the beginning of a gene on a DNA sequence or in RNA)
promoter is the site where transcription factor proteins bind to and recruit RNA polymerase (RNA polymerase recognizes the transcription factors, not the actual promoter)
-transcriptional unit is the unit that comes after the promoter (gene that codes for one mRNA)
-more mRNA synthesis means more protein synthesis
-to enhance protein synthesis: add enhancing factors to the promoter and make more mRNA when gene expression is induced
-to reduce protein synthesis: add inhibitors to the RNA polymerase of the specific mRNA gene that blocks the action of RNA polymerase this is called repression
-you control transcription by controlling what happens at the level of the promoter
Factors that act as the level of mRNA stability that alter transcript abundance.
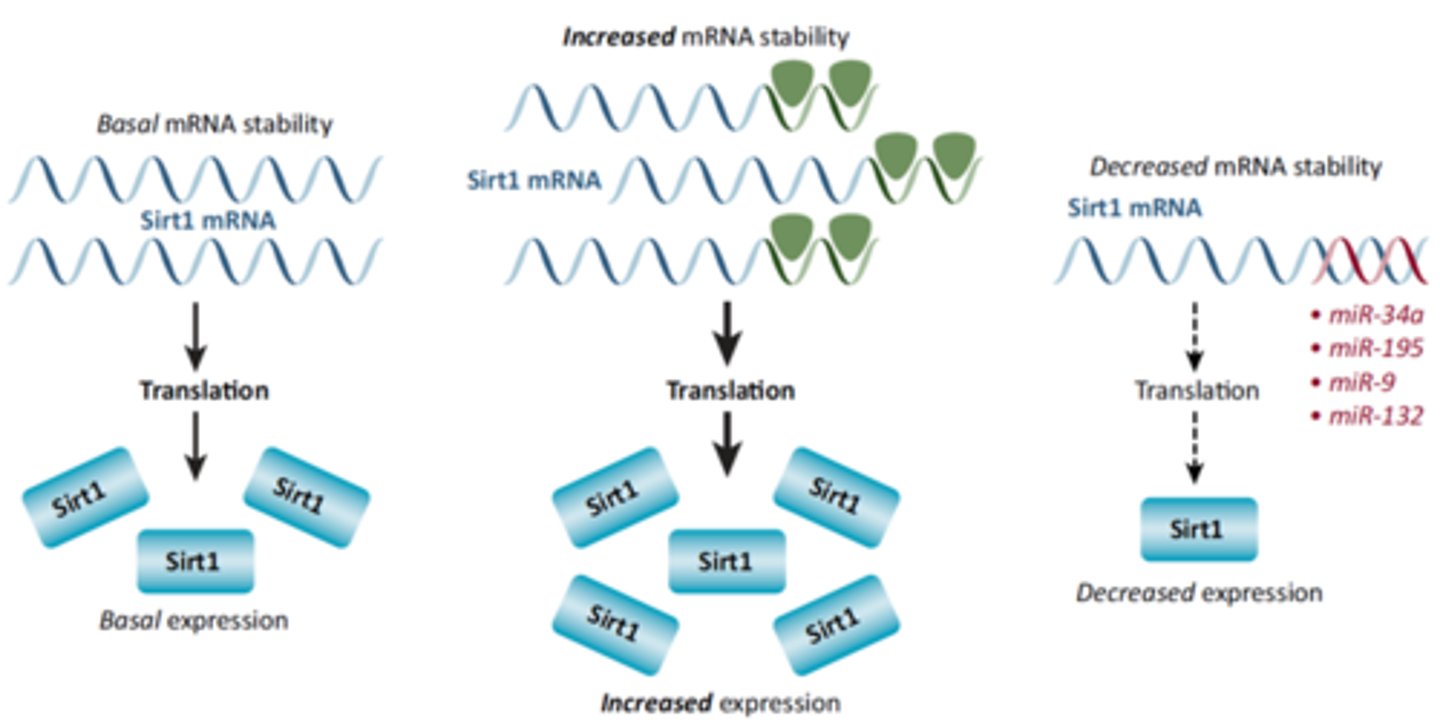
ANOTHER WAY OF REGULATING PROTEIN SYNTHESIS: REGULATING MRNA STABILITY
-the more stable mRNA is, the more it is going to survive inside the cell and the more it is going to be translated by ribosomes to make more proteins
-we add stabilization factors to pre-mRNA, like a 5' cap and 3' poly-A tail that increases stability (cycle 7 content)
-you could add protein factors that increase stability to increase protein synthesis (for example by adding a longer poly-A tail)
-you could decrease protein expression by destabilizing mRNA and making it survive a shorter time
-we can increase/ decrease the stability of the mRNA via protein factors and inhibitors as well
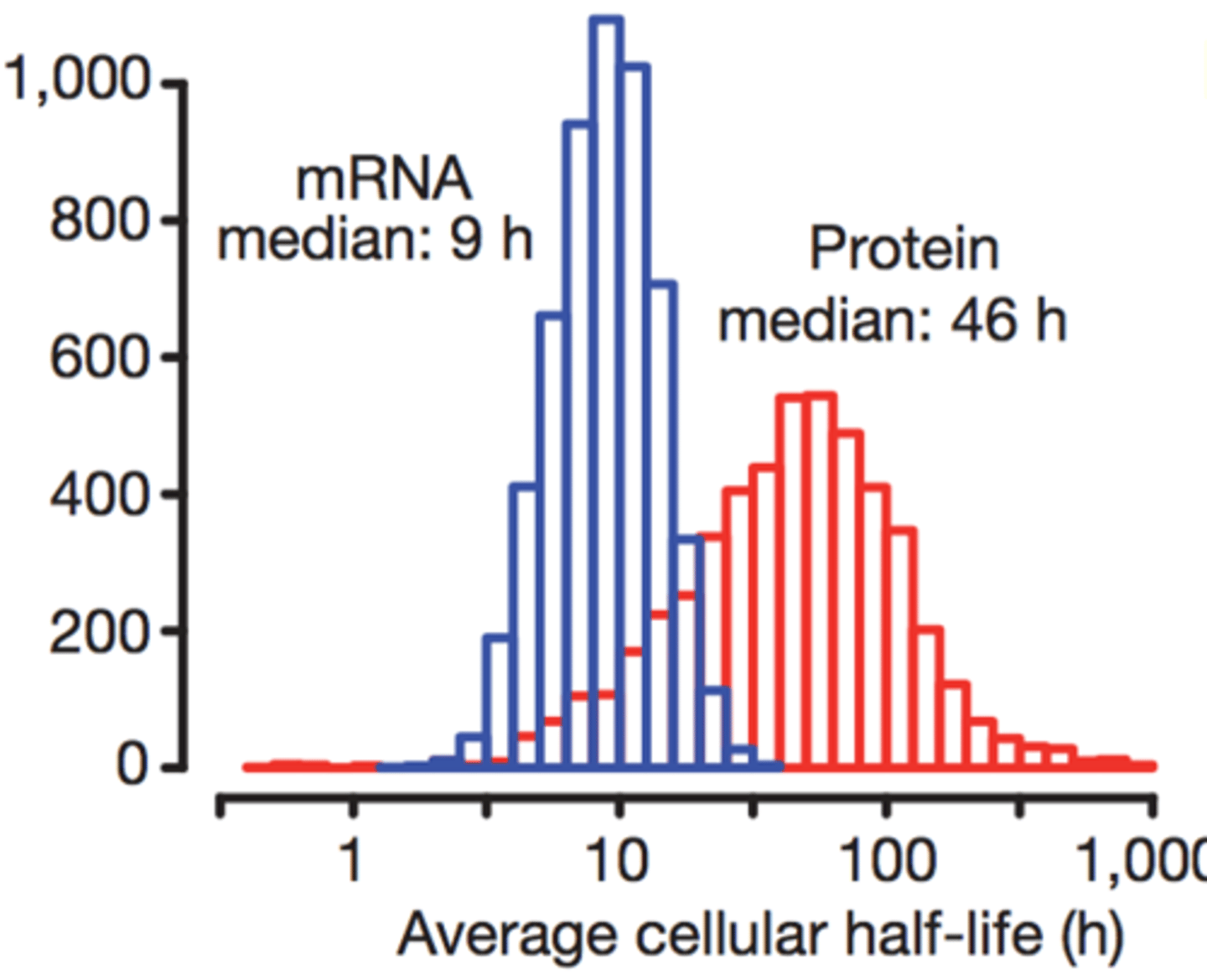
Rough idea of the median half-life of a mammalian mRNA, protein
-all molecules have half-lives (they eventually get degraded)
-mRNA half-life in humans: 9h
-protein half-life in humans: 46h
-protein-half is usually higher than mRNA because there are a lot of enzymes that cut down mRNA in order to prevent an overflow of mRNA (as it is constantly synthesized) and we also don't want mRNA to get mutated. Proteins, however, can't get mutated.
-the decay of mRNA also allows the cell to regulate the rate of protein creation/ trait expression, if mRNA never decayed, then the rate at which proteins would be transcribed would stay the same and would allow for the cells to adapt to different environments around themselves
Rough idea of the median number of molecules of transcript of a particular mammalian gene and the number of molecules of protein for a protein-coding gene
-mRNA median in one cell: 17
-protein median in one cell: 16000
-there are way more proteins than mRNA in one cell, because there are multiple proteins made from the same mRNA by the fact that multiple ribosomes work at the same time
-cells need a lot of protein to function, but they can function with not too much mRNA per gene because one mRNA gets translated multiple times
Understanding of how genes influence biochemical pathways (e.g. retinal, hormone biosynthesis)
study: TB12 mutation causes blindness in mice
-this is because the mutation makes the rhodopsin dysfunctional in mice
-this could be either
1. caused by mutation in the gene that codes for opsin, or because opsin did not fold properly into its 3D form
2. this problem could also be due to retinal: retinal is not coded by a gene and it is the product of a BIOSYNTHETIC PATHWAY (just like hormones), essentially, it must be created by the body, it's not naturally occurring
~there could not be a mutation in the retinal itself, HOWEVER, there could be mutations in the enzymes that play a role in each step of that biosynthetic pathway. SO, a defect in any enzyme involved in the pathway could produce defective retinal
~the biosynthetic pathway for retinal is essentially combining a cofactor with an apoprotein to create a functional protein such as retinal
What is are apoproteins
-a protein that requires a cofactor to be functional and is called an apoprotein when it is unbound to its cofactor
-not all proteins are apoproteins
-ex: opsin is an apoprotein before it binds to retinal (co-factor)
explain the concept of post-translational modification
-step after translation for SOME proteins
-binding of a cofactor to a protein to turn it into its functional form
-is essentially the modification of a functional protein after it has been translated, in the case of channelrhodopsin, it is modified by a pigment cofactor after the functional retinal is created
Meaning of potential, kinetic, chemical energy and correct examples of each.
potential energy: is how much energy an object has stored within itself due to the position that it's in or its chemical structure , realize that an object can have more than one form of energy at a time
-ex. when you are at the peak of a roller coaster drop you possess a lot of potential gravitational energy, you have a lot of potential to fall to the ground
-electron loses potential energy getting closer to nucleus
kinetic energy: the energy possessed by an object because it is in motion
-ex. when a photon is moving through space-time, it possesses a lot of kinetic energy as it is moving very quickly
chemical energy: energy stored between the bonds of atoms that make up a molecule (it's the potential energy of molecules)
-ex. as an electron moves closer to the nucleus, it loses potential energy
Distinction between Closed, Open and Isolated systems
Closed system: a closed system is when there can be a transfer of heat (energy) between the system and the universe but no transfer of matter
-ex. a closed water bottle, you can't pour water out of it, but the ice inside can still melt/ cool its surroundings
Open system: when there can be a transfer of heat (energy) and matter between the system and the universe
- think of this like a cup, you can pour water out (transfer of matter) and the water turns the temperature of the air in the room (transfer of energy)
Isolated system: when neither mass or heat(energy) can escape from the system into the surrounding universe -think of a thermos, nothing can spill out and in an ideal world, the food inside stays hot for very long periods of time (theoretically forever)
Definition and examples of: First Law of Thermodynamics, Second Law of Thermodynamics.
-First Law of Thermodynamics: energy can be transferred from one form to another or one place to another, but energy cannot be created or destroyed
-Second Law of Thermodynamics: the energy of a system tends to disperse, or spread out. In essence, a system wants to achieve the highest possible level of entropy (disorder)
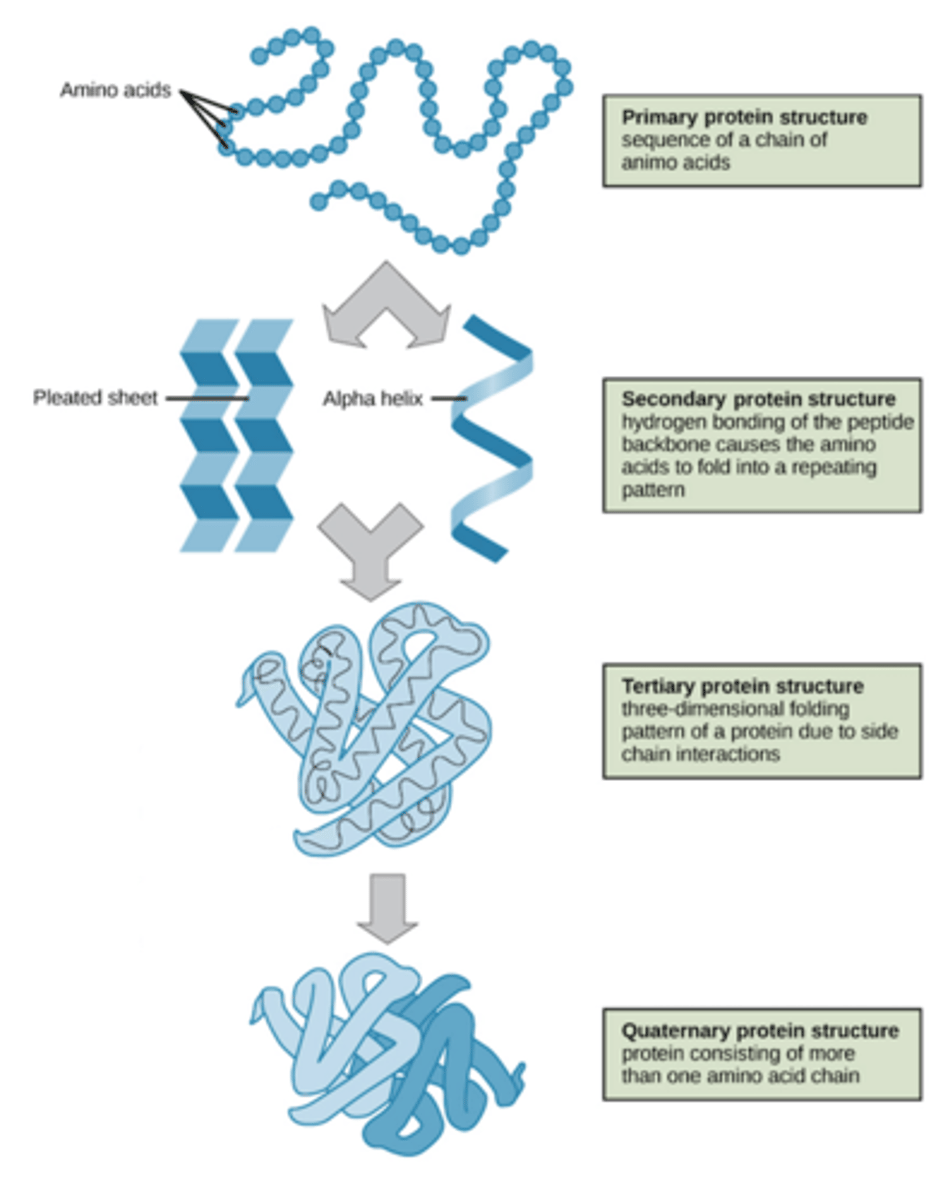
The Four levels of protein structure, what defines each (groups that interact and bonding that takes place).
Primary structure: a simple chain of amino acids (POLYPEPTIDE) bound together in a chain to create a protein, no self bonding has occurred to create a protein structure
Secondary Structure: the amino acid chain is folded onto itself to make specific arrangements (twisted polypeptide).. Hydrogen bonds between the atoms of the amino acid backbone
-Alpha helix: a coil shaped structure that's formed when hydrogen bonds occur with the N-H group of the backbone and the C=O group of the amino acid, looks like a cylinder or barrel
-Beta sheet: Formed by side-by-side alignment of beta strands, formed by H-H bonds between the atoms of each of the strands
Tertiary Structure: Created by four major interactions between R groups: ionic bonds, hydrogen bonds, hydrophobic interactions and disulphide bridges.
-It is flexible so it undergoes conformational changes (small alterations in 3D shape) which contribute to the function of many proteins
Quaternary Structures: When two or more tertiary structures (polypeptides) bond together to create a functional protein
-a protein does not NEED to have a quaternary structure to be functional (tertiary is enough for most proteins)
-chaperone proteins assist other proteins in their maturation and make this process more effective
What happens when a protein denatures.
-When a protein denatures it loses both it structure and function
-Occurs due to extreme conditions that break the bonds/interactions between itself causing it to lose its confirmations
-Denaturation typically occurs due to extreme heat or the use of chemicals
-denaturation reactions are not strong enough to break the peptide bonds (bonds of the polypeptide), so the primary structure remains the same after denaturation. So, some proteins are able to regenerate, while other can't.
With regard to life the explain the concept of: work and breakdown....use of energy brought in from the environment; maintaining low entropy, how the 2nd law applies to living systems , entropy as energy spreading or disorder.
-Cells are open systems, there is a constant transfer of energy and matter between the cell and the universe ( chlamy needs a constant stream of energy to stay alive)
~Why does it need this constant stream? This is because proteins have a tendency to breakdown, meaning that our bodies need to constantly rebuild these structures and decreasing entropy within the cell requires work
~This is why cells are considered islands of low entropy
~To even create these acids, takes energy and work as well
-Why do these cells work so hard to create states of low entropy within themselves? Why build glucose instead of keeping it as CO2? This is because these structures contain more bonds, and when these bonds are broken, energy is released from them allowing to power the life form
-Living systems follow the second rule of thermodynamics as although they are creating high order within themselves, they are releasing waste and heat, which creates a more positive entropy in the surroundings, meaning that energy is conserved
-life (cells) constantly fight entropy by building things aka doing work (SO CELL ENTROPY IS LOW, BUT THE RELEASE OF HEAT AND STUFF INCREASES ENTROPY OF THE SURROUNDING)
In broad terms, what's the difference between a heterotroph and an autotroph.
heterotroph: organism that cannot produce its own food/energy source and must obtain this energy from other organisms/ sources to stay alive ex. rabbits must eat grass to stay alive
autotroph: organism that can make its own food/energy source via complex metabolic pathways ex. plants create glucose from sunlight to stay alive
A photo-autpotroph is an organism that uses light and inorganic carbon (such as CO2) to create it own energy source ex. plants and chlamydomonas are photo-autotrophs as they use light to create energy
Definition of free energy and delta G, spontaneous...basic understanding of enthalpy and entropy.
Free energy(deltaG): the amount of energy in a system that is available to do work
-Delta G: G products - G reactants
Entropy (S): amount of disorder of energy that a system possesses
Enthalpy (H): absolute total amount of energy that exists, especially between bonds of molecules
-deltaG= deltaH- TdeltaS
-reactions that are exergonic are spontaneous and no energy input is needed for them to occur
-reactions that are endergonic can also be endothermic as long as delta S>delta H
-REMEMBER: SPONTANEOUS DOES NOT MEAN FAST!! THE ACTUAL RATE OF REACTION IS DETERMINED BY ENZYMES, SPONTANEITY ONLY SAYS IF WE NEED TO PROVIDE ENERGY FOR THE REACTION TO HAPPEN OR IS IT GONNA HAPPEN ON ITS OWN
Distinction among the terms: exothermic, endothermic, exergonic, endergonic
Gibbs Free Energy:
(+) then the reaction is ENDERGONIC/ absorption of free energy
(-) then the reaction is EXERGONIC/releases energy into its surroundings
Entropy
(+) things are more spread out
(-) things are less spread out
Enthalpy
(-) system releases energy (heat) and is EXOTHERMIC
(+) system uses energy from its surroundings to occur (absorption of heat) and this reaction is ENDOTHERMIC
-REMEMBER: nature wants to go towards stability(higher disorder) + all systems want to have low energy (release the most amt of energy). So any change in any of these factors that allow the system to go "with nature" =spontaneous.
The role enzymes play in increasing the rate of a spontaneous reaction and their role in non-spontaneous reactions.
-Enzymes do not provide energy for the reaction, they are proteins!!!
- HOWEVER, they can bring ATP molecules closer to the reactants (substrate) to make the reaction happen faster
-They are simply proteins that bring two substrates together to help facilitate a reaction
- If enzymes are used in a spontaneous reaction then they increase the rate of reaction (as spontaneous can be quite slow) without any additional energy intake (they won't bring any ATP closer to the substrate, because the reaction does not require it. They only speed it up by normal enzyme processes)
- If enzymes are used in a non-spontaneous reactions they require additional energy input from say, ATP to catalyze the reactions as again, enzymes themselves cannot provide energy for a reaction (in non-spontaneous, they bring ATP closer to substrate because the product has higher energy than reactants so these energy levels have to match by means of atp)
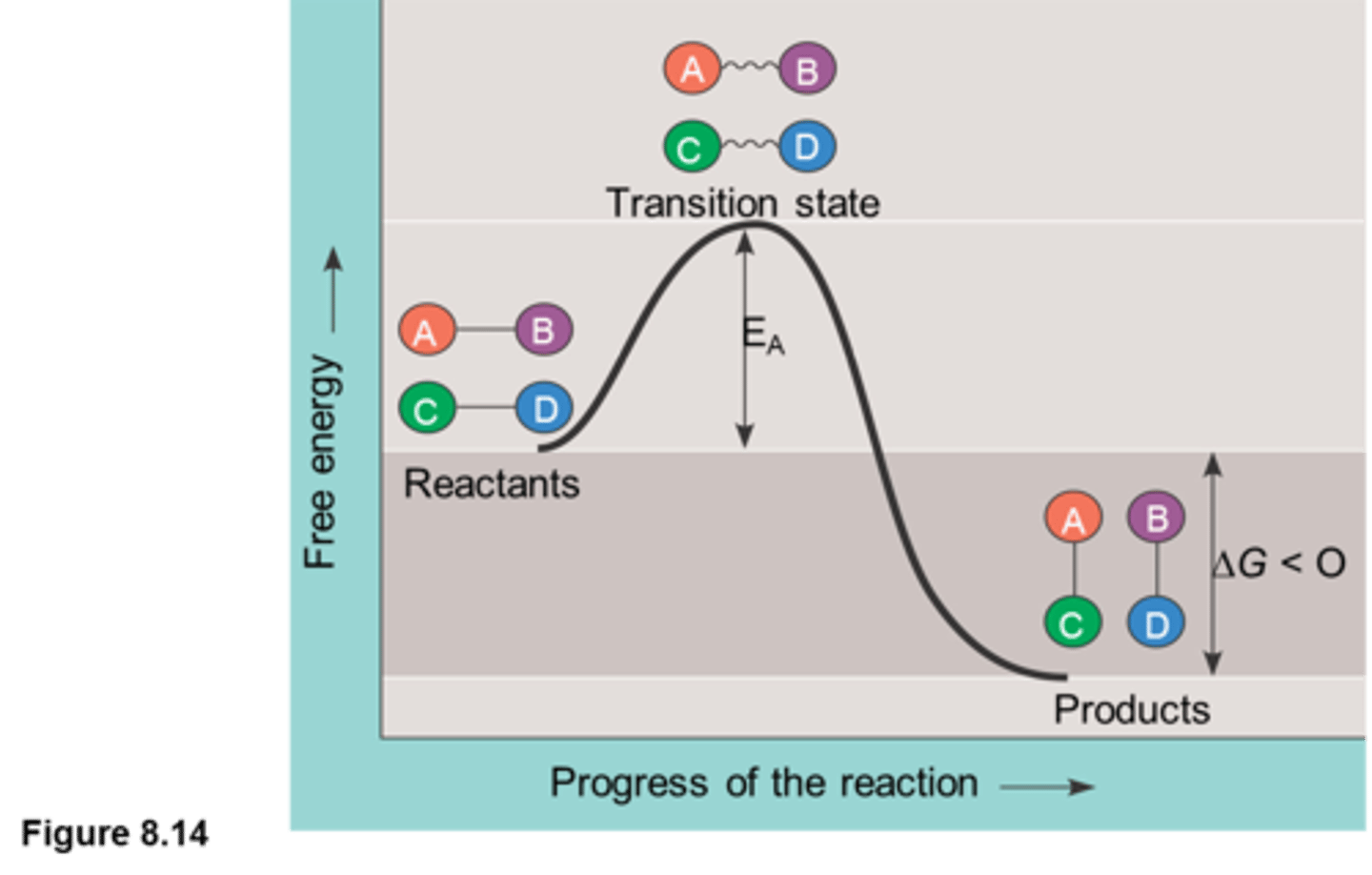
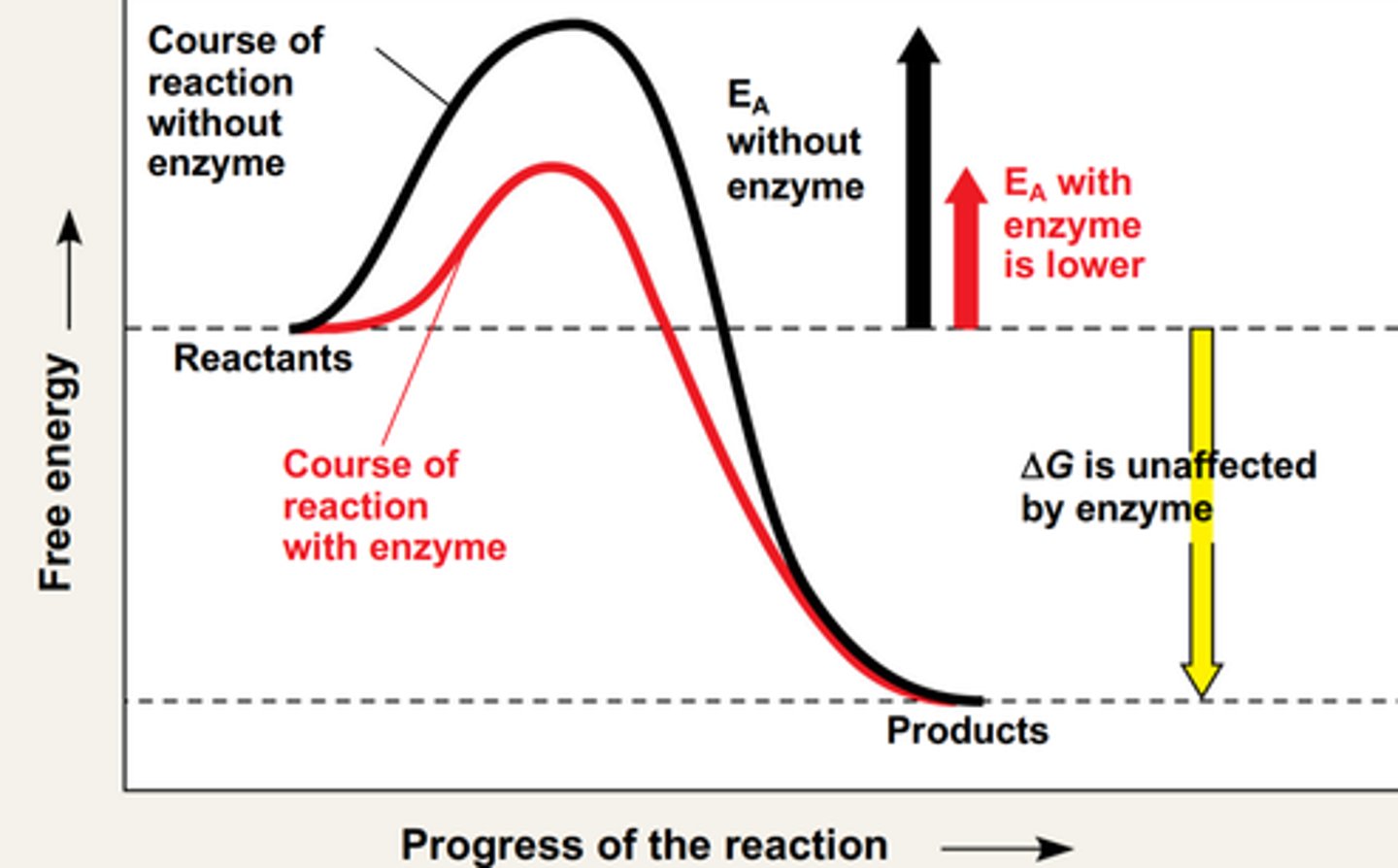
The features of the "exergonic reaction energy profile": delta G, transition state, activation energy (Fig. 3.17 & 19).
-The exergonic reaction profile is a visual representation of the free energy that is required to facilitate a substrate reaction
-in exergonic reactions: products have lower G than reactants
-at the beginning of substrate interaction, delta G is negative as it requires energy to combine the substrate and reach the transition state where substrate combine and rearrange to create the products
-then, delta G becomes positive and thus endergonic as the substrates finish recombining with each other and separate, due to increasing entropy as they split into many smaller molecules which increases their potential to do work and bond
-transition state: a higher free energy state where the conformation bonds are about to break
~requires activation energy (Ea): amount of energy needed to reach transition state. Ea is considered to be a kinetic barrier as not all molecules can arrive to this state (therefore, not all molecules can actually move through with the reaction!)
What an enzyme does to the "exergonic reaction energy profile"
-an enzyme lowers the activation energy for a substrate to enter the transitional state conformation
-Delta G remains unchanged (because the started and ending G are the same, it's just that less energy is required to reach transition state so more molecules of the reactants will be able to do that, hence faster reaction)
-by bringing the two substrates closer together, they require less free energy to react as the energy of bringing the two substrates together was already completed by the enzyme
-this causes enzymes to speed up a reaction process as the rate of reaction is proportional to the number of molecules that get to the transitional state
-enzymes were very important in terms of evolution and the development of life because in general, organisms have very low body temperatures (BIO IS COLD!!)
-organisms can't provide a lot of free energy to power these substrate reactions and they will fall apart at high temperatures
~since the enzyme lower the Ea, the substrates are able to react at much lower temperatures (this helped life evolve, because reactions were sustained at very low temperatures)
-Enzymes lower Ea by precisely orientating the substrates into their transition state confirmation, by charge interactions and by inducing conformational strain (bending)
Basics of protein folding: What is required for a protein to fold correctly.
-For a protein to fold correctly, nothing is required! This reaction occurs spontaneously!
-The only thing that dictates the final folding conformation is the amino acid sequence of the primary structure chain
-native conformation: the active and normal form of a protein
-it is impossible to predict how a protein will fold by simply looking at it as there are too many possible interactions between all of the sequences
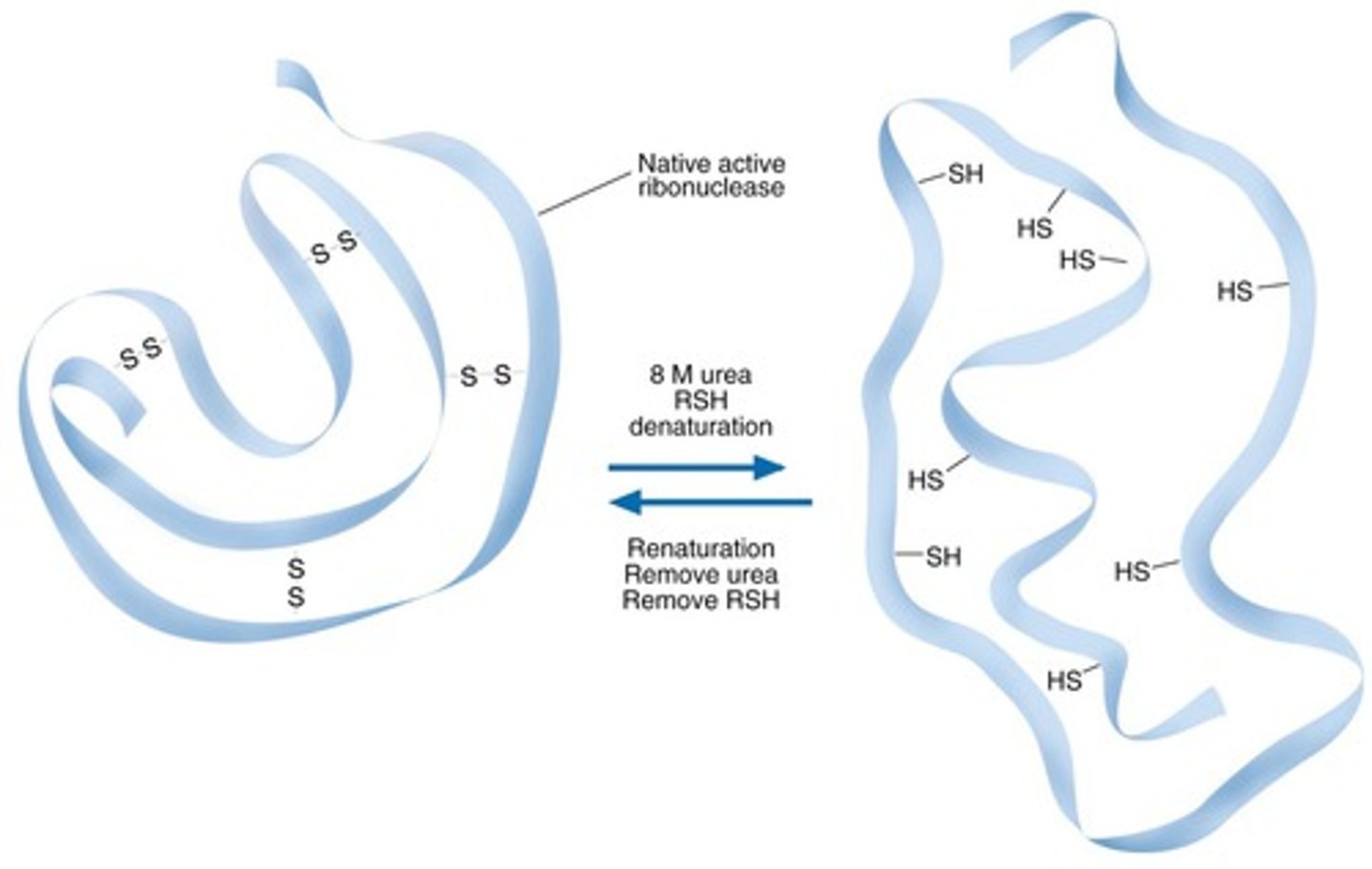
Properties of urea that enable it to denature proteins
-Anfinsen's Dogma: studied protein folding/unfolding by using urea to denature them
-urea is highly polar and causes proteins to very quickly hydrogen bond to it instead of folding onto themselves, because the polarity of urea overpowers its own polarity (it denatures proteins)
-because the proteins are hydrogen-bonded to urea instead of each other, it denatures the protein, causing it to become inactive
-OBSERVATION: after denaturing proteins, they were left to fold back to their form again, and they did!! they became 100% functional again
-CONCLUSION: protein folding is spontaneous
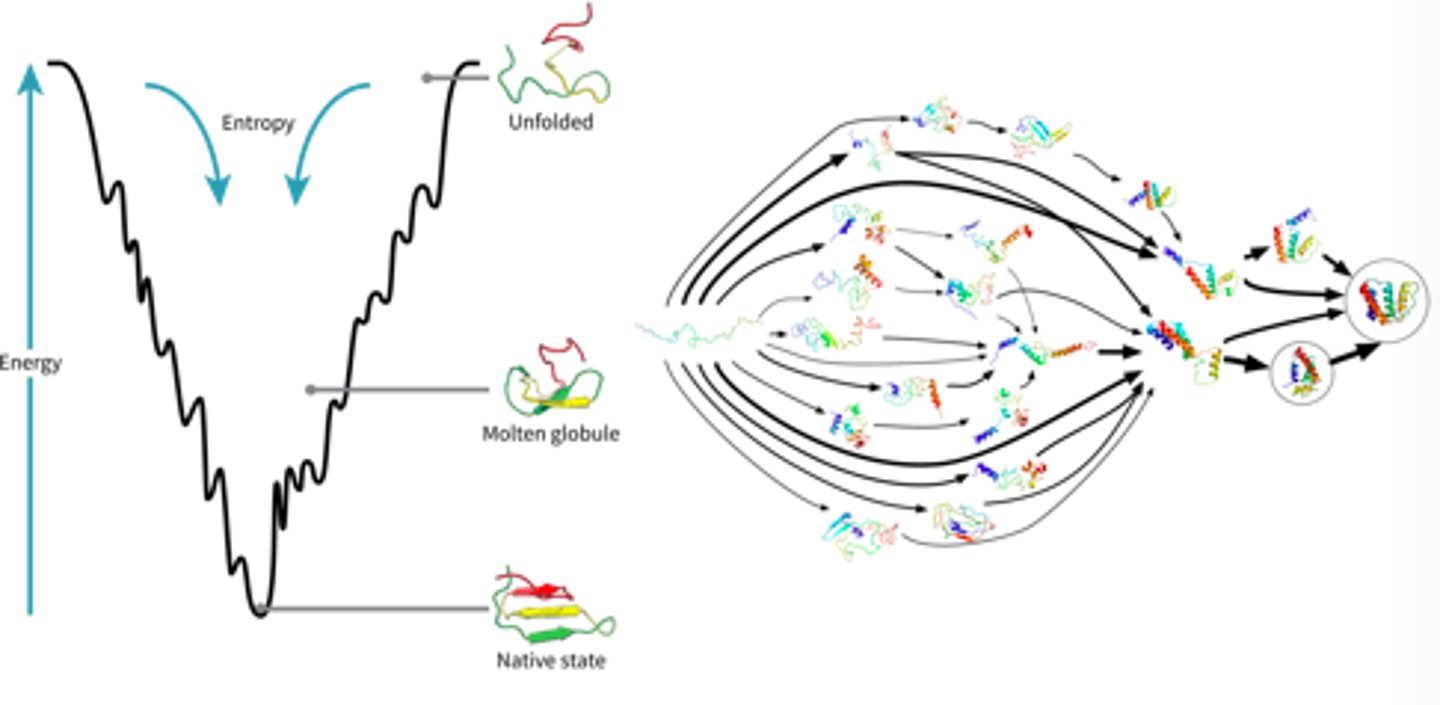
Free energy and the Energy funneling model of protein folding
-as a protein goes from its primary structure to its secondary structure and to its native conformational state, it looses free energy as when it is unbonded, it contains more potential to react and is less stable
-so, the native conformation of a protein always has the lowest energy state
-Energy funneling: when high energy states get pulled to lower energy states (system becomes more stable)
-the drive for the secondary state/native conformation of a protein occurs quickly, as does the hydrophobic effect: where the non-polar amino acids get buried inside the protein away from water, these two occurrences work hand in hand to fold proteins into native conformations
-HOWEVER, it is impossible to completely predict the final shape of a protein
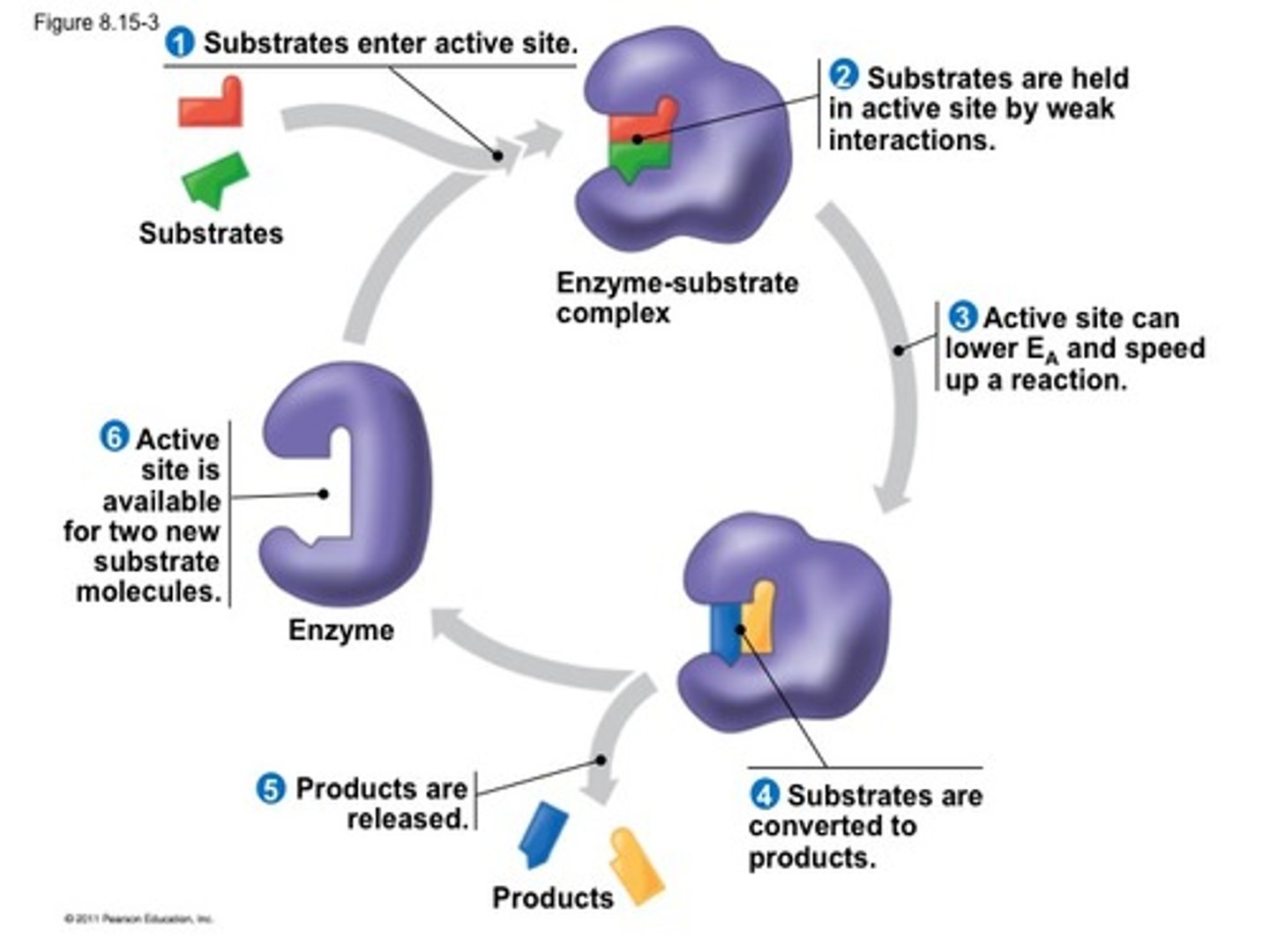
Basics of enzyme structure and the catalytic cycle.
-an enzyme is a flexible tertiary structure (can conform its structure to the substrate that binds to it)
-it contains an active site where substrate is catalyzed
-Contain an allosteric site where activators/ promoter can work to control the functionality of the substrate
-if a mutation occurs in the active site of an enzyme it can drastically affect the function of an enzyme
~neutral hypothesis: a mutation on the edge of an enzyme isn't as devastating ( because it is far from the active site) a mutation like this that doesn't affect the structure of a protein is called a neutral mutation
catalytic cycle:
(enzyme + substrate = enzyme substrate complex = enzyme + product)
-after this occurs the enzyme can be reused to catalyze more substrates, hence why it is called a cycle of catalysis
-temperatures affect reaction rates, the greater the temperature the greater the reaction rate
Basic structure of a phospholipid (what is hydrophobic and what is hydrophilic)
head:
-made of glycerol
polar
-hydrophilic (likes water)
tail:
-made of fatty acids
-hydrophobic
-one straight because saturated
-one tail bent because it has double carbon bond (unsaturated)
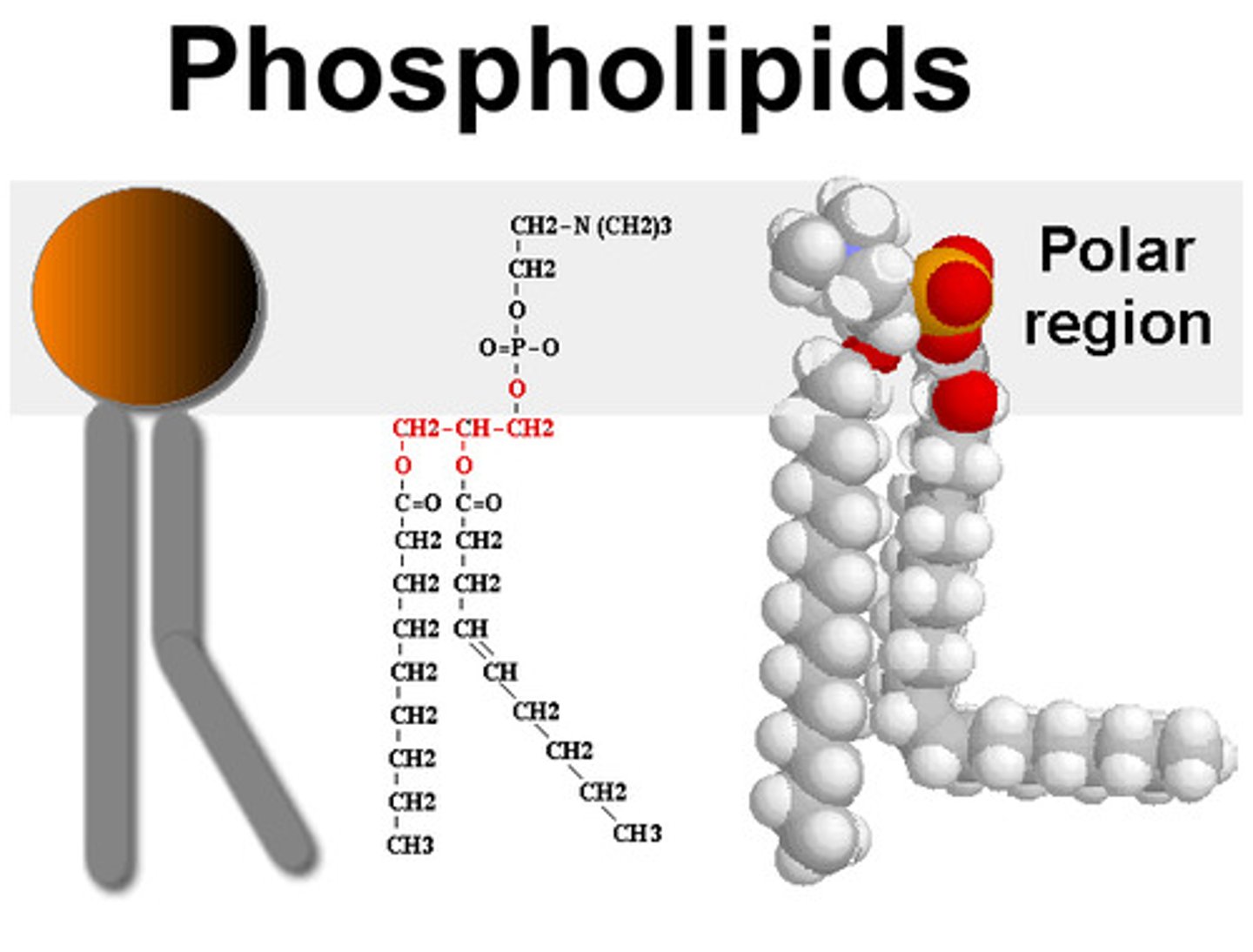
Transmembrane (integral membrane) proteins have characteristics that make them distinct from a normal soluble protein
-unlike other types of proteins, transmembrane proteins can be identified by their characteristic stretch of 17 to 20 amino acids that span the hydrophobic area of the membrane
-the ends of these transmembrane proteins are polar, charged and hydrophilic (so they can interact with water containing area of inside and outside of the cell)
-the middle ends of the amino acid chain that is found in the center of the membrane is hydrophobic and nonpolar( just like the tails, it is placed it a way that does not interact with aqueous environment of inside and outside of the cell)
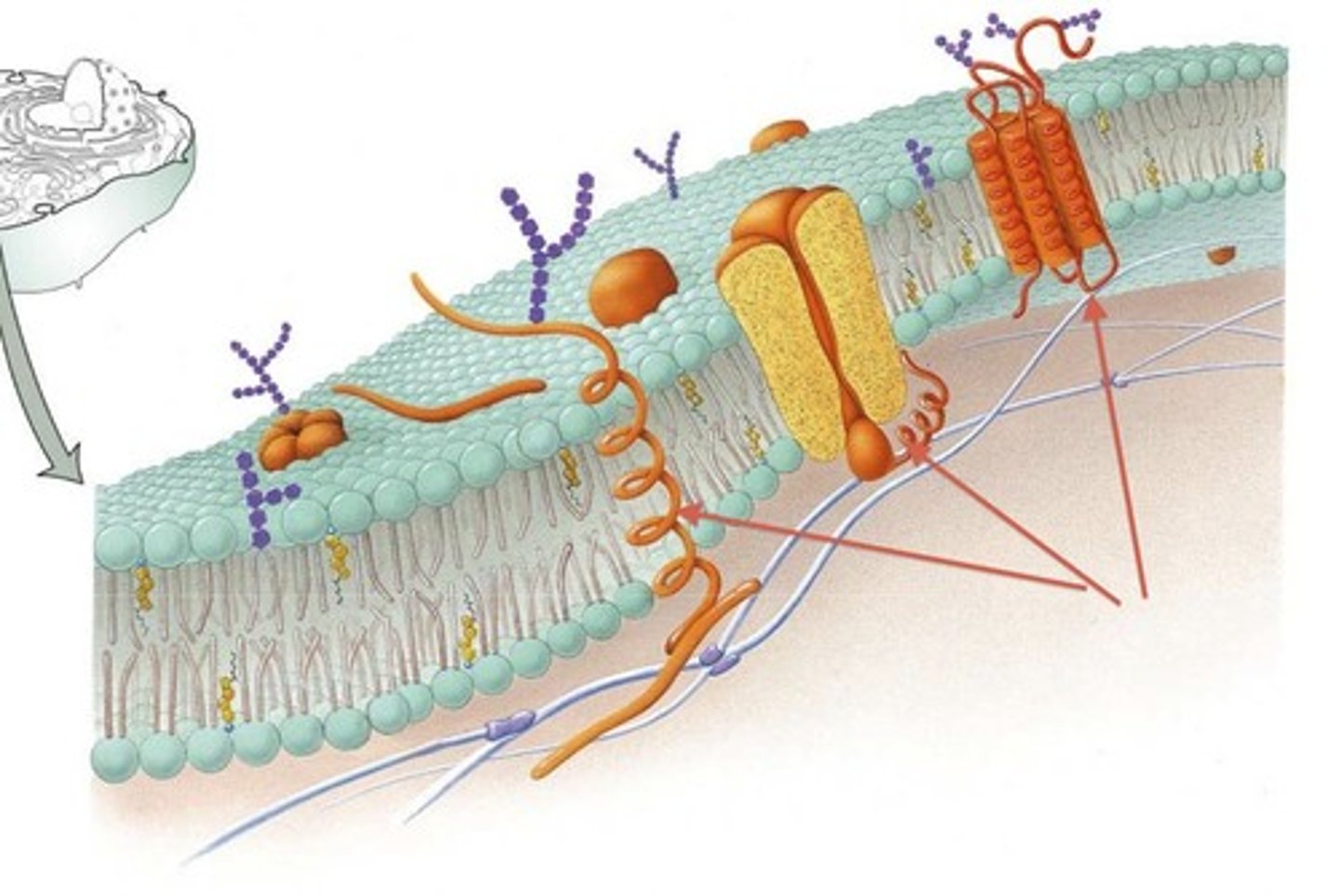
Basic characteristics of molecules that can easily diffuse across a membrane and those that cannot
Molecules that CAN diffuse:
-small, non-polar molecules such as O2 and CO2
-steroid hormones and drugs that are amphipathic (having both hydrophilic and hydrophobic parts.)
-small, uncharged molecules such as glycerol and water even if they're polar
Molecules that CAN'T diffuse:
-ions such as Cl^- and Na^+ , their entrance through the membrane is prevented due to their charge that causes the molecules to be surrounded by water molecules
-large, uncharged polar molecules
The secretory pathway (components of it...and what goes through it)
-The secretory pathway is made up of transmembrane/ integral proteins that interact with the hydrophobic core, that helps facilitate other proteins to leave the cell
-process by which the cell secretes proteins into the extracellular environments
-any protein that ends up on the plasma membrane must also go through this process ex. channelrhodopsin uses this process to become embedded
-proteins that have to go through organelles don't have to go through this pathway!!!
-proteins that are targeted to the ER have signal sequences
COMPRISED OF:
-ER (endoplasmic reticulum) that's connected to the nuclear envelope (nucleus) with ribosomes attached to it
-Proteins that are made in the ER end up in vesicles that connect to the golgi apparatus
-the golgi apparatus ends up releasing another vesicle full of the proteins that are being transported, this vesicle will end up merging with the plasma membrane
-proteins are either released from the cell via the plasma membrane or become embedded into the plasma membrane
-NH4^+, PO4^3- are nutrient transporters that facilitate the uptake of nutrients into the cell
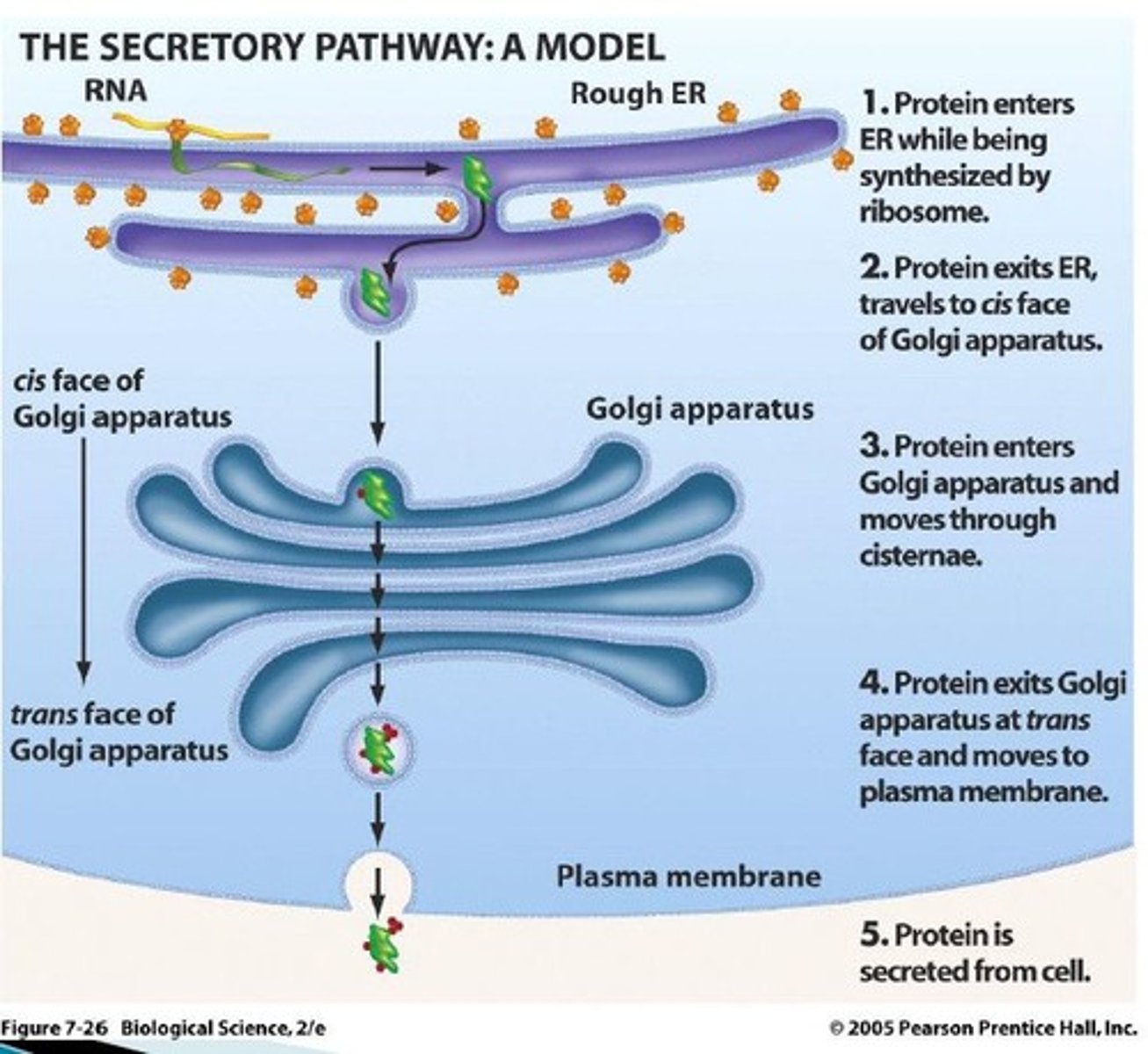
Protein targeting to the ER and the role of the signal sequence and signal recognition particle SRP).+ What is the function of the signal sequence (2 outcomes in one)
-How does a cell recognize that a protein needs to go through this secretory pathway? (how does it target some proteins and not others)
-the mRNA that encodes for these proteins has a signal sequence that encodes for a specific pattern of amino acids
-as this protein is being translated in the cytosol of the cell, signal recognition particles (SRPs) recognize these signal sequences and latch onto these proteins
-when these SRPs attach to the ribosomes and protein, translation is paused and the whole apparatus is pulled to the ER
~this is done via the SRP being drawn to the SRP receptor that's embedded into the ER membrane
-once attached to the SRP receptor, protein translation continues and the rest of the protein is translated into the ER lumen itself
-once translation of the protein is complete, the signal peptide removes itself from the membrane of the ER
-the ribosome removes itself from the ER
-The mature protein is now contained inside the ER!! (this whole process was to explain how proteins start their journey in the pathway, starting from the ribosome in the cytosol where they are translated to ER (which is the first component of the pathway)
Basics of simple and facilitated diffusion (passive transport)
simple diffusion:
-small molecules simply pass through the gaps in the membrane
-this action is driven by increase in entropy and the concentration gradient
-driven by concentration gradient (molecules like to go from high to low concentration to reach an equilibrium)
facilitated diffusion:
-Integral protein channels embedded in the plasma membrane allows large/charged protein molecules that can't fit between the gaps in the plasma membrane to diffuse
-using the concentration gradient and entropy as well...... channelrhodopsin uses this method! (only reason any molecule needs this protein facilitator is cuz they are phatttt)
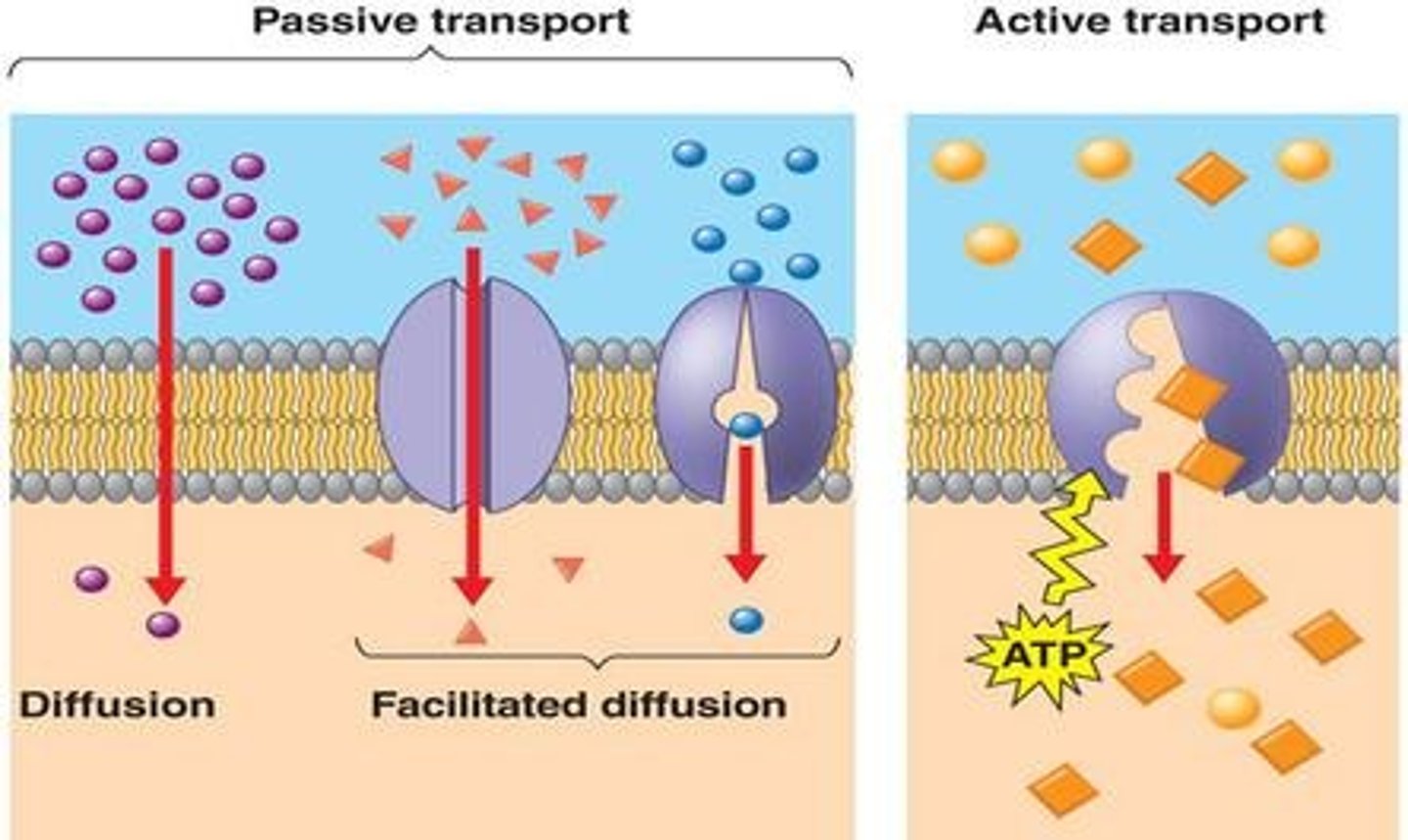
Transport against a concentration gradient (active transport)
Molecules move against the concentration gradient
they need energy aka ATP to do so (do work against concentration gradient)
use protein complex pump
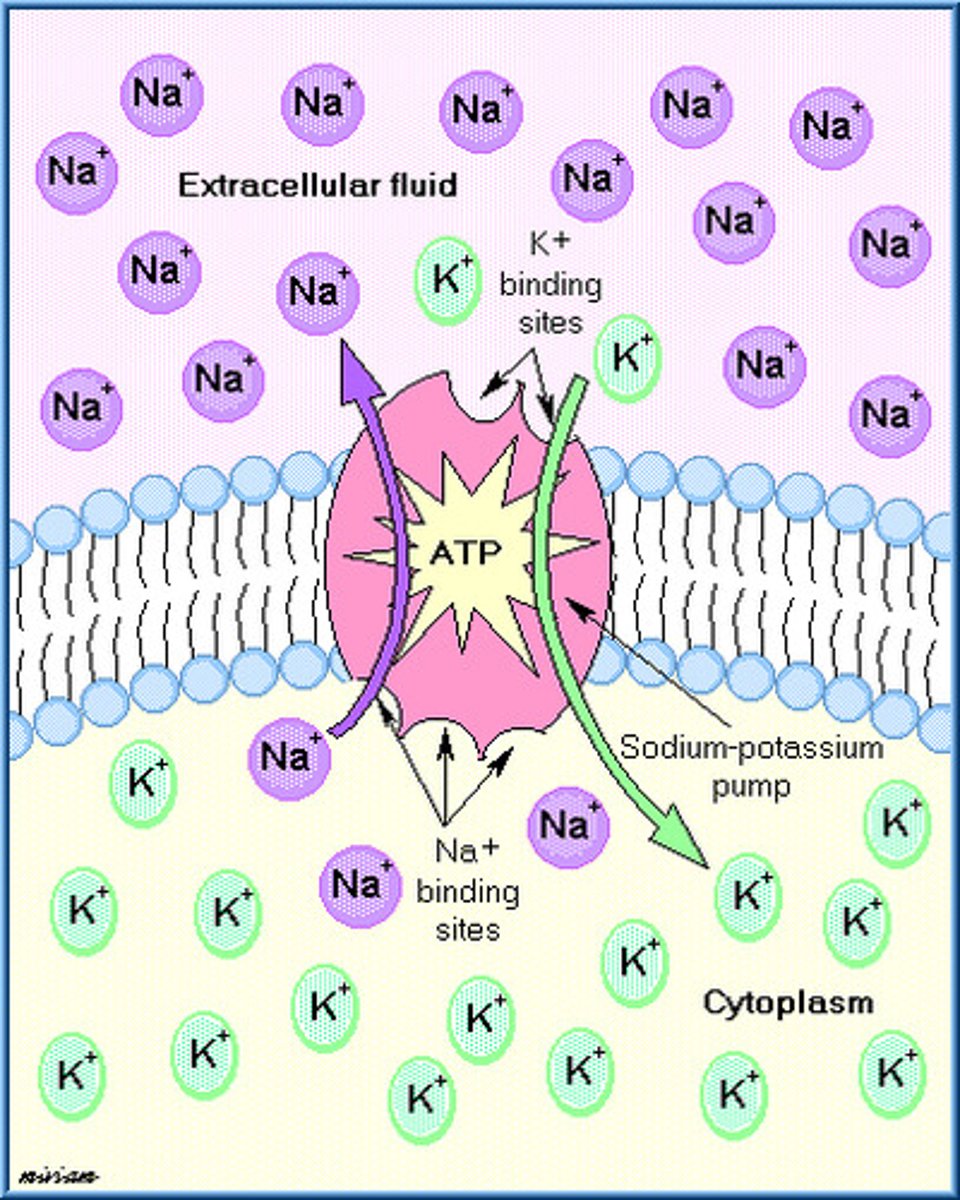
Linking transport with free energy change (delta G)....what drives transport
for active transport:
-transport of molecules actively is NOT spontaneous
-so, active transport causes positive change in energy (because energy is brought in)
for passive transport (diffusion):
-REMEMBER: free energy constitutes for a big part of potential energy and chemical energy
-when substances in highly concentrated area, they relative energy is higher
-Since everything wants to go towards disorder, the transport from highligly concentrated area to a less concentrated area makes the molecules occupy more space, which means entropy increases
-increase in entropy implies negative change in free energy
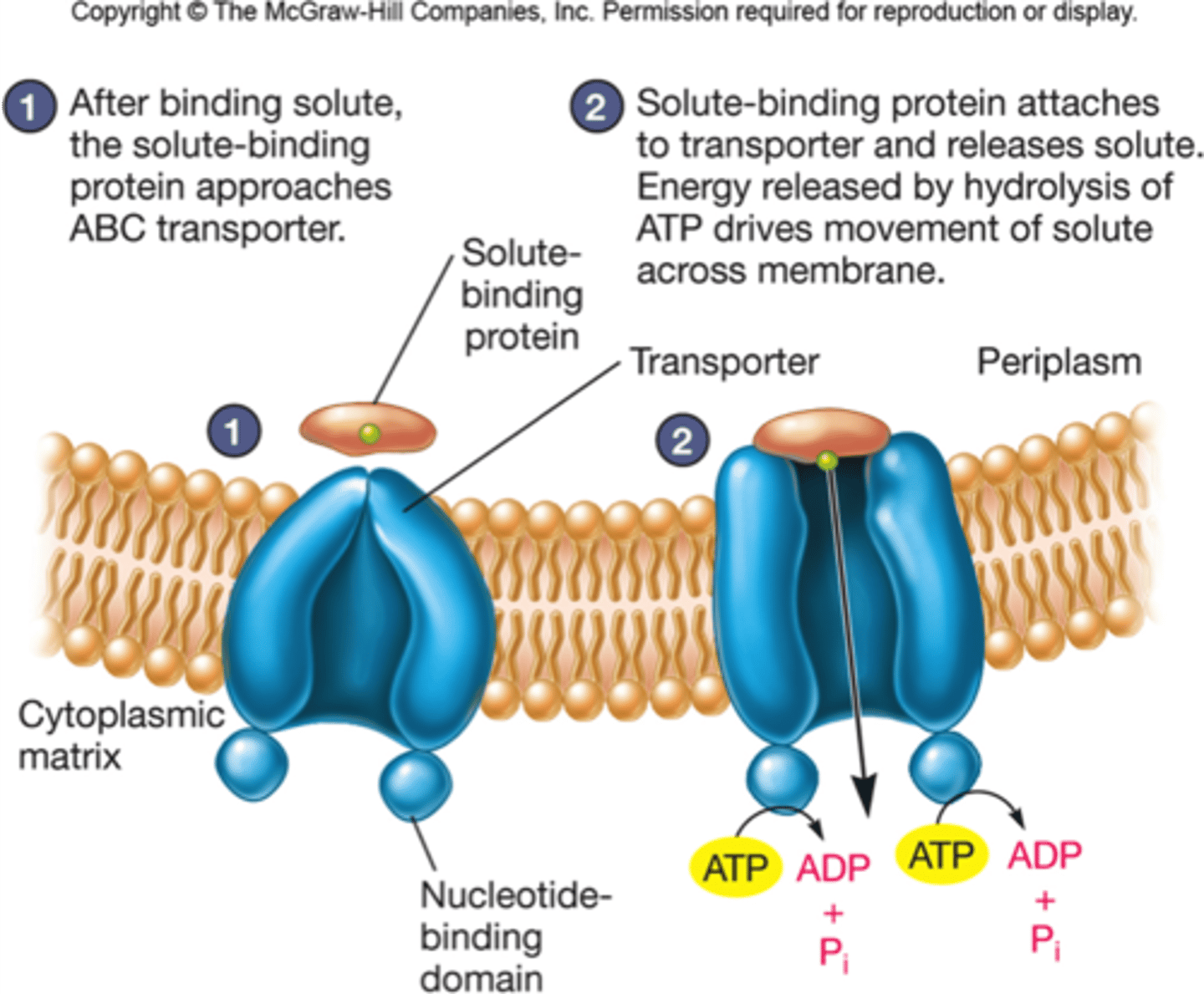
Basic structure of ABC transporter. What does each component do?
An ABC transporter (ATP-binding cassette) is comprised of two parts, the transmembrane domain and the ATP-binding domain (ABC transporter)
-The transmembrane domain makes a channel in the lipid core
-the ATP-binding domain is the motor that drives active transport, it is where ATP binds to and breaks down to provide energy to pump molecules against the gradient
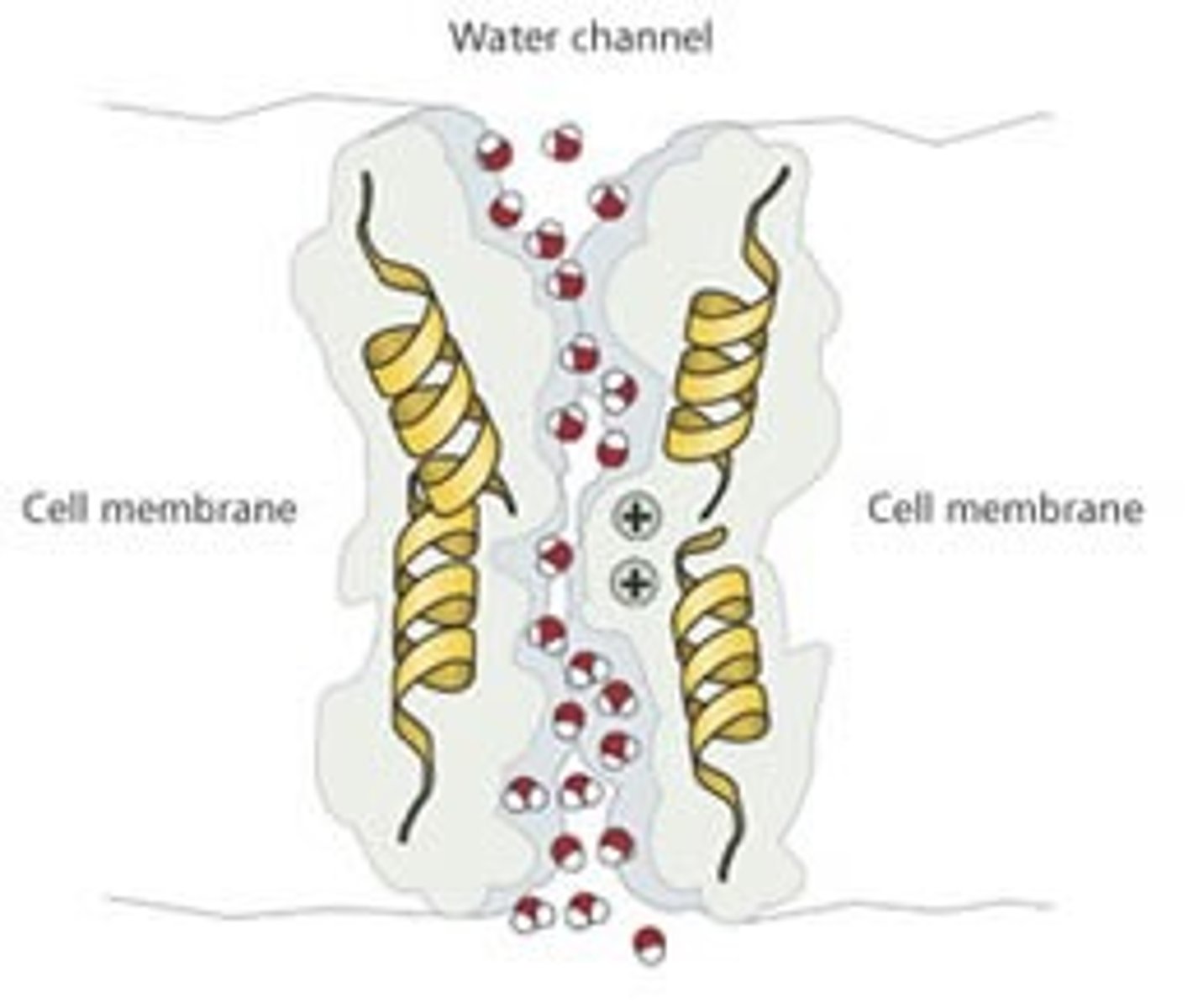
How can a pore/channel be specific for certain molecule (e.g. aquaporin).
an integral protein in the plasma membrane of a cell can be specific for a certain molecule by:
-conforming its shape: making the protein channels very long and convoluted that restricts entrance to molecules that are only a certain radius and shape, other molecules are not able to pass through
-conforming its charge: channels can repel certain charged molecules depending on their and the molecules charge.(water can still go in and out because although it is polar, it is overall neutral)
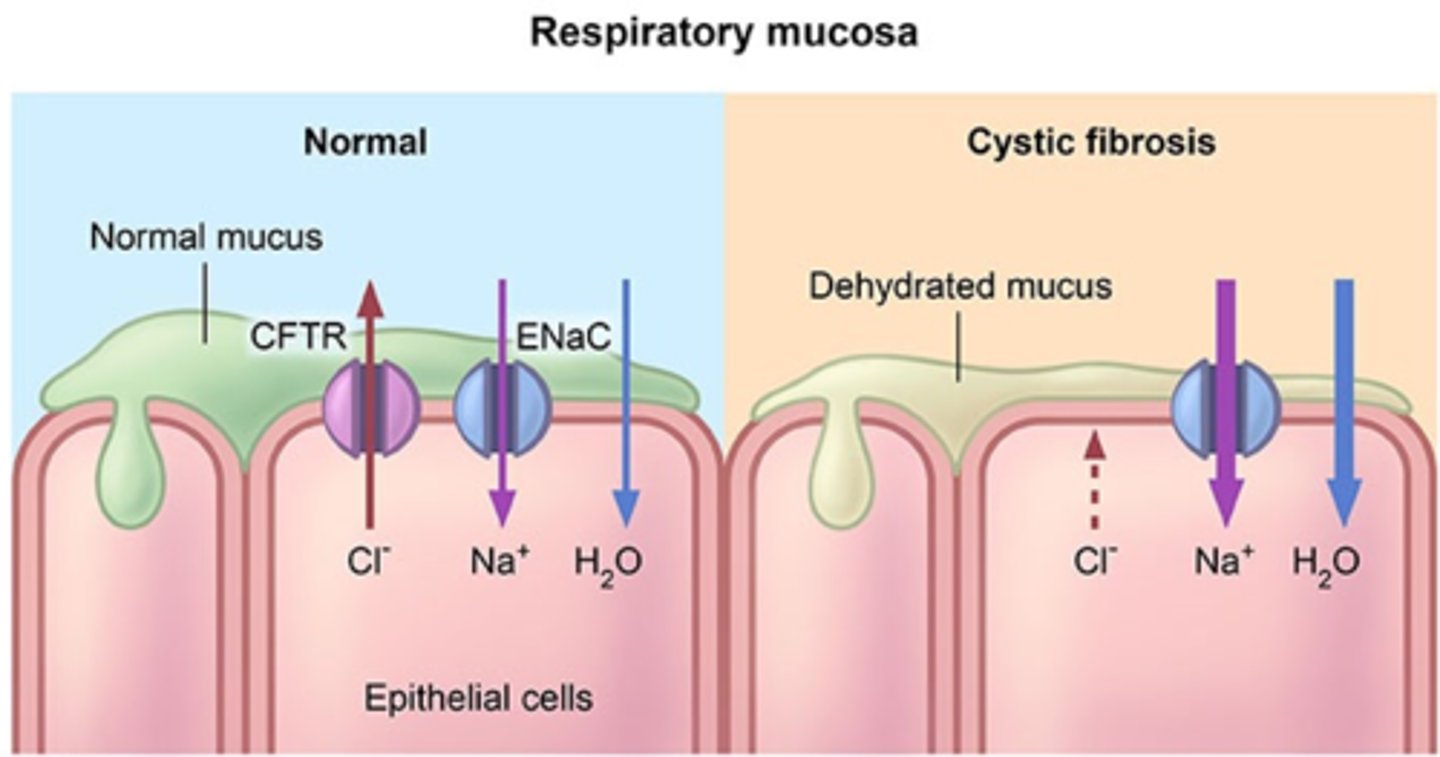
Cystic fibrosis and the function of CFTR
cystic fibrosis is when there is a misfold of the delta F508 gene that causes the CFTR transporter (protein and ABC transporter) to not function properly
normal lungs have to be moist to allow cilia to flow and a wet lung surface is required to facilitate gas exchange
in normal (non-mutated gene)
-CFTR pumps Cl- ions out of the epithelial membrane
-due to higher concentration of Cl- out of the membrane, water diffuse through facilitated diffusion
-water diffusion keeps the surface of the cell (mucus layer) moist
-healthy lungs, good gas exchange, the vibes !
IF MUTATION:
-CFTR does not work, so no Cl- gradient will be formed
-water won't follow the gradient through diffusion (bcz there is no gradient)
-mucus layer remains dry
-bad gas exchange
-higher chance of lung infections because you won't be able to cough the bad shit!
Fate of the deltaF508 form of CFTR + Role of chaperone proteins (HSP90) in folding of deltaF508 form and wildtype
Since CFTR sits on the plasma membrane, it must be secreted by the secretory pathway (to reach the plasma membrane), but the delta F508 mutation causes it to never get past the ER...why?
we compared western blotting of WT and F508 mutation genotypes:
-In WT, we see a very dark line on the section associated with plasma membrane, meaning most of the CFTR makes it to the plasma membrane + some amount still in ER where they used to be before
-In mutation, none in plasma membrane, some in the ER
This is because the mutated CFTR never makes it to the membrane, it stays in the ER
there is a quality control system in the ER, it detects misfolded proteins using chaperone proteins, in the case of cystic fibrosis, HSP 90 protein is the chaperone protein that is being used and controls the ER
HSP 90 is triggered and removes the misfolded proteins from the ER into the cytoplasm and is picked up by the proteomes of the cell and is degraded
because it is degraded, it never reaches the golgi apparatus to get transferred to the plasma membrane
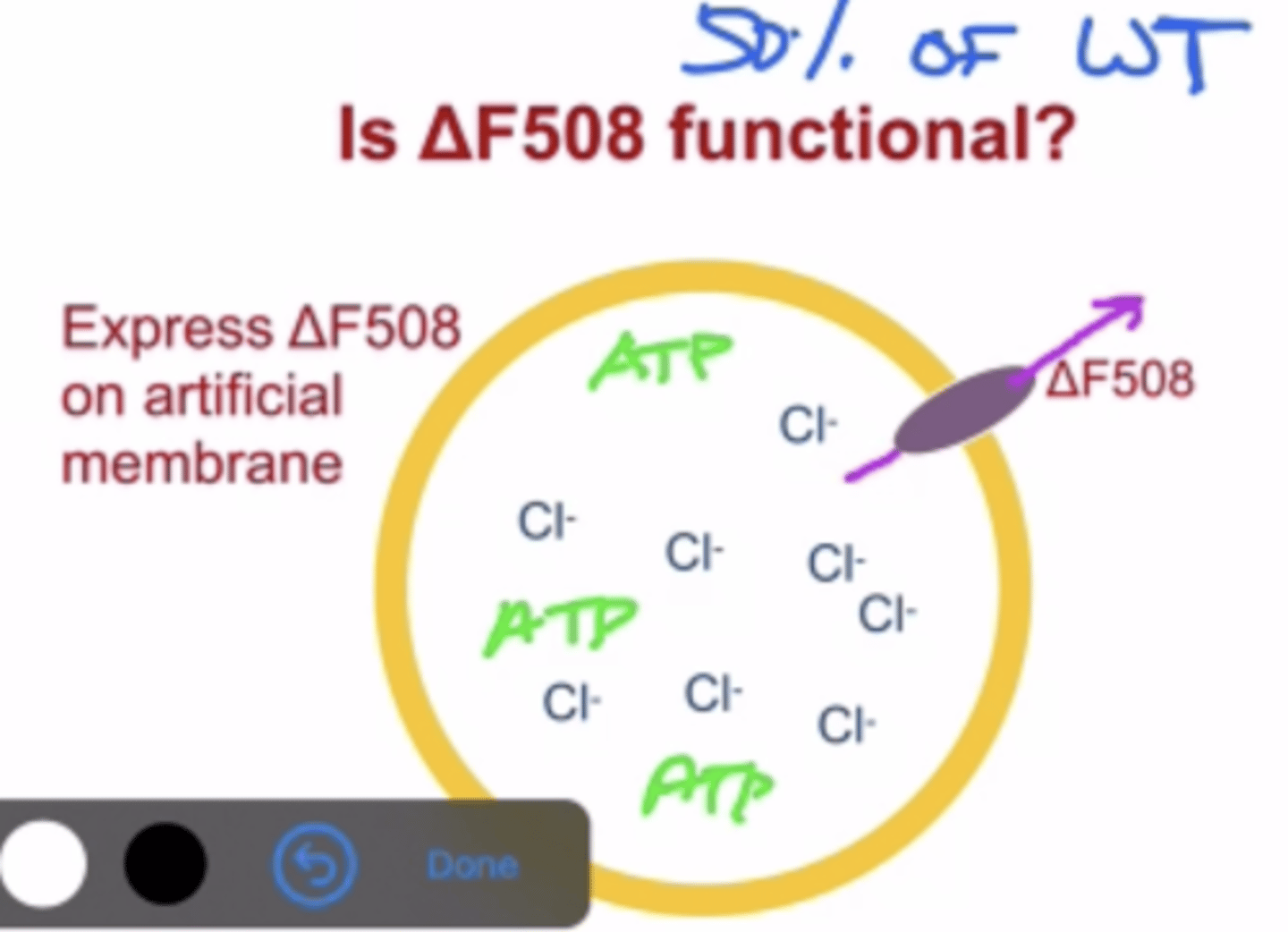
Experiment showing the deltaF508 mutation form is actually a functional CFTR
-Delta F508 is still functional and pumps protein to an efficiency rate of about 50%
-This was proven in a lab by creating an artificial membrane: filled with Cl- ion solution, deltaF508 and ATP for the Cl- pump to work
-by doing this we could see that the Cl- pump still worked to a 50% efficiency rate and changed the Cl- concentration inside this artificial membrane (because Cl- got transported out by the mutated CFTR)
-this again proves that the reason for cystic fibrosis is that CFTR does not get transported to the plasma membrane, not that it fully stops functioning
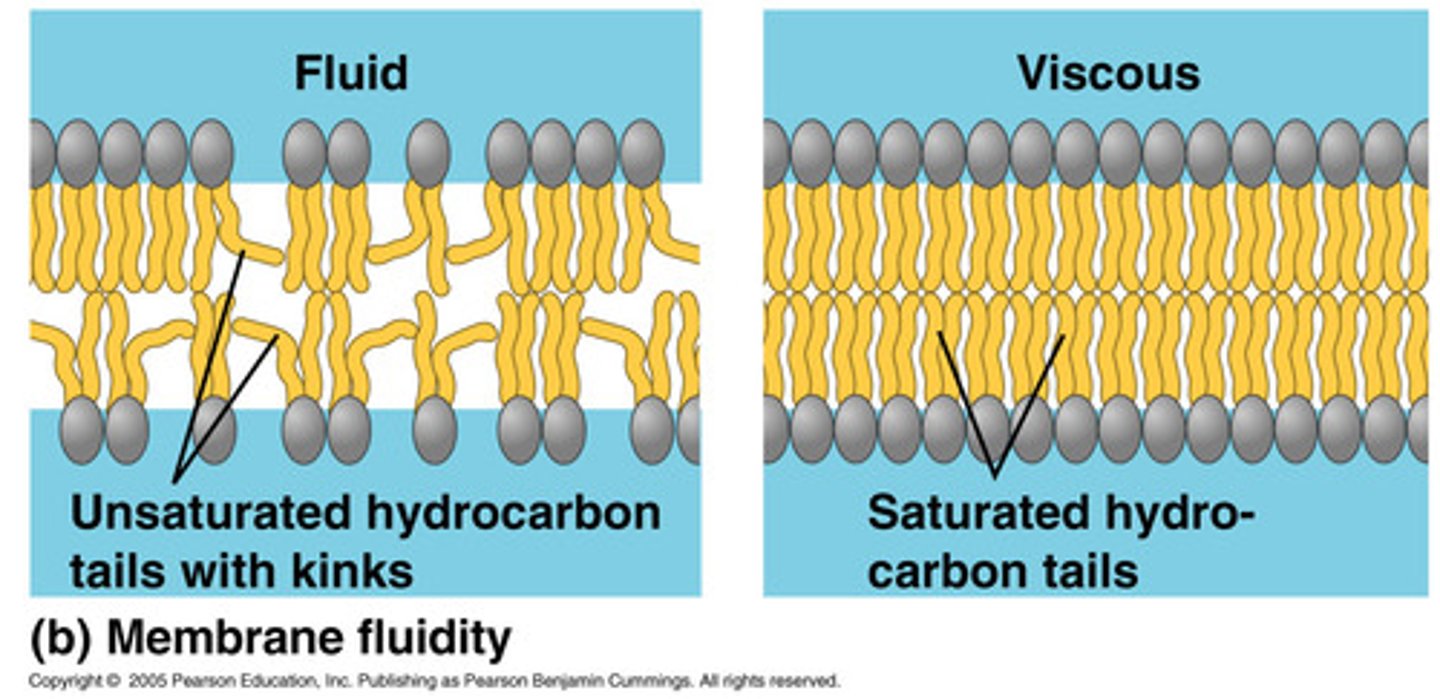
Relationship of fatty acid saturation levels on membrane fluidity.
fatty acids are completely straight when they are fully saturated, meaning that the phospholipid bilayer of the cell membrane is very stiff and tightly packed together when this occurs
when a fatty acid becomes unsaturated, it creates a double bond in the fatty acid which creates a kink in the fatty acid, making the cell membrane less packed between each of the phospholipids, allowing for the membrane to be able to move more freely and therefore become for fluid
in short, the more unsaturated the phospholipid bilayer is, the more fluid the cell membrane becomes
Relationship of temperature on membrane fluidity. relationship of fluidity to membrane functions such as transport.
as temperature increases, it creates the phospholipids to become unsaturated and thus more fluid, in the case of a temperature that is much higher than what it can handle, the cell membrane becomes too fluid with too many gaps
all this gapping allows too many molecules to passively diffuse between to the outside of the cell and vice versa, this means that cells can't maintain a concentration gradient within themselves, which leads to problems in completing active transport and cell death
Low temperature means that the phospholipids become saturated and the cell membrane becomes too rigid, not letting anything through, meaning that passive transport and facilitate transport cannot occur as small molecules cannot pass through any gaps in the membrane and the integral proteins can change conformation to let anything through to themselves
Properties of saturated vs. unsaturated fats.
saturated fats are straight chains of carbon chains ->(only single bonds)
unsaturated fats means a removal of a hydrogen which means the carbons in the chain double bond to themselves, causing a bend in the chain -> at least one double bond
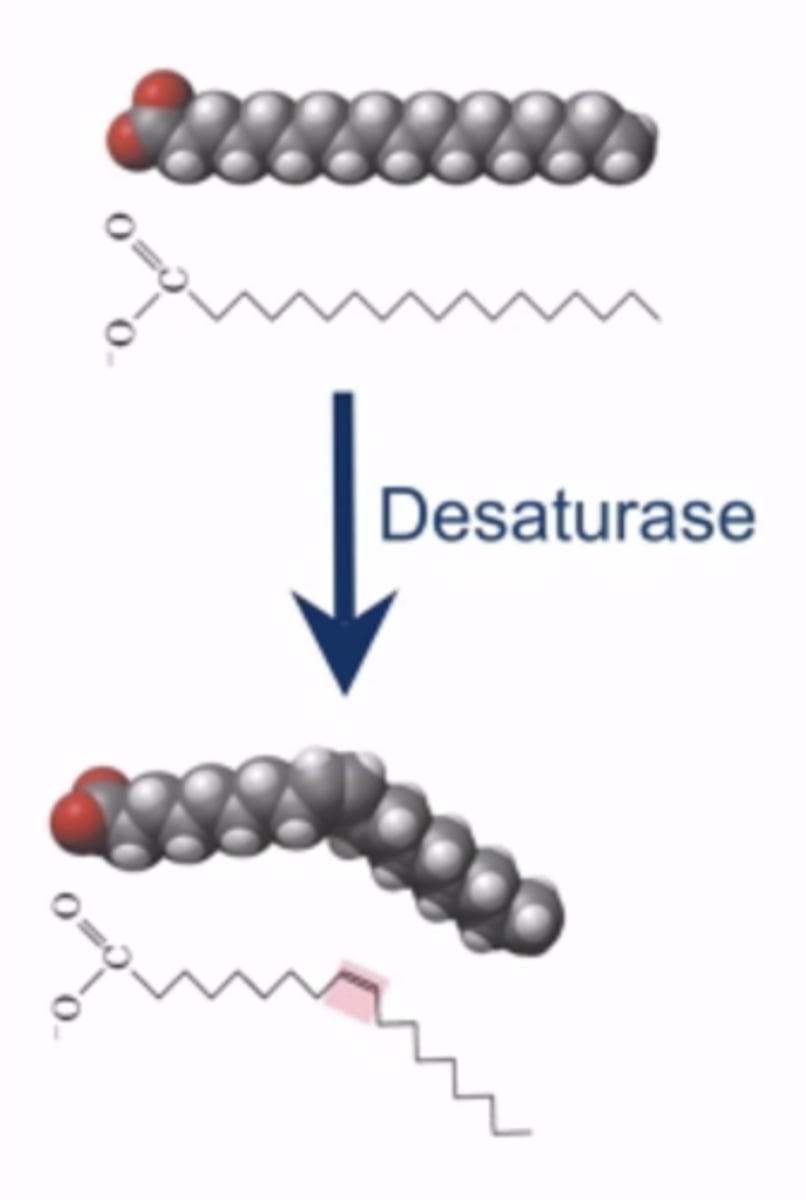
Role of desaturases (enzyme) in fatty acid biosynthesis
desaturases removes hydrogen from precise locations along the fatty acid chain -> unsaturated the chain (more unsaturation, more fluidity)
most organisms adjust their fluidity by regulating desaturization expression, as more unsaturated fatty acids means a more fluid membrane regardless of the temperature
Relationship of bacterial desaturase expression vs. temperature.
as most organisms can't maintain a constant temperature like mammals can, these desaturase proteins are very important when it comes to maintaining membrane fluidity in less than ideal temperature ranges
bacteria like warm temperatures, so they have adapted in a way that they have multiple desaturate enzymes that vary in expression depending on the temperature (some expressed more when it's hot, so bacteria survives lol)
if the membrane becomes too stiff because of the cold... desaturases! Then the membrane is effective and fluid again :)
Given that you can't remove the double bonds from a kinky (haha) fatty acid...how does an organism deal with that if they are moved from low temperature (where you want unsaturated fatty acids) to high temperatures (where you don't)?
the cells rely on the fact that everything has a half life, so when the temperature increases, the unsaturated forms of the fatty acids will return to their saturated forms
Understand the balanced chemical reaction of photosynthesis. Look at the two reactants (CO2 and H2O), what parts of the process do you find them?
6CO2 +6H2O -> C6H12O6 +6O2
this is the NET gain/loss of these molecules (there is much more that happens lollll)
H2O is used in the light reactions: photolysis of water will occur, the electrons freed from H2O will be used by photosystems through electron transport chain (O2 will be released as waste)
CO2 is used in light independent reactions: it will get fixed to eventually form enough G3P to make glucose in calvin's cycle (glucose will be the product)
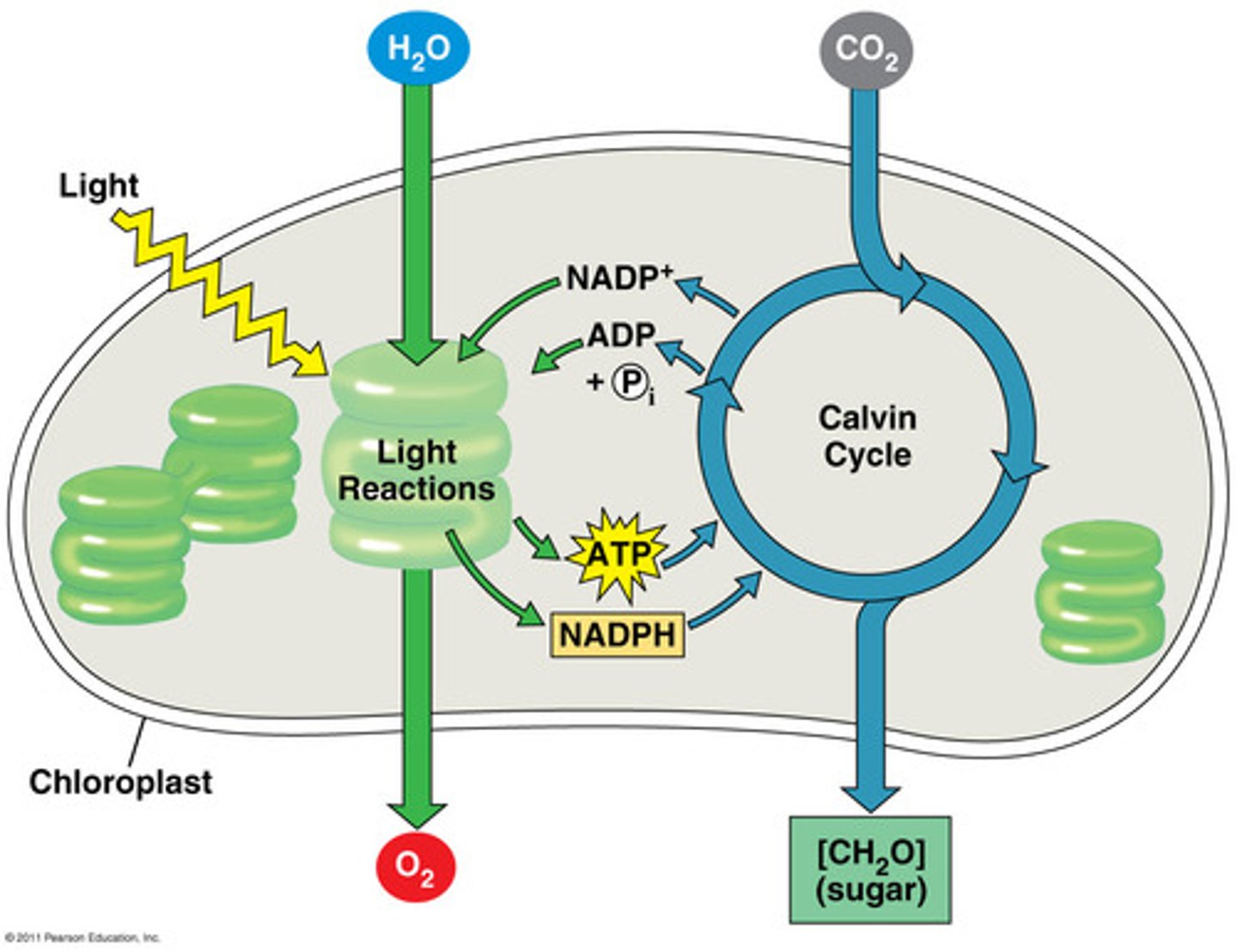
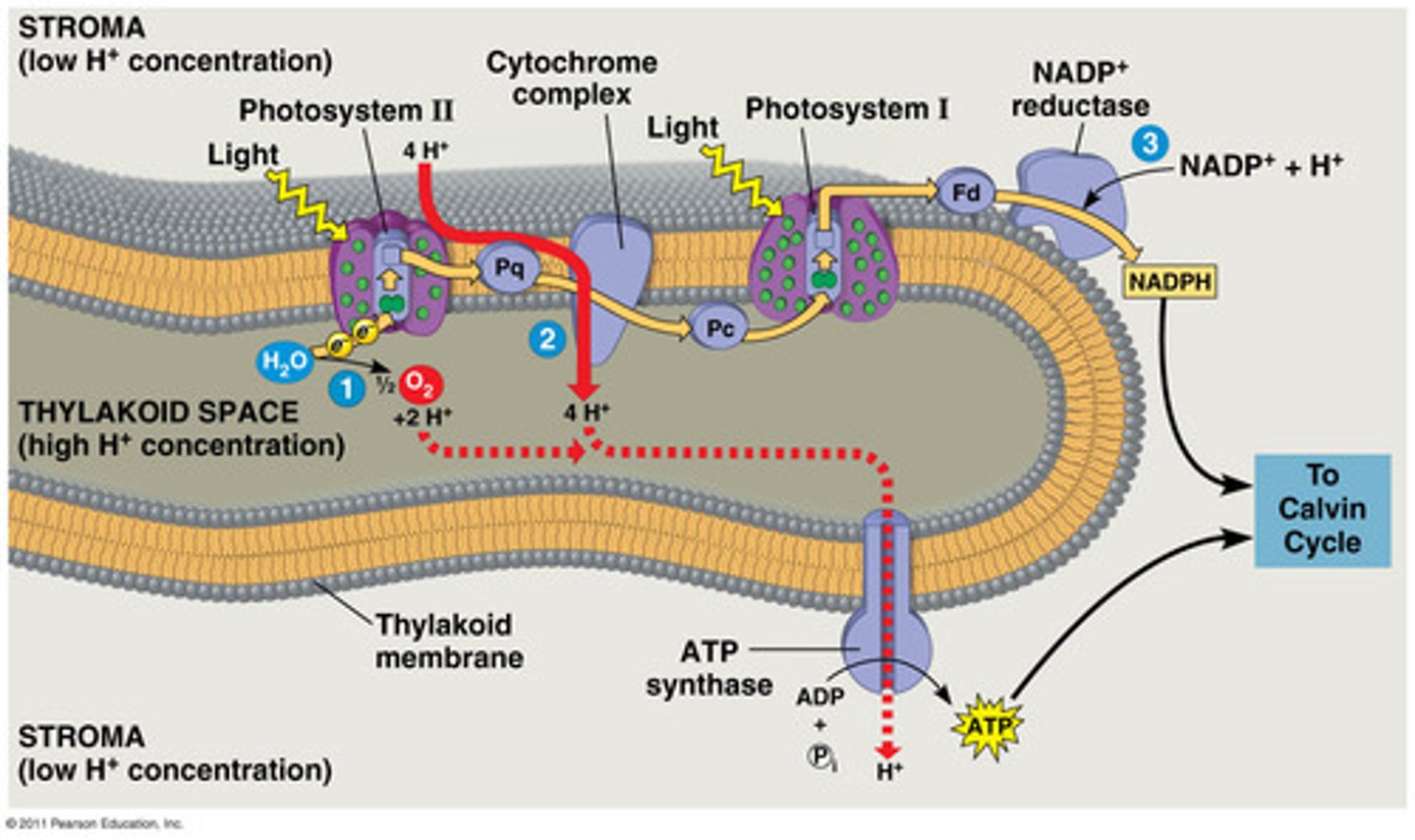
Explain light reactions (one of the two phases of photosynthesis) ...broadly
LIGHT REACTIONS
-light dependent (will not happen in the dark)
-purpose: making ATP and NADPH + H+ through ETC
-happens in the thylakoid membrane of the chloroplast
1. light energy absorbed by photosystem 2, the electrons in the reaction centre (chlorophyll b) of the photosystem get excited and get captured by an electron absorber
2. water photolysis: H2O -> 2H+ +2e + ½ O2
These two electrons serve to REPLACE the electrons lost by photosystem 2 when it gets excited (so it's not these two electrons that enter ETC directly, they get fixed in the photosystem and used up when the photosystem is excited). There is constant photolysis so there could be constant ETC
3.electron transport chain:
-the two electrons from PS2 pass over a chain of electron transporters (such as proteins and b6f)
-each electron transfer releases energy
-energy will be used by b6f to release H+ into the thylakoid lumen from the stroma
-electrochemical gradient formed: more H+ ions into thylakoid lumen than in the stroma
4.electrons will reach ps 1 and will excite chlorophyll a
5.FINAL ELECTRON ACCEPTOR: NADP is the final electron acceptor, which will be reduced into NADH. Not only does it accept the two electrons moving in ETC, it also accepts the 2 hydrogens left from water photolysis. enzyme at work is reductase
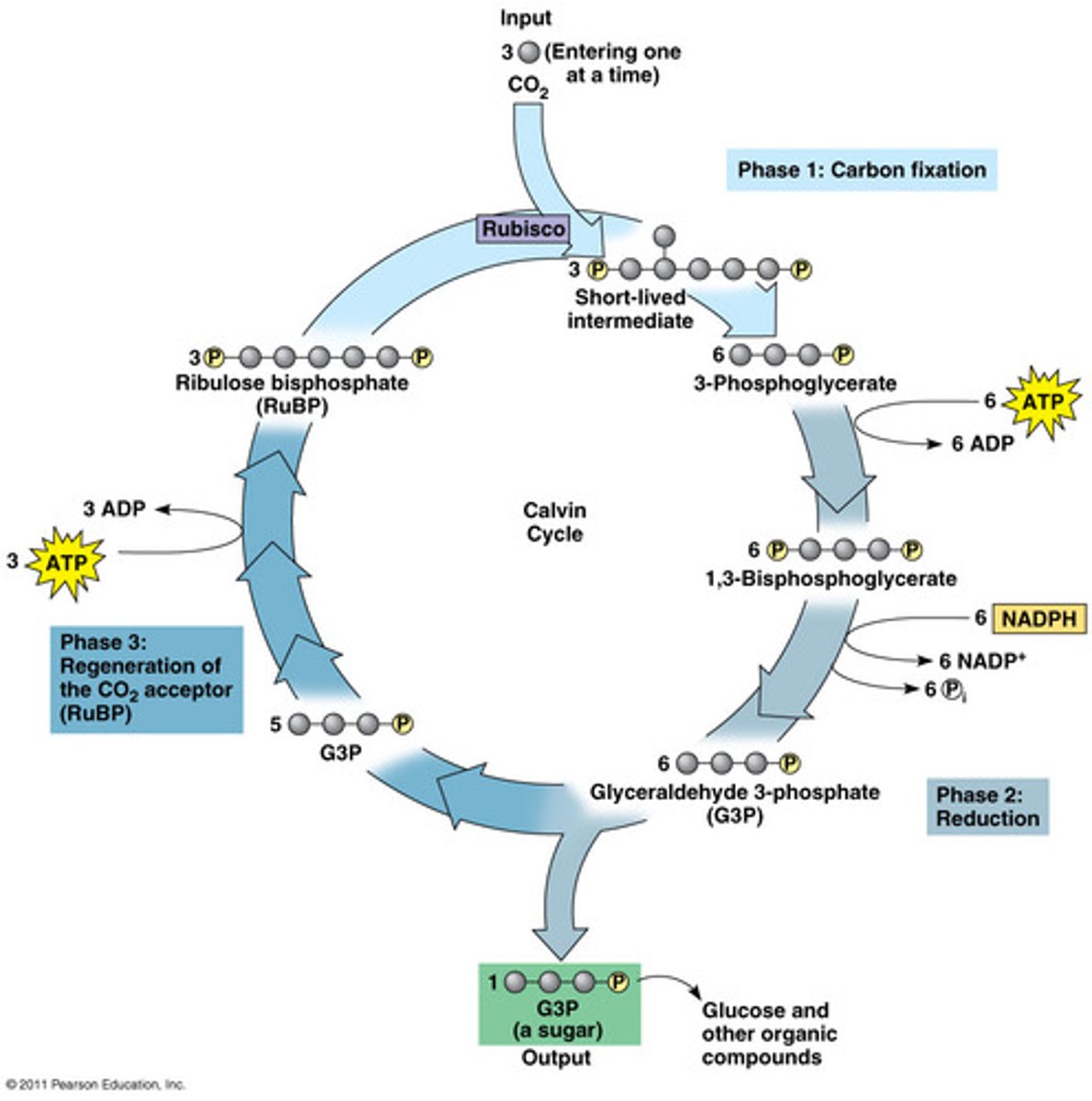
Explain the calvin cycle (one of the two phases of photosynthesis) ...broadly.
LIGHT-INDEPENDANT REACTIONS
-CALVIN CYCLE !
-we use the ATP and NADH made during light reactions to activate reactions of the calvin cycle
-happens in the stroma
-"dark reactions": does not mean in happens at night, it just does not directly use light like ETC does
1. carbon fixation
-Carbon binds to RuBP (5C) with help of Rubisco
-this carbon comes from the outside environment (how respiration and photosynthesis are related),c carbon dioxide has nothing to do with light reactions
-unstable 6 carbon molecule formed
-6 carbon compound turns into two 3 carbon molecules (PGA)
2. reduction
-the two PGA are energy low
-hydrolysis of ATP to ADP -> phosphorylation of PGA (we add phosphate group to them)
-oxidation of NADH into NAD+ to turn the PGA into 2 energy high molecules (G3P)
-some G3P will leave the cycle to produce glucose
-most G3P continue on with the cycle to repeat it
3. RuBP regeneration
-this step explains why we say its a Calvin CYCLE
-YOU NEED TO REGENERATE RUBP FOR THE CYCLE TO REPEAT
-RuBP is a 5 carbon compound, we have 2 3 carbon G3P to regenerate it
-5 carbons get used to regenerate RuPB, so only one left to make glucose
-SO, CALVIN CYCLE NEEDS TO HAPPEN 3 TIMES TO GENERATE 1 G3P (3C) AND NEEDS TO HAPPEN 6 TIMES TO GENERATE 1 GLUCOSE (6C)
-when we say we need 3 turns of the cycle, means we need 3 CO2 molecules (each CO2 molecules can make the cycle happen once)
Dark reactions don't directly depend on light, but they depend on the outputs (NADH and ATP) of the light reactions, so they technically do depend on them somehow (which is why photosynthesis could not happen in the dark using only the dark reactions)
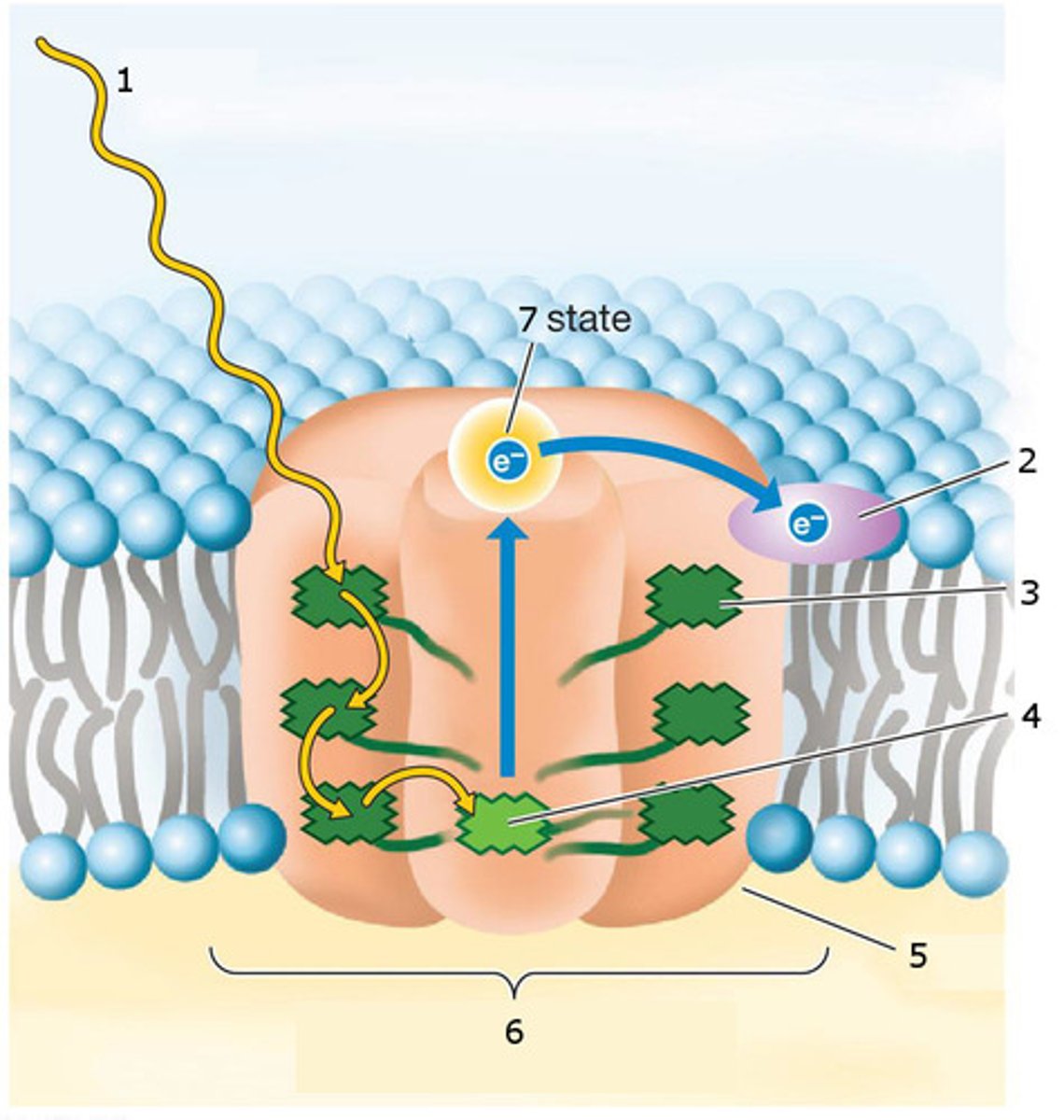
Structure and function of a photosystem - what do the different parts do?
protein complexes in the thylakoid membrane that are able to absorb light energy
each photosystem has a reaction centre that contains 2 chlorophyll a
they have light-harvesting antennae (mainly made of chlorophyll b) that take the light and bring them to the reaction centre
when chlorophyll a receives the energy brought by the antennae, both chlorophylls get oxidized (loss of electron as the electron gets excited)
P700: chlorophyll a of PSI, because it absorbs light at 700nm
P680: chlorophyll a of PSII, because it absorbs light at 600nm