Control of gene expression
1/75
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
76 Terms
What is a gene mutation?
The alteration of a base in the sequence of bases for one gene.
Likely to happen during DNA replication.
Spontaneous but mutagenic agents increase frequency.
Mutagenic agents
High energy and ionising radiation e.g. X-ray
Disrupt structure of DNA
Carcinogens e.g. cigarettes
Interfere with transcription
Addition mutations
Frameshift
When one base is added
Altered codons could potentially code for different amino acids→ different 3D tertiary structure → non-functioning protein.
Deletion mutation
Deletion of a base in a sequence
Frame shift
Also can lead to non-functioning protein.
Substitution mutation
Base has been changed for another base but number of bases remains the same.
No frame shift, only one codon changing and due to degenerate nature of genetic code, same amino acid and therefore have no impact.
Inversion mutation
A section of bases detach from DNA, but when they re-join they are inverted, so this section of code is back to front.
This results in different amino acids being coded for in this.
No frameshift but could still affect polypeptide as it can affect multiple codons.
Translocation
A section of bases on one chromosome detaches and attaches onto a different chromosome.
This is a substantial alteration and can cause significant impacts on gene expression and therefore the resulting phenotype.
What are stem cells?
Multicellular organisms have a diverse range of specialised cells that all originate as undifferentiated stem cells.
Stem cells are undifferentiated cells that can continually divide and become specialised (differentiation).
Totipotent
Stem cells that can divide and produce any type of body cell.
Found during early development of embryo for limited time.
During development, totipotent cells translate only part of their DNA, resulting in cell specialisation.
Pluripotent
Stem cells are found in embryos and can become almost any type of cell.
They are used in research with the prospect of using them to treat human disorders.
These stem cells could be used to regrow damaged cells in humans, such as replace burnt skin cells, or neurones in Parkinson’s disease sufferers.
There are issues with this as sometimes the treatment doesn’t work or stem cells continually divide to create tumours.
Additionally, ethically there is debate on whether it is right to make a therapeutic clone of yourself, to make an embryo to get a stem cells and then destroy it.
Multipotent and unipotent
These stem cells are found in mature mammals and can divide to form a limited number of different cell types.
Multipotent cells such as in bone marrow can differentiate into a limited number of cells whereas unipotent cells can only differentiate into one type of cell.
Sources of stem cells in mammals:
Early days of embryonic development development and slightly later but still early is pluripotent stem cells.
Umbilical cord contains multipotent stem cells.
Placenta also has multipotent.
Adult stem cells such as in bone marrow are a source of multipotent cells e.g B and T cells
Unipotent stem cells can be found in tissue e.g. in heart muscle cardiomyocytes-specialised muscle cells-can be found.
Induced pluripotent stem cells
Pros and cons of using stem cells to treat human disorders.
Induced pluripotent stem cells can be produced from adult somatic cells using appropriate protein transcription factors to overcome some of the ethical issues with using embryonic stem cells.
Turning on all genes in a cell means it’s no longer specialised.
How are pluripotent stem cells induced?
Adult unipotent cells and used and can be from almost any body cell.
Altered in the lab by turning off genes to make the cell specialised, so it is no longer specialised, using transcriptional factors, returning them to a state of pluripotency.
What is one way gene expression is controlled?
Epigenetics
What is meant by gene expression?
A protein is created: includes transcription and translation
What is epigenetics?
The heritable change in gene function without changing the DNA base sequence.
Changes are caused by environmental changes and can inhibit transcription.
Control of gene expression
Factors such as diet, stress and toxins can add epigenetic tags to the DNA, and this can control gene expression in eukaryotes.
A single layer of chemical tags on the DNA is called the epigenome and this impacts the shape of the DNA-histone complex and whether the DNA is tightly wound so won’t be expressed or unwound so it will still be expressed.
If the DNA is tightly wound, then transcription factors cannot bind and therefore the epigenome, which is due to changes in the environment can inhibit transcription.
What is heterochromatin?
When the histone is tightly coiled and transcription is inhibited due to increased methylation and decreased acetylation.
Euchromatin on the other hand is when genes will be expressed as there is decreased methylation or increased acetylation, making the DNA less tightly coiled.
Methylation of DNA
Increased methylation of DNA inhibits transcription.
When methyl groups are added to DNA, they attach to the cytosine base.
This prevents transcriptional factors from binding and attracts proteins that condense the DNA-histone complex.
In this way, methylation prevents a section of DNA from being transcribed.
Acetylation of histone proteins
Decreased acetylation of associated histone proteins on DNA inhibits transcription.
If acetyl groups are removed from the DNA then the histone become more positive and are attracted more to the phosphate group on DNA.
This makes the DNA and histones more strongly associated and hard for transcription factors to bind.
Epigenetics and cancer
Tumour Suppressor genes
These genes produce proteins to slow down cell division and to cause cell death if DNA copying errors are detected.
If a mutation results in the tumour suppressor gene not producing the proteins to carry out this function, then cell division could continue, and mutated cells would not be identified and destroyed. BRCAI and BRCA2 are two known mutated tumour suppressor genes that are linked to breast cancer.
This leads to abnormal methylation so tumour suppressing gene could become hypermethylated and this could result in the gene being inactivated and turned off.
The opposite could occur in oncogenes, as they may be hypomethylated, reducing the number of methyl groups attaches, meaning the gene is permanently switched on and is continually producing proteins that trigger DNA replication.
Where to transcription factors move from and to?
Cytoplasm into nucleus to turn genes on and off.
Where does transcription control occur?
In the nucleus
Transcription factors are proteins that can bind to different base sequences on DNA, initiating transcription of genes.
One part binds to DNA, and other part binds to a receptor.
Once bound, transcription begins, creating the mRNA molecule for that gene which can then be translated in the cytoplasm to create the protein.
Without this binding, the gene is inactive and the protein wouldn’t be made.
Oestrogen
Oestrogen is a steroid hormone that can initiate transcription by binding to a receptor site in the transcriptional factor.
When it binds to the transcription factor, it causes it to change shape slightly, and this makes it complementary and able to bind to DNA at promotor region to initiate transcription as RNA polymerase attaches after transcriptional factor attaches and the active site of RNA polymerase is only complementary to the DNA and transcriptional factor together.
Steroid means lipid soluble, can diffuse through plasma membranes simply into cytoplasm.
After binding, it then diffuses into nucleus.
What is RNA interference?
When an mRNA molecule that has already been transcribed gets destroyed before it is translated to create a polypeptide chain.
This is done by using small interfering miRNA, siRNA.
This is translational control.
RNAi- RNA interference with miRNA
What is cancer a result of?
Mutations in genes that regulate mitosis.
If these genes mutate and non-functioning proteins are made, then mitosis is not regulated, and, it results in the uncontrollable division of cells and the creation of a tumour.
Benign tumours
Can grow very large but at a slow rate.
Non-cancerous because they produce adhesion molecules, sticking them together and to a particular tissue, they are often surrounded by a capsule, so they remain compact and can be removed by surgery and rarely return.
The impact is localised and often not life-threatening, depending on the tumour location.
Malignant tumours
They are cancerous and grow rapidly.
The cells nucleus become large and the cell can become unspecialised again.
They do not produce the adhesive, so instead metastasis occurs, meaning the tumour breaks off and spreads to other parts of the body, forming secondary tumours.
The tumour is not encapsulated and instead can grow projections into surrounding tissues and develop its own blood supply.
It can be life threatening and the removal of the tumour needs supplementary treatment (radiotherapy and chemotherapy) and recurrence is more likely.
Tumour development
The development of a tumour is due to a gene mutation in either the tumour suppressor gene and or oncogene, the abnormal methylation of tumour suppressor genes and oncogenes or increased oestrogen concentrations.
What are oncogenes?
Oncogenes are the mutated version of a proto-oncogene, which creates a protein involved in the initiations of DNA replication in interphase and mitosis cell division when the body needs new cells.
Oncogene mutations can result in this process being permanently activated to make cells divide continually.
Tumour suppressor genes
These genes produce proteins to slow down cell division and to cause cell death if DNA copying errors are detected and proteins that repair mistakes in DNA base sequence.
If a mutation results in the tumour suppressor gene not producing the proteins to carry out this function, then cell division could continue, and mutated cells would not be identified and destroyed.
BRCAI and CRCA2 are the two known mutated tumour suppressor genes that are linked to breast cancer.
Abnormal methylation
This links to control of transcription-methylation can cause a gene to turn on or off.
Tumour suppressor genes could become hypermethylated, meaning an increased number of methyl groups attached to it.
This results in the gene being inactivated and become turned off.
The opposite could occur in oncogenes, as they may be hypomethylated, resulting in the gene being permanently switched on.
Increased oestrogen concentration
Oestrogen is produced by the ovaries to regulate the menstrual cycle, but after the menopause this stops.
Instead, fat cells in breast tissues can produce oestrogen and this has been linked with causing breast cancer in women post menopause because production is localised.
This has a knock on effect as the tumour then results in even more oestrogen production which increases the tumour size and attracts white blood cells which can increase the tumour size further.
This could be because oestrogen can activate a gene by binding to a gene that initiates transcription, and if this is a proto-oncogene the result is it is permanently turned on and activating cell division.
What does sequencing a genome mean?
Working out the DNA base sequence for all the DNA in a cell
Sequencing in simpler organisms
Prokaryotic cells do not contain introns in their DNA.
This means that the genome can be used directly to sequence the proteins that derive from the genetic code (proteome) of the organism.
This is useful for many reasons including identifying potential antigens to use in a vaccine.
Complex organisms
More complex organisms have introns and regulatory genes in their DNA.
Regulatory genes are part at beginning and end of gene that aren’t directly coding for an amino acid.
Due to this, the genome cannot easily be used to translate the proteome.
Recombinant DNA technologies flowchart
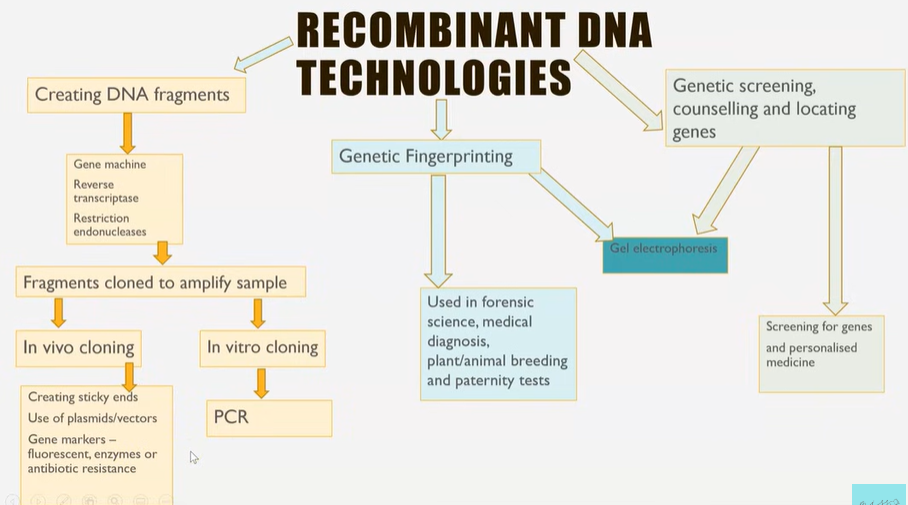

What can recombining different organisms DNA enable?
Scientists to manipulate and alter genes to improve industrial processes and medical treatments.
The first step in this technology is isolating the fragments of DNA, so it can be recombined with another piece of DNA.
There are 3 methods to create fragments of DNA.
What are the three methods?
Reverse transcription
Restriction endonucleases- enzymes
Gene machine
Reverse transcription
Enzyme reverse transcriptase makes DNA copies from mRNA.
Naturally occurs in viruses, HIV.
A cell that naturally produces the protein of interest is selected e.g. beta cell for human insulin.
These cells should have large amounts of mRNA for the protein.
The reverse transcriptase enzyme joins the DNA nucleotides with complementary bases to mRNA sequence.
Single stranded DNA is made- cDNA (c for complementary).
To make this DNA fragment double stranded, the enzyme DNA polymerase is used, and add more DNA nucleotides.
PCR can be used to make lots of copies of the DNA fragment that has just been made (later video).
Restriction endonucleases
Restriction endonucleases are enzymes that cut up DNA.
These naturally occur in bacteria as a defence mechanism.
There are many restriction enzymes that have an active site complementary to a range of different DNA base sequence, described as recognition sequences, and therefore each enzyme cuts the DNA at a specific location.
Some enzymes cut at the same location in the double strand at a blunt end, creating a blunt cut, other enzymes cut to create staggered ends and exposed DNA bases.
The exposed staggered ends are palindromic and referred to as ‘sticky ends’ because they have the ability to join to DNA with complementary base pairs.
Gene machine
DNA fragments can be created in a lab using a computerised machine.
Scientists first examine the protein of interest to identify the amino acid sequence, and from that work out what the mRNA and DNA sequence would be.
The DNA sequence is entered into the computer, which checks for biosafety and biosecurity that the DNA being created is safe and ethical to produce.
The computer can create small sections of overlapping single strands of nucleotides that make of the gene, called oligonucleotides.
These can then be joined to create the DNA for the entire gene.
PCR can be used to amplify the quantity to make the double strand.
Also intron free DNA, so can be transcribed in prokaryotic cells.
Advantages and disadvantages of reverse transcription
Introns are already removed as mRNA is being used, so cDNA is intron free, which is important for genetic engineering involving prokaryotic cells e.g. bacteria.
mRNA present in cells from actively transcribed genes, so lots of the mRNA of interest to make cDNA.
More steps, so more time consuming and technically more difficult.
Advantages and disadvantages of restriction endonucleases
Sticky ends on DNA fragment make it easier to insert to make recombinant DNA.
Does still contain introns though.
Advantages and disadvantages of gene machines
Doesn’t contain introns.
Can design exact DNA fragment you want, with sticky ends, labels and preferential codons.
Need to know the sequence of amino-acids or bases.
What are the two methods of amplifying DNA fragments via cloning?
Invitro and in vivo cloning.
Invitro means not inside of and happens via PCR- polymerase chain reaction.
PCR
Done in an automated machine.
Equipment list: thermocycler, DNA fragment to be amplified, DNA polymerase, primers and DNA nucleotides.
The temperature is first increased to 95 degrees C, breaking hydrogen bonds and splitting the DNA into single strands (denaturing).
The temperature is then decreased to 55 degrees C so that primers can attach and hydrogen bonds reform, holding primers in place (annealing).
The enzyme DNA polymerase then attaches complementary free nucleotides and makes a new strand to align next to each template (synthesis). The temperature is increased to 72 degrees C for this stage, the optimum for DNA polymerase.
Advantages of PCR
Automated- more efficient
Rapid- 100 billion copies of DNA can be made within hours.
Doesn’t require living cells- quicker and less complex techniques needed.
In vivo cloning: Different stages
Insert DNA fragment into a vector.
Transform a host cell with the vector.
Identify transformed cells.
Grow the host cell ( clone/make copies of the gene)
Promoter and terminator regions
Restriction endonuclease enzymes are used to cut out the gene (DNA fragment) of interest.
These enzymes cut at recognition sites leaving sticky ends.
The DNA fragments must be modified to ensure transcription of these genes can occur.
A promotor region must be added- at the start of the DNA fragment. This is a sequence of DNA which is the binding site for RNA polymerase to enable transcription to occur.
A terminator region is added at the end of the gene, causing RNA polymerase to detach and stop transcription, so only one gene at a time is copied into mRNA.
Need to get this isolated DNA fragment into a vector.
Inserting DNA into a vector
A vector is something that carries the isolated DNA fragment into the host cell.
Plasmids are the most common vectors.
The plasmid is cut open using the same restriction endonuclease.
This creates the same sticky ends.
Therefore, the DNA fragment sticky ends are complementary to the sticky ends on the plasmid.
Enzyme ligase sticks the DNA fragment and plasmid together by catalysing the condensation reaction to form phosphodiester bonds between nucleotides.
Transforming a host cell with the vector
The vector (plasmid with the recombinant DNA) next needs to be inserted into the host cell, where the gene will be expressed to create the protein required.
To do this, the cell membrane of the host cell must be more permeable.
To increase the permeability, the host cells are mixed with Ca2+ and heat shocked (sudden increase in temperature).
This enables the vector to enter the host cell’s cytoplasm.
Why do we need to identify which bacteria have taken up the recombinant plasmid before it is grown on mass?
Not all bacteria will take up the recombinant plasmid.
3 reasons/ issues:
Recombinant plasmid doesn’t get inside the cell.
The plasmid re-joins before the DNA fragment entered.
The DNA fragment sticks to itself rather than inserting into the plasmid.
Ways of identifying which bacteria successfully took up the recombinant plasmid and are transformed.
Marker genes can be used and 3 different ones:
Antibiotic resistant genes
Genes coding for fluorescent proteins
Genes coding for enzymes
Gene does this by coding for a protein allowing identification
Antibiotic resistant genes
Insert a gene that makes bacterial plasmid resistant to tetracycline gene and the ampicillin gene- they are antibiotics.
Insert DNA fragment that was isolated
Inserted into middle of tetracycline gene so it is disrupted and will no longer create a functional protein that makes the bacteria resistant.
Will still be resistant to ampicillin and bacteria is grown on agar.
To identify which of these colonies took up the recombinant plasmid, use a sterile velvet block and place this sheet over front of petri dish and transfer the colonies to a plate with ampicillin antibiotic in the agar.
Leave for days and colonies that do still grow must have plasmid inside of it as gene produces proteins making it resistant.
Use velvet block again and transfer colonies to a plate with tetracycline antibiotic within the agar.
If colonies can still grow then they are resistant to both and must be the original plasmid and those that don’t continue to grow contain the recombinant plasmid of interest and are taken and grown further.
Fluorescent markers
Jellyfish contain a gene which codes to create a green fluorescent protein.
This GFP gene can be inserted into the bacteria plasmid.
DNA fragment is inserted in the middle of the GFP gene, disrupting and preventing GFP production among bacteria that take up recombinant DNA plasmid.
This is grown on agar and this is exposed to UV light, colonies with GFP still intact glow.
Those that aren’t glowing are the colonies of interest as they took in the recombinant plasmid.
Enzyme markers
The enzyme lactase can turn a certain substance blue from colourless.
This gene is inserted into the plasmid and DNA is inserted in the middle of the gene to disrupt this so any bacteria that take in recombinant plasmid will not be able to produce lactase to turn substance blue.
This is grown on a colourless substance and colonies that cannot turn it blue contain plasmid of interest and will be collected and taken to grow further.
Grow the host cell
Once bacteria containing recombinant plasmid, and therefore section of DNA we want to clone is collected, a fermenter is used to grow multiple copies of the host cell.
This large, cloned population can then produce the protein coded for by the inserted DNA fragment e.g. bacteria producing insulin from the inserted insulin gene.
What is genetic fingerprinting?
The examination and analysis of VTNR FNA fragments.
95% of human DNA is made up of introns which consist of many variable number tandem repeats.
This refers to long sequences of DNA bases that are repeated but don’t code for anything.
The probability of two individuals having the sane VNTRs is very low, however the more closely related you are, the more similar the VNTRs are.
What can VNTRA’s be used to determine?
Genetic relationships e.g. paternity tests and the genetic variability within a population.
Genetic fingerprinting stages: overview
Collection
Extraction
Digestion
Separation
Hybridisation
Development
Analysis
Collection and extraction
The smallest sample of DNA can be collected for genetic fingerprinting.
This can be from blood, body cells or hair follicles.
If the sample of DNA is small, then PCR is used to amplify the amount of DNA.
Digestion
Restriction endonucleases are added to cut the DNA into smaller fragments.
Add those with active sites which are close to and therefore cut close to the target VNTRs, so entire length of VNTR can be maintained.
Separation
The DNA samples are loaded into small wells in agar gel.
One individual’s DNA in one well.
The gel is placed in a buffer liquid with an electrical voltage applied.
DNA is negatively charged, so the DNA samples move through the gel towards the positive end of the gel.
Gel electrophoresis:
The agar gel creates resistance for moving DNA, and the smaller pieces of DNA can move faster and further along the gel.
This is how the different lengths of DNA, VNTR’s are separated.
An alkaline is then added to separate the double strands of DNA.
Hybridisation
DNA probes are short, single stranded pieces of DNA complementary in base sequence to the VNTR’s.
The probes are radioactively or fluorescently labelled.
Different DNA probes are mixed with single stranded DNA VTNR’s on the agar gel for them to bind- hybridise.
Development
The agar gel will shrink and crack as it dries, and therefore the VNTR’s and DNA probes are transferred to a nylon sheet.
The nylon sheet can then be exposed to x-rays to visualise the position of radioactive gene probes, or UV light is fluorescence probes were used.
Analysis
The position of the DNA bands are compared to identify genetic relationships, the presences of a disease causing gene and to match unknown samples from a crime scene.
Uses of genetic fingerprinting
Forensic science to place suspects at crime scenes.
For medical diagnosis
To ensure plants and animals are not closely related before being breed.
What are DNA probes?
Short, single-stranded DNA that has been fluorescently or radioactively labelled so they can be identified and located.
This is used to locate specific alleles of genes and to screen patients for heritable conditions, drug responses or health risks.
DNA probes are created to have a complementary base sequence to the allele that is being screened for.
The patients DNA sample is treated to make it single-stranded and it is then mixed with the DNA probes.
If the patient has the allele, then the DNA probe will hybridise and the label indicates the presence.
DNA hybridisation
The patient’s DNA sample is heated to make it single-stranded.
The heat causes hydrogen bonds between bases to break, denaturing.
The patient’s single stranded DNA sample is mixed with the DNA probe and cooled to and any complementary base sequences can align and form hydrogen bonds (anneal).
Some of the patient’s DNA samples will anneal back together but some will anneal with the DNA probe.
Locating specific alleles of genes
To locate a specific allele, the DNA base sequence must be known to then create the DNA probe.
This can be determined using DNA sequencing techniques e.g. Sanger method.
This fragment can be amplified using PCR.
The label is then added, either a radioactive nucleotide containing the isotope 32 P or a fluorescent label which emits light under UV light.
After hybridisation, the DNA is washed so that any unbound DNA probes are washed away.
The presences of the radioactive or fluorescent or label indicates that the alleles of interest is present in the patient’s DNA.
Genetic screening
This method can be used to screen for potential disorders or for the presence of cancer causing oncogenes.
It is possible to screen for multiple diseases simultaneously using an array where multiple different DNA probes are attached.
Personalised medicine
This is one key reason that your DNA screened can be advantageous.
Certain drugs, such as painkillers are more or less effective depending on your genotype. It can also help determine the best dose, which increases the effectiveness, safety and can save money.
Vitamin E can be given to diabetics to reduce the risk of CVD for some, but for others with a different genotype it can actually increase the risk.
Genetic counselling
Having your DNA screened is a decision that must be thought through carefully and must be discussed either patients so they make an informed decision.
Genetic counselling is like a type of social work, where people can have their family history researched to consider the likelihood of them carrying any alleles linked to diseases before starting a family or for their general health. Patients are informed of any potential risks to themselves or future offspring if they were to carry the allele.
Some examples include screening for cystic fibrosis and sickle-cell anaemia prior to starting a family.
If there is a family history of breast cancer, you may want to be screened for alleles linked to this cancer.
If you find the allele is present you can be screened for tumours more frequently, reduce environmental risk factors or even opt for a mastectomy.