Bio 1
1/141
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
142 Terms
Cell theory fundamentals
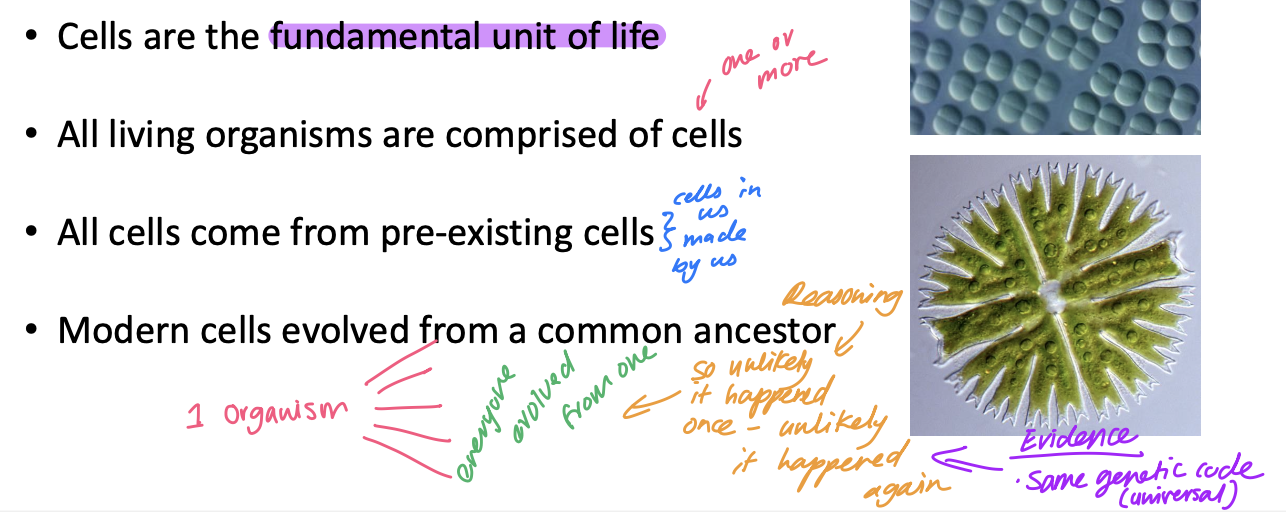
What do the similarities among living organisms indicate?
All life on Earth has a common ancestry
All organisms have arisen from earlier, more primitive forms over the past 4 billion years through the processesof evolution
Evidence that all organisms shared one common ancrstor
hey share the same
genetic code, chemical composition, and cellular structure
(“biochemical unity of life”)
Where did life come from?
Life formed spontaneously (BIOGENESIS) on early Earth (a reducing environment facilitated organic molecule formation, such as DNA bases)
Extra-terrestrial origin - life from another planet or comet (panspermia)
e.g. The Murchison Meterorite (w fatty acids + carbs)
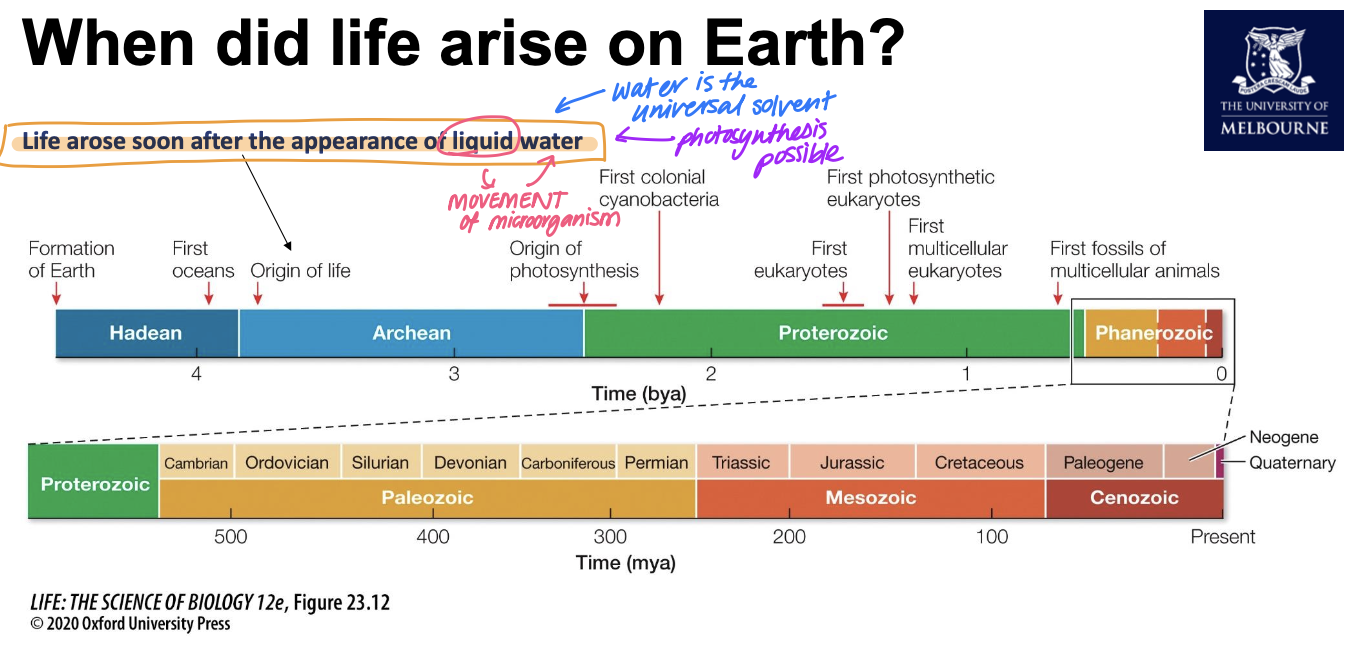
When did life arise on Earth?
around 3.5bil years ago
stromalites
Stromatolites thrive in the hyper-saline water of Hamelin Pools in Shark Bay,
Western Australia
scientists dated the water trapped in cyanobacteria (stromalites grew around cyanobacteria)
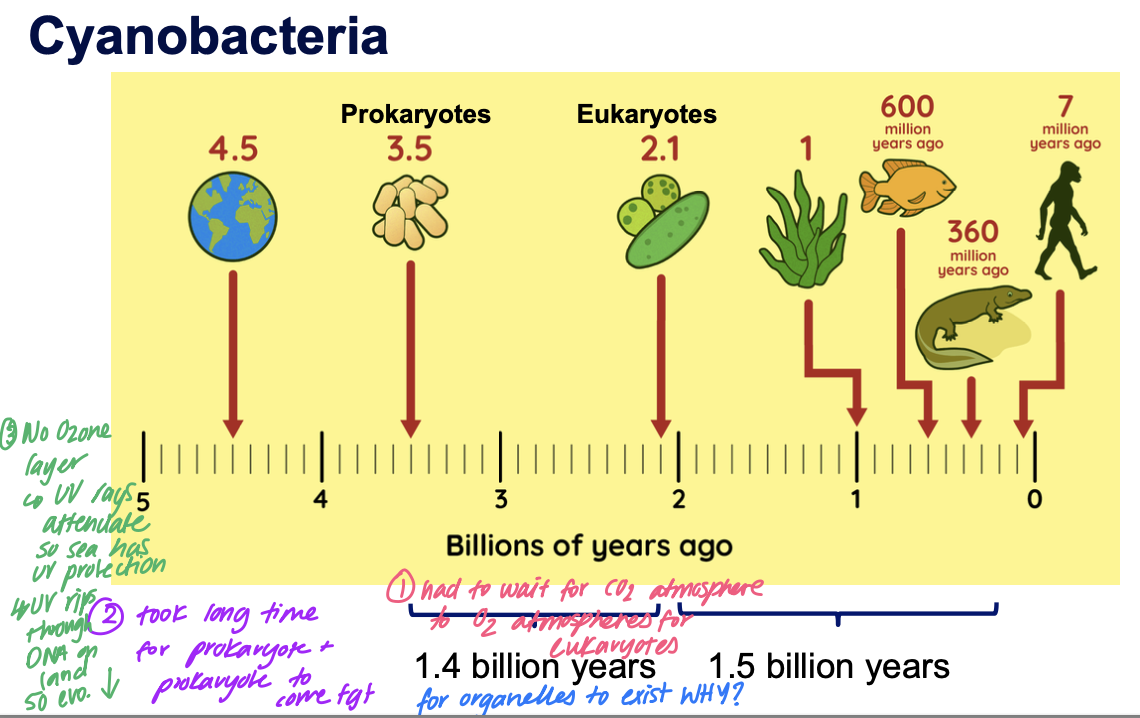
Cyanobacteria
Cyanobacteria arose around 3.5bil ya
Reasons for large time gaps bw bacteria, euk and terrestrial animals
The formation of the ozone layer
4 billion years ago, the atmosphere was primarily CO2 and CH4.
• As cyanobacteria performed photosynthesis, CO2 was converted to glucose and O2 gas.
• Ultraviolet light splits oxygen into two oxygen radicals, which can then react with other O2 molecules to create Ozone (O3).
• Ozone molecules congregate together to create a layer.
• This layer absorbs UV-C and partially absorbs UV-A and UV-B.
Before the ozone layer
UV rays attenuate so sea has UV protection but UV rips through DNA on land so can’t survive on land until oxygenation event
Consequences of oxygenation event
1.Alteration of the atmosphere that allowed for aerobic cellular respiration (Topic 2)
2.The Ozone layer provides a barrier to UV-C and partially to UV-A and UV-B, creating the conditions for the evolution of life on land.
Cambrian explosion
• A rapid diversification of life took place approximately 541 million years ago, which is commonly referred to as the Cambrian Explosion
• Most of the major animal groups living today appeared during the Cambrian
• For a long time, scientists had well preserved fossils from this time period but nothing earlier
• What did earlier life look like? - need hard parts of fossilise
Prokaryotic Domains or Superkingdoms
Two of the three domains of life are
Prokaryotic
• the Bacteria (Superdomain Prokarya)
• the Archaea (Superdomain Prokarya)
The Archaea are more closely related to the third domain, the Eukarya or Eukaryotes
Defining characteristics bw kingdoms of life
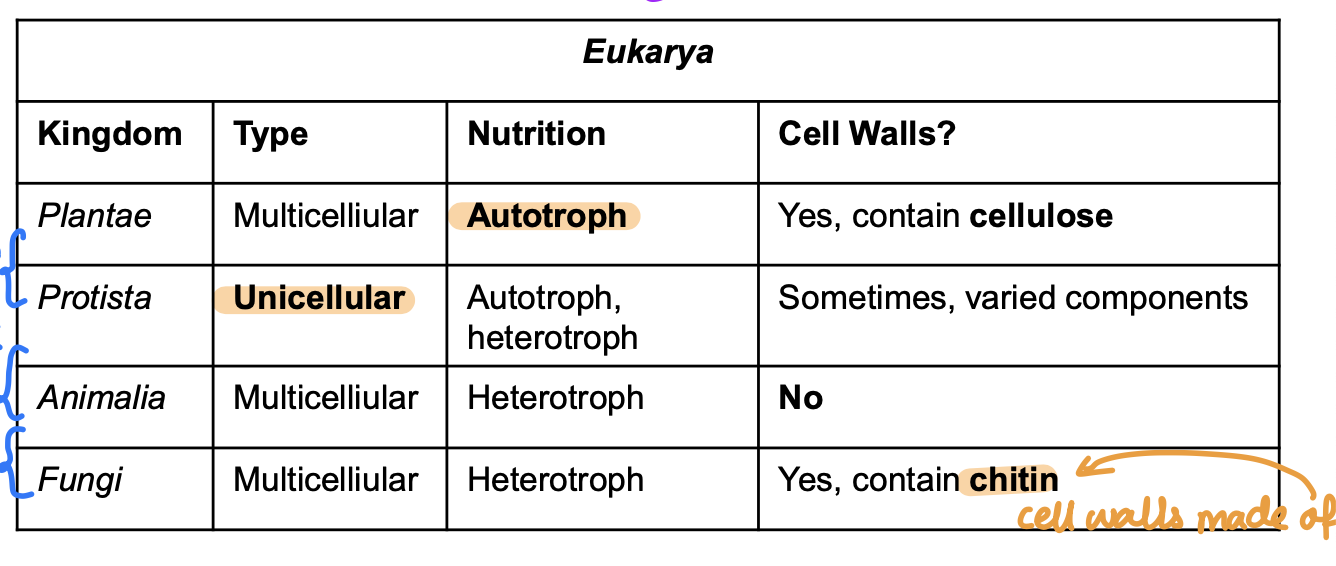
Autotroph vs Heterotroph
Autotroph: Makes its own food
Heterotroph: Must acquire food (eat)
Prokaryotic Cell
All prokaryote cells have:
• A cell membrane
• DNA located in a region called the nucleoid (NOT an organelle)
• Cytoplasm: the rest of the cell contents
• Ribosomes: sites of protein synthesis
Most bacteria have
Cell wall outside the plasma membrane (peptidoglycan)
Some bacteria have
• A capsule made of polysaccharides
• Cell wall and a capsule
• Provides protection from adverse environmental conditions
• Flagellum
• Pili (fimbriae)
• attachment structures, involved in exchange of DNA
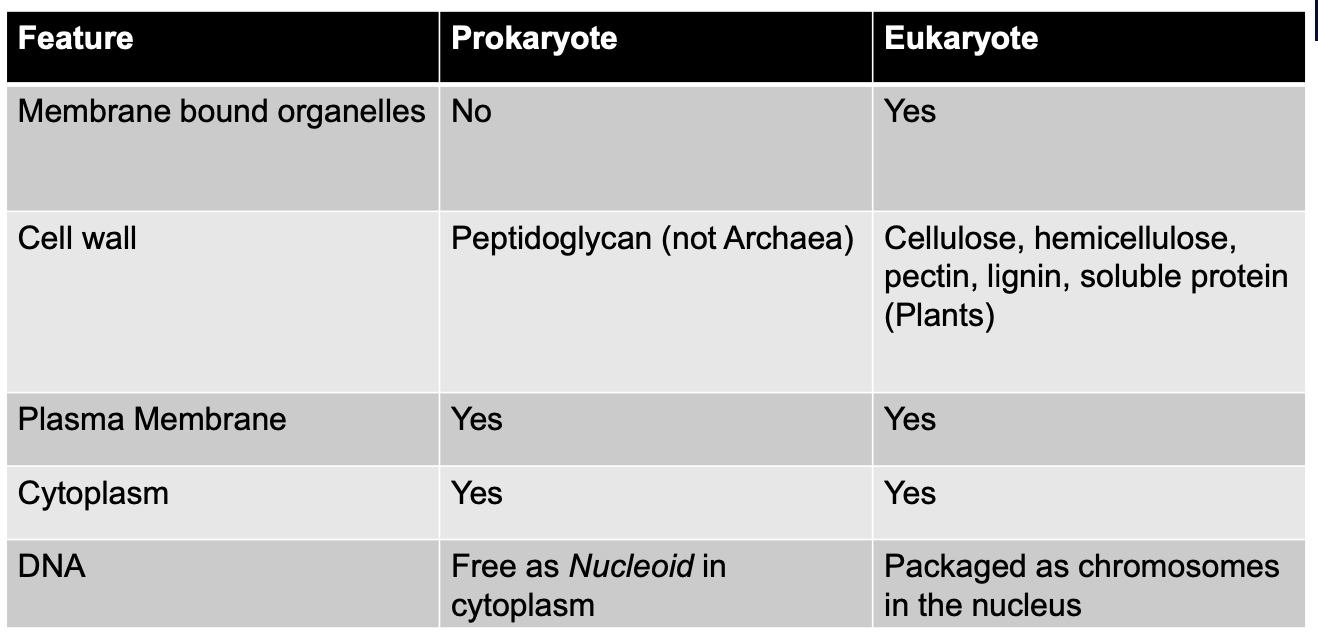
Prokaryote vs Eukaryote
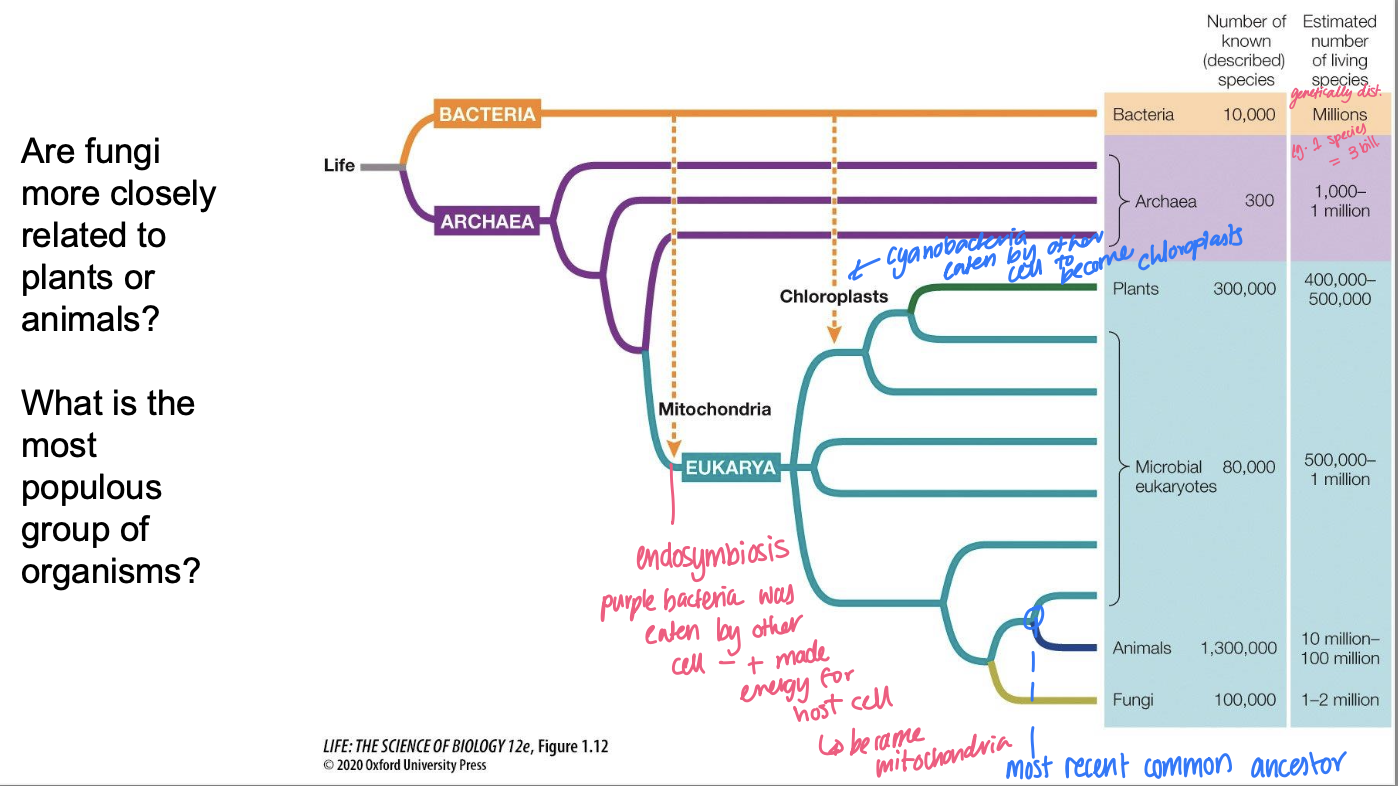
Phylogenetic trees
Electronegativity
The attraction of an atomic nucleus for outer/bonding electrons.
The most electronegative elements have the fewest shells, and highest positive nuclear charge.
They are VERY attractive to bonding electrons.
Things that effect electronegativity
distance from nucleus
strength of positive charge
Bond types
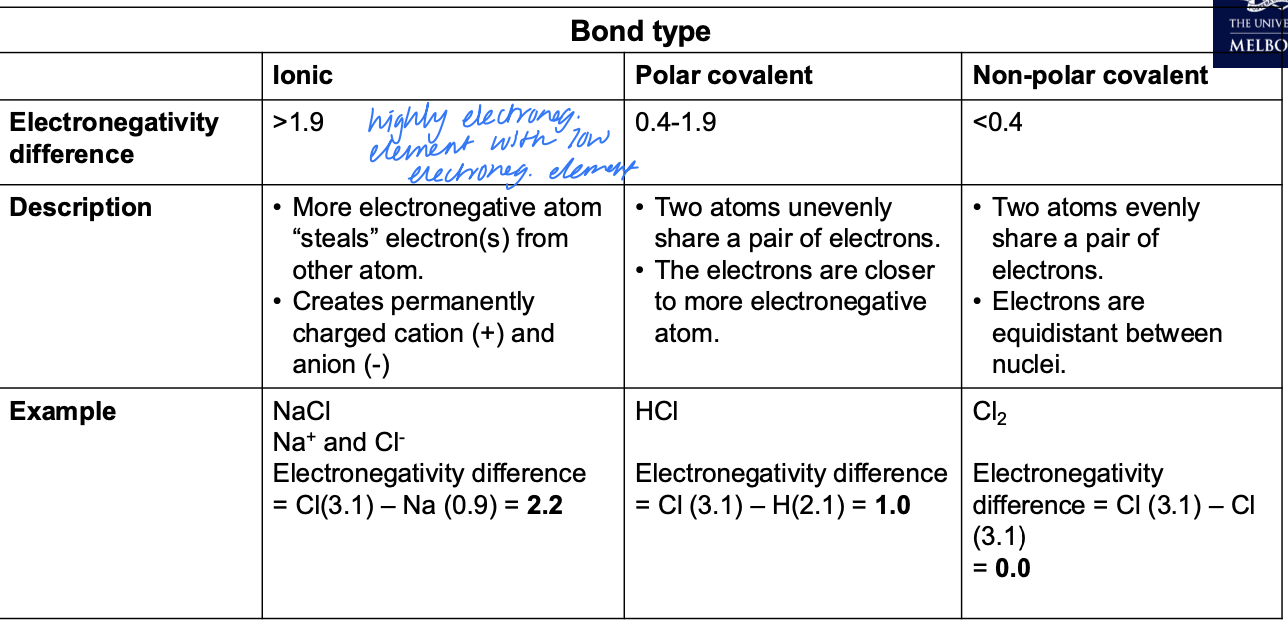
Functional Groups
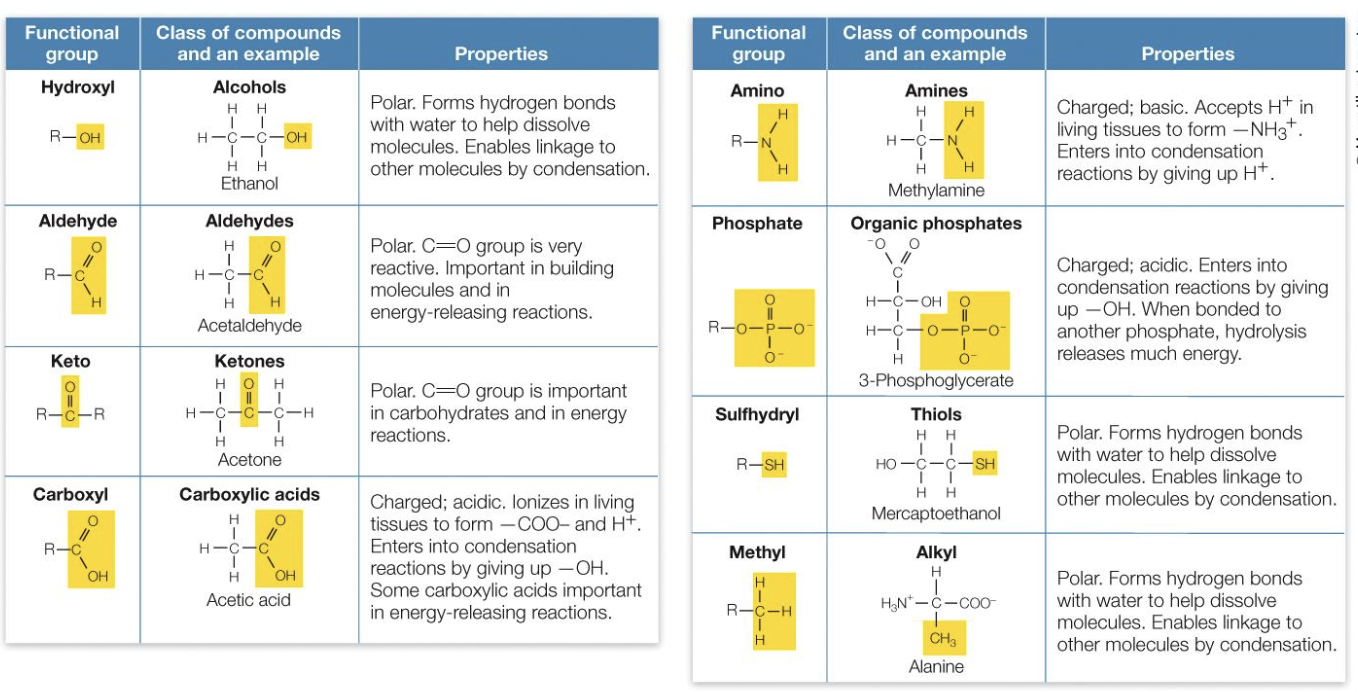
Macromolecules
Polymers containing thousands or more atoms. (Large lipids are treated as macromolecules).
All macromolecules except lipids are polymers of smaller molecules called monomers
Macromolecule function depends on the properties of functional groups. Each group has specific properties, such as polarity.
What makes molecule polar?
1) Has one more polar bonds.
2) Charge is asymmetrically distributed across the molecule.
Dipole dipole force
• The attractive force that exists between oppositely charged poles of two different molecules.
• Occurs between polar molecules.
• Also called hydrophilic interaction.
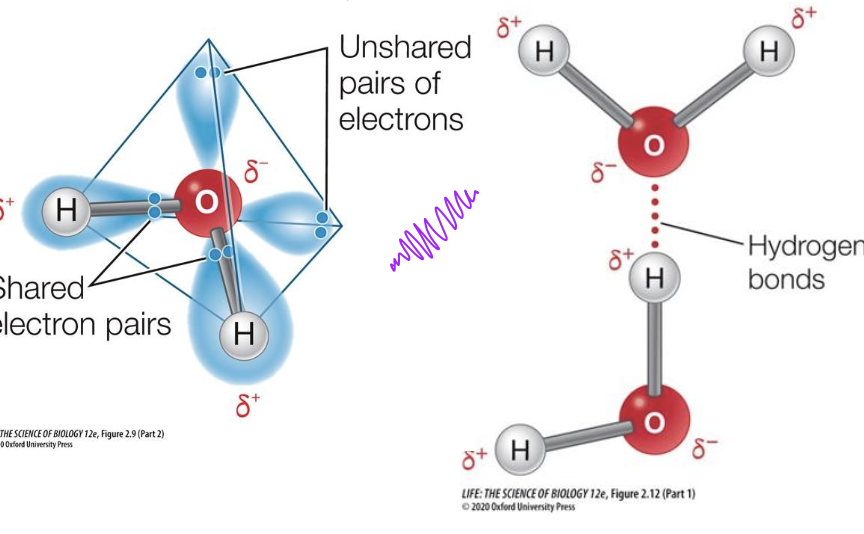
Hydrogen bonding
• Hydrogen has only 1 proton and 1 electron.
• When Hydrogen forms a covalent bond with Oxygen, Fluorine or Nitrogen the bond is extremely polar.
• The hydrogen nucleus becomes exposed - a naked positive charge.
• This naked proton is attracted to an unshared pair of electrons on a different atom, creating a hydrogen bond.
London Dispersion Forces
• Any molecule can have dispersion force, but it is the only force between non-polar molecules (e.g. fats, oils)
• Electrons are constantly shifting in the cloud around an atom. For a brief moment, more electrons are on one side of the molecule, causing it to polarize.
• The bigger a molecule is, the more electrons it has. This means the chance of Dispersion Force is higher.
• Also called a hydrophobic interaction. (LDE)
Liquid water
arose on Earth 3.8bill ya
most of water is ice - but liquid water allows movement of microbes so asexual reproduction can occur
The properties of water that are critical
Water molecules interact with each other through hydrogen bonding.
This gives water:
High specific heat (4.186 J g-1 °C-1).
High melting point (0 °C) and boiling point (100 °C) in comparison to other small molecules.
High heat of vaporization
Cohesion (hydrogen bonding between H2O molecules)
Adhesion (attraction of water to other molecules)
Chemical reactions of water

The effect of pH
pH influences rates of biological reactions and can change the 3-D structure of biological molecules, which impacts function.
Organisms use many mechanisms to minimise change in pH in their cells and tissues.
All organisms have an optimal pH for their internal systems.
Macromolecules
are biological polymers, composed of long chains of repeating subunits (monomers).
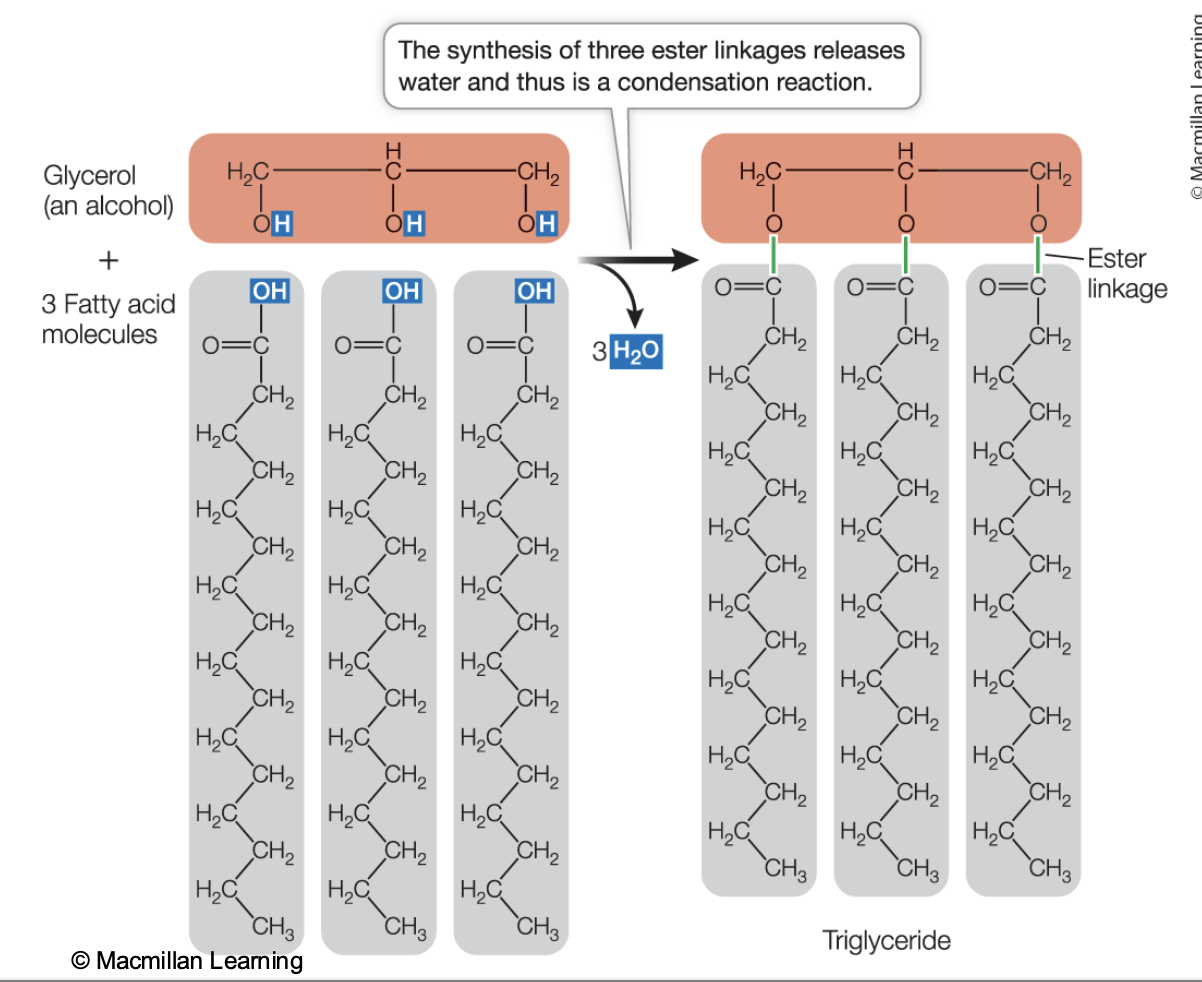
Condensation reaction
the hydroxyl (-OH) functional group of two monomers react with each other to create a new covalent bond, losing water in the process.
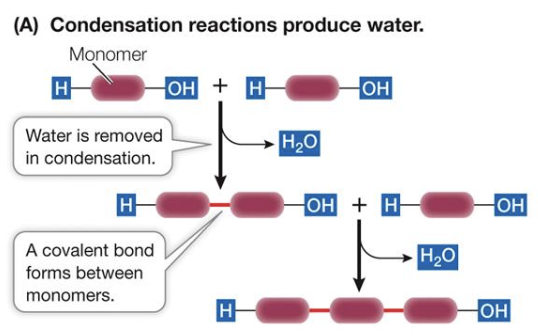
Hydrolysis
An enzyme-assisted reaction where water is added to a covalent bond, splitting it.
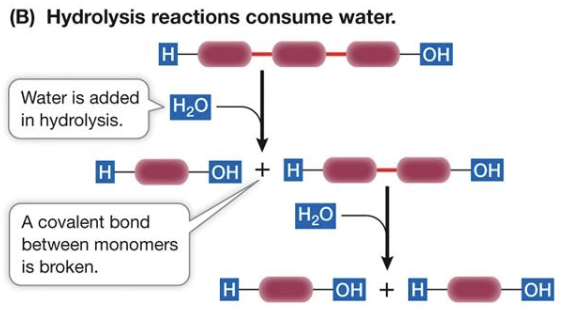
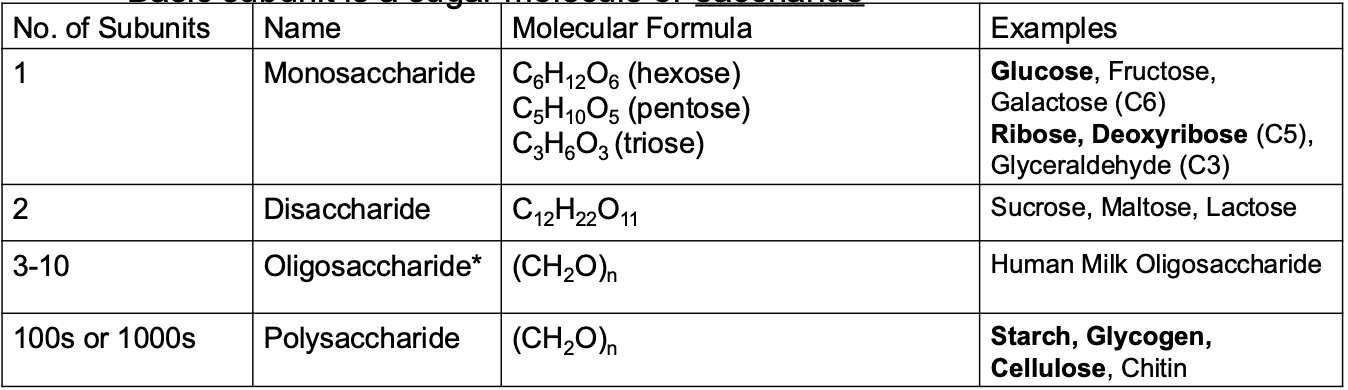
Carbohydrates
Carbohydrates are some of the most abundant organic compounds in nature
• Source of chemical energy
• Form structural components
Carbohydrate general formula
Have the general formula (CH2O)n
Carbohydrates fundamentals
• n is the number of carbon atoms. Generally n is between 3 – 8
• Often have the suffix –ose.
• Basic subunit is a sugar molecule or saccharide
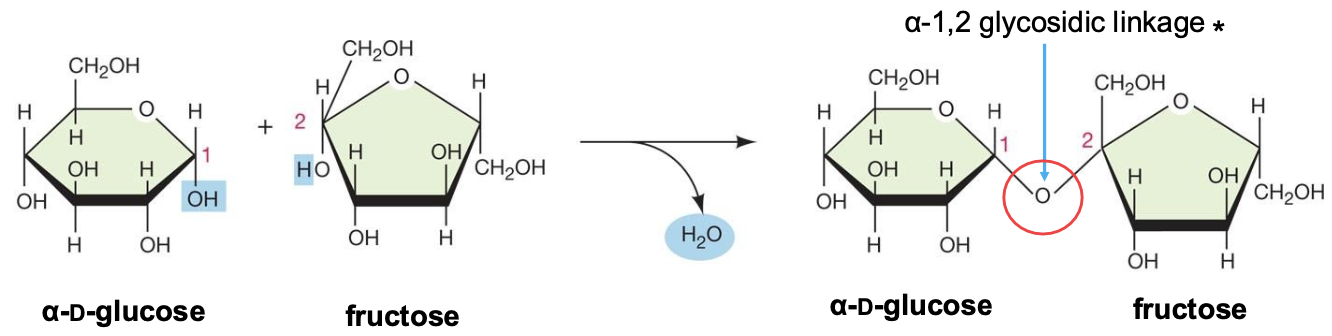
Glycosidic bonds
A disaccharide is produced from the condensation of two
monosaccharides
eg. maltose, lactose, sucrose
ether functional group but glycosidic is specific to carbs
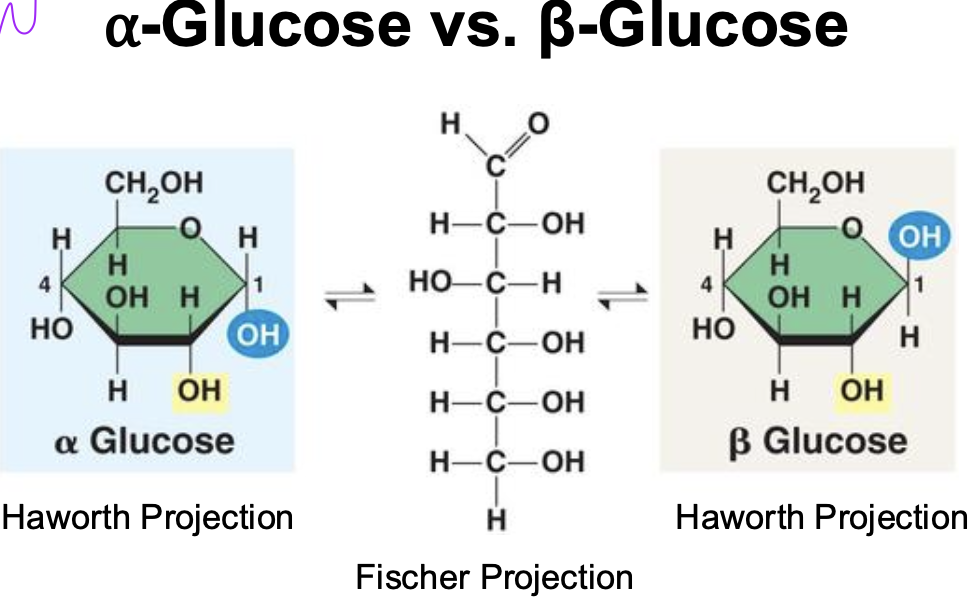
Alpha vs beta glucose
Position of OH varies bw alpha and beta due to carbon spinning so orientation changes
important bc that can change the shape of the bonds in polysaccharides
this is the fundamental difference between cellulose and starch
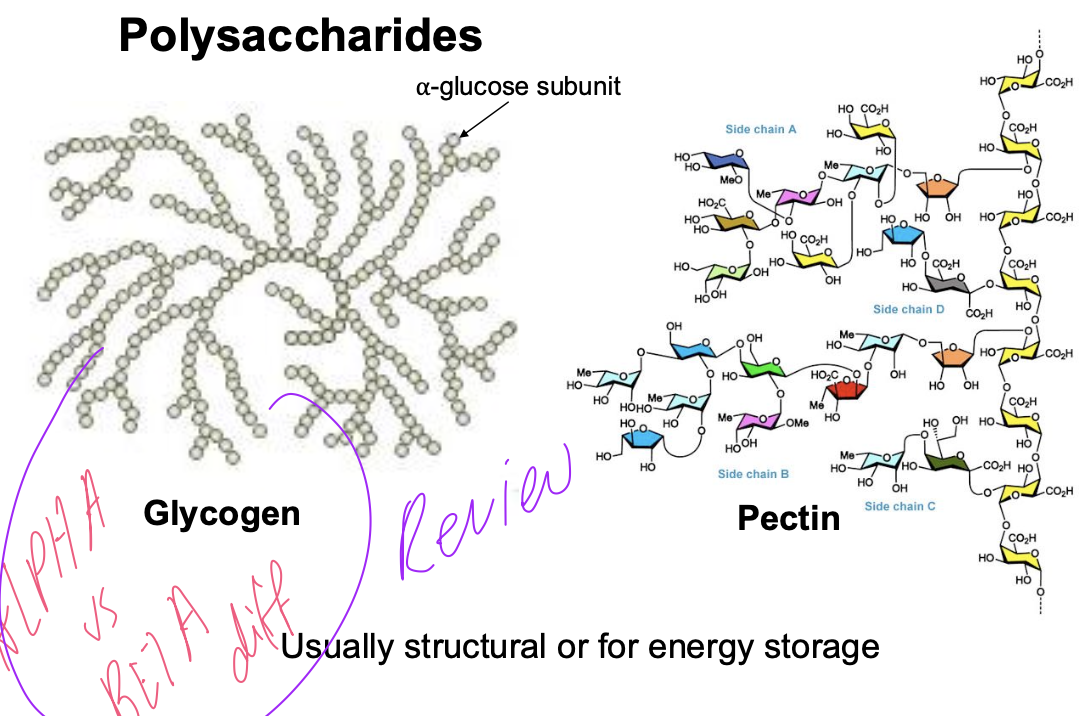
Polysaccharides
Glycogen is the focus - just made of glucose
Starch
Branched polysaccharide of ⍺-glucose
Easily degraded by enzymes (amylases)
Primary energy storage compound in plants
all of the glucose molecules have same orientation due to alpha glucose carbon 1 bonded to carbon 4 of the other alpha glucose
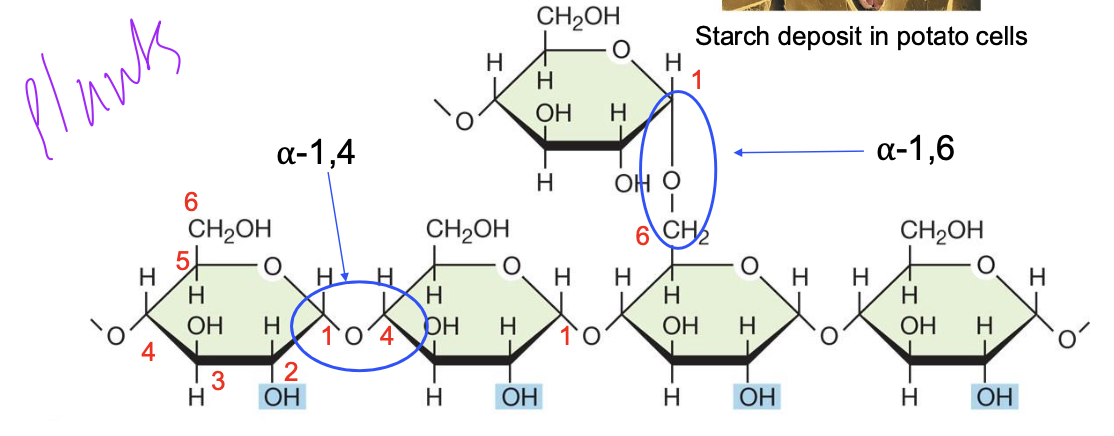
Glycogen
• Often called “animal starch” due to the same molecular structure and easily degraded
• Polymer of glucose with α-1,4 and α-1,6 glycosidic bonds. α-1,6 glycosidic bonds produce branching at carbon 6
• Primary energy storage compound in animals, fungi, and bacteria
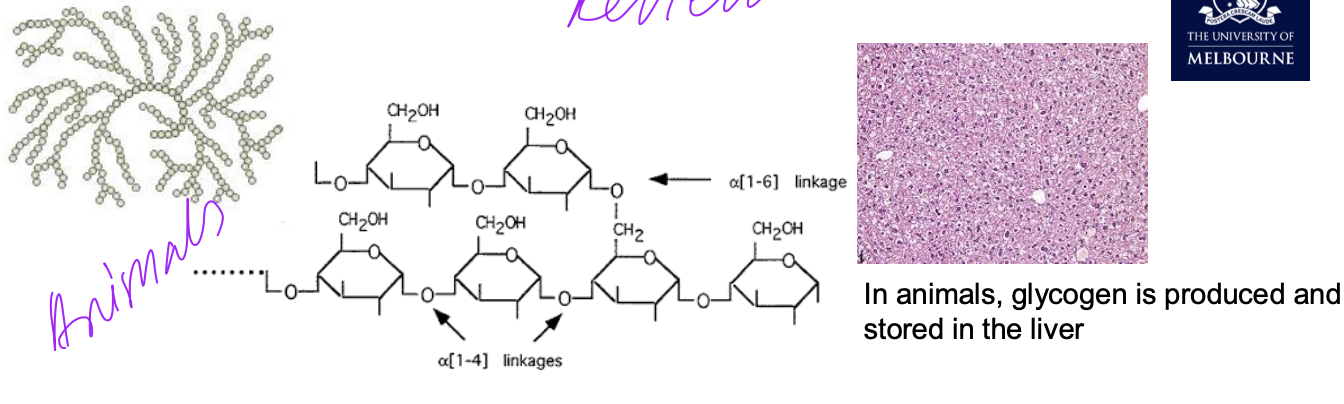
Cellulose
• Cellulose is the most abundant polymer on the planet.
• Linear polysaccharide
• Differs from starch (β-glucose rather than α-glucose)
• More chemically stable than starch
• Major structure of plant cell walls
• Chains are grouped together as microfibrils.
• Microfibrils bundle as fibrils.
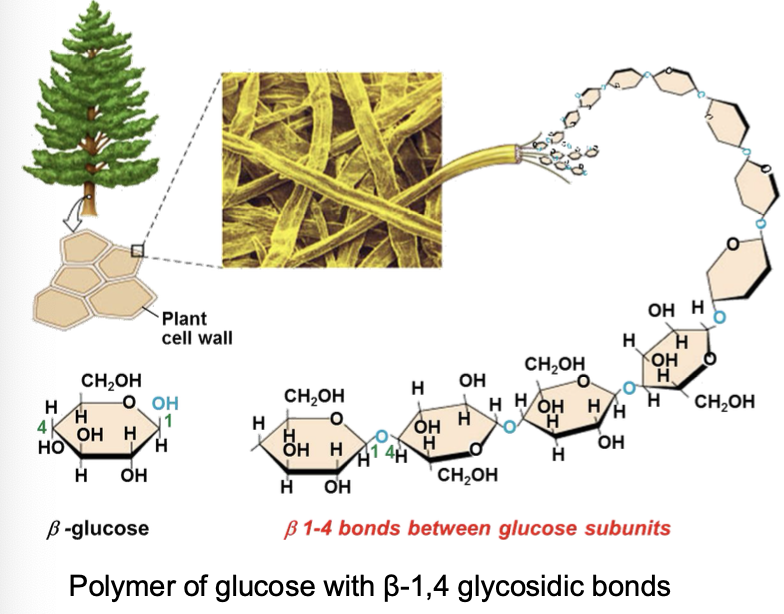
What chemical aspect of cellulose allows chains to bundle together as microfibrils?
strong H bonding and macromolecules which are huge so overall dispersion force which allows them to stay together
Lipids
• insoluble in water
• Non-polar. Held together by weak (but additive) London Dispersion Force only (but because they’re long - LDFs strong)
• dissolve readily in organic solvents
• composed mainly of C, H and O
• differ from carbohydrates due to a smaller proportion of oxygen
• may contain other elements (e.g. phosphorus and nitrogen)
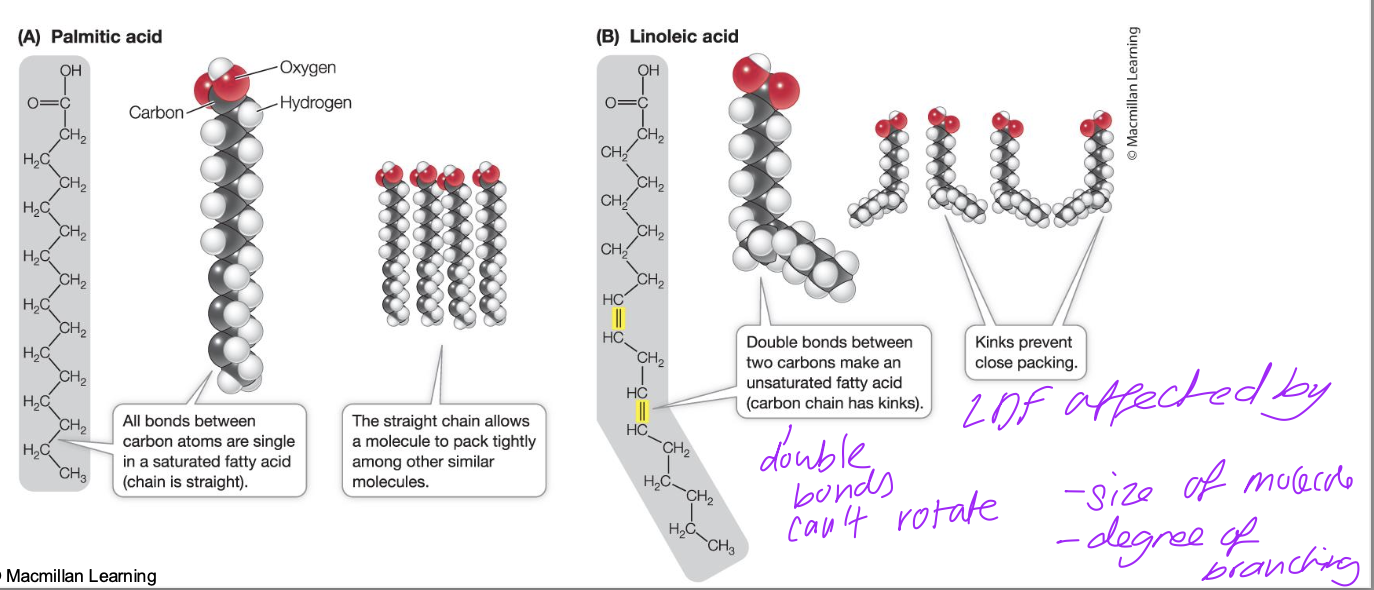
Saturated and Unsaturated Fatty Acids
double bonds can’t rotate so rigid and there’s kinks in fatty acid chain so they can’t pack closely
dispersion forces weaker
Synthesis of triglyceride
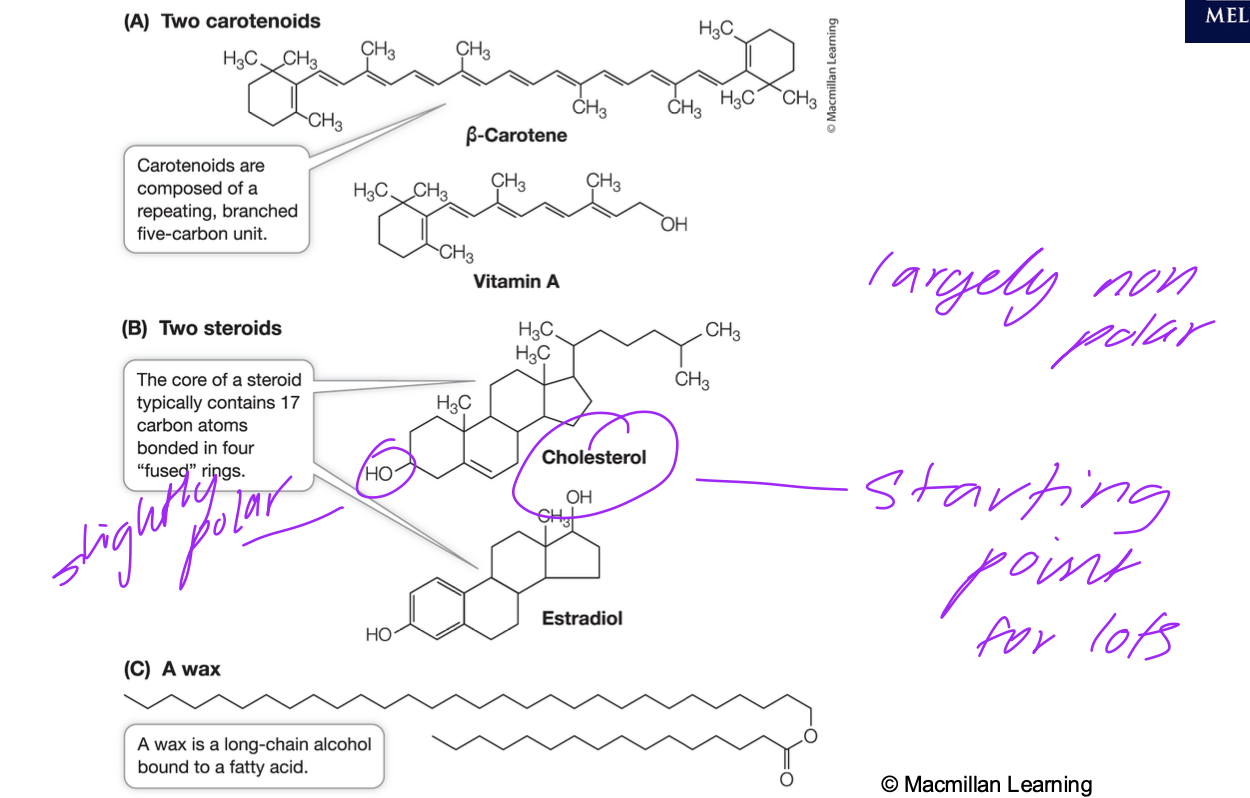
Common Lipids
Lipids for insulation (and energy)
• Blubber is important in marine mammals as it stores energy, insulates heat, and increases buoyancy
• Blubber lipid contains ca. 60% triglyceride (fat) and ca. 25–30% free fatty acid
Phospholipids
Amphipathic compound
Can interact with polar and non polar molecules, and ions. Can form ionic and ion-dipole interactions with head. and H-bonding
Can form LDF with other non polar compounds with tail.
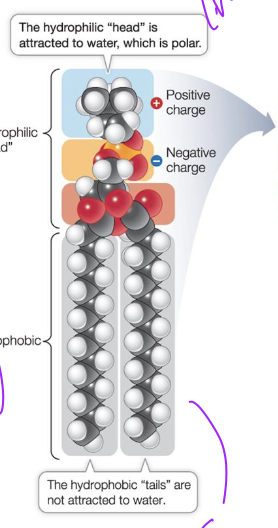
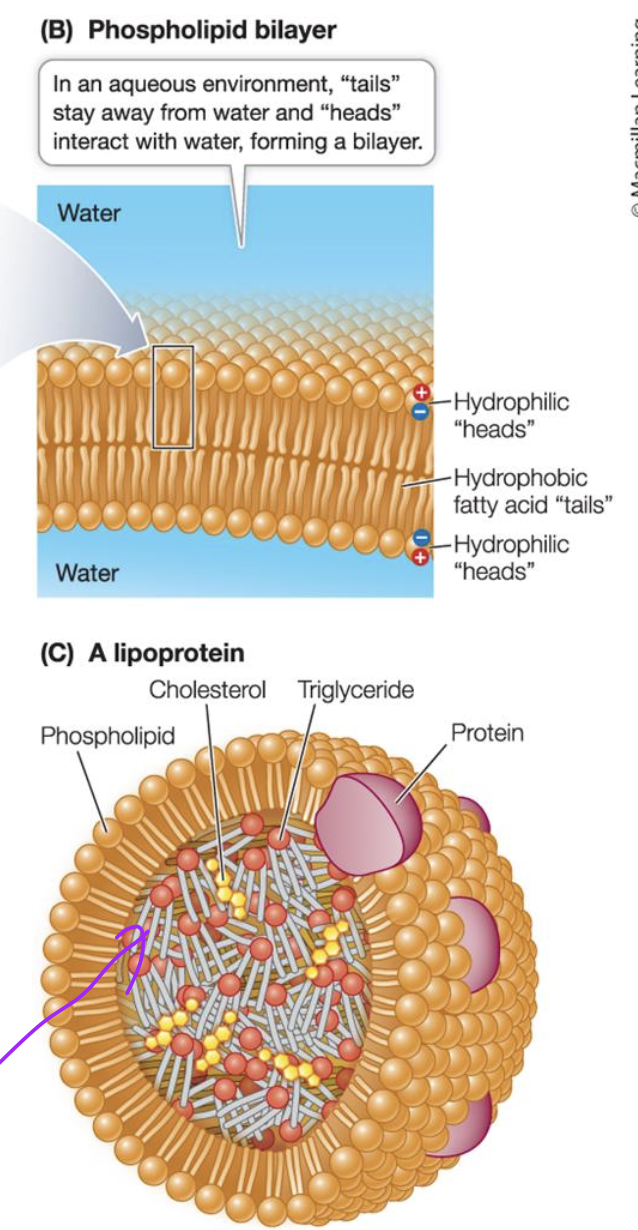
Phospholipid bilayer
This allows compartmentalisation with different environment on either side of bilayer.
allows diff pH inside vs outside and organelles in cytosol
Summary of Carbs vs Lipids
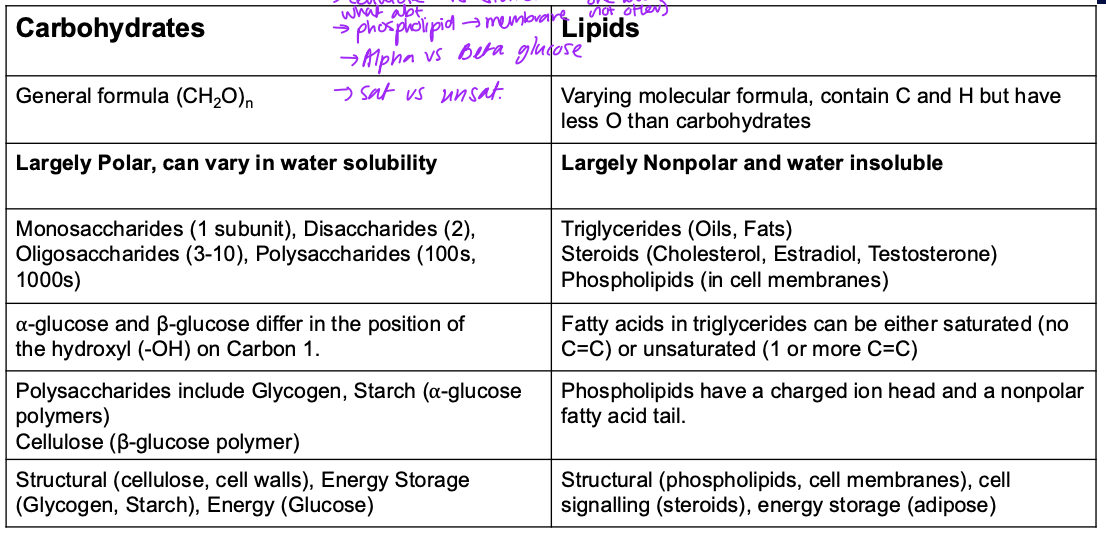
Basic role of nucleic acids
information storage
Types of nucleic acids
DeoxyriboNucleic Acid (DNA) and RiboNucleic Acid (RNA)
DNA nucleotides
• Adenine - (A) (comp. T)
• Cytosine - (C) (comp. G)
• Guanine - (G)
• Thymine - (T)
RNA nucleotides
• Adenine - (A)
• Cytosine - (C)
• Guanine - (G)
• Uracil - (U) instead of Thymine
Building up a nucleotide + ribose vs deoxyribose
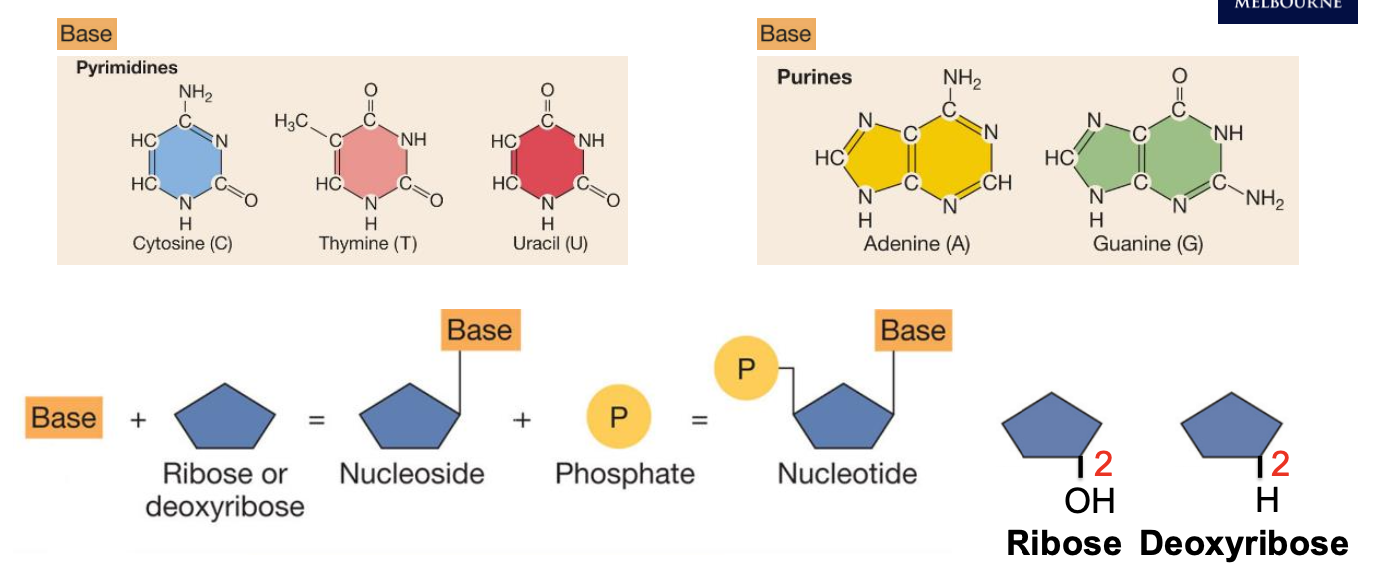
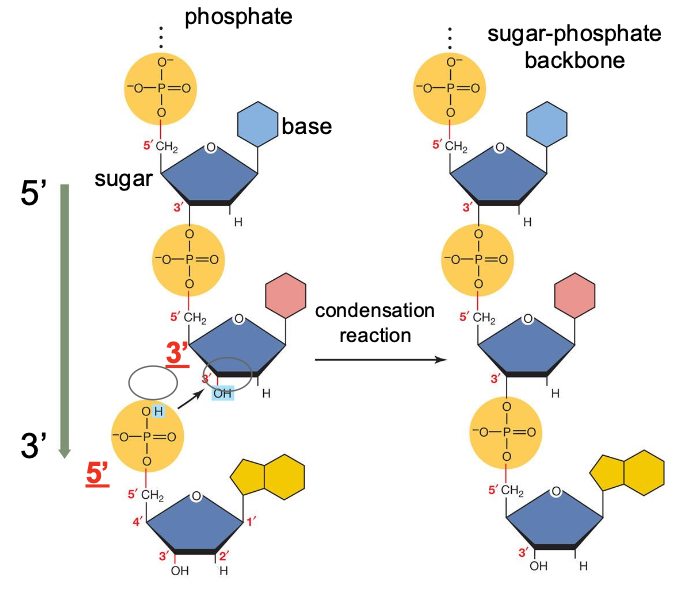
Nucleotides are linked together by:
phosphodiester bonds to form nucleic acids
Bases are held together by
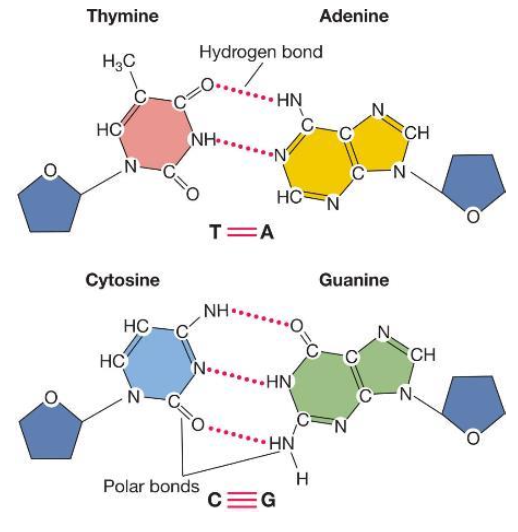
DNA’s double stranded nature + direction
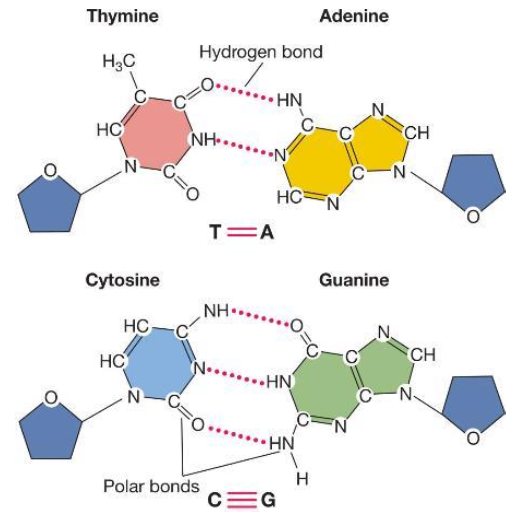
DNA strands names

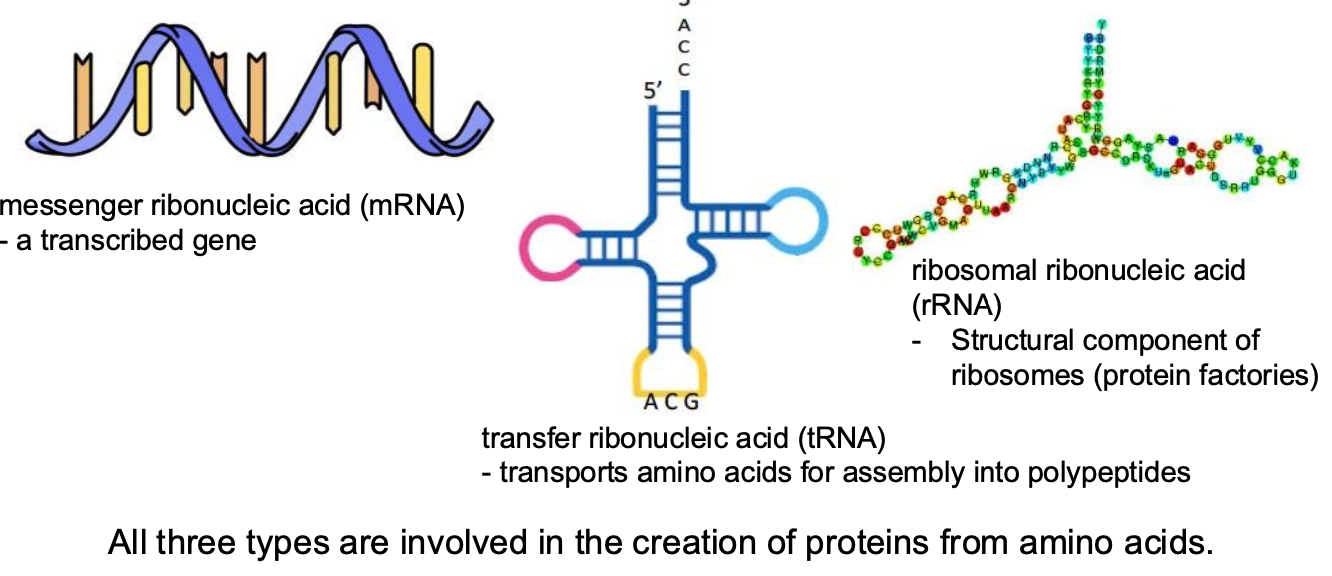
Types of RNA
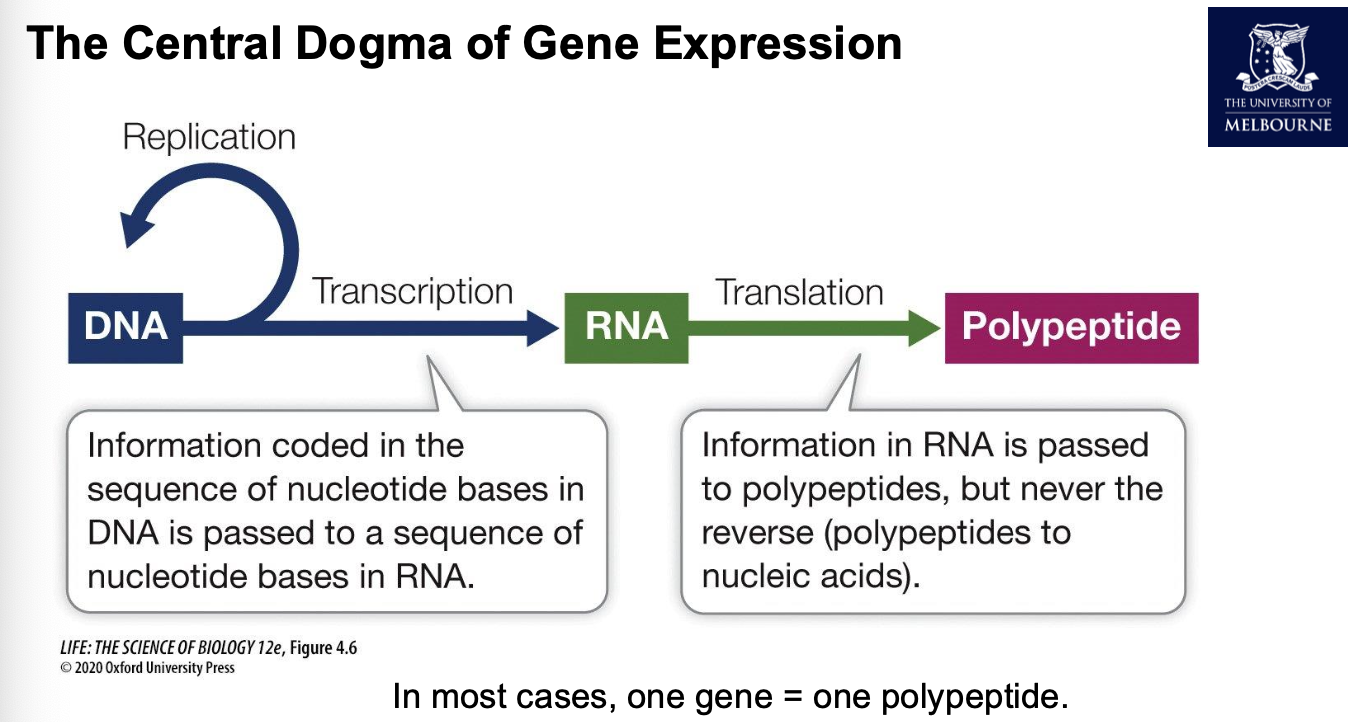
Central dogma of gene expression
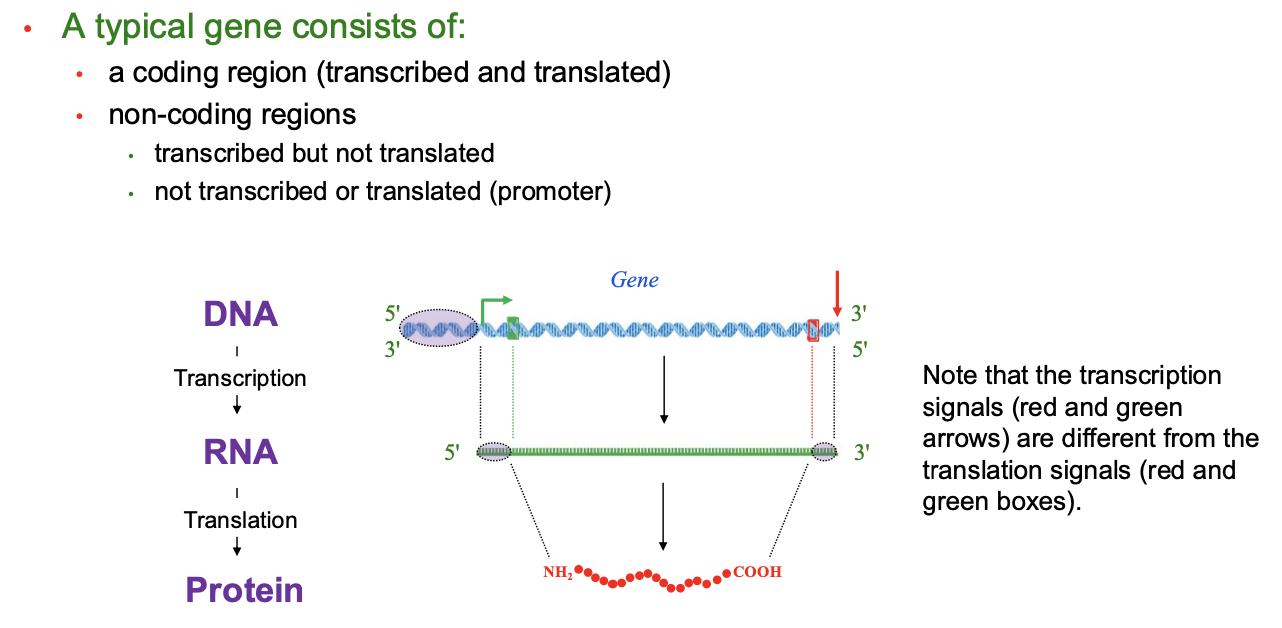
Gene Structure
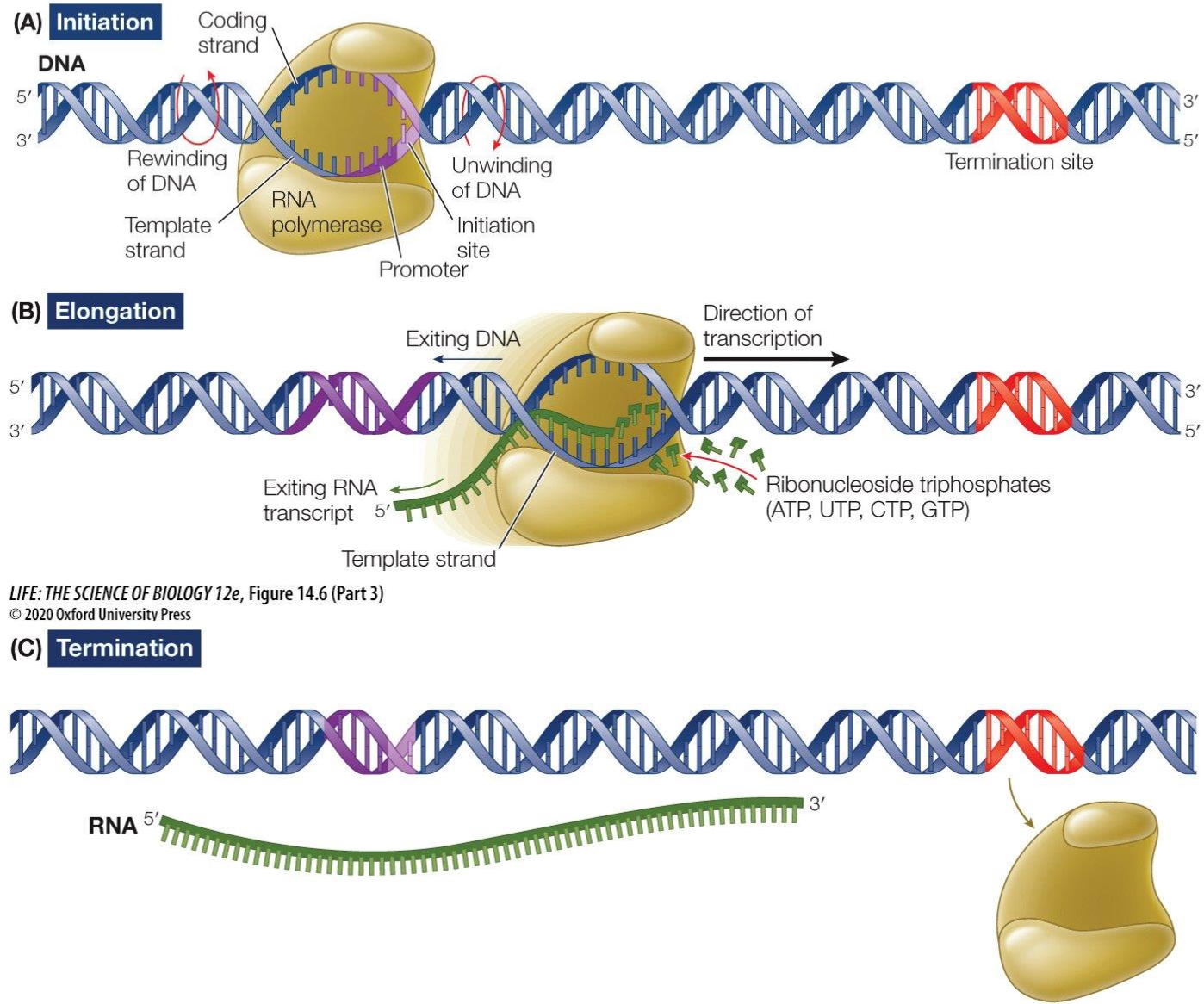
Transcription
Initiation: RNA Polymerase binds to the promoter and then unwinds the double strand of DNA section by section in the 5’ to 3’ direction until termination sequence
Elongation: Free moving complementary RNA nucleotides create a phosphodiester backbone and an mRNA strand which is complementary to the template strand is created
Termination: termination sequence reached and RNA polymerase detaches itself from the DNA strand
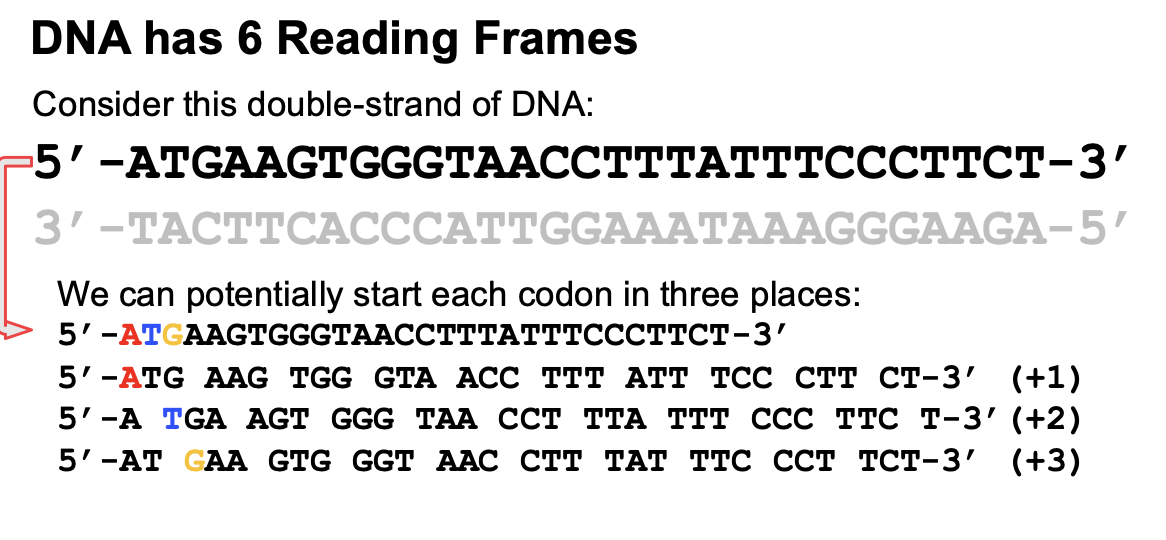
Reading frames
Where you start reading is important and each starting point is a different reading frame.
If we’re looking at the sense strand (template), then if you start at the first base, that’s reading frame 1 and second base is reading frame 2. It is important because methionine must be first. (+1, +2, +3)
If we’re looking at the non coding strand, then same concept but MUST start from 5’ end and the reading frames are "”-1,-2 etc”
A

Open reading frame
A successive run of non overlapping codons starting with AUG and ending with UAA, UAG or UGA
all open reading frames start with AUG and end with a stop codon
in some genes there will be multiple open reading frames you get from the reading frames, in this case the gene can code for more than one kind of polypeptide and it would be called an overlapping gene. 1 gene = more than 1 polypeptide.
Codon
Three nucleotide bases (triplet) that code for a specific amino
acid.
Start codon
AUG, codes for methionine.
• Always the first amino acid translated
Stop codon
UAA, UAG, UGA
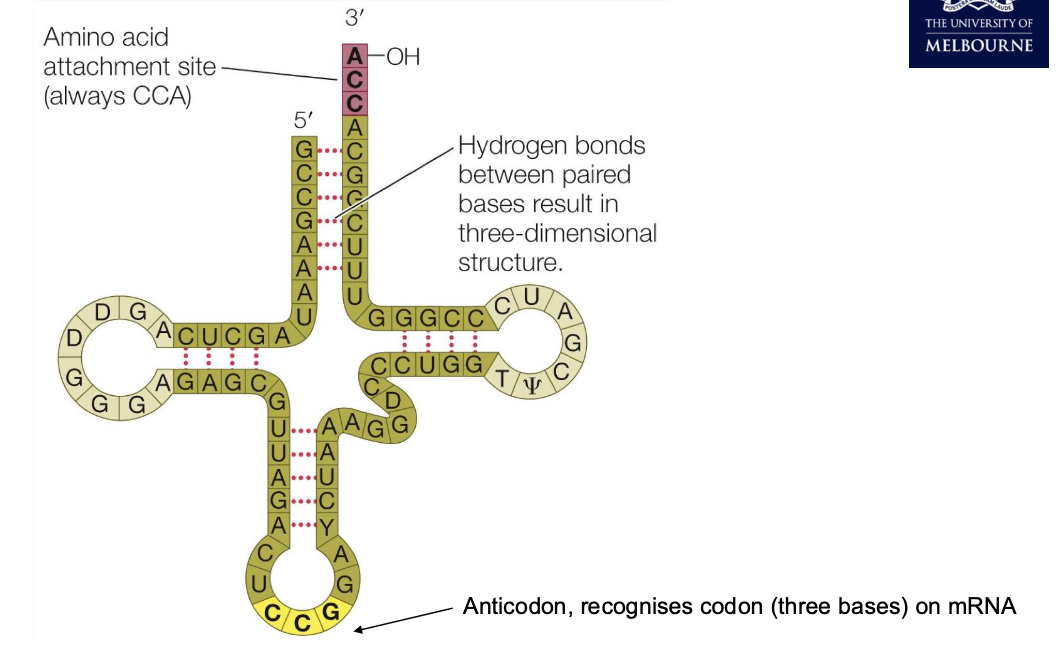
tRNA
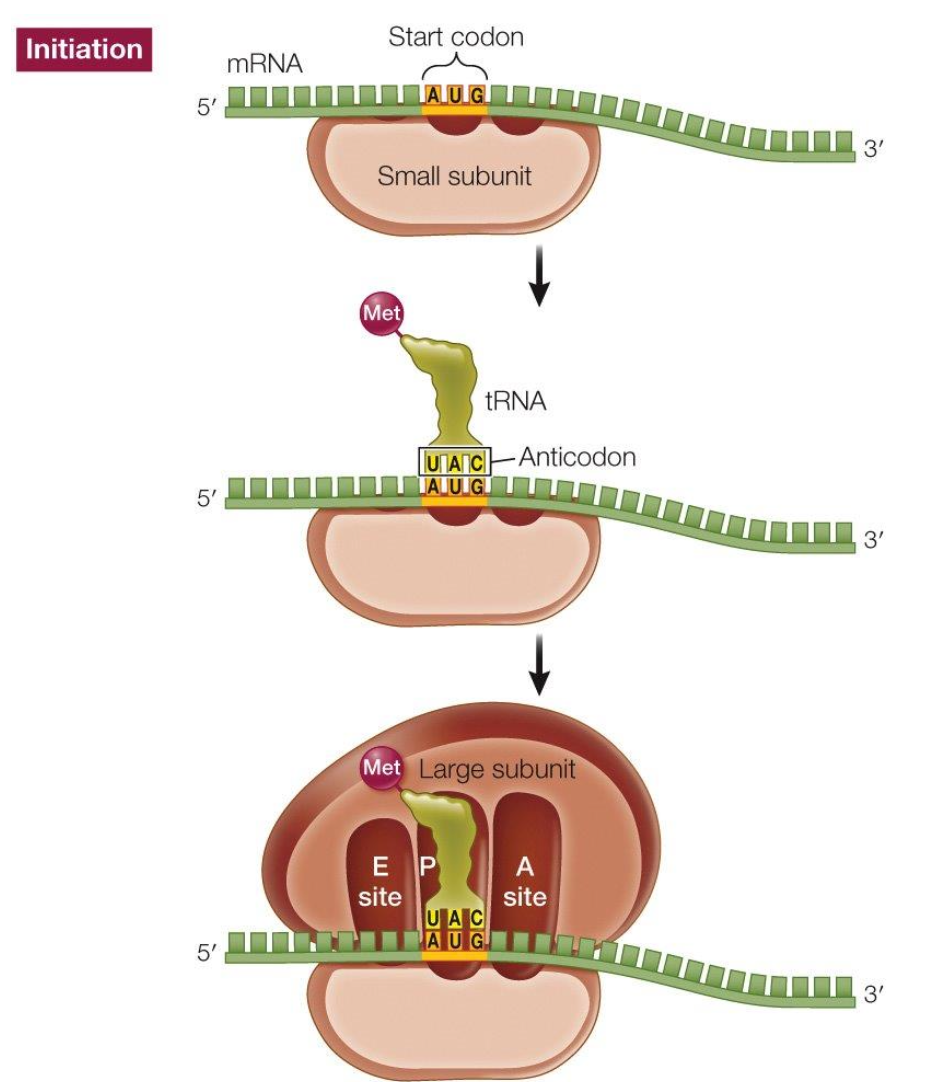
Translation: initiation
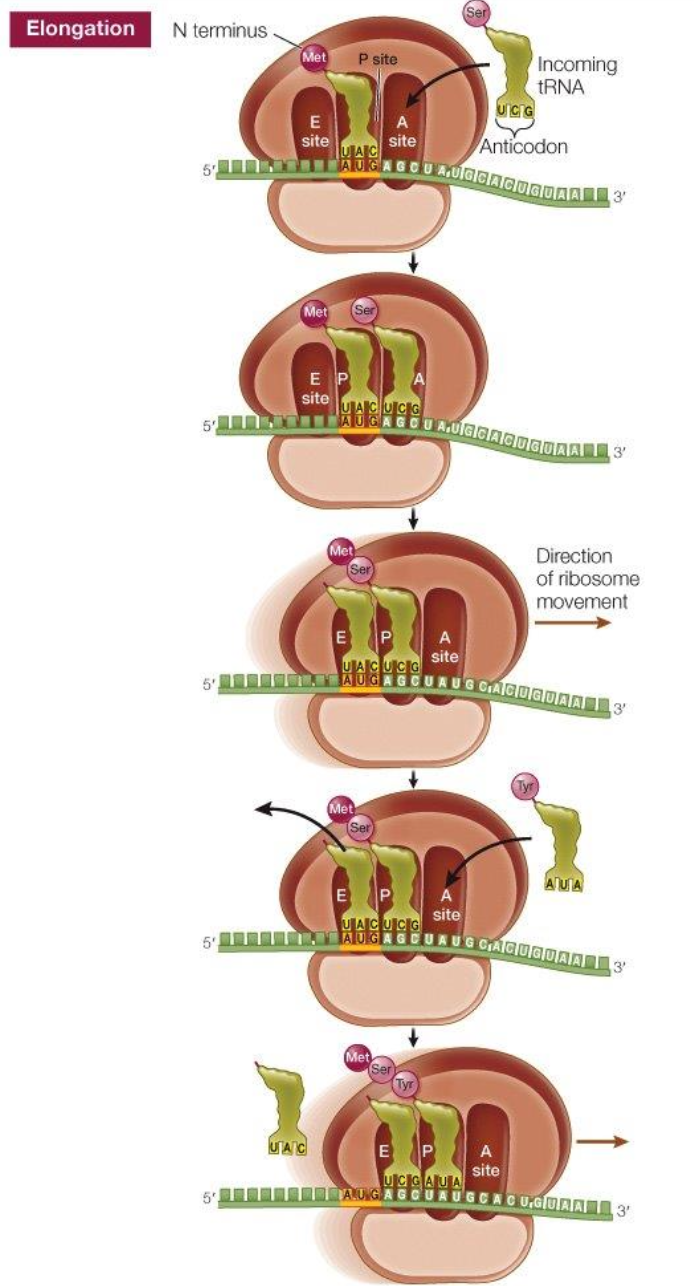
Translation: elongation
Translation: termination
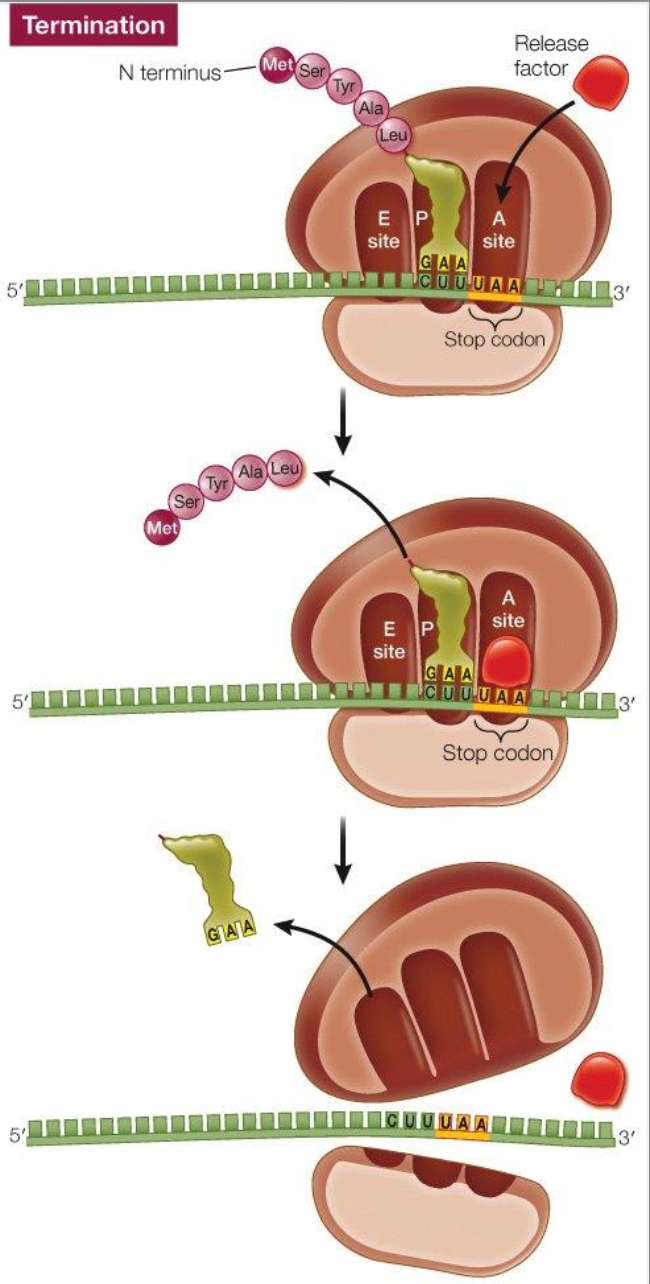
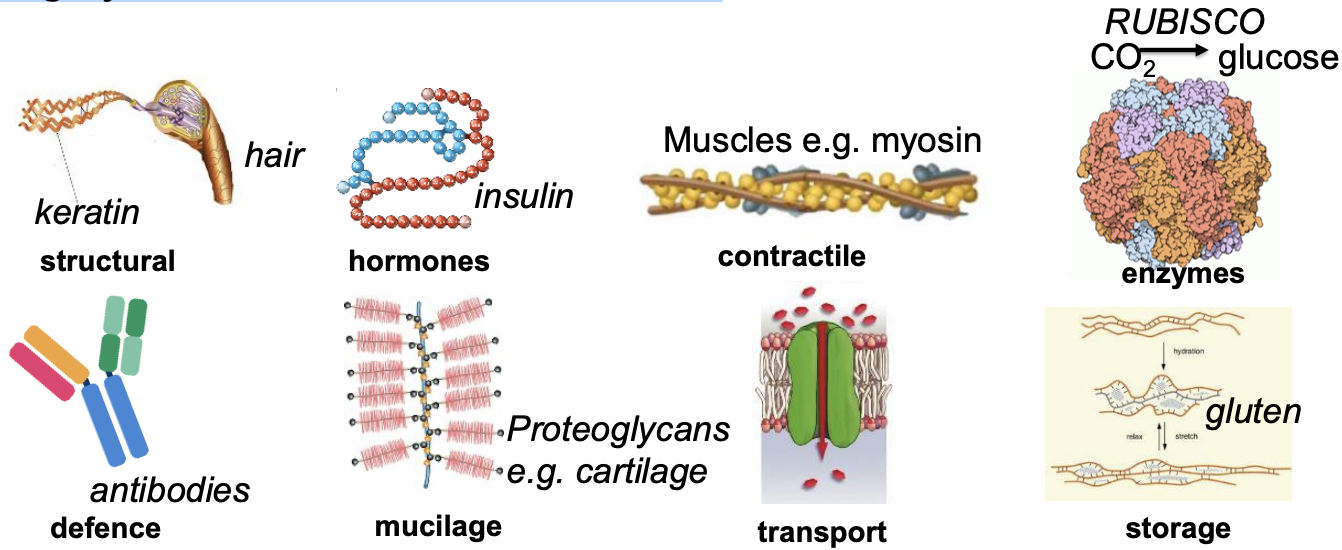
Basics of proteins
Highly variable structure & function
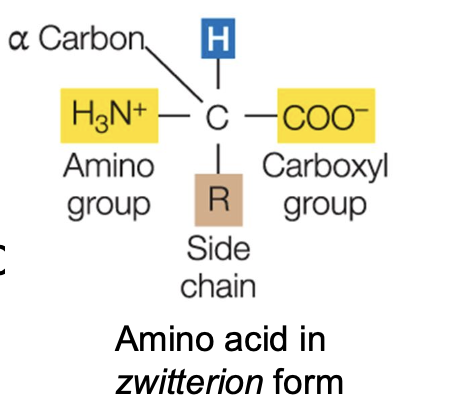
Amino acids
All amino acids have a central (⍺) Carbon with 4 attached groups:
• Hydrogen (-H)
• A basic amino (-NH2)
• An acidic carboxyl (-COOH)
• A side chain (-R)
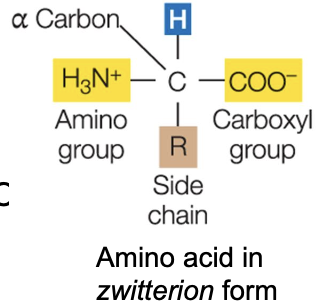
Zwitterions
Remember, bases can receive H+ (a proton), acids can donate H+
In low pH, NH2 becomes NH3+
In high pH, COOH becomes COO-
Basic and Acidic amino acids
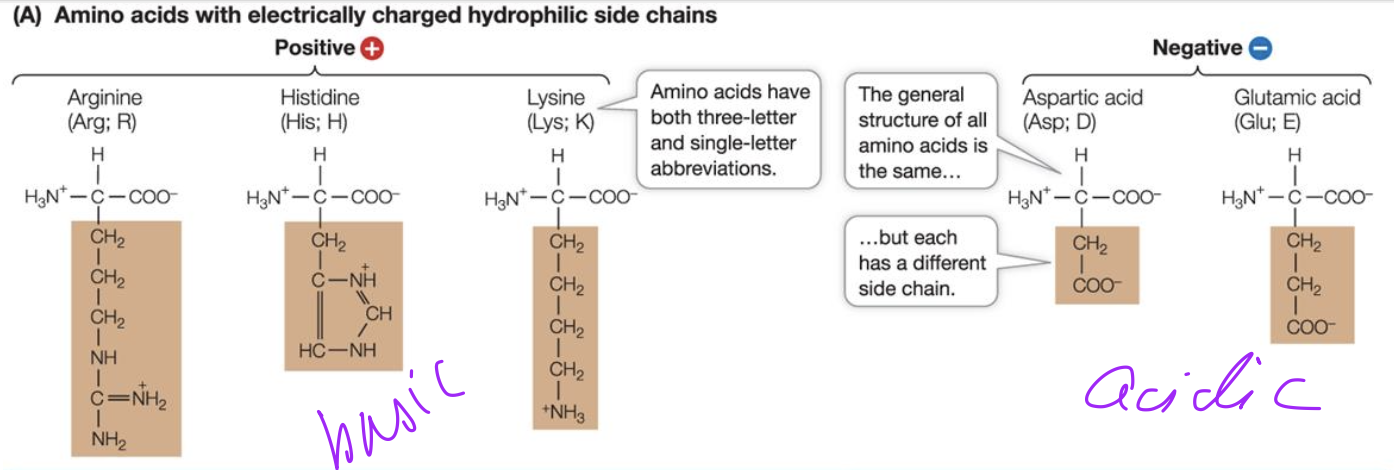
Peptide bond formation
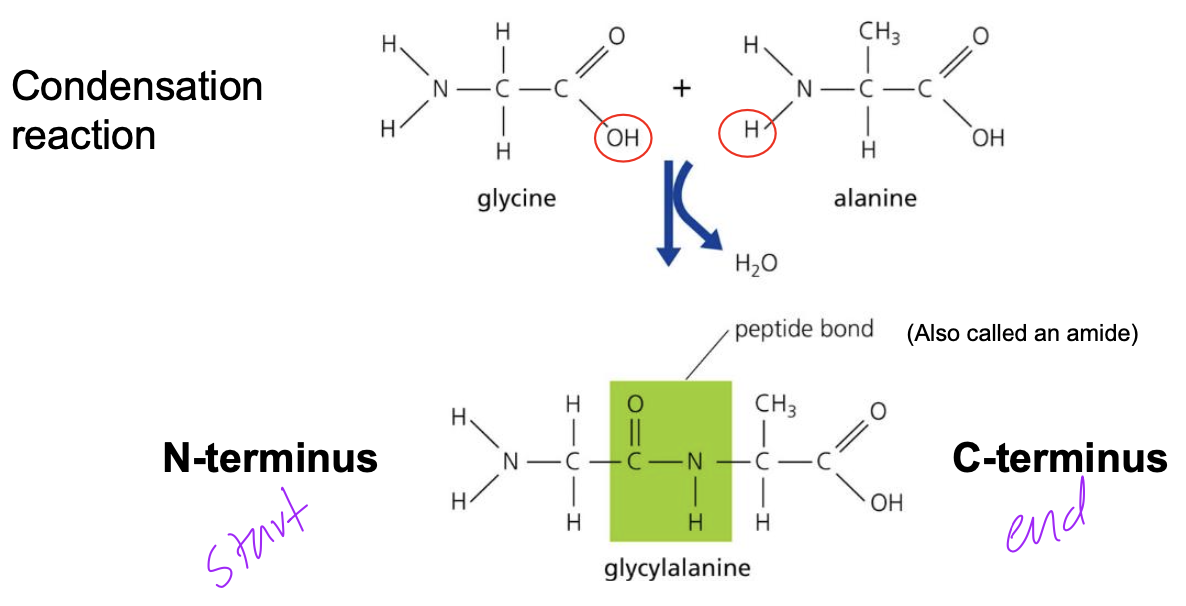
Polypeptide chains
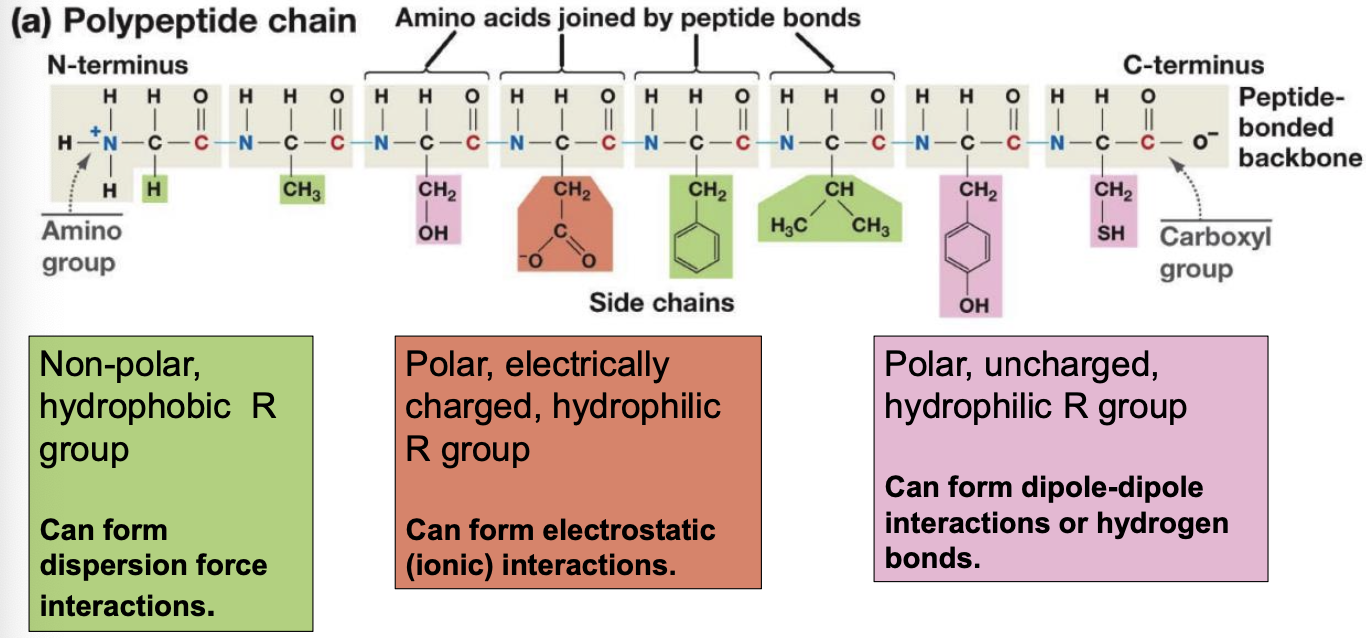
4 Levels of Proteins
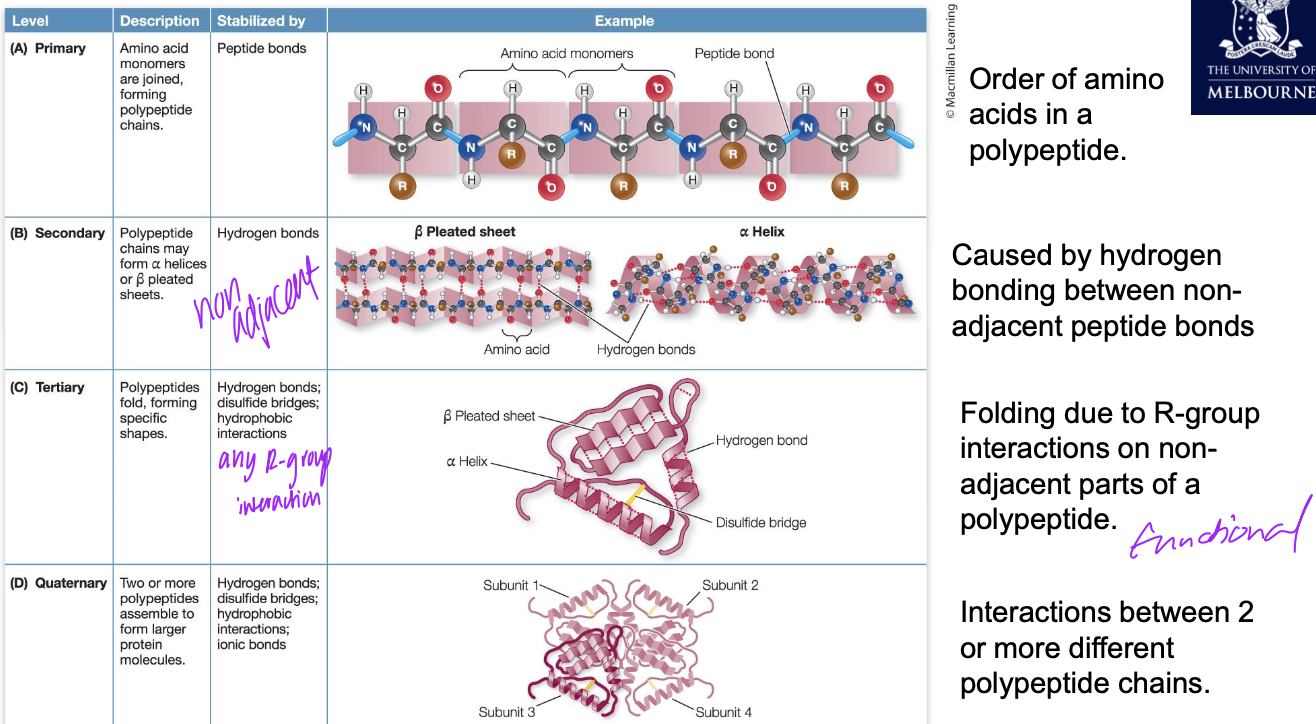
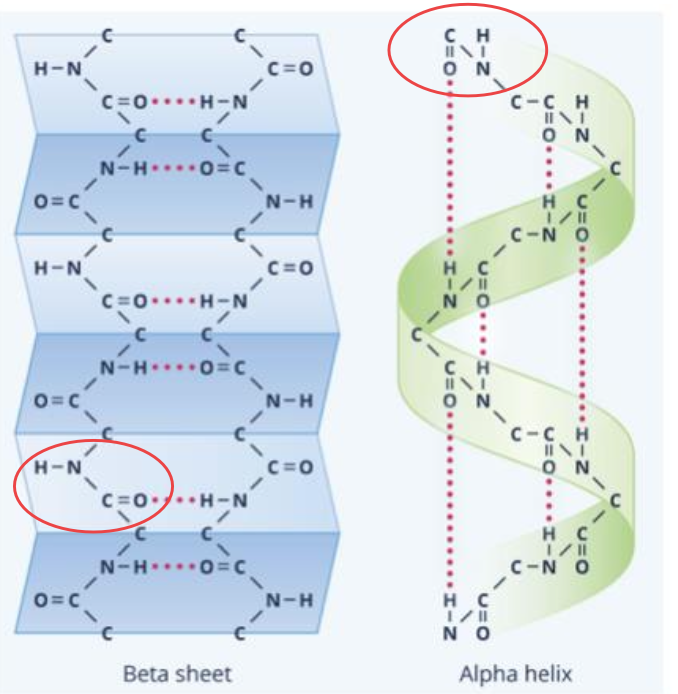
Secondary structure
Alpha helix or Beta sheet is formed through hydrogen bonding between non-adjacent peptide groups (-CONH-)
Tertiary structure
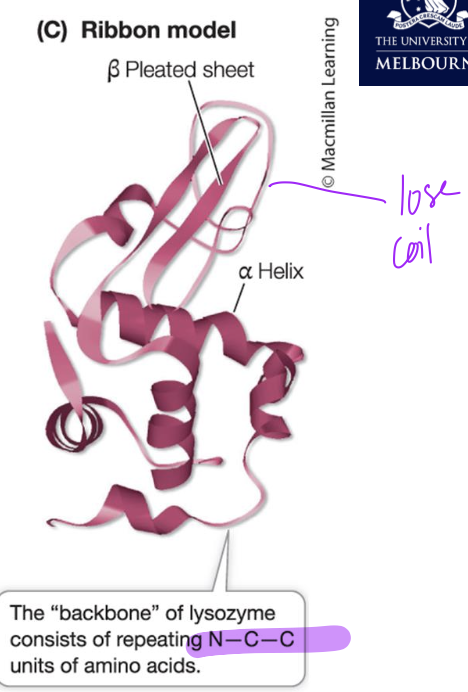
Lysozyme structure
Ribbon shows secondary structures and primary structure (secondary not formed) with the loose coil but altogether makes functioning tertiary structure
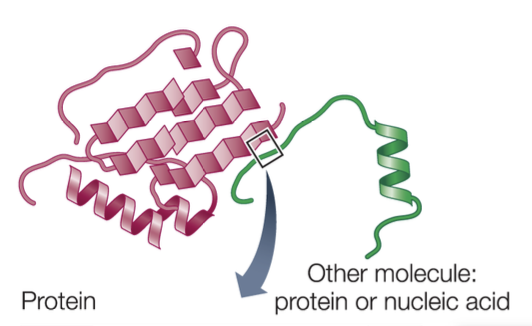
Specificity of proteins influenced by
Shape and chemistry
Protein shape
There must be a general “fit” between the protein and the other molecule.
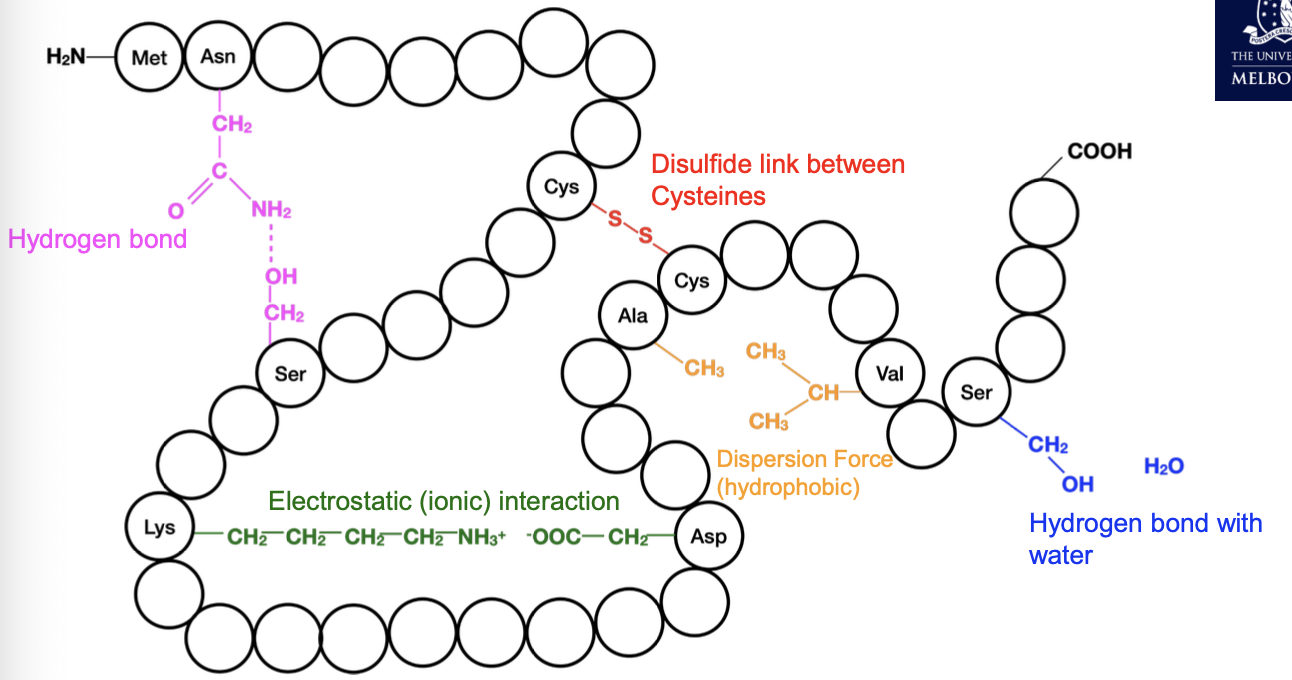
Protein chemistry
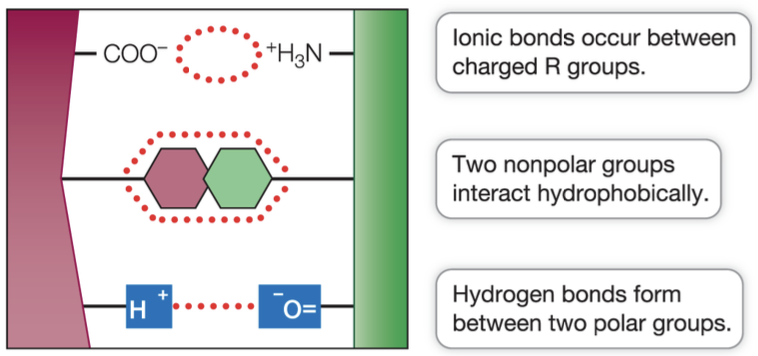
Surface R groups interact with other molecules via:
Ion-ion, ion-dipole
Hydrogen bonds
Dispersion Force
Dipole-dipole
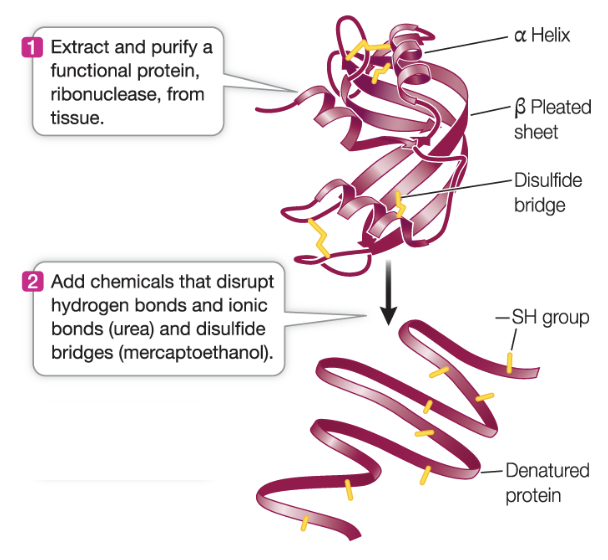
Protein denaturation
Proteins, although highly specific, can easily have their structure changed, through disruption or alteration of their side-chains. When secondary or tertiary structure of a protein is altered in this way, it is said to be denatured.
change in tertiary structure (also secondary sometimes) or change in side chain interactions → lose function
Define denaturation
Denaturation is the change to the secondary or tertiary structure of a protein.
It causes the protein to either completely, or partially, stop functioning.
pH change denaturation explanation
pH change: Addition or removal of H+
This causes changes to COOH and NH2 containing R-groups. COOH←→COO- NH2 ←→ NH3+
Disrupts ionic interactions
Temp change denaturation
Temperature change: Increasing temperature provides energy to disrupt
intermolecular interactions between R-groups.
addition of polar denaturation
Addition of polar: Disrupts all polar R-groups by interfering with dipole dipole force and hydrogen bonding
addition of non polar denaturation
Addition of non-polar: Disrupts all non-polar R-groups by interfering with dispersion force.
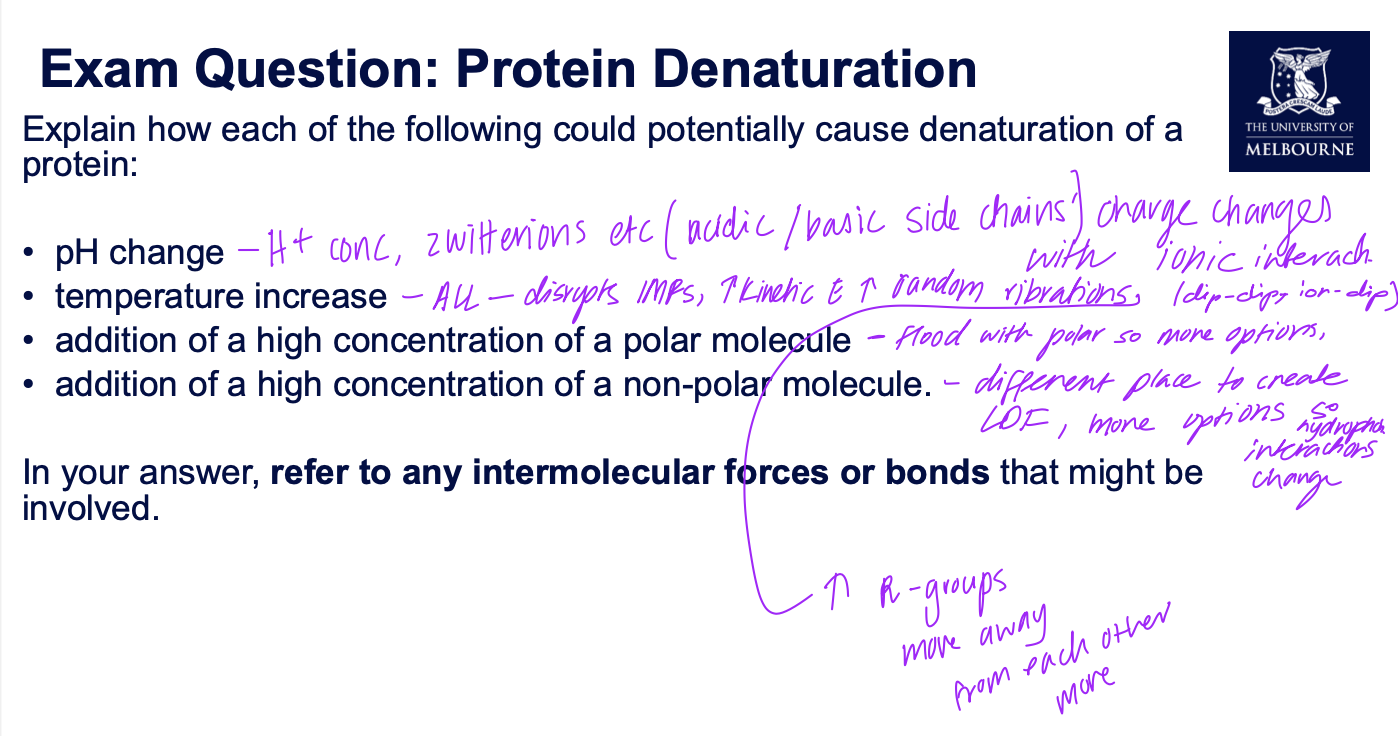
Exam question on denaturation

Fluid mosaic model
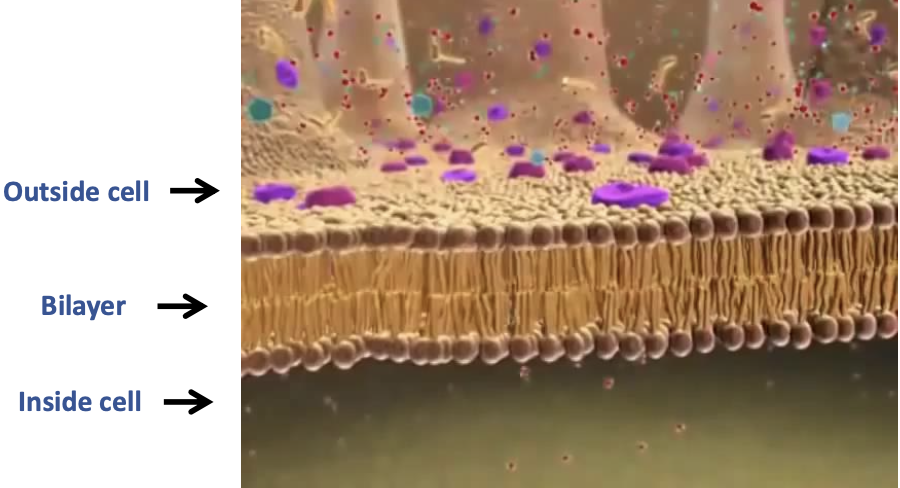
Plasma membrane structure
The phospholipids form a bilayer, with the two groups of tails sandwiched towards the interior, one group of Phosphate heads towards the cytoplasm (aqueous), other towards the cell’s exterior (aqueous)
Cell membranes are not rigid
The phospholipids are mobile
they move laterally all the time, along the plane of the membrane
Membranes must be fluid to work properly
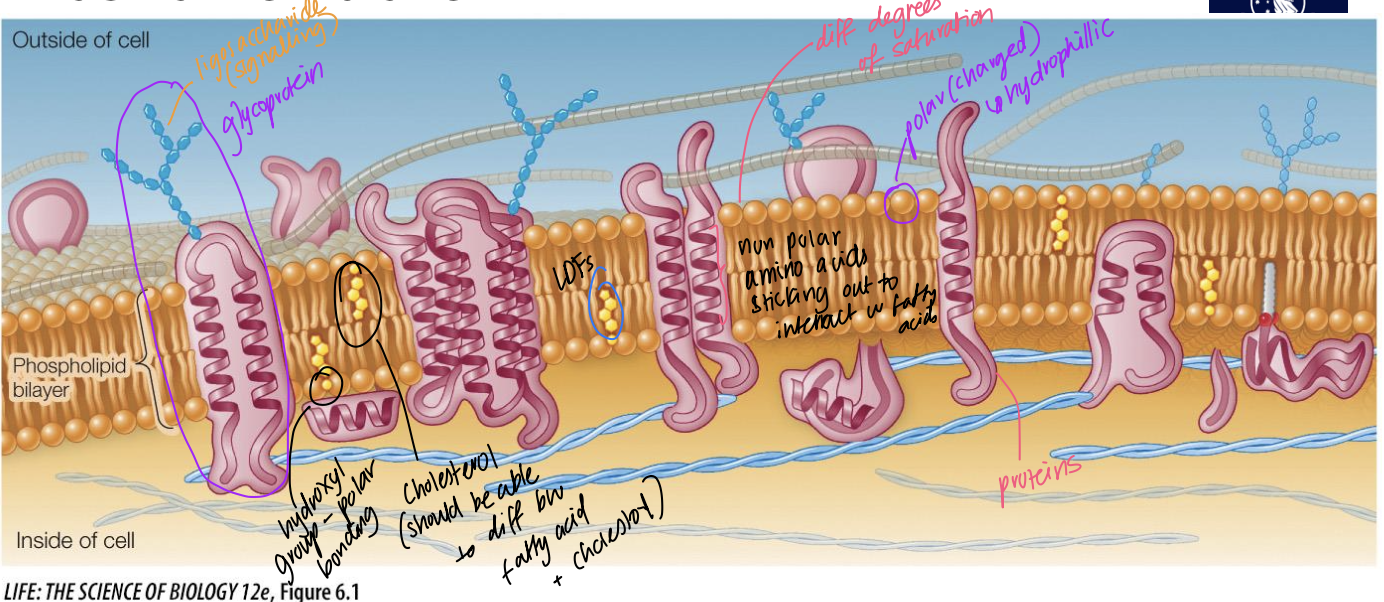
Plasma membrane role
prevents free flow of liquids into/out of cells due to the hydrophobic middle layer, and protects cell’s interior
Membrane fluidity
unsaturated hydrocarbon tails of phospholipids have kinks that keep the molecules from packing together, enhancing membrane fluidity
less packing = more membrane fluidity