BIOMI 2900 Quiz 3
1/129
Earn XP
Description and Tags
after midterm material (lecture 20- )
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
130 Terms
requirments of life
liq water
carbon
nutrients
free energy
major atoms in microbes
CONHPS
C 50%
O(not O2) 20%
N 14%
H 8%
P 3%
S 1%
What evolutionary adaptation has the greatest potential to benefit a bacterium living in an environment with very low P?
Reduce ribosome number
Macronutrients
– Required by all cells: C, O, N, H, P, S
– Required by most cells: K, Na, Ca, Mg, Cl
Trace Elements
inorganic micronutrients, they are typically found in enzymes where they are found in Cofactors
– Include: Fe, Se, B, Cu, Mn, Mo, Zn, F, Si, As, Cd, Sr, Ba, V, Co, W, Ni
– Requirements vary from species to species
Growth Factors
organic micronutrients
– Include: vitamins, amino acids, purines, pyrimidines, etc
– Vitamins: often function as Coenzymes (non-protein molecule thatcontributes to enzyme function)
– Requirements vary from species to species
Culture media (plural = media, singular = medium)
– Nutrient solutions used to grow microbes in the laboratory
Liquid or solid
– Solid media requires solidifying agent (e.g. agar)
Two broad classes
– Defined media
– Complex media: composed of digests of chemically
undefined substances (e.g., yeast, meat or plant extracts)
Defined media:
precise chemical composition is known
Complex media:
composed of digests of chemically undefined substances (e.g., yeast, meat or plant extracts)
Composition of Defined Media
C: can be an organic C source (for heterotrophic microbes) can be CO2 (for autotrophic microbes)
N: NH4+, NO3-, N2, or organic N (such as protein)
P: phosphate PO42- (or organic P)
S: sulfate SO42- (or organic S)
Other macronutrients: in the form of Inorganic salts
Trace elements
Growth Factors: typically vitamins
pH buffer
Diagnostic Media: used to identify certain bacteria
-Selective Media: selects for target bacteria
(e.g. use of inhibitors, specific nutrients)
-Differential Media: have in indicator (e.g. a dye) that
changes color when it interacts with a target organism
Selective Media:
selects for target bacteria (e.g. use of inhibitors, specific nutrients)
Differential Media:
have in indicator (e.g. a dye) that changes color when it interacts with a target organism
Blood agar diagnostic media
differential media
Identifies bacteria able to lyse blood cells
Pattern of hemolysis is diagnostic
Bile Salts agar diagnostic media
Selective media
Selects for bacteria resistant to bile salts
MacConkey agar diagnostic media
Selective & Differential
targets enteric G- bacteria that ferment lactose
Crystal violet and bile salts inhibit G+ bacteria (selective)
Lactose is the only C-source (selective)
pH indicator reveals acid production (differential)
Enrichment Culture
use a medium that provides a target microbe some selective advantage allowing it to be enriched in abundance over time (results in a mixture of bacteria where one type is common)
Direct Isolation:
use selective and/or differential agar to isolate a single colony (using streak or spread plate technique) – used to obtain a pure culture composed of a single strain
streak plate, direct isolation method
diluting to a single colony by streaking from previous streaks on the plate, sterilizing tool each time
spread plate - direct isolation method
spreading a liq with bacteria of an entire plate to get a spread of colonies
Microscopic Counts
counting bacteria on gridded plate
• Limitations of microscopic counts
Laborious
Likelihood of human error, visually cannot distinguish cells vs. random impurity
can’t tell which ones are living or dead cells
Viable Counts (Plate Counts)
• Measure Colony Forming Units (CFU) (slight underestimate)
• Serial Dilution: Sample must be diluted to obtain a countable number of colonies
• Spread Plating: Cells are spread on a plate, plates are incubated, and then colonies are counted
Assumption: One cell, one colony (assumption violated for filamentous and ‘sticky’ cells)
number of bacteria after dilution?
plate count x dilution factor = 1.59 x 10^5
spectrophotometry
measure turbidity: light scatterying by cells
fast,easy, nondestructive
optical density
Growth Rate:
change in cell number over time
Growth Yield:
mass (or number)of cells relative to an input
Batch Culture:
a closed-system culture of fixed volume
Continuous Culture:
an open-system culture of fixed volume
Biofilm Culture:
a system that favors microbial growth on a surface
Growth Curve
Exponential Growth in Batch Culture
Lag phase:
Physiological changes to prepare for growth in new environment, length varies
Exponential phase:
Balanced exponential growth of all cells
Stationary phase:
Growth terminates when the environment no longer supports growth (e.g. nutrient limitation, pH, waste buildup), period of no net growth, physiological changes to prepare for starvation & dormancy
Death/Decline Phase:
Decline in cell number
how many cellsl in exp growth euqation:
Nt = N02n
n = number of generations
to solve for n:
n = (ln Nt - ln N0) / 0.693
numbers of generations (n)
generation time = time per gen (t/n)
growth rate constant (k) equation
dN/dt = kN → k = (ln Nt - ln N0) / t
dN - change in cells over t
k = grwoth rate constant
N = cell number
Nt = cell number at time t
N0 - initial cell number
division rate
1/g
units for growth rate equations
g: doubling time
t: time
Nt: cells at time t
N0: initial cell number
n: generation number
k: growth rate constant
D: division rate
chemostat
a continuous culture device for growth bacteria
growth of continous culture
in chemostat:
growth rate is controlled by dilution rate
growth yield is controlled by the concentration of limiting nutrients
Aerobes interaction with oxygen
require O2 to grow (typically air, 20% O2)
Microaerophiles interaction with oxygen
grow only at low O2 (often 0.5 – 5%)
Facultative organisms interaction with oxygen
can live with or without O2
Aerotolerant anaerobes interaction with oxygen
can tolerate O2 and grow in its presence even though they cannot use it
Obligate anaerobes interaction with oxygen
killed by exposure to O2
Mesophiles
organisms that have midrange temperature optima; found in
• Microbes associated with warm-blooded animals
• Terrestrial and aquatic environments
• Temperate and tropical latitudes
Extremophiles
Organisms that require very hot or very cold conditions
Psychrophiles
Growth optima < 20°C
inhabit permanently cold environments
Life possible as long as there is liquid water (observed to -12°C)
Psychrotolerant
Growth optima > 20°C, but can grow at 0°C
More common than true psychrophiles
Thermophiles
growth optima > 45°C
Live in hot environments: composts, solar heated, geothermal
Molecular adaptations that support thermophily
• Changes protein structures for stability
• Changes membrane structure fore stability
Saturated fatty acids increase membrane stability
Isoprene lipids and lipid monolayers increase membrane stability in Archaea
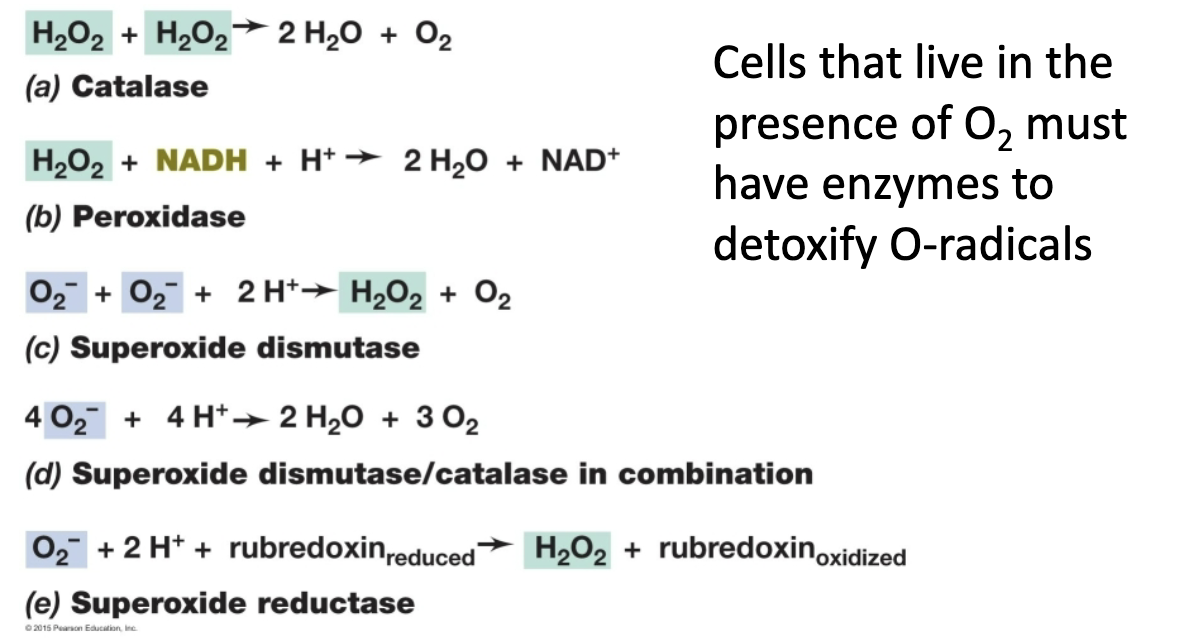
peroxidase
H2O2 → H2O + O2
catalase
H2O2 → H2O
types of enzymes for detoxification
Water activity (aw) = water availability
Water activity is related to solute concentration, water affiliating with solutes is not available to cells
Solutes include all dissolved molecules (salts, sugars, small molecules, amino acids, etc)
Water activity is measured as the ratio of the vapor pressure of air in equilibrium with a solution to the vapor pressure of pure water
Molecular adaptations related to pH
• change membrane structure for stability
• pH of the cytoplasm can change
typically remains close to neutral, but can dip as low as 4.6 in extreme acidophiles, and as high as 9.5 in extreme alkaliphiles
• At very high pH there are few available protons (H+), so use Na+ motive force instead of a pmf
Hyperthermophiles
growth optima > 80°C
Live in geothermal and hydrothermal environments
optima examples (note)
No Eukarya can grow at > 65°C,
No Bacteria can grow at > 95°C
the theoretical limit for growth is ~150-180°C
Sterilization
complete removal of all microbes
Heat sterilization
routine use, requires >120°C
Autoclave
Chemical sterilization
formaldehyde bath
Bleach bath
-Ethylene oxide gas – sensitive medical devices
Radiation - sterilization
Gamma rays/x-rays – medical/food industries
UV light – for surfaces (low penetration)
Filtration
0.2μm removes bacteria (bacteria usually larger than .2)
HEPA filters – for gases/air (High-Efficiency Particulate Air filter)
Membrane filters – for solutions
what is an autoclave and what is it used for
sealed device that uses steam under pressure to sterilize items and solutions
Pressure allows heating water to 121°C without boiling
not pressure, but high temperature kills microbes
Pasteurization
process of using precisely controlled heat to reduce the microbial load in heat-sensitive liquids
Does not sterilize (ultra-pasteurization comes close), so it is different from sterilization
heat killing (math)
function of temperature v time
Decimal Reduction Time (D) : time required for 10x reduction in cell number
Decimal Reduction Time (D)
time required for 10x reduction in cell number
Which of the following terms describes a chemical that prevents microbial growth but does not cause cells to die?
D. Bacteriostatic
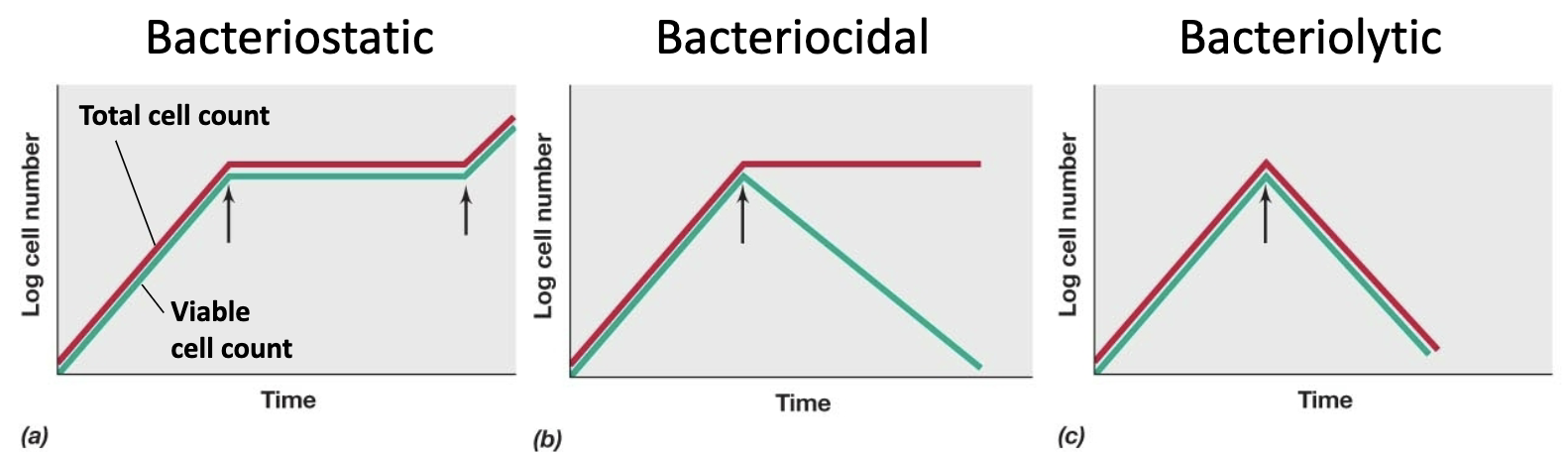
Bacteriostatic
Inhibit growth, bind weakly to target
Bacteriocidal
Kill cells, bind strongly to or irreversibly damage target
Bacteriolytic
Lyses cells
causes osmotic shock, attacks cell walls etc.
How to determine the chemical type of antimicrobial
red is microscope data, green is spread plating data
Sterilants
destroy all cells including endospores
e.g. ethylene oxide, bleach bath, flame
Disinfectants/Sanitizers
kill most cells (including human cells), but not endospores, do not sterilize
e.g. bleach on surfaces, alcohols, harsh detergents
Antiseptics (germicides)
kill many cells but safe enough to be used on living tissues (not injestion)
e.g. mild detergents, Betadine (iodine)
antibiotic common traits
Low toxicity to humans (few or no side effects)
Target structures unique to bacteria
Water soluble – need to disseminate in body
Generally bacteriocidal or bacteriostatic
types of antimicrobials
Antibiotics- typically target bacteria
Antifungals- target fungi
Antivirals- target viruses
Most common targets for antibiotics (structures unique to bacteria),
• Bacterial Cell Wall Synthesis
• Bacterial Protein Synthesis
• Bacterial DNA Replication
• Folate Synthesis
• Bacterial Transcription
• Bacterial Cytoplasmic Membrane
beta-lactam
antibiotic that attacks transpeptidase (helps form peptidoglycan)
sealing
seal food more because of cross contamination, not for preventing bacterial growth
food preservation
Heat
Hot Holding for cooked foods (>57 °C)
Pasteurization (~65 °C)
Canning (boiling)
Ultrapasteurization (up to 135 °C)
Refrigeration and Freezing
– Bacterial growth slowed at 4 °C, but still grow (e.g. Listeria)
pH for food preservation
acidic pH tends to preserve foods
Acidic additives: sorbic acid, benzoic acid, propionic acid
Microbial acid production: lactic acid bacteria (yogurt, cheeses, sausages, saurkraut, pickles, etc...)
Water availability for food preservation
Drying
Freeze Drying
Osmotic Pressure e.g. with salt or sugar
Chemical Preservatives
• Smoking (dries and deposits aldehydes, acids and phenols)
• Nitrites, nitrates
• Sulfites
• Etc . . .
irradiation
food preservative
free energy (G)
energy available to do work
ΔG°′
change in free energy at standard condition (°) and pH 7 (′)
exergonic
if ΔG°′ < 0
endergonic
if ΔG°′ > 0
OIL RIG
oxidation is lose electrons (ox state increase),
reduction is gain electrons (oxidation state is reduced)
Reduction potential (E0′):
– Measured in volts (V)
– We start with ‘half reactions’
– ‘Half reactions’ indicate
oxidized/reduced products
– Transfer of e- from more negative donor
to more positive acceptor is exergonic
– Redox couples are reversible
Redox tower:
O2 at the bottom, great electron acceptor and is beneficial to do so with coupled reactions

ΔG0′ = -nFΔE0’
ΔG0′ change in free energy (std cond, pH7)
n = number of electrons transferred
F = Faraday Constant (96.48 kJ/V)
E0’ change in reduction potential
(accepting couple) – (donor couple)
Glycolysis (aka: Embden–Meyerhof pathway)
• a common pathway for catabolism of glucose
• glucose is the electron donor
• anaerobic (no O2 required)
• used by chemoorganoheterotrophs capable of either fermentation, aerobic respiration, or anaerobic respiration
• ATP is generated by Substrate Level Phosphorylation
• Generates 2 ATP, 2 NADH, and 2 pyruvate (C3H3O3)
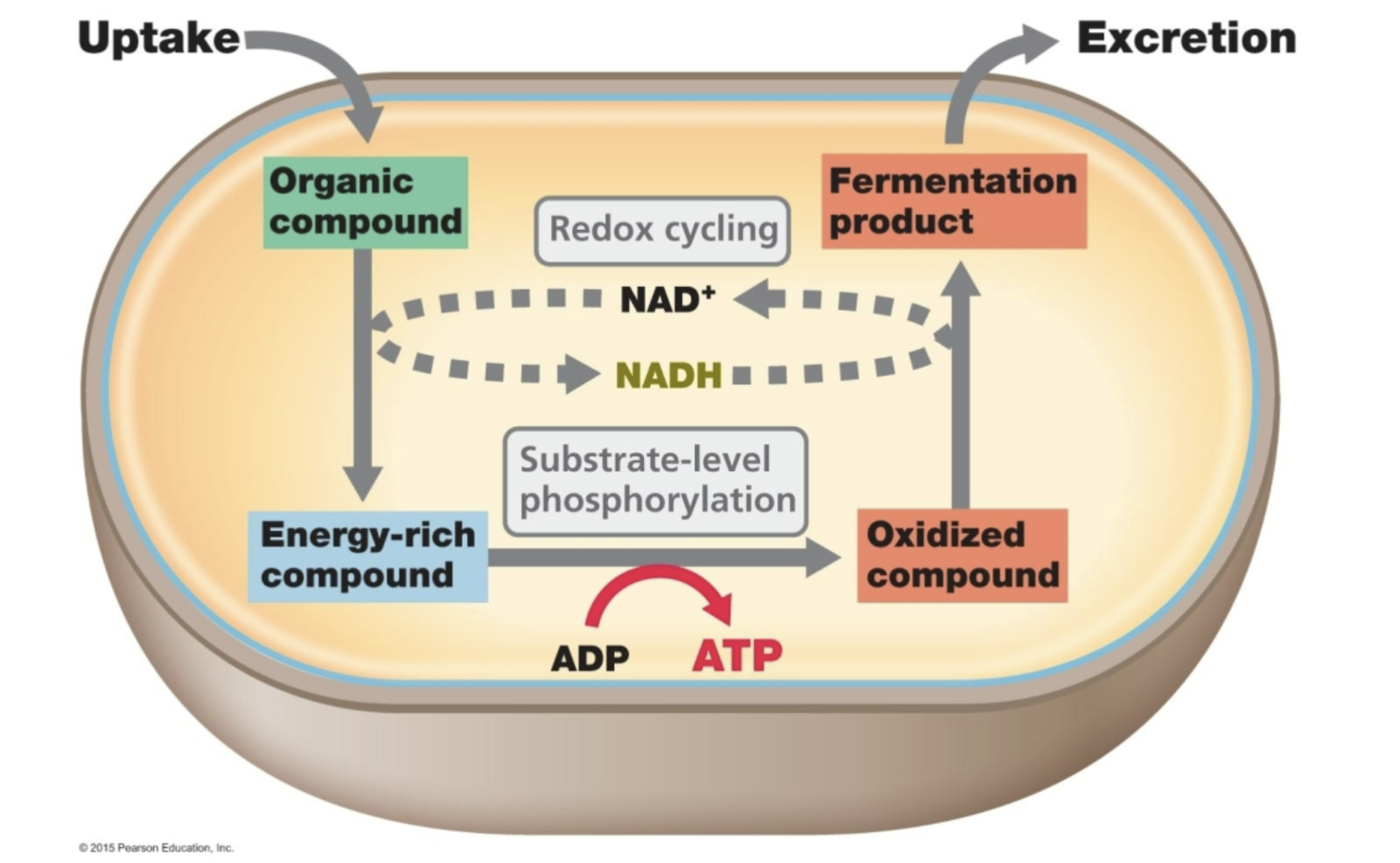
Problem:
- Fermentations do not have an external electron acceptor.
- During glycolysis you reduce NAD+ to NADH.
- The electrons need to be removed from NADH so that you can
regenerate NAD+ . If you run out of NAD+ then glycolysis will stop.
Hence, YOU NEED AN ELECTRON ACCEPTOR to get rid of the
excess electrons. If there is no external electron acceptor what can
you do. . .
Solution:
- Put the e- from NADH back onto a breakdown product of the original substrate
2) Fermentation
2 Pyruvate 2 Lactate + 2 H+ 2 NADH 2 NAD+
e.g. Homofermentative lactic acid bacteria
2 Pyruvate 2 Ethanol + CO2
2 NADH 2 NAD+
e.g. Yeast
Citric acid Cycle (a.k.a Krebs Cycle)
• A central and highly conserved
metabolic cycle
• Required by many anabolic processes
as intermediates can be siphoned off
and used in biosynthesis
• Required for catabolism by many
heterotrophs
• Pyruvate (C3) enters
• Generates:
3 CO2, 1 ATP, 1 FADH2, 4NADH
Electron transport in Oxidative Phosphorylation (used in Respiration)
- System of integral membrane proteins that perform redox reactions which oxidize electron donors, shuttle electrons along the membrane, generate a Proton Motive Force, and ultimately reduce a terminal electron acceptor
Electron transport in aerobic respiration
1: electrons enter the chain from primary electron donor: NADH → NAD+
2: H+ pumped out (pmf) and 1e- flows through chain
flavin takes e-, H+ out → quinone takes both H+ and e- → Iron picks up e-, H+ out
3: electron exits by reducing terminal electron accepter: O2 → H2O