L9- DNA structure + analysis
1/17
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
18 Terms
Who coined the term nuclein
Miescher
discovered nucleic acid (contains C,H,O,N,P) found in nucleus of all cells→ Nuclein
Griffiths and Avery expts
To determine whether protein/DNA contains genetic material
Griffiths transformation expt (mice die S+R, heat killed S+R)
R→S
Avery’s expt→ DNA=transforming component
bc DNA from dead S destroyed w/ DNase→ no transformation
Bacteriophage T2
phages are 50% protein and 50% DNA
Hershey and Chase
Phages labelled w/ 32P-DNA 35S- protein
E.coli infected w/ phages→ radioactivity recovered in host and passed on to phage progeny
32P in pellet
With 35S→ radioactivity in phage ghosts + NOT passed on to progeny
³²P radioactivity was found in the pellet, meaning the labeled DNA entered the bacterial cells.
³⁵S radioactivity stayed mostly in the supernatant, meaning the protein coats remained outside.
How does viral genetic material vary
Depending on virus
Some use DNA/RNA
cricular/linear chroms
small/large segments
single/ds
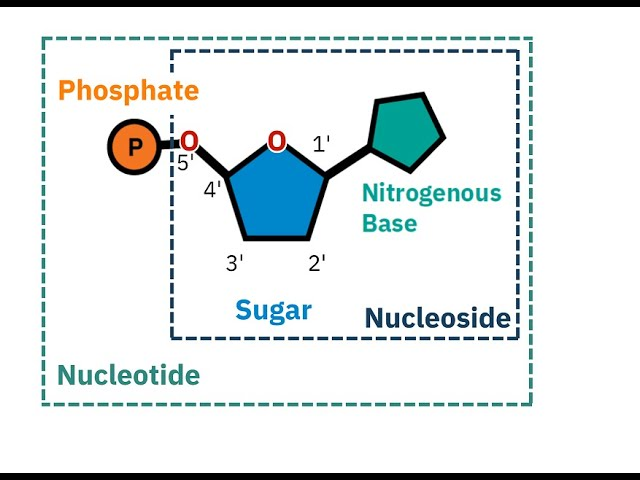
Nucleoside vs nucleotide
nucleoside= sugar + n base NO phosphate
nucleotide= sugar (pentose), N base, P group
Polynucleotide chains have
polarity
Phosphodiester linkages bw
3’ OH and 5’ Phosphate group
(synthesis occurs 5’-3’ but adds on to 3’ end)
Chargaff
50% base= purines, 50% bases= pyrimidines
A=T
C=G
X ray diffraction studies
Franklin + Wilkins
regularities of 0.34 nm and 3.4 nm
DNA is a helical structure
Watson and Crick model of DNA
2 polynucleotide chains wound around each other in a right handed helix
2 chains antiparallel
Sugar-P backbones on outside of double helix
Bases connected by H bonds
Bases 0.34 nm apart, 10 bases= 1 full turn
Grooves of unequal size form bw backbones
double helix diameter= 2 nm
RNA structure
single stranded
will bind to itself when possible (folds on itself)
Many diff forms of RNA in cell
rRNA, tRNA, mRNA
miRNA, snoRNA, etc
Different DNA structures
B-DNA→ watson and crick model right handed helix
A-DNA→ dehydrated B DNA adopts this conformation (dry envt) right handed helix
Z-DNA→ left handed helix, artifical maybe in our genome
DNA applications
Can store hrs of video or pictures
Origami (make shapes)
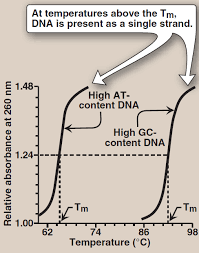
Melting Temp (Tm)
DNA can be denatured by heat/pH stress
Tm
Diff UV absorbency single vs ds (ds bases hidden so absorb less)
Hyperchromatic shift (change in absorbance (increase) as ds turns into ss)
Temp where absorbance is in middle of ds and ss DNA
Factors affecting Tm
GC content (3 H bonds more stable harder to break than 2 AT)→higher Tm
Base stacking ( smaller role but within a strand GCs next to each other more stable, increasing Tm)
Degree of complementarity (how well strands stuck to each other)
Hybridization (shorter vs longer seqs and annealing)
After denaturation DNA strands can reanneal
shorter anneals faster than longer
Highly complementary (repetitive) seqs anneal faster than diverse ones
What is FISH used for
identifying chromosomal location of a DNA sequence
ex) Downs syndrome bc will see 3 signals instead of 2
What’s electrophoresis used for
Separating DNA and RNA fragments by size
Smaller fragments migrate through gel faster than larger fragments