Bioinformatics + Homology Modelling
1/35
Earn XP
Description and Tags
Things you need to know! • Databanks vs. databases • The difference between a primary and a secondary databank • Examples of primary and secondary databanks • The definition of homology / orthology / paralogy and their importance • The difference between a databank (like SwissProt) and a search tool (like BLAST) • Local and global alignment • Basic ideas of sequence searching and the meaning of the e-value • Problems of annotation
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
36 Terms
Define bioinformatics
Aiding the Biologist in creating, storing, searching and analyzing biological data (particularly sequences and structures, but also 'omics data), presenting it in a way Biologists can use and applying the analysis to make predictions
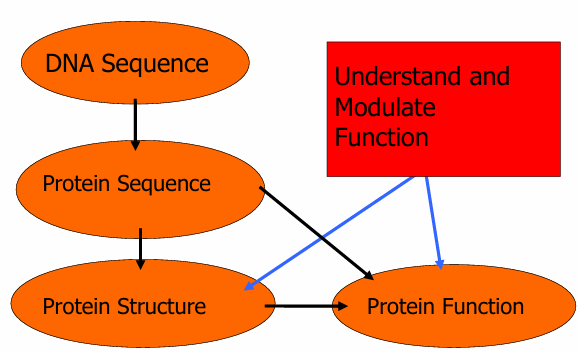
What are we trying to achieve with bioinformatics?
What is a key point both homologous proteins/genes share?
Have descended from a common ancestor
Homologues have derived from a common ancestor hence they will have similar characteristics
“It is WRONG to refer to ‘percentage homology’”, why?
An old term used before the evolutionary angle took over the word homology
Percentage sequence identity is used instead to infer homology
What is the criteria for a protein sequence to be always homologues? (and vice versa)
100AA with >35% sequence identity = Homologues and same function still
Lower sequence identity = May be homologues
Which source can you use to compare structures of homologues and determine which type of homologue?
Structure comparison
PDB Database
GPCRdb
Identifying type
NCBI Homologene
EBI
What are the two types of homologues?
Orthologues
Paralogues
What are orthologues?
Homologues resulting from a speciation event
i.e. the same gene in two different species
What are paralogues? Give an example
Result from gene copying within a species
E.g. globin genes (beta and alpha globin)
What is protein modelling, why use it, and why not just rely on software predictions?
Protein modelling = Using instrumentation to work out the actual protein structure, i.e. cryo-EM
You need to validate things experimentally to verify that what you’ve computed is correct (support the computational prediction), and you get the results you expected
What are the two types of storages of data?
Databank & database
Define databank
A collection of data (normally in simple text files) without a fixed associated query tool
You can download all the raw data and handle it however you wish
What are the types of databanks?
Primary
Raw sequence/structure data
Possibly with detailed annotations
Secondary = Curates the primary data
Derived data - sequence profiles, etc
Generally highly annotated
Give examples of primary databanks, and what they (+may) contain
DNA: Genbank, EMBL-ENA, DDBJ
Protein: UniProtKB/SwissProt
Simply contain sequence data (DNA or protein)
May also have ‘feature’ information (splice sites, signal sequences, disulphides, active sites, etc., etc.)
DNA databanks may also contain translations (known or predicted)
What is the main primary databanks for structural data and enzyme classifications (EC numbers)?
PDB
Enzymes
Give examples of primary databanks, and what they (+may) contain
PROSITE, PRINTS, BLOCKS, INTERPRO
Contain derived information
Patterns that characterize a protein family
Detailed annotation
NOTE: Can be used for finding homologues
Where would you find annotations of proteins/genes from the source?
References
Methods
Cross-links to other databases
‘Feature tables’
Segments of biological significance (e.g. coding regions, active sites)
More general descriptions
Computer parsable?
May use a ‘restricted dictionary’ or ‘ontology’
Authors
NOTE: Just because an information isn’t there doesn’t necessarily mean it is not known
How are most sequences predicted and why are gene-prediction methods imperfect?
Predicted from genomic data
Most of the the time the software only makes a sensible prediction after more biological information is put into the database
“A protein identified from genome data is hypothetical until verified by experiment”
Define database
A structured collection of data hidden behind a fixed search tool
What is protein/peptide sequencing largely replaced by?
DNA sequencing (now automated)
What are the two important things to remember about quality?
The quality of raw data is as good as the methods that produce it
The quality of annotations is as good as the curators
How were annotations in the pre-genome world different to annotation in the genome world?
PRE
DNA sequence came from a single group with particular interest in that gene
Annotations grounded in experiment
Written by experts in that gene
POST
Rarely experimental confirmation of expression of putative genes or characterization of products
Annotation based on computer analysis
The weakest link of genomics!
Getting it right is labour intensive and underfunded
Poor annotation negates the benefit of having genome sequence data!