11. Distinction Between Self and Non-Self: T-Cell Receptors I
1/25
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
26 Terms
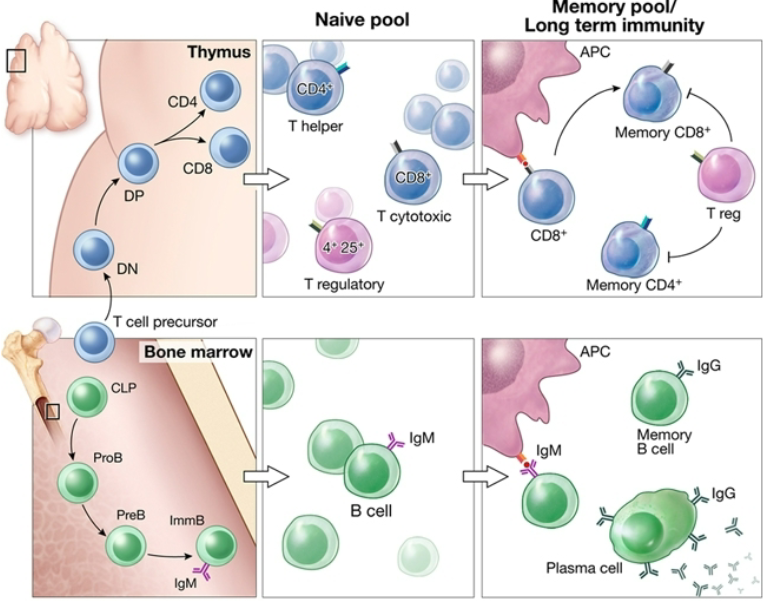
Describe the process of T-cell development.
T-cells originate in the bone marrow, their precursors migrate to the thymus.
In the thymus, positive and negative selection occurs, producing fully functional naive T-cells that react to MHC but do not react to self-peptides.
Naive T- cells are released into the peripheral lymphatic system, to be activated upon cognate ligand binding.
What is positive selection in T-cell development?
Tests whether T-cell receptors (TCRs) can moderately bind self-MHC molecules.
T cells with TCRs that moderately recognise self-MHC receive survival signals. This ensures maturation of T cells that can recognize antigens presented by self-MHC.
Determines co-receptor fate (CD4⁺ or CD8⁺) based on whether they recognize MHC class II or I.
What is negative selection in T-cell development?
Eliminates thymocytes whose TCRs bind strongly to self-peptide:MHC complexes.
Prevents development of autoreactive T cells and helps establish central tolerance. Occurs primarily in the thymic medulla.
What is the affinity hypothesis of T-cell development?
The affinity hypothesis proposes that T-cell fate is determined by the strength of TCR binding to self-peptide:MHC:
• No/very weak interaction → death by neglect
• Moderate interaction → positive selection
• Strong interaction → negative selection (apoptosis).
What distinguishes T cell antigen recognition with B cell antigen recognition?
T cell antigen recognition requires antigen presentation with APC and MHC molecules. This involves a cell-cell interaction between the TCR and peptide-MHC interface.
B cells can recognise free (soluble or surface-bound) antigens directly without the need for MHC presentation. B cell receptors (BCRs) bind to native antigens in their natural conformation.
Describe how the TCR complex is involved in T cell activation.
TCRs recognises foreign peptide antigens presented by MHC molecules on APC surfaces. This antigen recognition is MHC-restricted and involves direct cell-cell contact.
TCR engagement activates intracellular signalling cascades (via kinases like Lck and ZAP-70), resulting in T cell activation, proliferation, and differentiation.
This mode of antigen recognition reflects the role of T cells in cell-mediated immunity, which is essential for detecting and eliminating intracellular pathogens (e.g., viruses, some bacteria).
What is antigen recogntion?
The activation of a T-cell in response to a binding events of TCR clone to a cognate presented antigen.
TCR releases peptide-MHC ligand otherwise.
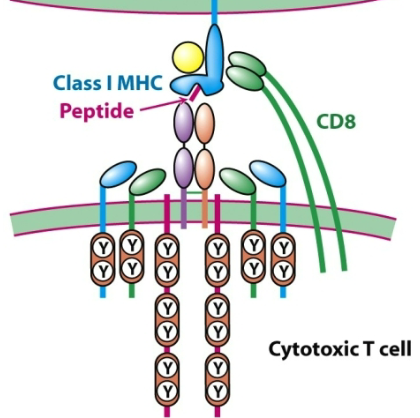
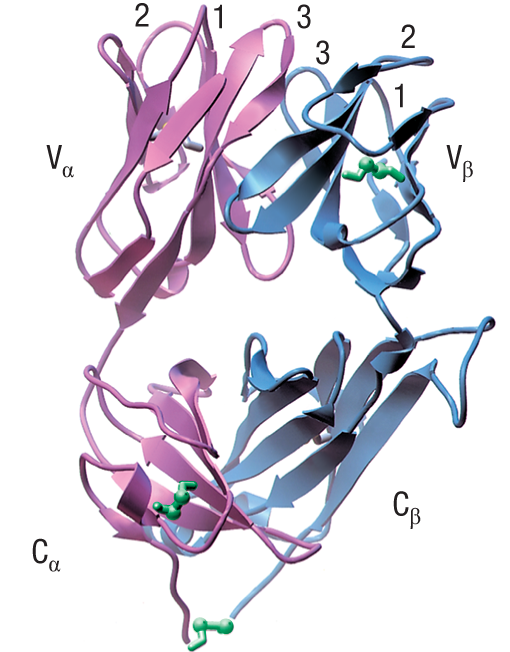
What is the structure of a TCR?
The TCR is a heterodimer of two transmembrane glycoprotein chains: α and β.
Each chain has two extracellular Ig-like domains: one variable (V) and one constant (C).
A short stalk connects the Ig-like domains to the membrane and includes a cysteine residue that forms an interchain disulfide bond.
The transmembrane regions contain positively charged (basic) residues, which are unusual in hydrophobic membrane segments.
α chain: 2 basic residues
β chain: 1 basic residue
These residues are essential for assembly with CD3 complex proteins, forming the full TCR complex.
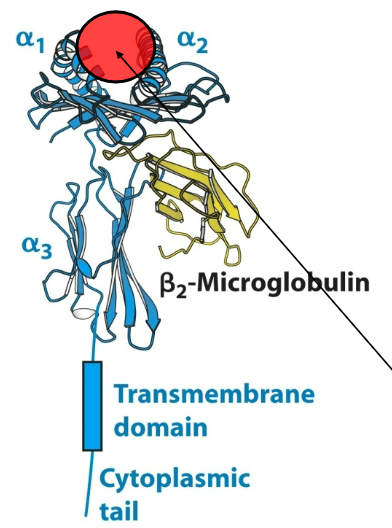
Describe the architecture of the MHC (Class I) molecule.
Short cytoplasmic tail
Single transmembrane helix
α3 (Ig fold)
α1, α2 domains form peptide binding platform
β2 microglobulin (β2m) associates non-covalently with domains α1-α3 , but plays no role in recognition.
What are differences between antibody-Ag and TCR-Ag interactions?
Antibodies bind native antigens directly, without the need for antigen processing or presentation.
TCRs (specifically αβ-TCRs) recognise processed peptide antigens only when presented on self-MHC molecules (MHC restriction).
TCR-antigen interaction involves both the foreign peptide and the self-MHC molecule on an antigen-presenting cell (APC).
MHC molecules are self proteins; T cells are trained to recognise antigen only in this self context.
TCRs that recognize non-self MHCs are termed alloreactive.
TCR binding affinity is relatively weak (Kd ~ 1–90 µM) compared to high-affinity antibody-antigen interactions.
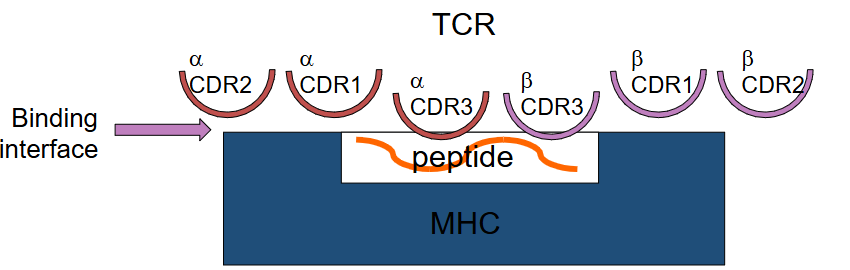
What is MHC restriction?
TCR binding to cognate peptide antigen only when presented by the correct MHC molecule.
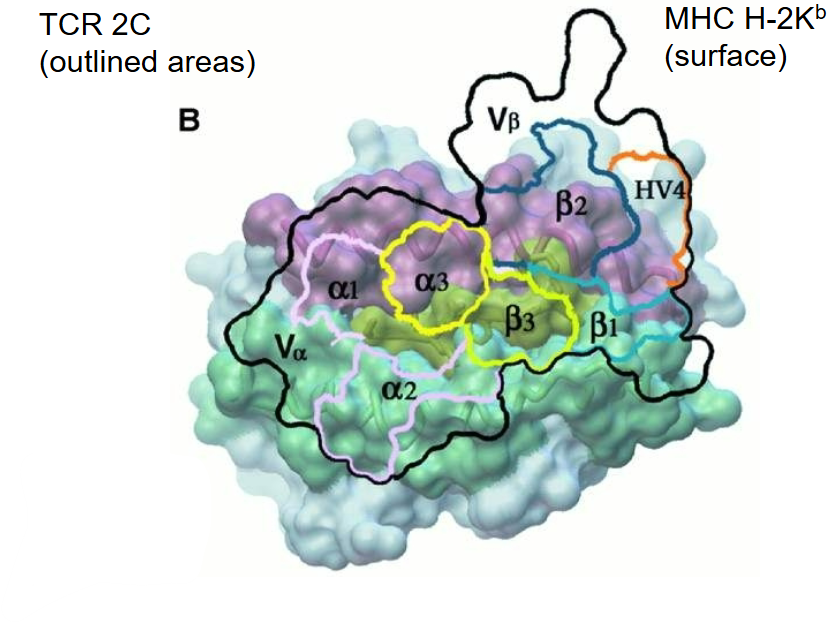
What does the TCR-pMHC complex provide insight into?
The TCR–pMHC complex reveals:
The canonical orientation of TCR docking on MHC: diagonal alignment over the peptide-binding groove.
Which structural elements of the TCR contact the MHC and the peptide.
The CDR3 loops of both TCR chains interact exclusively with the peptide, making them key for antigen specificity and recognition.
CDR1 and CDR2 loops primarily contact the MHC α-helices, mediating MHC restriction.
The interaction surface is relatively small, contributing to the relatively low-affinity binding.
What are the key features of cytolytic T cells (CTLs)?
Express αβ-TCR + CD8 co-receptor (dimer)
Bind peptides presented by MHC class I
Have direct effector function: kill target cells by inducing lysis
Exhibit memory function after activation
What are the key features of helper T (Th) cells?
Express αβ-TCR + CD4 co-receptor (monomer)
Bind peptides presented by MHC class II
No direct effector function, instead activate effector functions through cytokine release
Exhibit memory function after activation
Differentiates into helper subtypes: Th1, Th2, Th17, Treg
What are some other types of T cells and their recognition features?
γδ T cells express TCR, often recognising non-peptide antigens like lipid carbohydrate ligands; less MHC-restricted.
MAIT (mucosal-associated invariant)T cells recognise bacterial metabolites presented by MR1 (non-classical MHC)
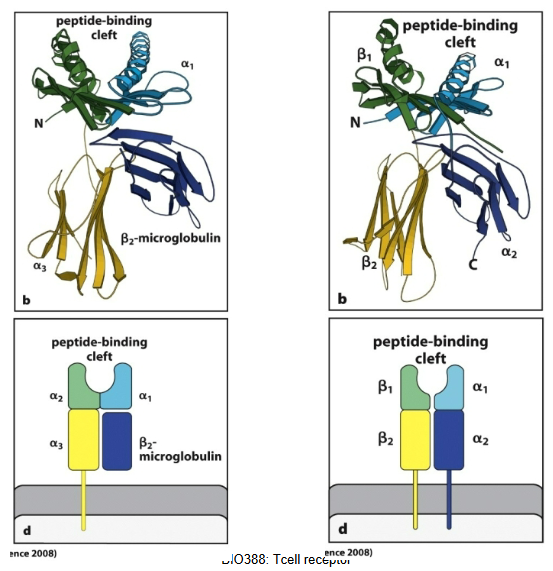
How does chain architecture differ between MHC classes?
MHC class I consists of a heavy α chain (with three extracellular domains: α1, α2, α3) and a non-covalently associated β2-microglobulin.
Presents peptides to CD8⁺ T cells
CD8 binds to the α3 domain
MHC class II is composed of two transmembrane chains, α and β, each with two extracellular domains (α1/α2, β1/β2).
Presents peptides to CD4⁺ T cells
CD4 binds to the β2 domain
Both have a peptide-binding groove formed by the α1 and α2 domains (class I) or α1 and β1 (class II), but differ in chain architecture and co-receptor binding sites.
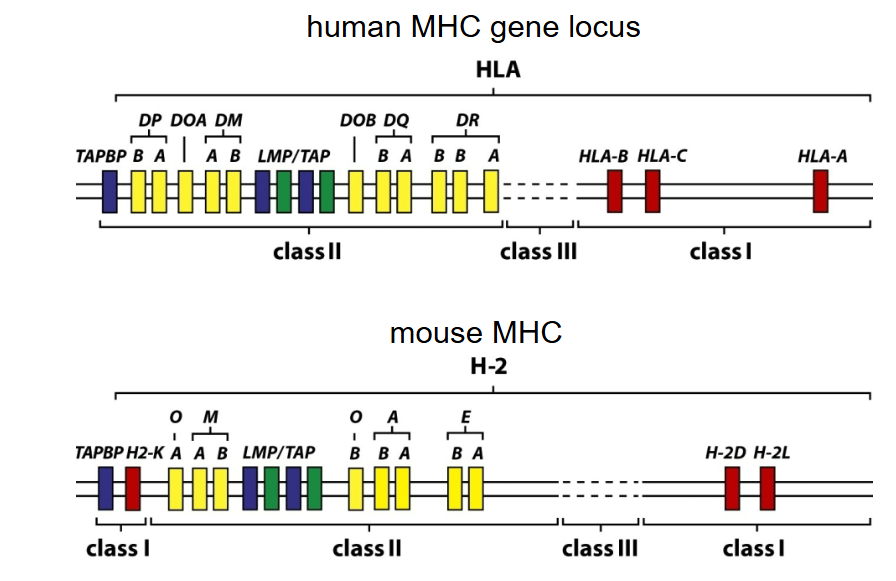
What can be observed following loci comparisons between human and mouse MHC?
MHC is polygenic (encoded by several genes)
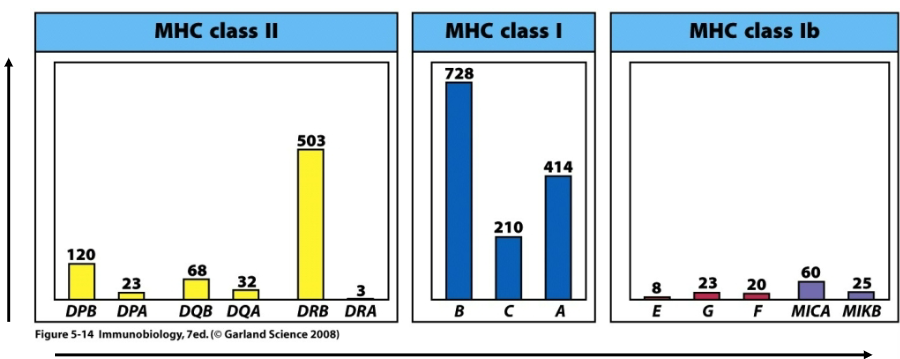
What can be observed following sequence comparison between individual MHC classes?
MHC is polymorphic (high sequence variation)
How do antigen processing pathways differ for MHC class I and II presentation?
MHC class I presents peptides from the cytosolic (endogenous) pathway:
Proteins from inside the cell (e.g. viral or self-proteins) are degraded by the proteasome
Peptide fragments are loaded onto MHC I in the ER and presented to CD8⁺ T cells
MHC class II presents peptides from the exogenous pathway:
Extracellular proteins taken up by endocytosis are degraded in lysosomes
Peptides are loaded onto MHC II in endosomal/lysosomal compartments and presented to CD4⁺ T cells
How does the MHC gene repertoire influence peptide binding diversity?
The diversity of MHC alleles determines which peptides can be presented to T cells.
Each MHC molecule has a peptide-binding groove that selects for peptides with specific anchor residues at defined positions.
These anchor residues fit into binding pockets on the MHC surface.
As a result, only certain peptides can bind to a given MHC allele. Binding is highly selective.
The expressed set of MHC molecules limits the repertoire of peptides that can be presented to the TCR.
What are anchor residues? What is their significance in TCR binding?
Anchor residues on peptides which bury into the peptide binding groove of the MHC.
They play no role in antigen recognition but confer much of the affinity between the peptide and MHC complex.
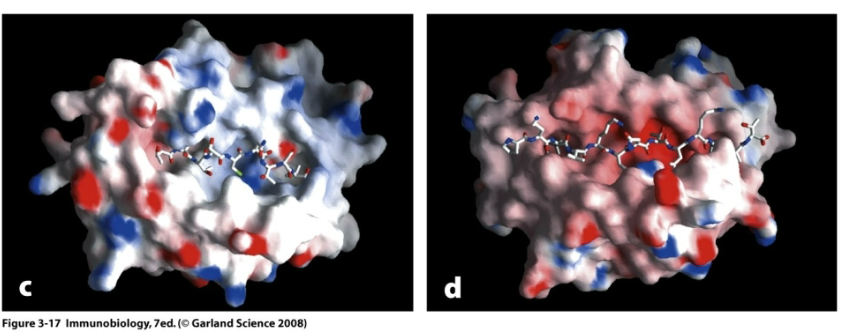
How do peptides bind MHC Class I?
Peptides bind helices of the binding groove.
Closed ends of the groove restricts the length of the peptide. The residues exposed to the solvent mediate specificity of binding to TCR.
8-10 residues
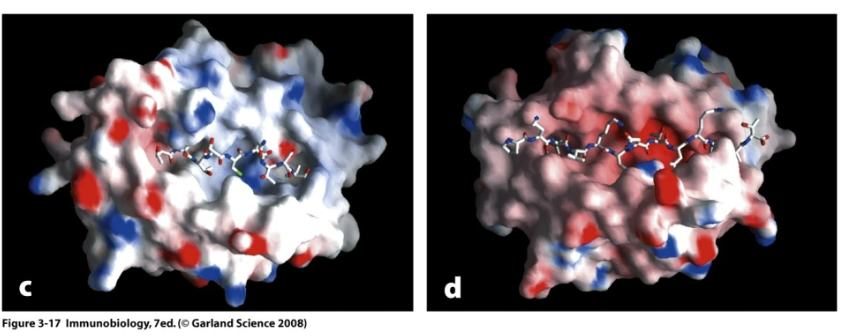
How do peptides bind MHC Class II?
Peptides bind helices of the binding groove.
Open ends of the groove allows a flexible length of peptides. The residues exposed to the solvent mediate specificity of binding to TCR.
10-13 residues
What is meant by a "variable footprint" of the TCR on the MHC surface?
TCRs can engage pMHC complexes using different binding angles and contact patterns.
Structural studies show a variable binding mode and often poor shape complementarity, indicating that TCR-pMHC recognition is not rigidly uniform.
How do TCR CDR loops behave upon binding pMHC?
TCR binding to pMHC often induces conformational rearrangements in the CDR loops, particularly CDR3, allowing the TCR to adapt to the presented peptide.
This structural flexibility contributes to cross-reactivity and specificity.
How can TCRs achieve specificity despite weak binding affinity and limited contact surface?
The TCR-pMHC interaction surface is small, limiting peptide contact.
Specificity is maintained through precise contact by CDR3 and CDR loop adaptability, even when affinity is low (Kd ~1–90 μM).