Chapter 15 Signal Transduction and G Protein- Coupled Receptors
1/78
Earn XP
Description and Tags
Bolded terms and pathways from slides
Name | Mastery | Learn | Test | Matching | Spaced |
|---|
No study sessions yet.
79 Terms
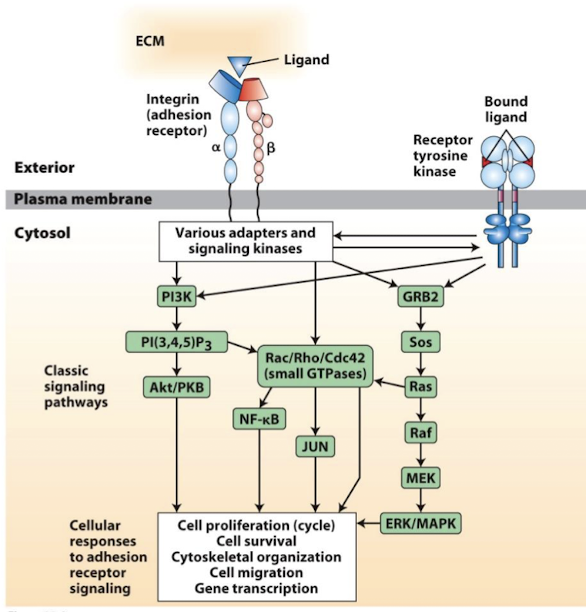
Cell-adhesion molecules mediate signal transduction
Can respond to changes in the ECM, tansduce signals, and promote altered cell activity
regulate:
migration
proliferation
apoptosis
enzymatic activities
gene transcription
Receptor Protein
A protein that binds a ligand, which could be a small molecule such as adrenalin, a growth factor such as epidermal growth factor, a hormone such as insulin, or another macromolecule, usually with engagement of a signaling cascade as a consequence.
Ligands
Signaling molecules that bind specific receptors triggering a specific cellular response
Any molecule, other than an enzyme substrate, that binds tightly and specifically to a macromolecule, usually a protein, forming a macromolecule-ligand complex.
Agonist
A molecule, often synthetic, that mimics the biological function of a natural molecule (e.g., hormone)
Activating
Antagonist
A molecule, often synthetic, that blocks the biological function of a natural molecule (e.g., hormone)
Blocking
Signal Transduction
The process by which cells receive and respond to external signals.
Involves change in the position and type of information transmitted through a cell.
Signal Transduction Pathway
The set of proteins or small molecules involved in relaying a chemical message from the receptor at the cell surface to the target molecule(s) inside the cell.
Second messengers
Small, non-protein, intracellular molecules that relay, amplify, and transduce signals
Transduction enzymes
Highly specific promote transduction through allosteric regulation of the next protein
Transduce signaling via covalent modifications
kinases
phosphatases
cleavage enzymes
enzymes that catalyze synthesis of second messengers
Effector Proteins
An enzyme, transcription factor, or cytoskeletal protein whose activation leads directly to changes in cellular activities.
Transduction regulation of effector proteins
Transduction pathways regulate terminal effector proteins.
Enzymes that alters cell behavior or physiology- rapid response
specific transcription factors that alters gene expression- slower long-term response
Often a combination of both
Specific transcription factors
Bind regulatory DNA sequences either in the promoter or discrete regulatory DNA regions (cis-regulatory elements)
Binding transcription factors to these elements can either activate or inhibit RNA polymerase and therefore regulate gene transcription.
Enhanser
A regulatory sequence in eukaryotic DNA that may be located at a great distance from the gene it controls or even within the coding sequence. Binding of specific proteins to an enhancer modulates the rate of transcription of associated gene.
Cis-Regulatory elements (CREs)
discrete regulatory DNA regions
Binding transcription factors to these elements can either activate or inhibit RNA polymerase and therefore regulate gene transcription.
Feedback inhibition
The final effector protein also blocks continued transduction of the pathway- ensuring a transient, but appropriate amount of response.
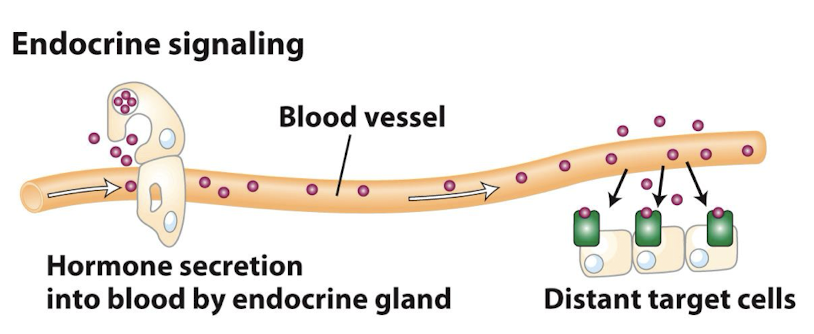
Endocrine Signaling
Ligands called hormones are secreted by specialized cells of the various endocrine glands and travel through the circulation system to act on target cells in distant tissues
Examples:
insulin/glucagon
testosterone/estrogen
cortisol and thyroid hormones
Some ligands can act as both endocrine and paracrine signals. Epinephrine, for example, acts hormonally as part of the fight or flight response, but is also a neurotransmitter.
Hormone
Generally any extracellular substance that induces specific responses in target cells; specifically, those signaling molecules that circulate in the blood and mediate endocrine signaling.
Paracrine Signaling
Secreted ligands diffuse a short distance through the extracellular matrix before binding receptors on nearby target cells (1 to 20 cells away)
Examples:
Neurotransmitters that signal adjacent, but unconnected cells at their synapse
Diverse growth factors that control rates of proliferation and or differentiation: immature cells adopting terminal fates
Some ligands can act as both endocrine and paracrine signals. Epinephrine, for example, acts hormonally as part of the fight or flight response, but is also a neurotransmitter.
The secreted ligands diffuse from the source and generate a concentration gradient.
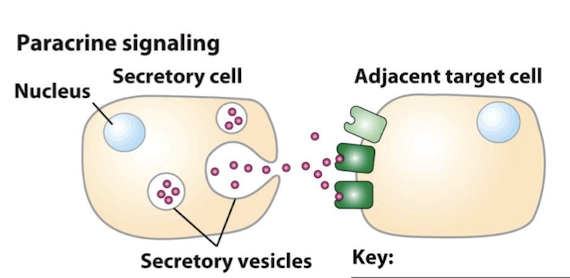
Growth Factors
An extracellular polypeptide molecule that binds to a cell-surface receptor, triggering an intracellular signaling pathway generally leading to cell proliferation
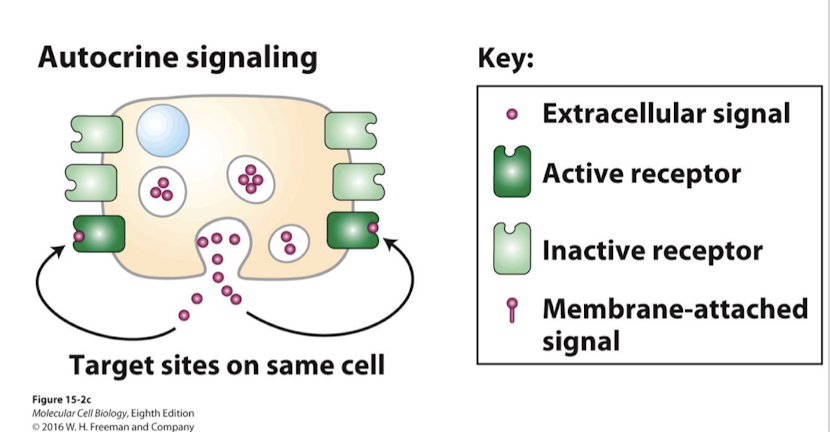
Autocrine Signaling
Synthesis and secretion of ligands that sustain some physiological activity of the cell itself
Most autocrine ligands are growth factors
For example, the constant replacement of skin cells is driven by proliferation induced via autocrine signaling
May types of cancer overproduce growth factors, enhancing their growth rate
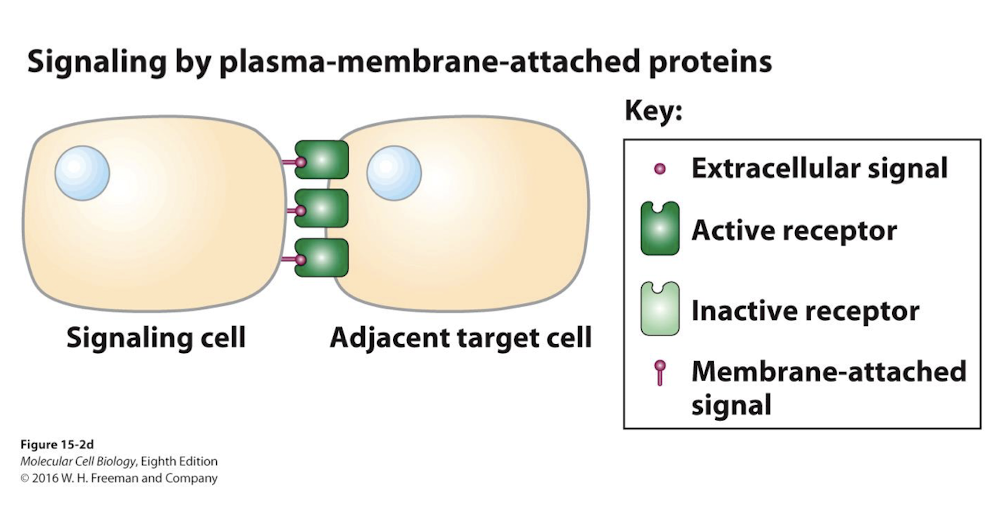
Juxtacrine Signaling
Both the receptor and ligand are membrane bound
Therefore, signaling only occurs between immediately adjacent cells
Signal transduction
The conversation of an impulse or stimulus from one physical or chemical form to another; the processes by which cells respond to extracellular signals.
The activated receptor relays this information to cytoplasmic peoteins and other molecules that:
Amplify the signal
Integrate it with other pathways
Generate a cell response
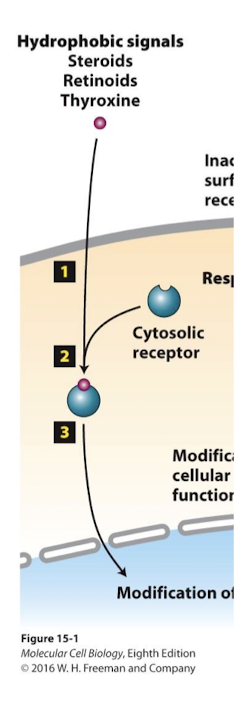
Lipid derived hormones
enter cells, via endocytosis or diffusion
cytosolic receptors
Some ligands-like lipid derived hormones- enter cells, via endocytosis or diffusion. They bind and acrivate soluble cytosolic receptors.
These cytosolic receptors are usually specific transcription factors.
Cell-Surface Receptors
Protein embedded in the plasma membrane that has an extracellular domain that binds an extracellular molecule(s), called a ligand. Many cell-surface receptors bind to extracellular signaling molecules; such binding includes conformational changes in the receptor, altering the activity of the receptor’s intracellular domain, which transmits the signal to the interior of the cell.
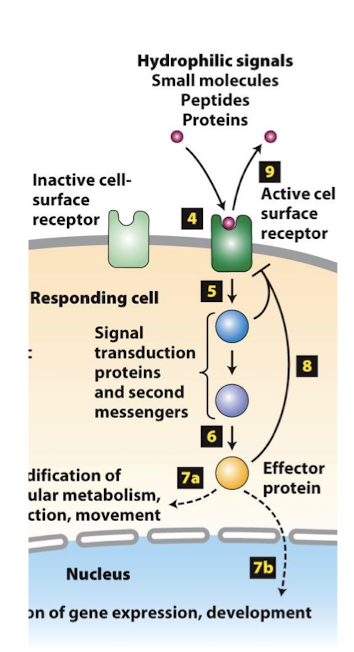
Basal State
When ere is no signal cells capable of receiving that signal are said to be a basal or unstimulated state. A condition in which cells perform only basic, housekeeping functions, common to all cells of that type.
Principles of signal transduction
The same ligand may activate different responses in different cells because:
They bind different receptor on the cells
They bind the same receptor, but it activates a different transduction pathway
Expression of different regulators of a shared pathway
Expression of different terminal effector proteins
Expression of different targets of a shared effector protein
Different combinations of ligands activating different combinations of pathways
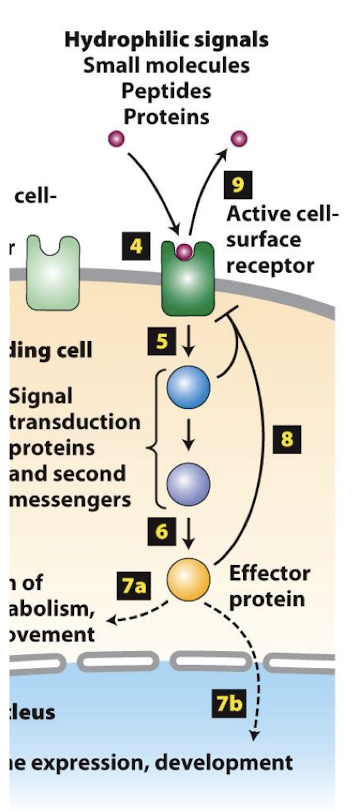
Mechanisms of signal transduction:
Some activated cell-surface receptors directly regulate a cytoplasmic enzyme that catalyzes synthesis of second messengers.
Other cell-surface receptors recruit cytoplasmic proteins to their intracellular domains and initiate a series of protein-protein interactions that transduction the signal
often mediated by kinase proteins
In both cases, signal transducction leads to the activation of one, or a small number of, specific effector proteins.
Effector proteins general functions
Enzymes that immediately alter some cellular process by regulating a key event or
Transcription factors that translocate to nucleus and regulate expression of specific target genes
Switches
Two major types of switches:
Those that regulate phosphorylation state:
Kinases and Phosphatases
G proteins that allosterically regulate targets proteins. Two families:
Small monomeric G proteins- we’ve seen several examples of these
Heterotrimeric G proteins- the focus of CH 15
Phosphorylation
The covalent addition of a phosphate group to a molecule such as a sugar or a protein. The hydrolysis of ATP often accompanies phosphorylation, providing energy to drive the reaction and the phosphate group that is covalently added to the target molecule. Enzymes that catalyze phosphorylation are called kinases.
Protein kinase
An enzyme that chemically adds a phosphate group, usually the gamma phosphate from ATP, to serine, threonine, or tyrosine reside of another protein.
Protein phosphatase
An enzyme that chemically removes a phosphate group from another protein by hydrolysis.
Monomeric G protein
A monomeric GTPase with a structure similar to that of the Ras protein that changes conformation when a bound GTP is hydrolyzed to GDP and phosphate
Heterotrimeric G proteins
A class of GTPase switch proteins, composed of alpha, beta, and gamma polypeptides, that bind to and are activated by certain cell-surface receptors. When activated, hertotrimeric G proteins release GDP and bind GTP.
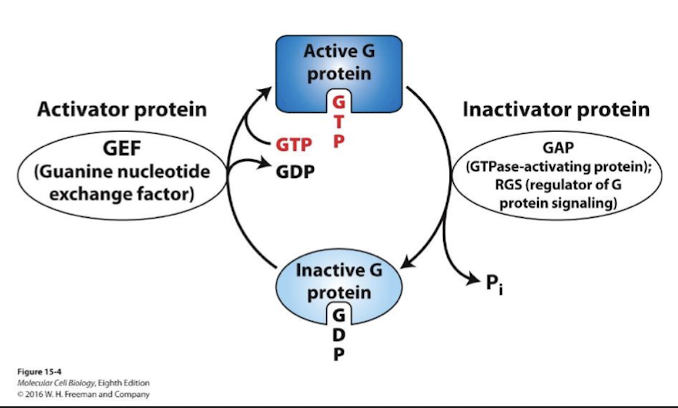
GTPase switch proteins cycle between active and inactive state
GTPase protein superfamily members often serve as signal transduction on-off switches:
ON conformation- GTP bound
OFF conformation- GDP bound
GEFs (guanine nucleotide exchange factors) catalyze dissociation of bound GDP and replacement by GTP- active G proteins
GAPs (GTPase-activating proteins; aka regulators of G protein signaling) catalyze hydrolysis of GTP to GDP and therefore promote inactivation of G proteins
Guanine nucleotide-dissociation inhibitors (GDIs)- prevent G proteins from exchanging GDP for GTP and therefore keep G proteins inactive
G protein- coupled receptor (GPCR)
Member of a large class of cell-surface signaling receptors, including those for epinephrine, glucagon, and yeast mating factors. All GPCRs contain seven transmembrane alpha helices. Ligand binding leads to activation of a coupled trimeric G protein, thereby initiating intracellular signaling pathways.
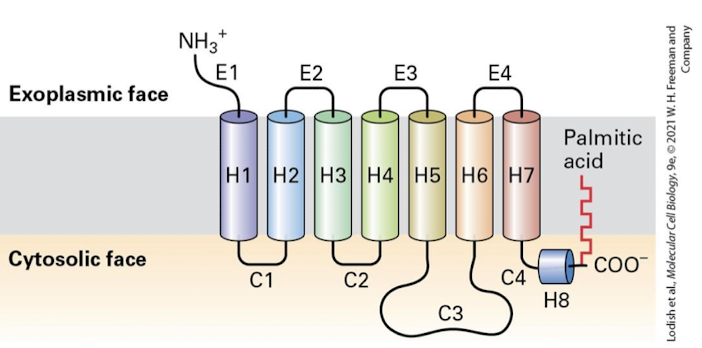
G⍺ subunit
The catalytic G-protein that is regulated by its GTP/GDP bound state. Homologous to the small monomeric G proteins we have already discussed. All G⍺ subunits are lipid anchored and remain bound to the membrane even after activation.
Gβ𝛾 subunit
A dimer that binds G⍺- GDP and keeps it inactive until a GPCR activates it. The G𝛾 subunit is also lipid anchored so Gβ𝛾 also remains membrane tethered even after separation from G⍺ . Gβ𝛾 & G⍺ united unless inactive.
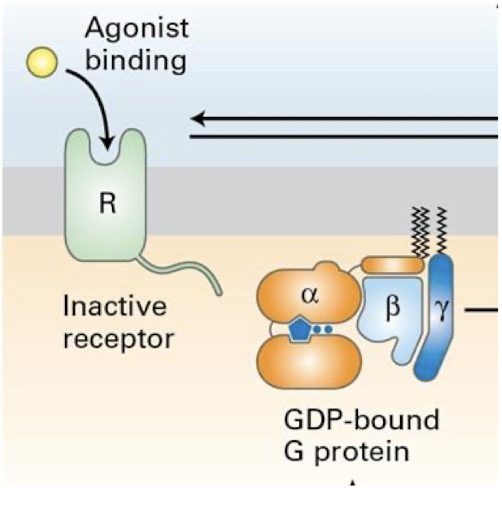
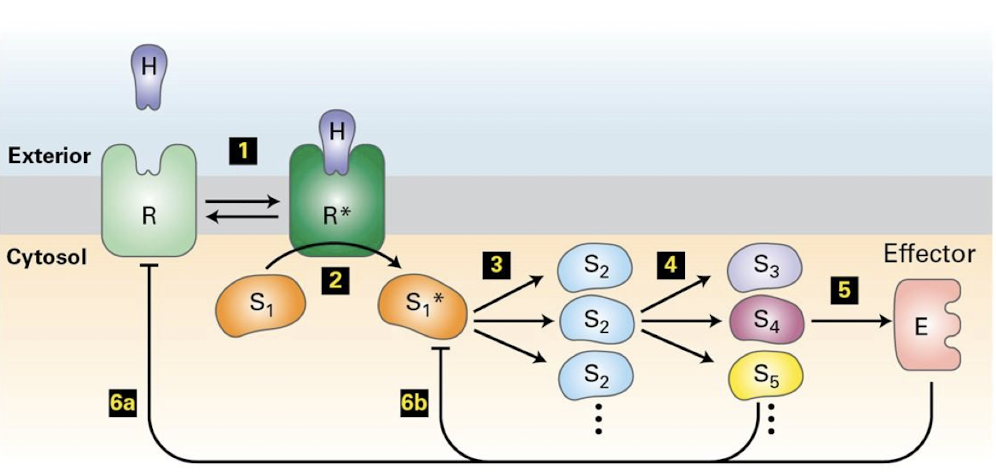
Generalized GPCR signal transduction pathway
Ligand binds GPCR, altering its conformation and ability to engage heterotrimeric G-protein
Bound receptor activates hetero-trimeric G protein, via catalyzing ⍺-subunit exchange of GDP for GTP
⍺-subunit disengages from β𝛾 dimer
⍺-subunit binds and regulates activity of a target enzyme
β𝛾 dimer also usually has regulatory targets- like an ion channel shown below
The enzyme regulated by G⍺-subunits regulates production of a second messenger
Some G⍺-subunits are positive activators of their target effector enzymes, other inhibit second messenger synthesis
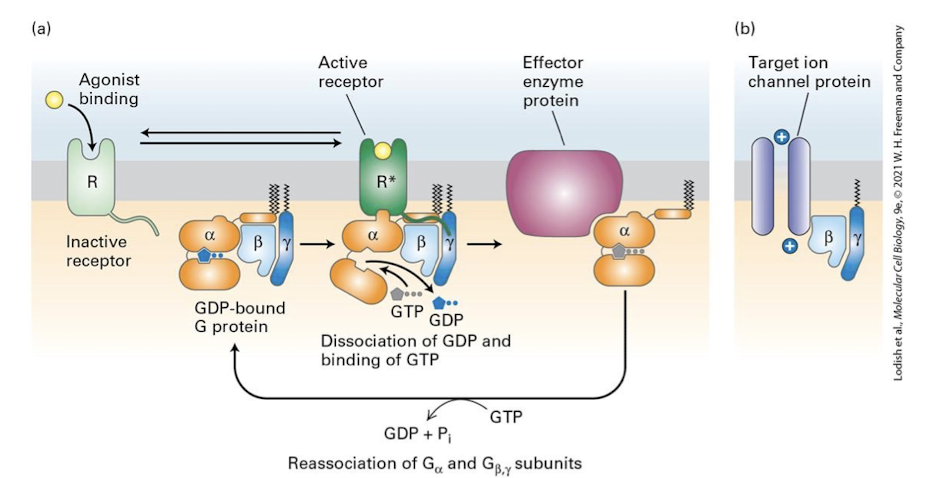
The major G⍺ families and their target effector enzymes are:
G⍺s: Stimulatory- Activate the effector enzyme adenylyl cyclases (cAMP)
G⍺i: Inhibitory- Inhibit the effector enzyme adenylyl cyclase (cAMP)
G⍺q: Activate the effector enzyme phospholipase C
G⍺t: Activate the effector enzyme phosphosiesterase
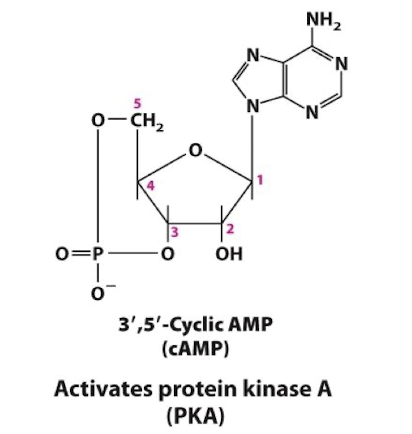
cAMP
Generated from ATP by the enzyme adenylyl cyclase
Allosteric activator of the enzyme Protein Kinase A
¼ common intracellular second messengers
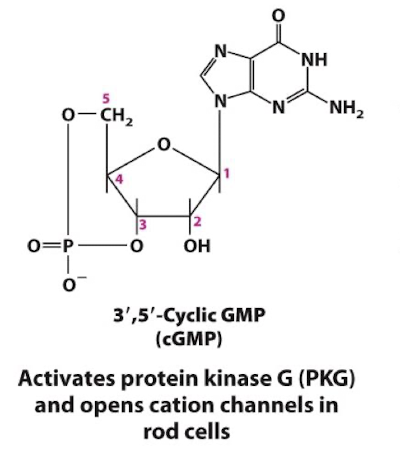
cGMP
Generated from GTP by the enzyme guanylyl cyclase
Allosteric activator of the enzyme Protein Kinase G and specific cation channels
¼ common intracellular second messengers
cyclic guanosine monophosphate
A second messenger that opens cation channels in rod cells and activates protein kinase G in vascular smooth muscle and other cells.
Adenylyl cyclase
One of the several enzymes that is activated by binding of certain ligands to their cell-surface receptors and catalyzes formation of cyclic AMP (cAMP) from ATP; also called adenylate cyclase.
Phospholipase
One of serval enzymes that cleave various bonds in the hydrophilic end of phospholipids.
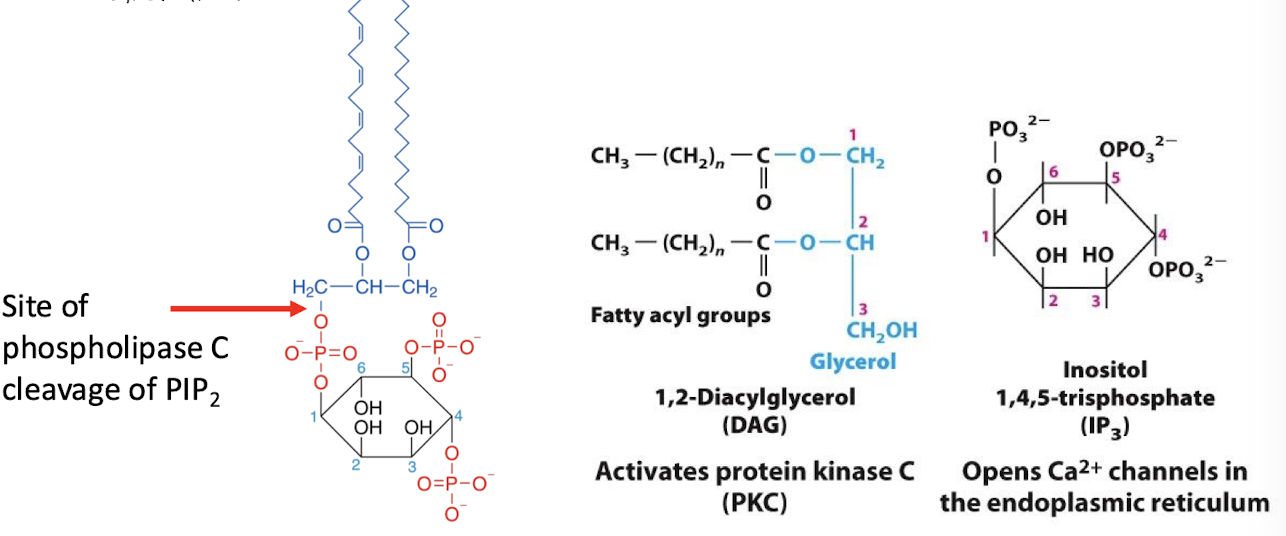
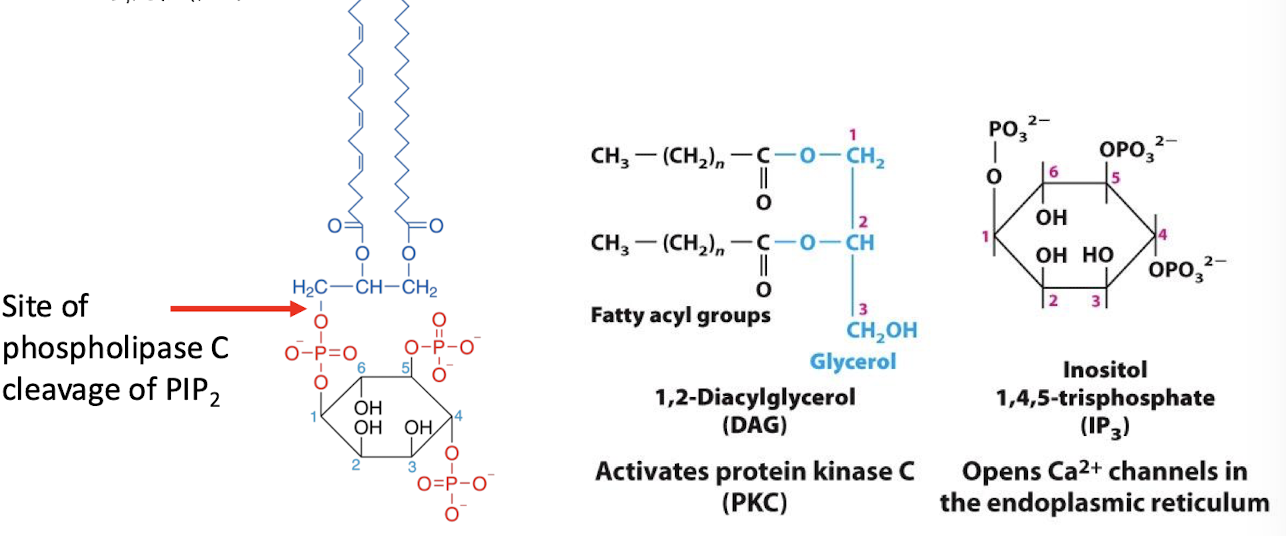
Phospholipase C
An effector enzyme which catalyzes cleavage of the phospholipid phosphatidylinositol 4,5-bisphosphate (PIP2) into the two molecules:
inositol 1,4,5-triphosphate (IP3) (second messenger)
Diacylglycerol (not considered a second messenger b/c stuck in the membrane)
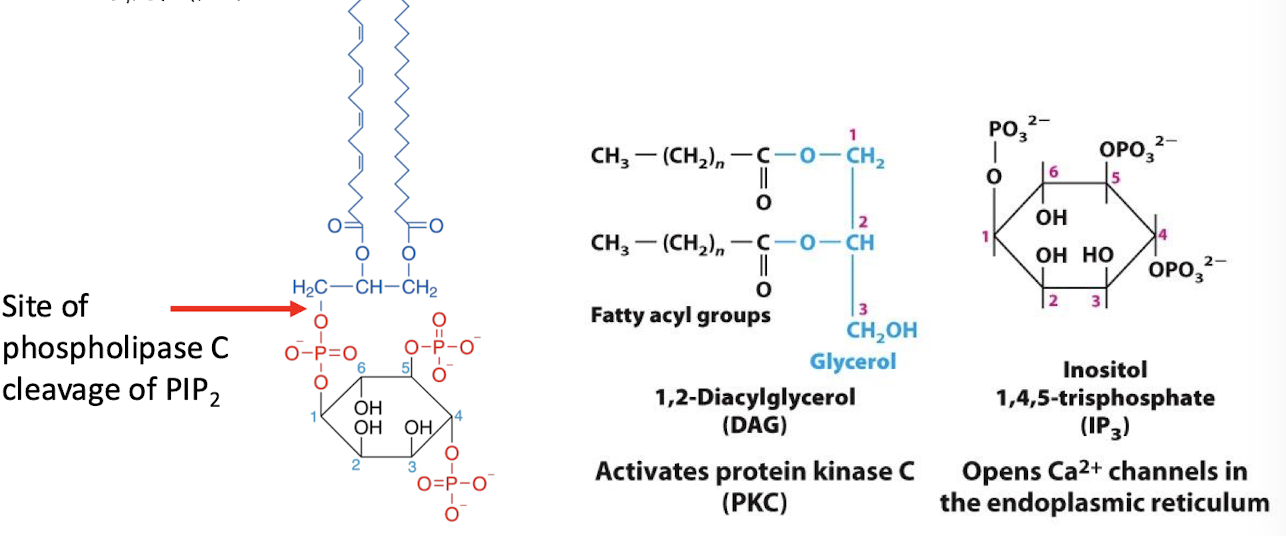
Inositol 1,4,5-triphosphate (IP3)
Functions as a second messenger. It diffuses into the cytoplasm and binds the IP3 receptor on ER membrane
¼ common intracellular second messengers
The IP3 receptor is a lignad-gated Ca2+ channel- IP3 binding ops the channel and releases Ca2+ into the cytoplams
Ca2+ bind and regulates several Ca2+ dependent proteins
Calcium ions ( Ca2+ )
Released from the ER, or imported from extracellular space:
¼ common intracellular second messengers
Activates:
Calmodulin, a ubiquitous switch protein
A subset of protein kinase C proteins (PKC)
Diverse other regulatory proteins (troponin in muscle cells, for example)
1,2-diacylglycerol (DAG)
remains membrane associated- so it is not a true second messenger
it cooperatively activates with Ca2+ a subset of protein kinase c proteins
Epinephrine
regulates body’s fight or flight stress response pathway
A catecholamine secreted by the adrenal glad and some neurons in response to stress; also called adrenaline. It functions as both a hormone neurotransmitter, mediating fight-or-flight responses (e.g., increased blood glucose levels and heart rate)
Glucagon
expressed in response to low blood sugar level
A peptide hormone produced in the cells of pancreatic islets that triggers the conversion of glycogen to glucose by the liver; acts with insulin to control blood glucose levels.
cAMP phosphodiesterase (PDE)
This enzyme is activated by other pathways to stop this transduction. It catalyzes hydrolysis of the cyclic bond-releasing AMP
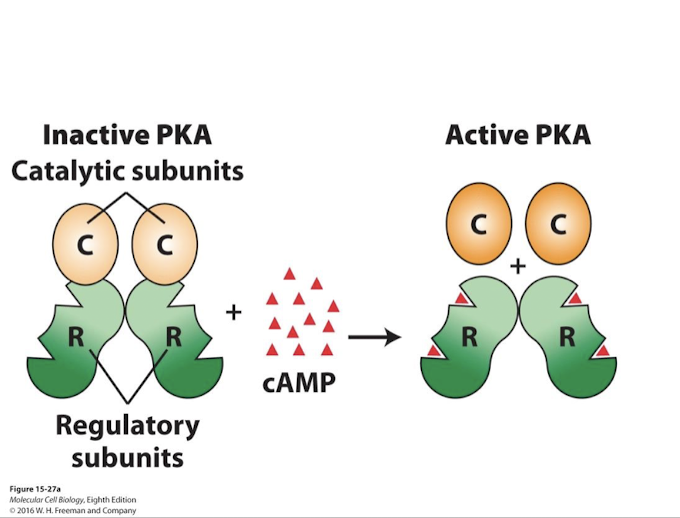
Protein Kinase A (PKA)
cAMP allosterically activates Protein Kinase A (PKA)
A family of soluble serine/threonine kinases
two catalytic kinase subunits catalyze transfer of terminal phosphates from ATP to hydroxyl groups of specific Serine or Threonine residues on target proteins
two regulatory subunits (inhibitory) whose conformation is altered by cAMP
cAMP activates protein kinase A by negative allosteric regulation of the regulatory subunits
cAMP regulates diverse activities depending on:
the cell type in which cAMP is produced
which PKA is expressed
which PKA target proteins are expressed
different cell types express different PKAs and PKA targets- this is what is responsible for distinct cellular responses.
GPCR pathway
G protein coupled receptor pathway
signal transduction pathways initiated by the binding of a ligand to a GPCR, triggering a cascade of events that ultimately lead to a cellular response, often involving G proteins and second messengers.
G⍺s flowchart
ligand → GPCR → G⍺s (stimulatory) → Adenylyl cyclase → +ATP cAMP → PKA activation by binding and inhibiting PKA negative regulatory subunits → PKA phosphorylates cell-type specific target proteins to generate appropriate response
G⍺i flowchart
ligand → GPCR → G⍺i (inhibitory) → adenylyl cyclase → cAMP synthesis blocked
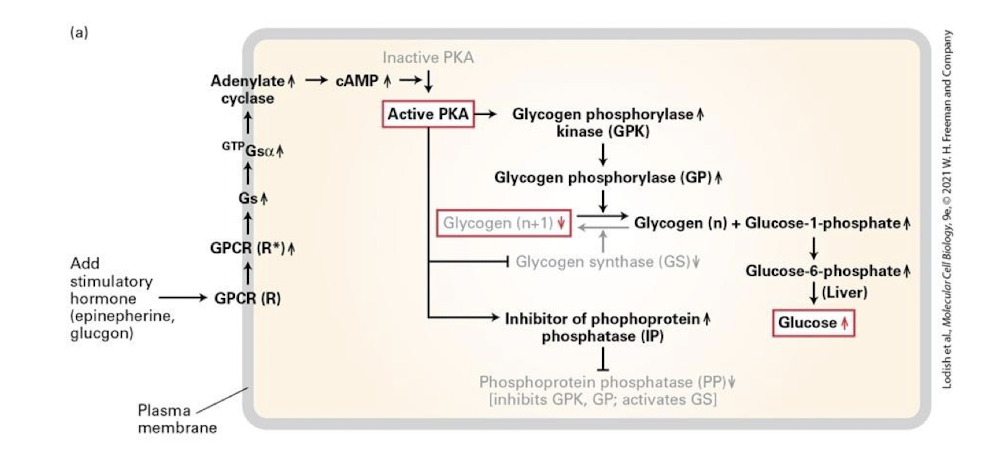
Enzymatic response
Inhibition of glycogen formation and glycogenolysis- hydrolysis of glycogen to G6P
½ responses that lead to elevated Glucose-6-phosphate (G6P) to aid in ATP synthesis
active adenylyl cyclase synthesizes cAMP from ATP which binds and activates PKA
PKA phosphorylates and inactivates glycogen synthase halting glucose polymerization
PKA also phosphorylates and activates glycogen phosphorylase kinase which in turn phosphorylates and activates glycogen phosphorylase - glycogen is cleaved into glucose-1-phosphate monomers which can be converted into glucose-6-phosphate for immediate use in muscles, while a liver enzyme removes the phosphate to the sugar can be released into the blood stream
PKA promotes glycogen hydrolysis by regulating activity of an additional enzyme called phosphoprotein phosphatease (PP)- a protein that removes phosphate groups from residues on target proteins
PP removes phosphates from GPK and GP, thereby inhibiting these proteins and blocking glucose release
PP removes phosphates from Glycogen synthase- which enhances its activity and therefore increases glycogen production
activated PKA phosphorylates and activates an inhibitor of PP, leading to phosphorylation and activation of GPK and GP and the inhibition of GS
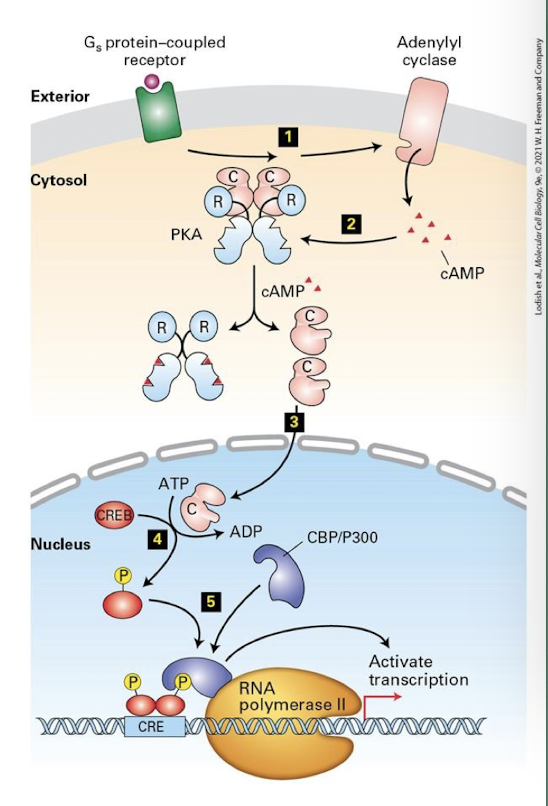
Transcriptional response
Transcription of gluconeogenesis genes that promote the synthesis of glucose from precursor metabolites like lactate and pyruvate- the reverse of glycolysis.
This includes genes encoding the three enzymes that catalyze the irreversible steps of glycolysis are turned off and genes encoding enzymes that catalyze the reverse reactions are transcribed.
½ responses that lead to elevated Glucose-6-phosphate (G6P) to aid in ATP synthesis
PKA is also transported into the nucleus, where it controls expression of specific genes- in the liver, those invloved in gluconeogenesis
CRE: cAMP Response Element- regulatory DNA (enhansers) adjacent to genes activated by PKA
CREB: cAMP Response Element Binding Protein- transcription factor that binds CRE and activates expression of associated genes
CBP/P300: CREB binding protein- links CREB to RNA polymerase promoting transcription.
PKA phosphorylates and activates CREB, leading to assembly of the RNA transcription complex and activation of gene expression
The same transcription factor CREB is activated by PKA is diverse cell types downstream of cAMP and different GPCRs
Different sets of target genes are transcribed in these different cell types because of chromatin structure and the accessibility of genes for transcriptional regulation
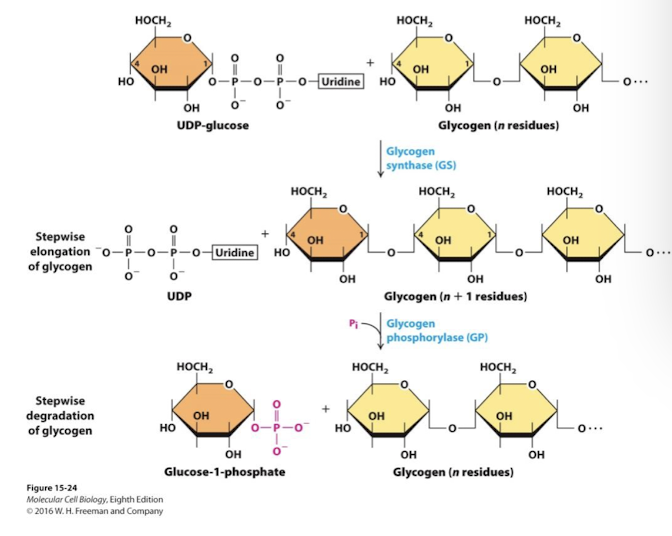
Gluconeolysis
Major way glucose is made avaliable to the body when blood sugar levels are low (glucagon) or when excited/ stimulated (epinephrine)
two key enzymes glycogen phosphorylase and glycogen synthase, are controlled by GPCR signaling
Glycogen phosphorylase is activated in response to glucagon and epinepherine (low blood sugar; energy needed). Glucose is released from glycogen.
GLycogen synthases is activated in response to insulin (elevated blood sugar). Glucose is polymerized into glycogen
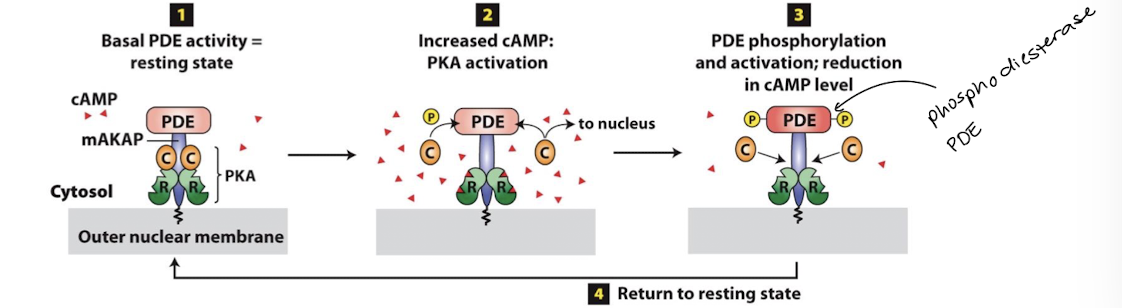
A-Kinase-Associate Proteins (AKAPs)
Membrane anchored and have both PKA and PDE binding domains- tightly restricting where and how much cAMP is produced
Plasma membrane vs nuclear membrane AKAP localization in different cell types can control whether the primary response is enzymatic or transcriptional
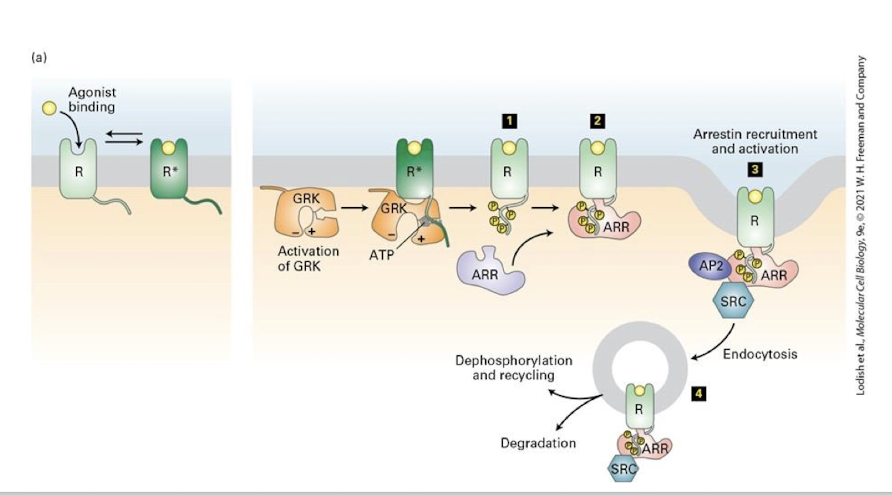
Inhabitation of activated GPCRs
active receptors only are phosphorylated by GPCR Kinases (GRKs) - this inhibits the ability of the receptor to engage G proteins- signaling is suppressed, a phenomenon called desensitization
This then recruits proteins called β-arrestins, which bind the GPCR and enhance the block of transduction
Ultimately β-arrestin recruits clathrin and promotes endocytosis of the activated receptor
The receptor may be recycled back to the cell surface to once again receive signal or be directed to lysosomes for degradation making the cell unable to receive signal
β-arrestin has also been shown to recruit a variety of intracellular signaling kinases (like SRC, shown here) that may activate additional transduction pathways from the membrane of endosomes before being degraded
G⍺q IP3 flowchart
ligand → GPCR → G⍺q → Phospholipase C → + PIP2 (phosphtidylinositol (4,5) bisphosphate) → IP3 inositol (1,4,5) triphosphate: second messenger binds to IP3 receptor on Er releasing Ca2+ → 1. Ca2+ regulates cell-type specific target proteins to generate appropriate response: usually indirectly via binding calmodulin or → DAG: diacyl glycerol not a second messenger, binds and activates some PKC proteins → PKC phosphorylates cell-type specific target proteins to generate appropriate response
G⍺q DAG flowchart
ligand → GPCR → G⍺q → Phospholipase C → + PIP2 (phosphtidylinositol (4,5) bisphosphate) → DAG: diacyl glycerol not a second messenger, binds and activates some PKC proteins → PKC phosphorylates cell-type specific target proteins to generate appropriate response
G⍺t flow chart
ligand → GPCR → G⍺t (transduction) → phosphodiesterase (PDE) → +cGMP or cAMP depending on the PDE → GMP or AMP → cG/AMP signaling is halted
PHospholipase
One of serval enzymes that cleave various bonds in the hydropholic end of phospholipids
GPCRs signaling through the effector enzyme phospholipase C
G⍺q - activates phospholipase C
the G-protein effector enzyme phospholipase C cleaves PIP2 into IP3 (soluble and DAG (membrane bound)
IP3 opens ligand-gated CA2+ channels in the Er releasing Ca2+ into the cytosol which activates Protein Kinase C (PKC) and calmodulin
DAG binds and cooperatively activates PKC
Phosphoinositides
A group of membrane-bound lipids containing phosphorylated inositol derivatives; some function as second messengers in serval signal transduction pathways
are phosphorylated forms of the lipid phosphatidylinositol
PI kinases
Phosphorylate specific -OH groups of inositol generating various PI forms - note difference in number and placement of phosphates
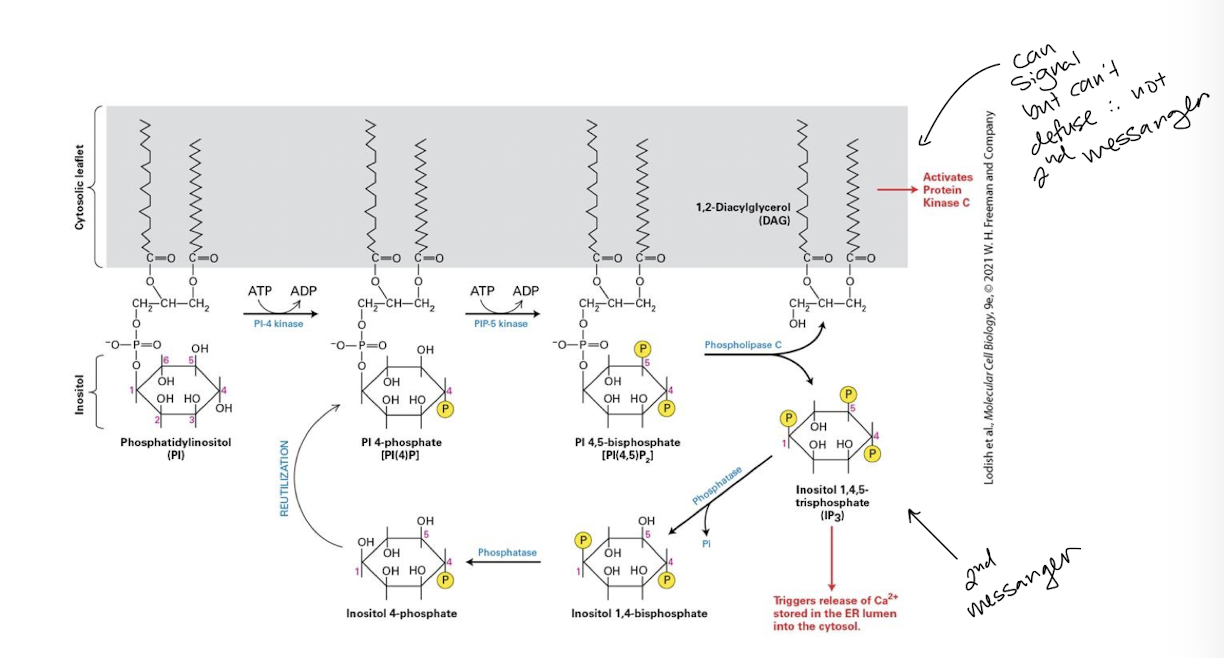
PI-4,5-bisphosphate [PI(4,5)P2] (PIP2)
in the case of GPCR signal transduction, the phosphinosotide PI-4,5-bisphosphate [PI(4,5)P2] (PIP2) is substrate for the effector enzyme phospholipase C itself activated downstream of certain GPCRs
![<p>in the case of GPCR signal transduction, the phosphinosotide PI-4,5-bisphosphate [PI(4,5)P<sub>2</sub>] (PIP<sub>2</sub>) is substrate for the effector enzyme phospholipase C itself activated downstream of certain GPCRs</p>](https://knowt-user-attachments.s3.amazonaws.com/b68b30f2-c6de-45cf-9999-4aa75caf7595.png)
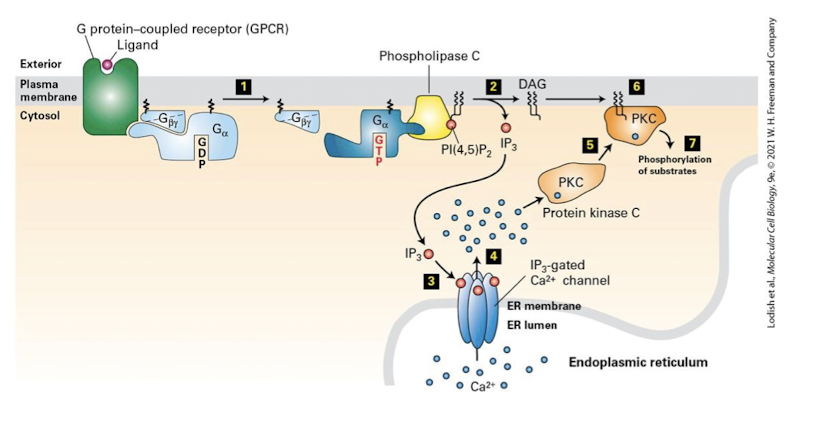
G Protein-Coupled Receptors and Their Second Messengers: IP3 and DAG
the effector enzyme phospholipase C is activated by a subset of heterotrimeric G proteins via either G⍺0 or G⍺q proteins
PLC cleaves PI(4,5)P2 generating insoitol triphosphate (IP3) and the membrane tethered diacylglycerol (DAG)
the sugar phosphate IP3 is water soluble and can rapidly diffuse throughout the cell
IP3 binds the IP3 receptor on the SER surface. the receptor is a ligand-gated Ca2+ channel that once activated releases Ca2+ into the cytoplasm
depending on the cell type these ions bind and regulate diverse proteins that regulate different cellular activities
DAG remains membrane bound and allosterically activates the enzyme protein kinase C which phosphorylates serine and threonine residues on diverse target proteins
these proteins play varied roles in cell growth, differentiation, cell death, metabolism, and the immune response
PKC activation also requires Ca2+ release. only when bound to Ca2+ can PKC translocate to the membrane where it is activated by DAG
CaM Kinases
Ca2+ / calmodulin dependent protein kinases
important family of calmodulin targets
Depolarized State
When potential across the plasma membrane is less negative than that of the normal resting state of the cell
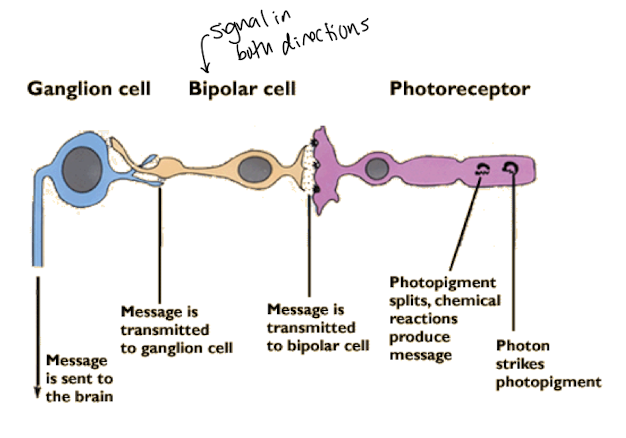
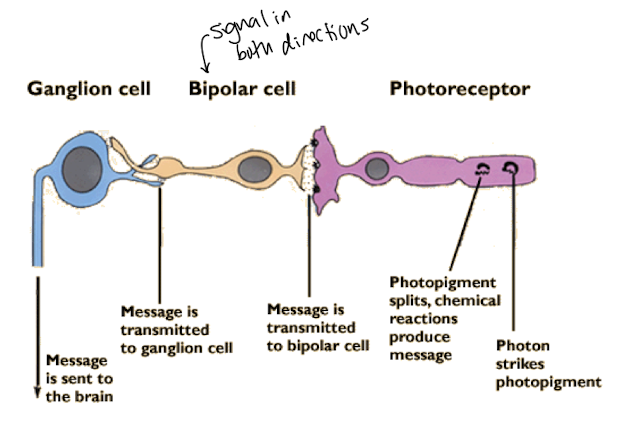
In the absence of light Rod cells are depolarized ~-30mV and secrete a neurotransmitter that stimulates neurons called bipolar cells
Rod cells
photoreceptors responsible for sensing light when it’s dark
Opsin
Once dark adapted, however, our eyes can begin picking up faint light through the action of GPCR opsin, located in flattened membrane disks in the outer segment of rod cells.
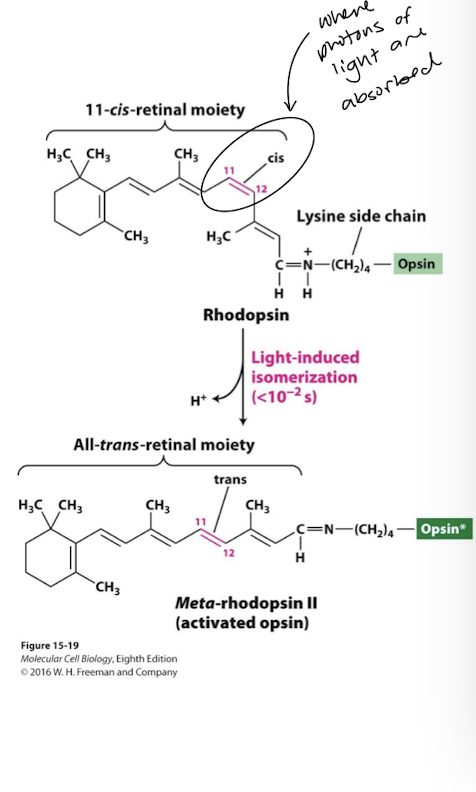
Rhodopsin
Composed of the GPCR opsin covalently attached to the light-absorbing pigment 11-cis-retinal
when 11 cis-retinal absorbs a photon it rapidly isomerizes to the all-trans isomer
this induces a conformational change in rhodopsin to the activated unstable intermediate meta-rhodopsin II
meta-rhodopsin II binds and activates its cognate heterotrimeric G-protein generating G⍺t-GTP (transducin)
the all-trans-retinal moeity quickly dissociates from opsin (within milliseconds)
It is converted by a series of enzymes back to the cis isomer and reattaches to opsin
Guanylyl Cyclase
In the depolarized state the enzyme guanylyl cyclase catalyzes production of high levels of the second messenger cGMP
Glutamate
neurotransmitter blocking neurotransmitter release by bipolar cells
Dark adapted rod cells and closing of cation channels
In the dark, our retinal are able to regenerate 11 cis-retinal. It can take up to 30 minutes for human eye to become fully dark adapted. One adapted however:
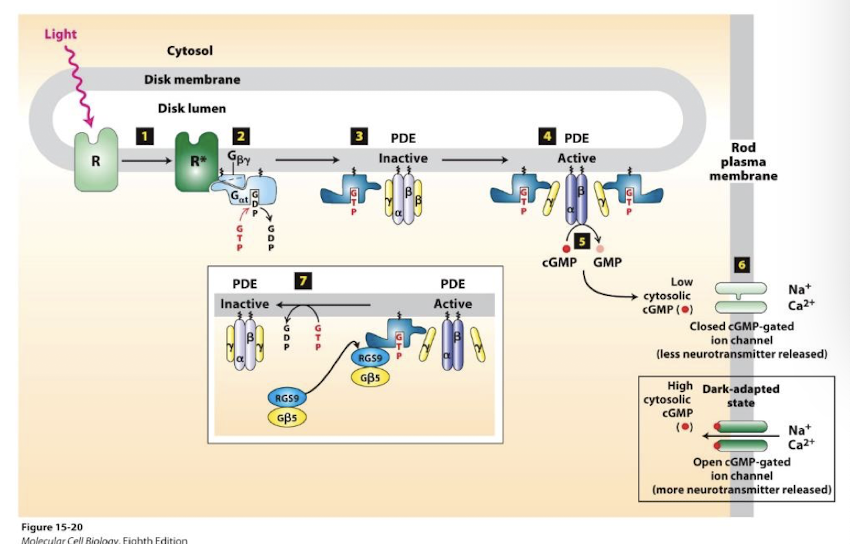
Rhodopsin absorbs photons and is converted to meta-rhodopsin II
meta-rhodopsin II functions as a GEF to catalyze Gαt exchange of GDP for GTP.
Free GαtGTP binds and activates the enzyme phosphodiesterase (PDE)
PDE hydrolyzes cGMP to GMP - dramatically lowering cGMP concentration
Cation channels close and Na+ and Ca2+ influx ceases
Membrane is transiently hyperpolarized.
Glutamate release is blocked.
Bipolar cells are no longer stimulated
They secrete their own neurotransmitters
Which are received by the ganglia that transmit the perception of light to the brain.
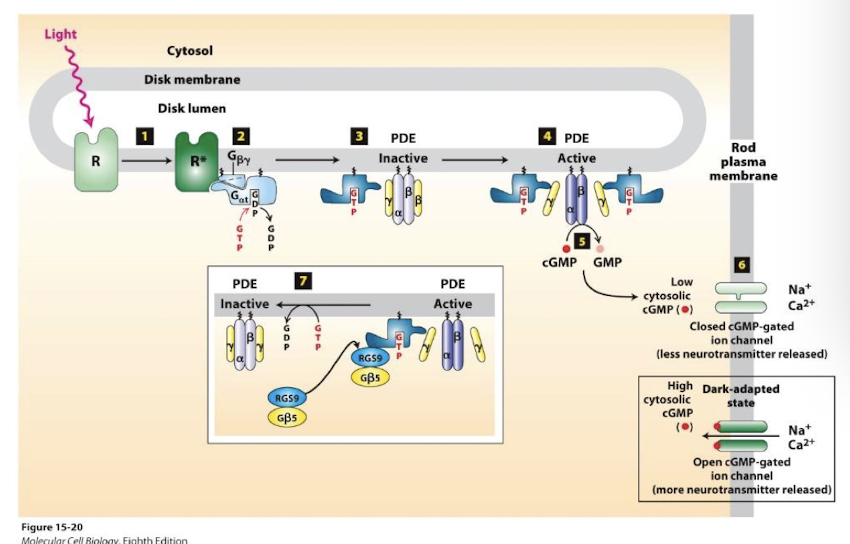
Rapid termination of rhodopsin signaling
signal termination: (~0.2 sec, essential for the temporal resolution of vision)
G⍺T bound by a GAP complex (inset box)
Which stimulates rapid G⍺t GTP hydrolysis to GDP
This reestablishes the inactive heterotrimeric G-protein
PDE is deactivated and cGMP accumulates- cation channels reopen
Also, the drop in Ca2+ activates guanylyl cyclase activating proteins. These are Ca2+ sensing proteins that are inactive when bound by Ca2+ .
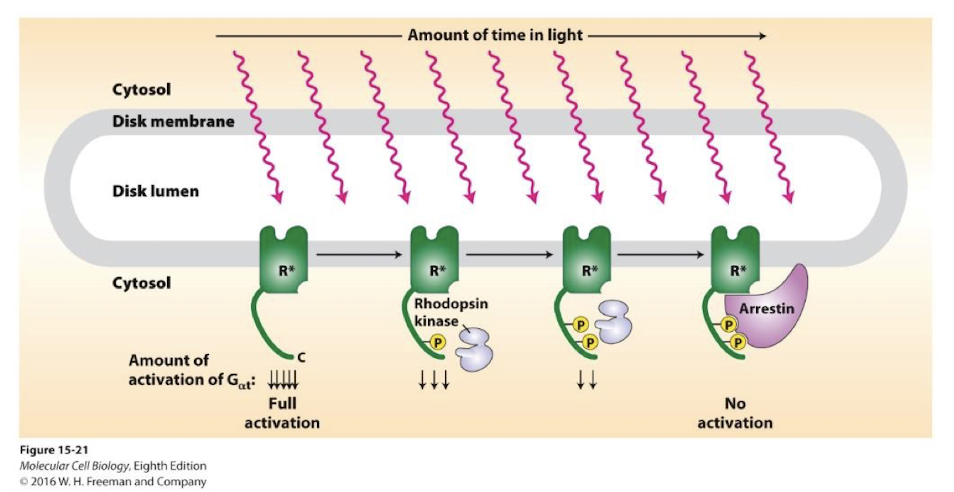
Feedback repression of activated rhodopsin
Rhodopsin kinase- A GRK kinase that phosphorylate activated GPCRs Phosphorylation inactivates and desensitizes them to further stimulation
Arrestin proteins bind phosphorylated GPCRs preventing them from interacting with and activating G⍺T - stimulation is transiently halted
phosphatases removes GPCR phosphorylation, arrestin is released and rhodopsin can again be excitied- there is no endocytosing of these GPCRs