Unit 8
1/40
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
41 Terms
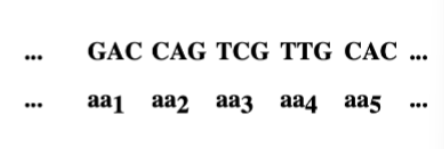
The coding ratio. Given that there are four
different bases, what is the minimum number of
nucleotides per amino acid required to specify all
20 amino acids ?
minimum = 3 nucleotides per amino acids = triplet code
Define "codon"
A sequence of three nucleotides in mRNA that specifies a particular amino acid or a stop signal during protein synthesis.
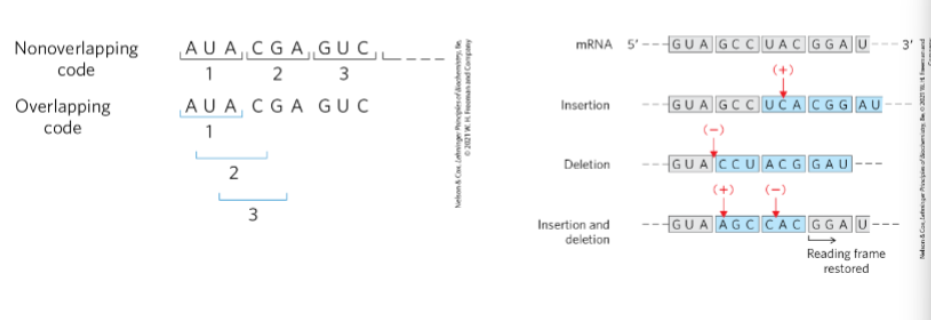
Is the code overlapping or non-overlapping? Use Figs. 27-3 and 27-4 (p. 1007) to
distinguish between each.
Non-overlapping code
Each nucleotide is part of only one codon.
Codons are read sequentially in groups of three.
Overlapping code (not how real genetic code works)
A nucleotide would be shared between multiple codons.
Example:
AUA
UAC
ACG
A single base change would affect several amino acids.
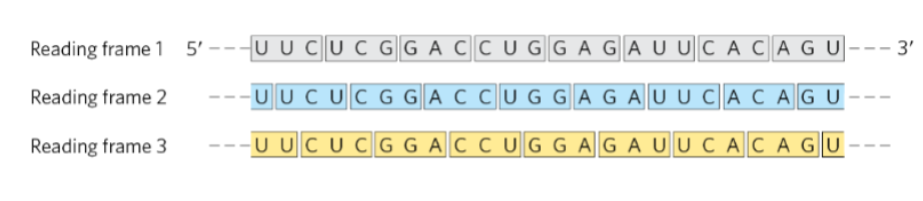
define a reading frame
The way mRNA nucleotides are grouped into codons (sets of three bases) during translation.
mRNA is read 5′ → 3′.
There are three possible reading frames depending on where translation begins.
Each frame produces different codon groupings and therefore different amino acid sequences.
The correct reading frame is set by the start codon (AUG).
Are individual codewords delimited, as for example words in a sentence are
delimited by spaces
No. Individual codewords are not delimited.
The genetic code is comma-less (no spaces or punctuation between codons).
mRNA is read continuously in groups of three nucleotides.
Codons are determined only by the reading frame set by the start codon (AUG).
What establishes the reading frame in RNA (pp. 1007 - 1008)?
The start codon (AUG) on mRNA sets the reading frame.
AUG codes for methionine (Met).
The ribosome begins translation at AUG and then reads nucleotides in triplets (codons) from that point onward.
This determines how all downstream codons are grouped.
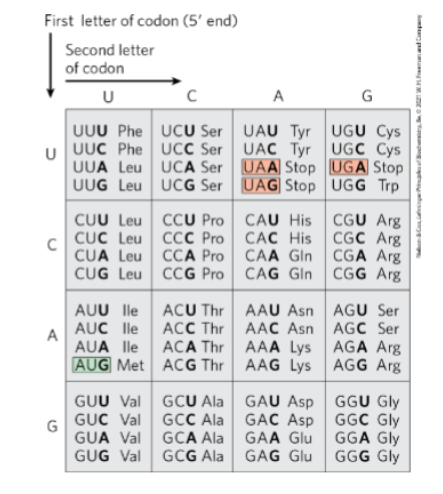
Given a sequence of RNA, an indication of
the correct reading frame, and the genetic
code (Fig. 27-7, p. 1009), deduce the
relevant polypeptide sequence encoded in
that reading frame
an RNA sequence
the correct reading frame
the genetic code table
Do this:
Read the RNA 5′ → 3′
Separate the sequence into codons (triplets) using the given reading frame
Translate each codon with the genetic code table
Start at AUG if the question is asking for the translated protein
Stop when you reach a stop codon: UAA, UAG, or UGA
Important notes
AUG = Met and usually starts translation
Stop codons do not code for an amino acid
A different reading frame gives a different amino acid sequence
What are the expected consequences of a one or two base addition or deletion within the
above sequence
A one- or two-base insertion or deletion causes a frameshift mutation.
Consequences:
The reading frame shifts, changing how codons are grouped.
All codons downstream of the mutation change.
This usually produces a completely different amino acid sequence.
Often results in a premature stop codon, producing a short, nonfunctional protein.
Why are such additions or deletions (when they occur in the DNA) called frameshift
mutations?
They are called frameshift mutations because the addition or deletion of nucleotides shifts the reading frame of the codons.
Explanation:
Codons are read in groups of three nucleotides.
If 1 or 2 bases are added or deleted, the grouping of all downstream nucleotides changes.
This shifts the reading frame, so different codons are read, producing a different amino acid sequence and often a premature stop codon.
initiation codon , temrination codon , open reading frame
1) Initiation codon
The start codon (AUG) that signals the beginning of translation.
Codes for methionine (Met) and establishes the reading frame.
2) Termination codon
A stop codon that signals the end of translation.
UAA, UAG, UGA.
These do not code for an amino acid.
3) Open reading frame (ORF)
A continuous sequence of codons beginning with a start codon (AUG) and ending with a stop codon.
Represents the region of mRNA that can be translated into a polypeptide.
![<p>What does degeneracy refer to in the context of coding (Table 27-3 [p. 1009], and</p><p>p. 1009)?</p><p></p>](https://assets.knowt.com/user-attachments/6841fd3d-4a7e-4f47-b9f5-4f704c34cfb6.png)
What does degeneracy refer to in the context of coding (Table 27-3 [p. 1009], and
p. 1009)?
Degeneracy means that more than one codon can specify the same amino acid.
Although there are 64 possible codons, they encode only 20 amino acids, so several codons correspond to the same amino acid.
These codons are called synonymous codons.
Example:
Leucine (Leu) → 6 codons
Arginine (Arg) → 6 codons
Glycine (Gly) → 4 codons
Key point for exams:
Degeneracy usually occurs because the third base of the codon can vary (wobble position) without changing the amino acid.
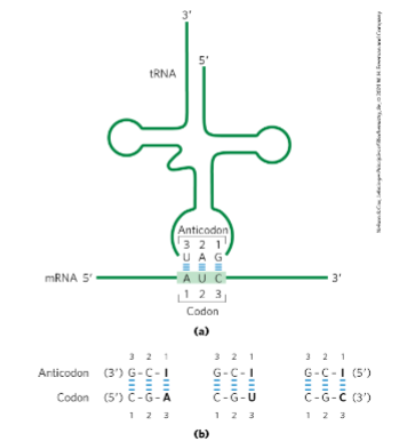
Use Fig. 27-8 (p. 1012) to describe how aminoacyl tRNAs interact with mRNA.
Point out the anticodon on the tRNA and the codon on the mRNA.
How aminoacyl-tRNAs interact with mRNA:
An aminoacyl-tRNA carries a specific amino acid attached to its 3′ end.
The anticodon on the tRNA (a sequence of 3 nucleotides) base-pairs with a complementary codon on the mRNA.
The codon is the 3-nucleotide sequence on the mRNA read 5′ → 3′.
The anticodon on tRNA binds antiparallel and complementary to the codon.
Example (from figure):
mRNA codon: AUC (5′ → 3′)
tRNA anticodon: UAG (3′ → 5′)
Result:
Correct tRNA binds the codon.
The amino acid carried by that tRNA is added to the growing polypeptide chain.
Is the genetic code almost universal
Yes, the genetic code is almost universal.
Nearly all organisms use the same codons to specify the same amino acids.
This means a gene from one organism can often be correctly translated in another organism.
Exceptions:
Some mitochondria
Certain protozoa and microorganisms
In these cases, a few codons are assigned different amino acids or stop signals.
The genetic code has been conserved through evolutionary time because it is resistant to the
deleterious effects of mutations. Discuss, and give an example of, how the genetic code protects
against missense mutations at both the 5' and 3' nucleotide positions in the codon (pp. 1012 - 1013).
The genetic code protects against missense mutations because similar codons often encode the same or chemically similar amino acids.
3′ position of the codon (third base)
The third base is highly degenerate (wobble position).
A mutation here often does not change the amino acid → silent mutation.
Example:
UUU → UUC
Both code for Phe (phenylalanine).
A mutation at the 3′ base does not change the amino acid.
5′ position of the codon (first base)
A change at the first base may change the amino acid, but often to one with similar chemical properties, reducing functional damage.
Example:
GAU → Asp (aspartate)
AAU → Asn (asparagine)
Both are polar amino acids with similar properties, so the protein function may be less affected.
Key idea
Degeneracy and chemical similarity of encoded amino acids help minimize harmful effects of mutations, which is why the genetic code has been conserved through evolution.

Use Table 27-4 (p. 1012) to discuss the meaning of "wobble" (pp. 1010 - 1012)? Do not confuse
degeneracy with wobble. Degeneracy is a feature of the genetic code (some amino acids are encoded
for by more than one codon), while wobble is a feature of a tRNA (some tRNAs can bind to more than
one codon).
Use Table 27-4 (p. 1012) to discuss the meaning of "wobble" (pp. 1010 - 1012)? Do not confuse
degeneracy with wobble. Degeneracy is a feature of the genetic code (some amino acids are encoded
for by more than one codon), while wobble is a feature of a tRNA (some tRNAs can bind to more than
one codon).
Wobble refers to flexible base pairing between the 3′ base of the mRNA codon and the 5′ base of the tRNA anticodon.
Because of this flexibility, one tRNA can recognize more than one codon.
Key points
The first two bases of the codon pair normally (Watson–Crick pairing).
The third base of the codon can pair less strictly, allowing alternative base pairing.
This reduces the number of different tRNAs needed.
Degeneracy: multiple codons encode the same amino acid (property of the genetic code).
Wobble: a single tRNA can bind multiple codons (property of tRNA).
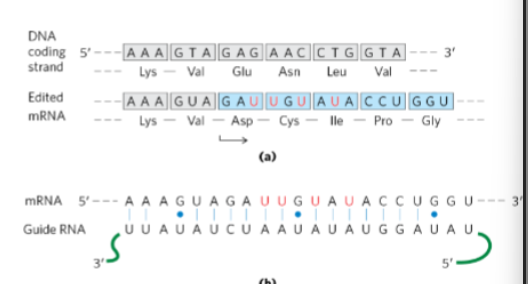
Use Fig. 27-10 (p. 1014) to show how guide RNAs can be used as templates to direct the
addition or removal of specific nucleotides from an mRNA transcript.
Guide RNAs (gRNAs) base-pair with a pre-mRNA transcript.
The gRNA acts as a template showing where nucleotides must be inserted or removed.
Process
gRNA binds to the mRNA by complementary base pairing.
Mismatches between gRNA and mRNA identify editing sites.
Enzymes then:
Insert nucleotides (commonly U residues) or
Delete nucleotides from the mRNA.
The edited mRNA now matches the guide RNA template.
Result
The codon sequence changes, which can change the amino acid sequence of the protein produced.
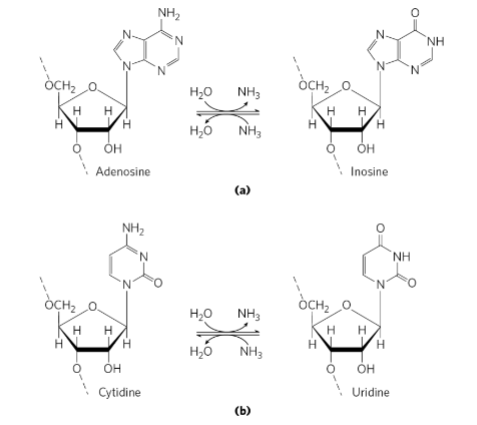
Use Fig. 27-11 (p. 1014) to show how deaminases can alter specific nucleotides in an mRNA
transcript. Use Fig. 27-12 (p. 1015) to show how RNA editing by this mechanism is used to
regulate the production of two related but different products from a single transcript in a tissue-
specific manner.
1. How deaminases alter nucleotides in mRNA
Deaminase enzymes chemically modify bases by removing an amino group, changing the nucleotide identity.
Examples:
Adenosine → Inosine (A → I)
Inosine is read by the ribosome as G.
Cytidine → Uridine (C → U)
This changes the codon sequence, which can change the amino acid specified during translation.
2. Example: tissue-specific RNA editing (ApoB protein)
RNA editing can produce different proteins from the same mRNA transcript.
Human liver:
mRNA codon = CAA (Gln)
Translation continues → produces ApoB-100 (full-length protein).
Human intestine:
C → U editing converts CAA → UAA
UAA is a stop codon
Translation stops early → produces ApoB-48 (shorter protein).
✅ Key idea for exam:
RNA editing by deaminases can change codons (including creating stop codons), allowing one gene to produce different proteins in different tissues.
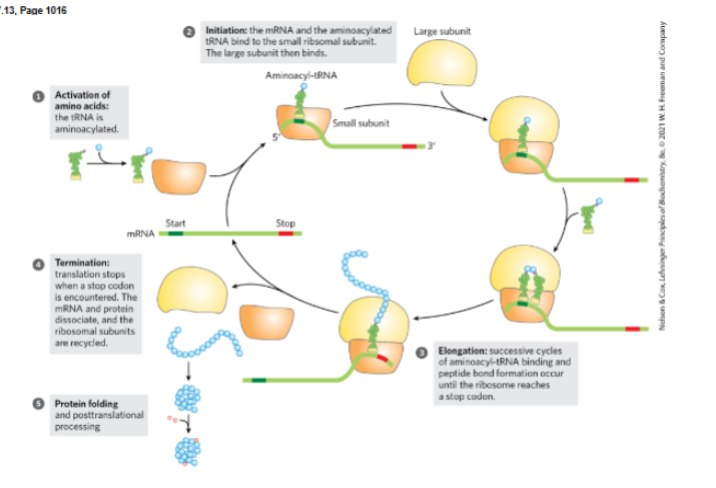
activation of amino acids, initiation of protein synthesis, elongation , termination and ribosome recylcing , folding and post translational processing.
a. Activation of amino acids
Each amino acid is attached to its specific tRNA by aminoacyl-tRNA synthetase.
Uses ATP to form an aminoacyl-tRNA (charged tRNA).
This prepares the amino acid for incorporation into the growing polypeptide.
b. Initiation of protein synthesis
Small ribosomal subunit binds mRNA near the start codon (AUG).
Initiator tRNA carrying Met binds the start codon via its anticodon.
Large ribosomal subunit joins, forming the complete ribosome.
c. Elongation
Aminoacyl-tRNA enters the A site of the ribosome matching the codon.
Peptide bond forms between the growing chain and the new amino acid.
Ribosome translocates along mRNA (5′ → 3′) and the cycle repeats.
d. Termination and ribosome recycling
A stop codon (UAA, UAG, UGA) enters the A site.
Release factors bind, causing the polypeptide to be released.
Ribosomal subunits, mRNA, and tRNA dissociate and are recycled.
e. Folding and post-translational processing
Newly made polypeptide folds into its functional 3-D structure (often with chaperone proteins).
May undergo post-translational modifications such as:
Cleavage
Phosphorylation
Glycosylation
Disulfide bond formation.
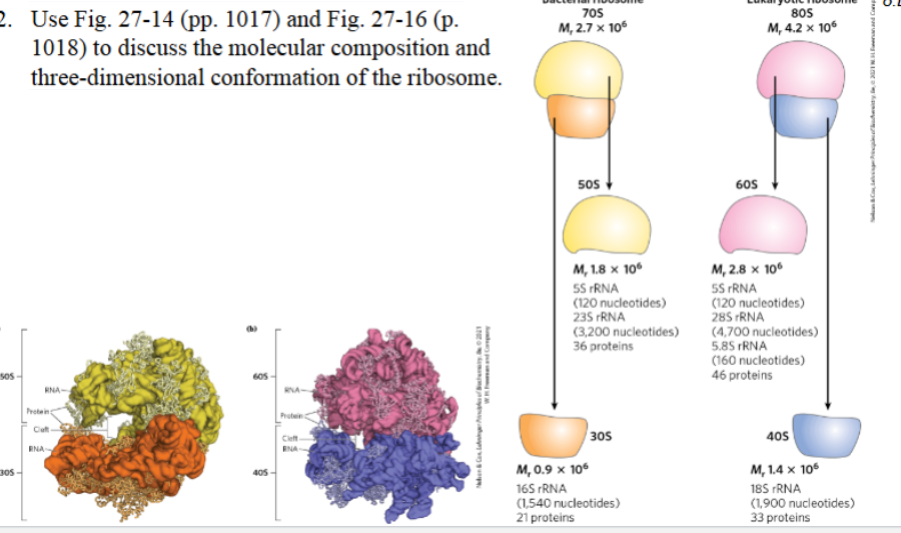
Use Fig. 27-14 (pp. 1017) and Fig. 27-16 (p.
1018) to discuss the molecular composition and
three-dimensional conformation of the ribosome.
Bacterial ribosome (70S):
Large subunit (50S):
rRNAs: 23S rRNA, 5S rRNA
~36 proteins
Small subunit (30S):
rRNA: 16S rRNA
~21 proteins
Eukaryotic ribosome (80S):
Large subunit (60S):
rRNAs: 28S rRNA, 5.8S rRNA, 5S rRNA
~46 proteins
Small subunit (40S):
rRNA: 18S rRNA
~33 proteins
Three-dimensional conformation of the ribosome
The ribosome has two subunits that fit together to form a cleft where mRNA binds.
tRNAs bind in three sites:
A site (aminoacyl site) – incoming aminoacyl-tRNA
P site (peptidyl site) – tRNA holding the growing polypeptide
E site (exit site) – tRNA leaving the ribosome
The large subunit contains the peptidyl transferase center that forms peptide bonds.
The small subunit binds and decodes the mRNA.
Use Fig. 27-15 (p. 1018) to illustrate that RNA (in addition to proteins) contributes
to this three-dimensional structure of the ribosome.
RNA contributes directly to the three-dimensional structure of the ribosome.
rRNA forms the structural core of the ribosome, creating the main framework of the large and small subunits.
Ribosomal proteins are mostly located on the outer surface, stabilizing the rRNA structure rather than forming the core.
The complex folding of rRNA into helices and loops generates the ribosome’s 3-D architecture.
rRNA also forms the peptidyl transferase center, which catalyzes peptide bond formation.
Key idea:
The ribosome is a ribozyme, meaning rRNA (not protein) provides both much of the structure and the catalytic activity of the ribosome.
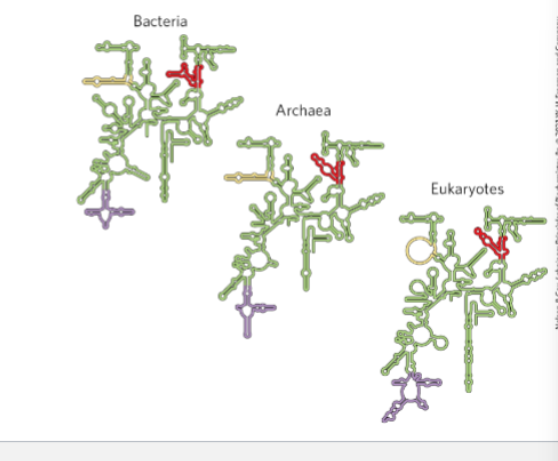
Discuss the ribosome in the context of the RNA world hypothesis
Ribosome and the RNA world hypothesis
The RNA world hypothesis proposes that early life relied primarily on RNA for both genetic information storage and catalytic activity before DNA and proteins evolved.
Evidence from the ribosome:
The core structure of the ribosome is made mainly of rRNA, not protein.
The peptidyl transferase center (where peptide bonds form) is catalyzed by rRNA, making the ribosome a ribozyme.
Ribosomal proteins mainly stabilize the rRNA structure and are located on the outer surface.
Implication:
Because the ribosome’s essential catalytic activity is carried out by RNA, it is considered a molecular relic of the RNA world, supporting the idea that RNA-based catalysts existed before protein enzymes evolved.
Why are tRNAs referred to as "adaptors"
tRNAs are called “adaptors” because they link the genetic code in mRNA to the correct amino acid during protein synthesis.
Each tRNA has an anticodon that base-pairs with a specific codon on mRNA.
The other end of the tRNA carries the corresponding amino acid.
This allows the codon sequence in mRNA to be translated into the amino acid sequence of a protein.
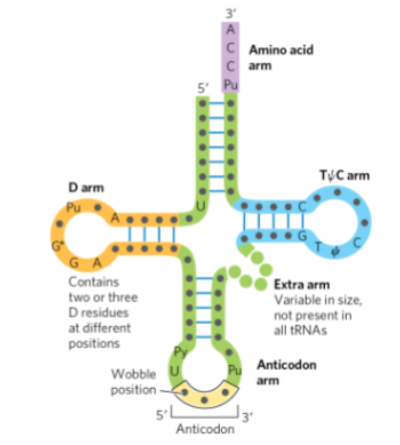
the anticodon arm (or loop)
b. the TC arm
c. the dihydrouracil arm (D arm)
d. the 3' (CCA) and 5' ends of the tRNA molecule
e. the position at which the amino acid is attached
f. regions that are hydrogen bonded
a. Anticodon arm (loop)
The bottom loop of the tRNA.
Contains the anticodon (3 bases) that base-pairs with the codon on mRNA during translation.
b. TΨC arm
The right arm of the tRNA containing the sequence T–Ψ–C (ribothymidine–pseudouridine–cytidine).
Important for interaction with the ribosome.
c. D arm (dihydrouracil arm)
The left arm of the tRNA containing dihydrouridine residues (D).
Helps with recognition by aminoacyl-tRNA synthetase.
d. 3′ (CCA) and 5′ ends of the tRNA
3′ end: contains the CCA sequence, located at the top of the molecule.
5′ end: opposite side of the acceptor stem.
e. Position where the amino acid is attached
The amino acid attaches to the 3′ hydroxyl of the terminal A in the CCA sequence at the 3′ end (acceptor stem).
f. Regions that are hydrogen bonded
The stem regions of the cloverleaf structure (acceptor stem, D stem, anticodon stem, and TΨC stem).
These regions form double-stranded sections stabilized by hydrogen bonding between complementary bases.
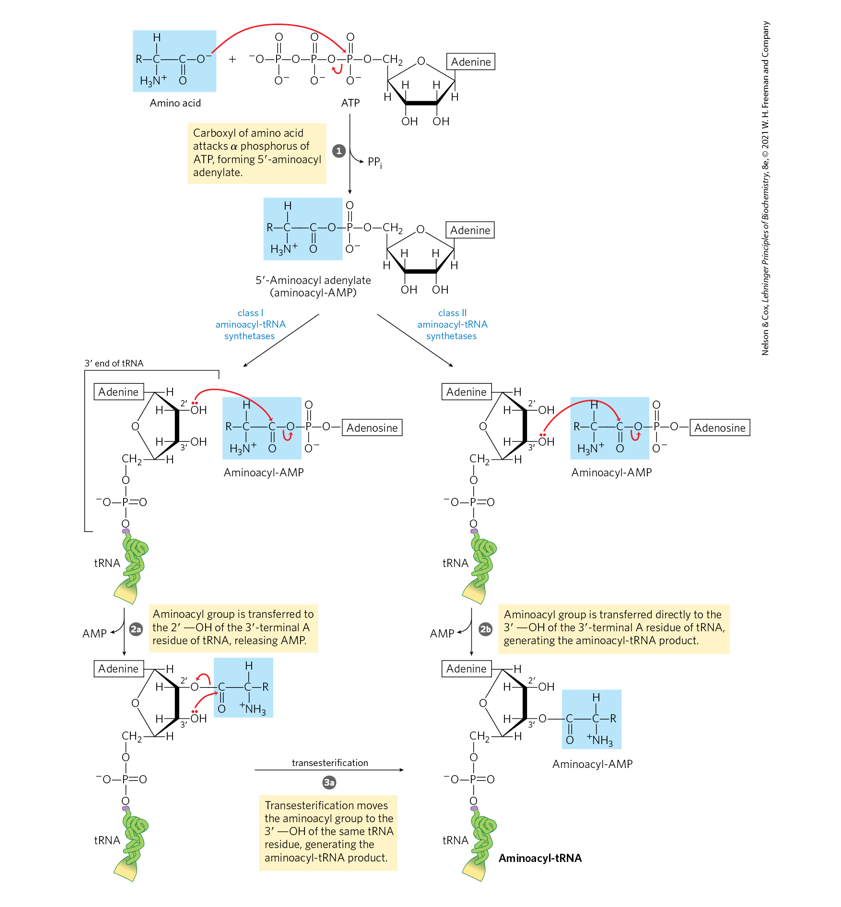
Amino acid activation (pp. 1020 - 1021).
Using structures, describe in two steps how
the activation of an amino acid for protein
synthesis takes place (Fig. 27-19, p. 1021).
Show the structures of the reactants,
intermediates, and the products
Step 1: Formation of aminoacyl-AMP (activation)
Reaction:
Amino acid+ATP→Aminoacyl-AMP+PP
The carboxyl group of the amino acid forms an anhydride bond with AMP.
Pyrophosphate (PPᵢ) is released.
Step 2: Transfer of the amino acid to tRNA
Reaction:Aminoacyl-AMP+tRNA→Aminoacyl-tRNA+AMP
The amino acid is transferred to the 3′ hydroxyl group of the terminal adenosine of the CCA end of tRNA.
Final products
Aminoacyl-tRNA (charged tRNA)
AMP
PPᵢ
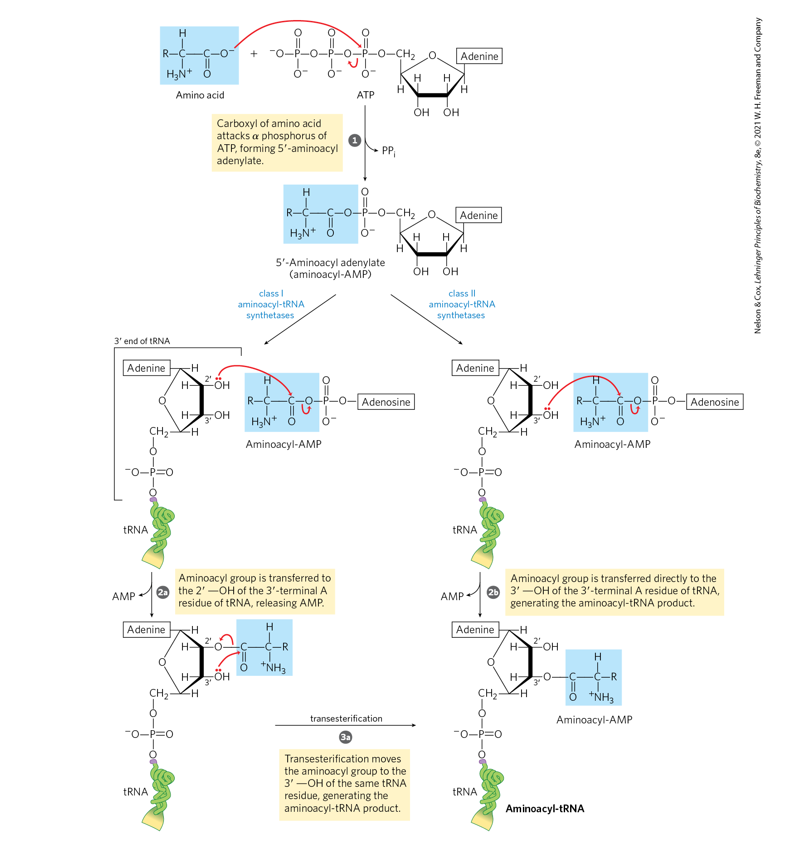
Name the class of enzyme that loads amino acids
onto tRNAs.
b. Name the linkage that is broken in ATP (refer to Fig.
1-14, p. 12)?
c. Name the linkage that is formed in the activated
intermediate which is then broken in the final stage
of the reaction (refer to Fig. 1-14, p. 12)?
d. Name the linkage that is ultimately formed between the
amino acid and the tRNA (refer to Fig. 1-14, p. 12)?
e. What is the other product of this series of reactions?
What happens to this product (p. 1020)?
a. Class of enzyme that loads amino acids onto tRNAs
Aminoacyl-tRNA synthetases.
b. Linkage broken in ATP
The phosphoanhydride bond between the α and β phosphates of ATP is broken when ATP → AMP + PPᵢ.
c. Linkage formed in the activated intermediate (then broken later)
A mixed anhydride bond between the amino acid carboxyl group and AMP in aminoacyl-AMP (aminoacyl adenylate).
d. Linkage ultimately formed between the amino acid and the tRNA
An ester linkage between the carboxyl group of the amino acid and the 3′-OH of the terminal adenosine (CCA) of tRNA.
e. Other product of the reaction and what happens to it
Pyrophosphate (PPᵢ) is produced.
It is rapidly hydrolyzed to 2 inorganic phosphates (2 Pᵢ) by pyrophosphatase, which drives the reaction forward (makes it essentially irreversible).
Write a balanced word equation for the sum of the activation and transfer steps, including hydrolysis of PPi (p. 1020).
g. There is a cost of two high energy anhydride bonds
in the synthesis of an aminoacyl-tRNA? Account
for this cost (pp. 1020).
h. Name two other synthetic processes you have
studied in which pyrophosphate is generated.
What is the significance of the fact that all cells
possess pyrophosphatases that catalyze the
hydrolysis reaction, PPi + H2O 2Pi?
f. Balanced word equation (including PPᵢ hydrolysis)
Amino acid + tRNA + ATP + H₂O → aminoacyl-tRNA + AMP + 2 Pi
g. Why two high-energy bonds are used
ATP is cleaved to AMP + PPᵢ, breaking one phosphoanhydride bond.
The pyrophosphate (PPᵢ) is then hydrolyzed to 2 Pi, breaking another high-energy phosphoanhydride bond.
Thus the reaction consumes the equivalent of two high-energy bonds.
h. Two other synthetic processes that generate PPᵢ
Examples:
DNA synthesis (polymerization of nucleotides)
RNA synthesis (transcription)
Activation of fatty acids (formation of fatty acyl-CoA)
Significance of pyrophosphatase:
PPᵢ + H₂O → 2 Pi rapidly occurs in cells.
This drives biosynthetic reactions forward and makes them essentially irreversible by removing PPᵢ.
Discuss how Ile-tRNA synthetase distinguishes between Ile and Val at high fidelity
(p. 1020 - 1022).
How Ile-tRNA synthetase distinguishes Ile from Val (high fidelity)
Isoleucine (Ile) and valine (Val) are very similar in structure (Val is slightly smaller), so the enzyme must ensure correct amino acid selection.
1. Initial selection (active site discrimination)
Ile-tRNA synthetase active site is shaped to preferentially bind isoleucine.
Because valine is slightly smaller, it can sometimes still fit into the active site and be activated.
2. Proofreading/editing mechanism
The enzyme has a second editing site that acts as a quality-control pocket.
Two possible corrections occur:
a. Pre-transfer editing
If Val-AMP (valine attached to AMP) forms, the enzyme hydrolyzes it before transfer to tRNA.
b. Post-transfer editing
If valine is mistakenly attached to tRNA (Val-tRNAᶦˡᵉ), the enzyme moves the tRNA to the editing site.
The incorrect Val is hydrolyzed off the tRNA.
Double-sieve model
First sieve (activation site): excludes larger amino acids than Ile.
Second sieve (editing site): removes smaller amino acids like Val.
Result
This double-sieve proofreading system ensures very high accuracy in attaching Ile to tRNAᶦˡᵉ and prevents incorporation of Val during protein synthesis.
What is the "second genetic code" (Fig. 27-21 [p. 1023] and p. 1022)
The “second genetic code” refers to the rules that determine how the correct amino acid is matched with its specific tRNA by aminoacyl-tRNA synthetases.
Key points:
Aminoacyl-tRNA synthetases recognize specific structural features of each tRNA.
These features include:
Anticodon sequence
Bases in the acceptor stem
Other identity elements in the tRNA structure
These recognition signals ensure the correct amino acid is attached to the correct tRNA.
Importance:
This system guarantees that the codons in mRNA are translated into the correct amino acids during protein synthesis.
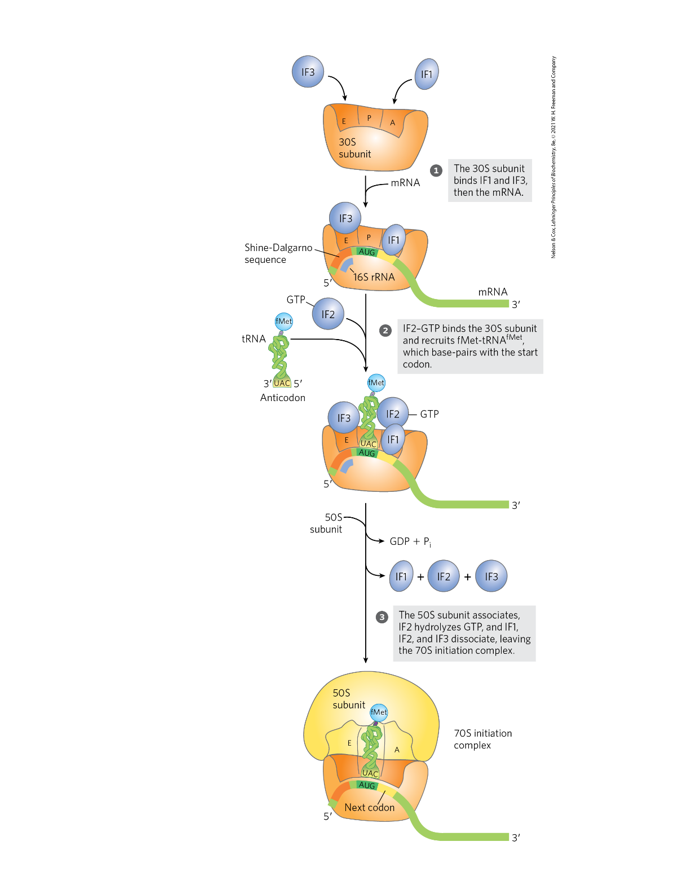
Use Fig. 27-24 (p. 1028) to discuss the key events in translation
initiation.
What is the Shine-Dalgarno sequence (p. 1028)? Use Fig. 27-25 (p. 1029) to
explain its significance in prokaryotic translation initiation.
Shine–Dalgarno sequence:
A short purine-rich sequence in prokaryotic mRNA located ~5–10 nucleotides upstream of the start codon (AUG).
Consensus sequence: AGGAGG.
Significance in translation initiation
The Shine–Dalgarno sequence base-pairs with a complementary sequence in the 16S rRNA of the 30S ribosomal subunit.
This aligns the ribosome correctly on the mRNA so that the start codon (AUG) is positioned in the P site.
Proper positioning allows the initiator tRNA (fMet-tRNAᶠᴹᵉᵗ) to bind the start codon and begin translation.
Key idea:
The Shine–Dalgarno sequence recruits and positions the ribosome on prokaryotic mRNA for accurate initiation of protein synthesis.
Account for the cost of one high energy bond used when IF-2 delivers
an aminoacyl tRNA to the P site during translation initiation (p. 1028).
IF-2 binds GTP and forms a complex with the initiator tRNA (fMet-tRNAᶠᴹᵉᵗ) and the 30S ribosomal subunit.
IF-2 delivers the initiator aminoacyl-tRNA to the P site where it pairs with the start codon (AUG).
Energy cost:
After correct assembly and binding of the 50S subunit, IF-2 hydrolyzes GTP → GDP + Pi.
This GTP hydrolysis provides the energy required for:
Release of initiation factors
Proper assembly of the 70S initiation complex
✅ Thus, one high-energy phosphoanhydride bond from GTP is consumed during delivery of the initiator tRNA to the P site during translation initiation.
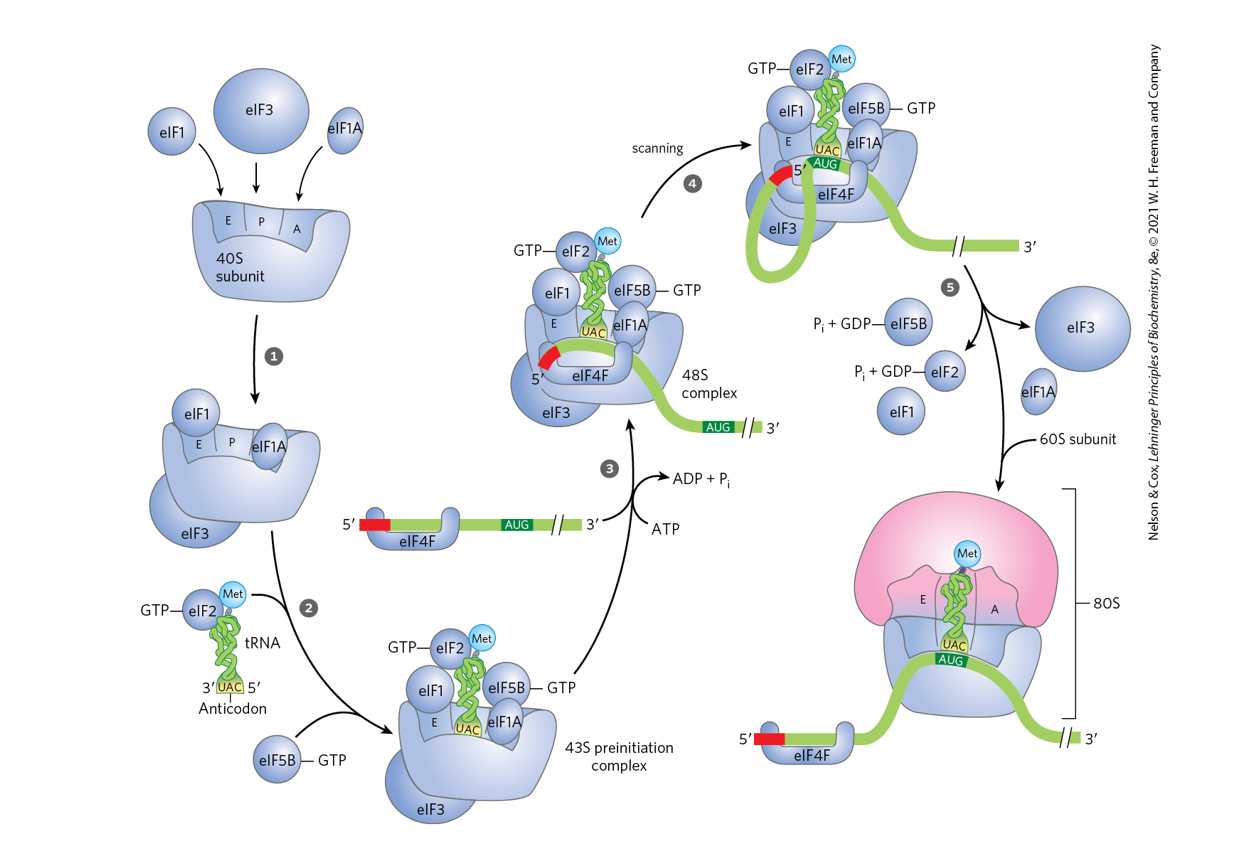
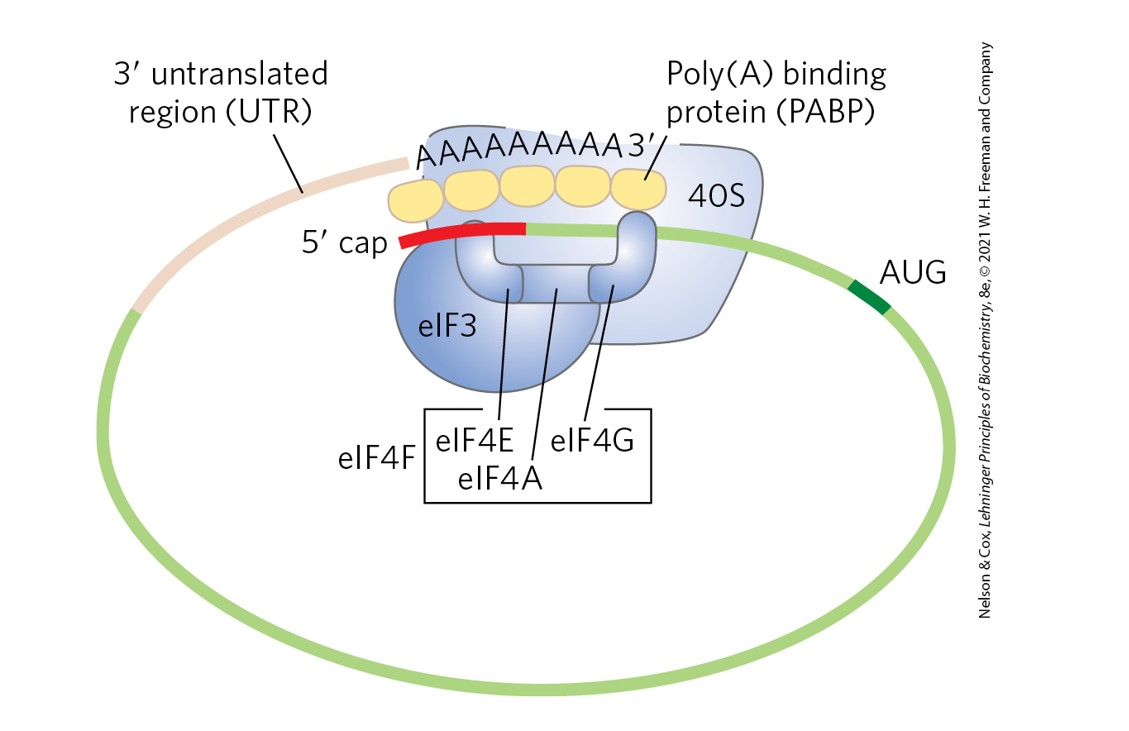
Compare the initiation of translation in prokaryotes and eukaryotes (pp. 1029 -
1030, and Figs. 27-26 and 27-27, p. 1030).
Prokaryotes: ribosome is positioned by Shine–Dalgarno pairing.
Eukaryotes: ribosome binds the 5′ cap and scans to the start codon with help of many initiation factors.
Feature | Prokaryotes | Eukaryotes |
|---|---|---|
Ribosome | 70S (30S + 50S) | 80S (40S + 60S) |
Initiator tRNA | fMet-tRNAᶠᴹᵉᵗ (N-formylmethionine) | Met-tRNAᵢᴹᵉᵗ (methionine) |
mRNA recognition | Ribosome binds directly to Shine–Dalgarno sequence upstream of AUG | 40S subunit binds 5′ cap and scans mRNA to find AUG |
Positioning of start codon | Shine–Dalgarno sequence base-pairs with 16S rRNA to position AUG in P site | Scanning mechanism moves along mRNA until first suitable AUG |
Initiation factors | IF1, IF2, IF3 | Many eIFs (e.g., eIF1, eIF2, eIF3, eIF4E, eIF4G, eIF4A) |
mRNA structure involved | Shine–Dalgarno region | 5′ cap and poly(A) tail interact through proteins |
mRNA organization | Often polycistronic (multiple proteins from one mRNA) | Usually monocistronic (one protein per mRNA) |
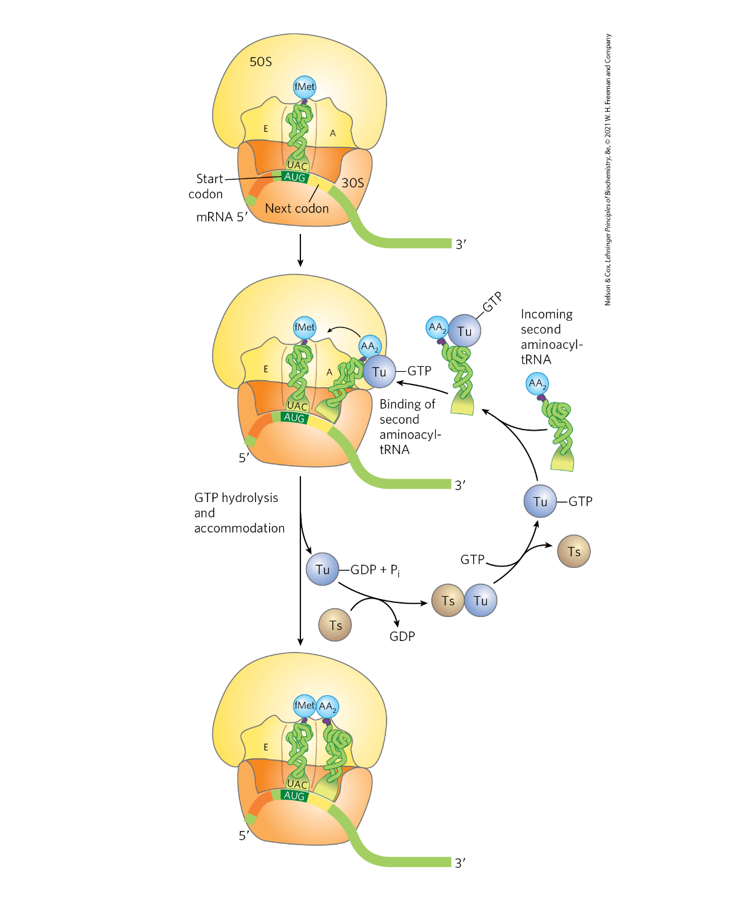
Use Fig. 27-28 (p. 1031) to discuss the role of EF-Tu in
delivering AA-tRNAs to the A site during elongation.
Account for the one high energy bond used by EF-Tu.
What is the role of EF-Ts?
Role of EF-Tu in elongation:
EF-Tu bound to GTP forms a complex with an aminoacyl-tRNA (aa-tRNA).
This complex delivers the aa-tRNA to the A site of the ribosome.
If the codon–anticodon pairing is correct, GTP is hydrolyzed → GDP + Pi.
EF-Tu then releases the tRNA, allowing it to fully enter the A site.
Energy use:
One high-energy phosphoanhydride bond is used when EF-Tu hydrolyzes GTP → GDP + Pi.
Role of EF-Ts:
EF-Ts regenerates EF-Tu-GTP by exchanging GDP for GTP, allowing EF-Tu to participate in another cycle of aa-tRNA delivery.
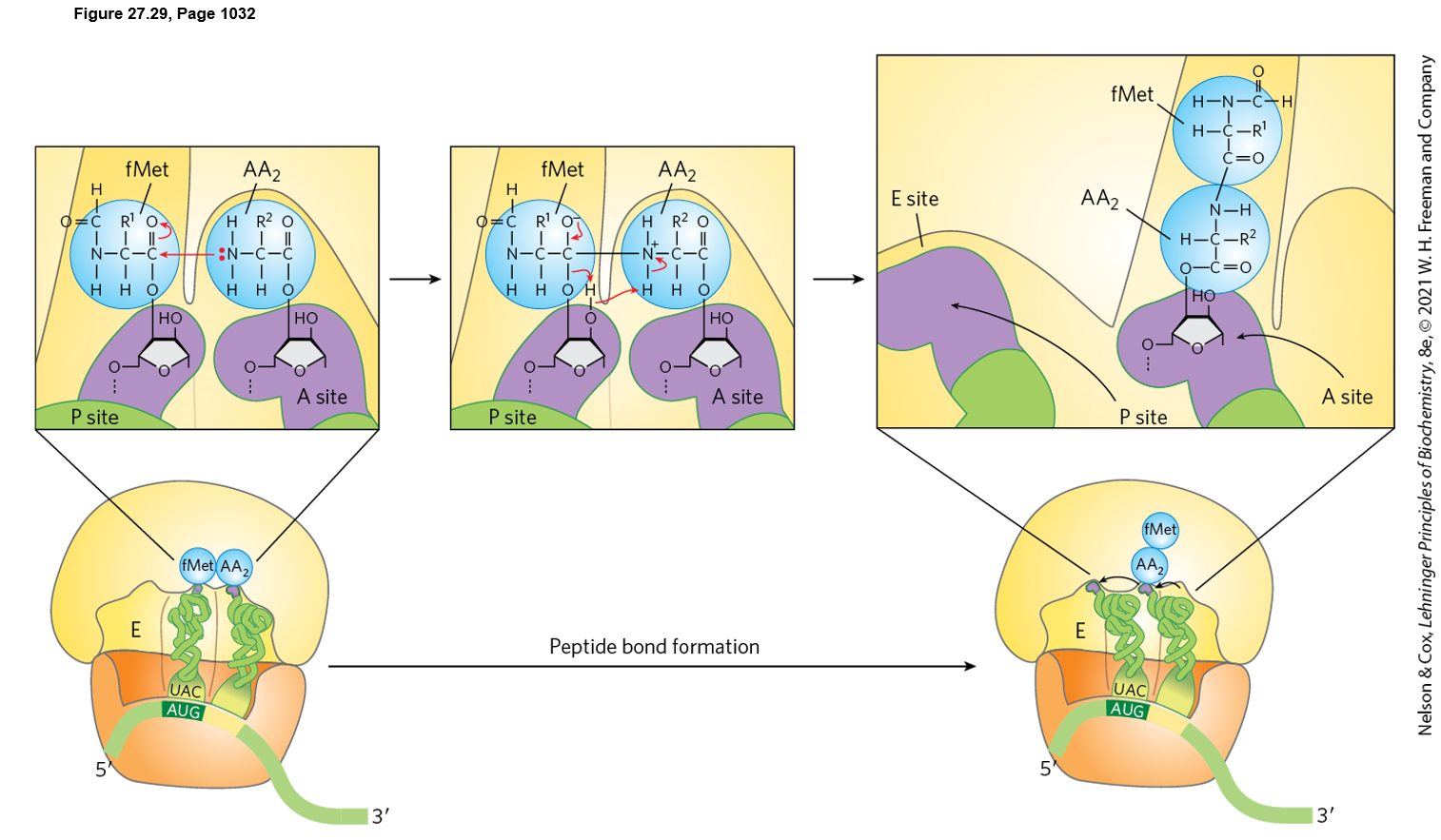
Name the enzyme and point out the
nucleophilic attack mechanism catalyzed by
this enzyme. Is this reaction catalyzed by a
protein enzyme or a ribozyme?
2) Does protein synthesis run from N to C
terminal end -or- C to N?
3) Note the position of the peptidyl tRNA after
peptide bond formation.
4) Note: No additional input of energy is
required for peptide bond formation because
the required energy is released when the
peptidyl-tRNA ester bond is broken.
1. Enzyme and nucleophilic attack mechanism
The enzyme activity is peptidyl transferase, located in the large ribosomal subunit (23S rRNA in bacteria).
The reaction is catalyzed by rRNA, so the ribosome functions as a ribozyme (not a protein enzyme).
Mechanism:
The α-amino group of the aminoacyl-tRNA in the A site acts as the nucleophile.
It attacks the carbonyl carbon of the ester bond linking the growing polypeptide to the tRNA in the P site.
This forms a peptide bond and transfers the growing peptide to the A-site tRNA.
2. Direction of protein synthesis
Protein synthesis proceeds from the N-terminus to the C-terminus (N → C).
3. Position of the peptidyl-tRNA after peptide bond formation
After the peptide bond forms, the growing polypeptide is attached to the tRNA in the A site (peptidyl-tRNA now in the A site).
4. Energy requirement
No additional energy input is required for peptide bond formation.
The energy comes from the high-energy ester bond of the peptidyl-tRNA that is broken during the reaction.
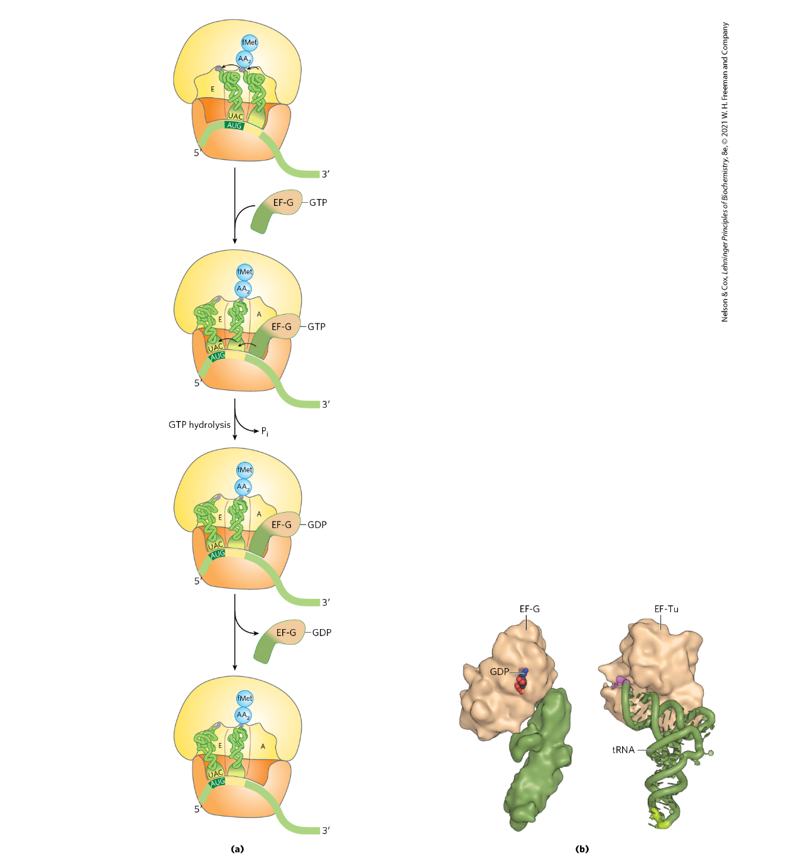
Use Fig. 27-30 (p. 1034) to discuss translocation.
Account for the one high energy anhydride bond
that is used to translocate the peptidyl tRNA from
the A site to the P site?
Energy accounting
One high-energy phosphoanhydride bond from GTP is hydrolyzed by EF-G.
This GTP hydrolysis provides the energy for ribosome movement along the mRNA and tRNA repositioning.
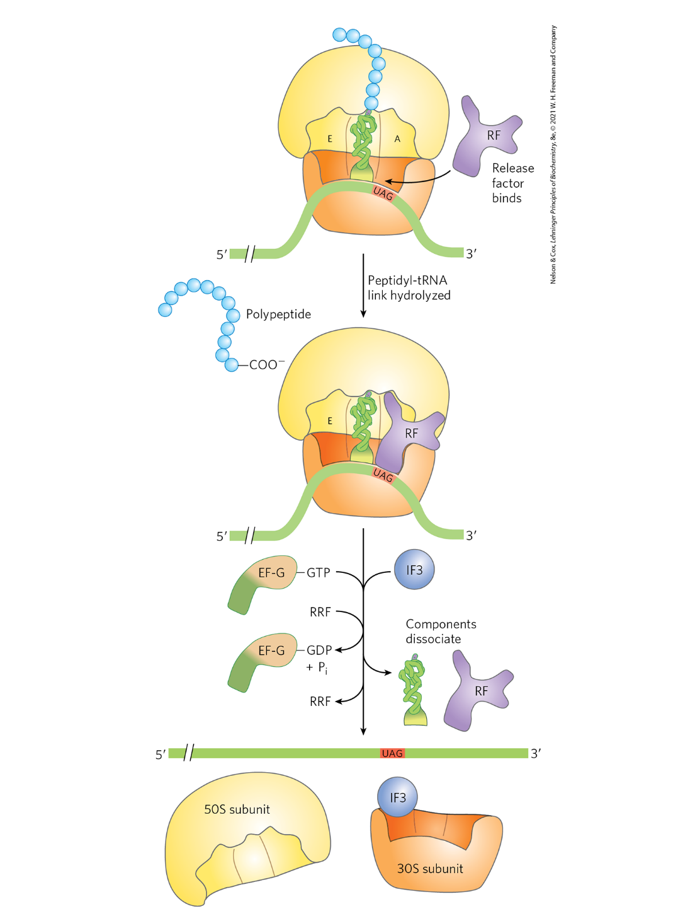
Use Fig. 27-31 (p. 1035) to account for the one high
energy bond used in termination.
Folding and processing: Describe some ways that newly synthesized
polypeptides are processed following their synthesis (pp. 1036 - 1039).
1. Protein folding
The polypeptide folds into its correct three-dimensional structure.
Chaperone proteins (molecular chaperones) assist folding and prevent aggregation.
2. Proteolytic cleavage
Some proteins are synthesized as inactive precursors (proproteins or zymogens).
Specific peptide segments are removed to produce the active protein.
Example: proinsulin → insulin.
3. Covalent modifications
Proteins may undergo chemical modifications such as:
Phosphorylation (addition of phosphate groups)
Glycosylation (addition of carbohydrate groups)
Acetylation or methylation
Lipid attachment
These modifications can affect activity, stability, localization, or interactions.
4. Disulfide bond formation
Cysteine residues form disulfide bonds that stabilize protein structure.
5. Assembly into multi-subunit complexes
Some proteins associate with other polypeptides to form functional complexes (quaternary structure).
Key idea
These folding and modification steps convert the newly synthesized polypeptide into a stable, properly localized, and biologically active protein.
Consider a bacterial gene for a hypothetical polypeptide comprised of 100 amino
acid residues.
a. Approximately how many high-energy phosphate anhydride bonds are used to make the necessary
mRNA molecule that codes for this protein?
b. How many high-energy phosphate anhydride bonds are used to synthesize the hypothetical polypeptide
molecule from free amino acids, given the preformed mRNA?
c. Assuming that each mRNA molecule is translated 20-50 times, does the cell spend more energy in
transcription or translation?
a.
A 100-amino-acid protein needs about 100 codons + 1 stop codon = 101 codons
So the mRNA coding region is about:
101 × 3 = 303 nucleotides
Each nucleotide added during transcription uses the equivalent of 2 high-energy phosphate bonds (NTP → NMP + PPi, then PPi hydrolysis).
Approximate cost:
303 × 2 = ~606 high-energy bonds
b.
For translation of a 100-aa polypeptide from free amino acids, approximate cost is:
Amino acid activation: 100 × 2 = 200
Delivery of aa-tRNA to A site: 99 × 1 = 99
Translocation: 99 × 1 = 99
Initiation + termination: about 2
Total ≈ 400 high-energy bonds
(Exam shortcut: ~4 high-energy bonds per amino acid, so 100 aa → ~400)
c.
The cell spends more energy in translation.
Why:
One mRNA costs about ~606 high-energy bonds to make.
But one 100-aa protein costs about ~400 high-energy bonds to translate.
If that mRNA is translated 20–50 times, translation costs:
20 × 400 = 8000
50 × 400 = 20,000
So translation >> transcription in total energy cost.
Many of the antibiotics used clinically are inhibitors of protein synthesis. Discuss why
some antibiotics are specifically used in the treatment of bacterial infections while
others are used to treat yeasts and fungi (pp. 1039 - 1041; Note: Mitochondrial and
bacterial ribosomes are similar and both are therefore sensitive to bacterial inhibitors.)
Bacterial antibiotics:
Target 70S ribosomes (30S + 50S) found in bacteria.
Human cells have 80S ribosomes, so these drugs mainly affect bacteria.
Yeast/fungal treatments:
Yeasts and fungi are eukaryotes with 80S ribosomes, so different drugs are used that target fungal-specific features.
Note:
Mitochondrial ribosomes are similar to bacterial ribosomes, so some bacterial antibiotics can affect mitochondria and cause side effects.
What is a "signal sequence" and what is its function
Signal sequence:
A short sequence of amino acids at the N-terminus of a newly synthesized protein.
Function:
Directs the protein to the correct cellular location (e.g., endoplasmic reticulum for secretion or membrane insertion).
Often removed after the protein reaches its destination.