CELL BIO EXAM 2
1/28
Earn XP
Description and Tags
THUR MARCH 19
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
29 Terms
METHODS
MOLECULAR BIOLOGY -
A. Terms and background
Group of techniques designed to manipulate, analyze, expand, and express nucleotide sequences (RNA,DNA) so to better understand genes + associated proteins
The power of genomics
Gene transfection
Gene editing
Controlling gene expression
DNA sequencing
Genomics - Study of genes and expression
Gene transfection - Inserting intact functioning genes
Regulated GFP reporter
Gene editing - Adding or removing a few bases or full genes
CRISPR
Organ xenotransplantation
Controlling gene expression
Gene knock in/knock out
siRNA - DSRNA that will silence genes complementary to its sequencing via RNAi (RNA interference)
shRNA - Same idea as siRNA but is in a hairpin shape
DNA sequencing
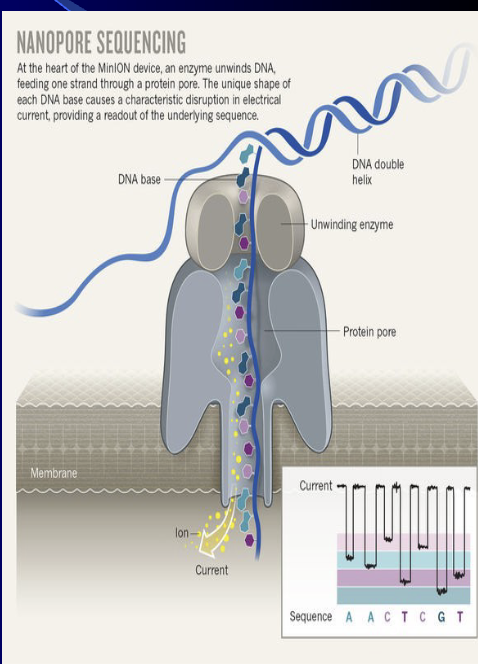
DNA Sequencing - oxford nanpore technology
Major advances that have led to “$1000 human genome” Oxford Nanopore Technologies MinION system uses bacterial pores
(1) Enzyme will pull DNA through a pore in a naturally occurring membrane protein
(2) The enzyme will unwind the DNA and the nucleotide sequencing can be read as the DNA is brought through the pore
(3) Ion current can be used to read the nucleotide sequence
Who came up with the idea of nanopore sequencing?
Daniel Branton of Harvard University
He developed Freeze Fracture that led to the idea. In 1995, with David Deamer, Branton proposed that biopolymers can be characterized and sequenced by drawing them through a nanopore. This idea led to the development of nanopore sequencing of DNA and RNA.

Epigenetics
Inheritable non-base change of DNA and histones (nuclear proteins) due to environmental influences)
Reversable
DNA methylation
Histone phosphorylation
can be caused by a number of factors
stress
environment
diet
Transcriptome
the set of all RNA molecules in a cell or cells
umbrella term for RNA
mRNA
tRNA
Proteomics
Study of all proteins
Metabolomics
The study of other molecules related to metabolism
Small molecules found in a biological sample
Cancer cells take the easy road during metastatic migration
Metastatic cancer cells spread in the body in a complex microenvironment. A group of researchers discovered they pick the route the demands the least amount of energy - this implies that targeting the metabolome may be important. They prefer large spaces rather than narrow passages. They are LAZY!
Secretome
Analysis of secreted proteins, cfDNA, exosomes, and microvesicles (anything secreted by a biological sample.)
protein
cfDNA
exosomes - originate from the cytoplasm
Exosomes are being looked at to replace stem cell therapies.
Stem cells secrete growth factors and important proteins in exosomes so we can simply isolate the exosomes for treatments.
Exosomes can be a possible biomarker for cancer
advantage : stem cells are forever, exosomes have a half life
Microvesicles → sprout from the cell membrane
Mutations
absolutely critical in molecular genetics
W/o mutations we would not understand gene function
can be caused by ultraviolet light, chemicals, etc.
techniques
UV light - thymine dimers
Chemicals - e.g. ethylmethanesulfunate G-C ---> A-T
B. Yeast Cells
L. H. Hartwell — 1960s-1970s
Advantages to using yeast cells for research:
Simple to grow - in a simple medium flask
Exist in haploid or diploid form
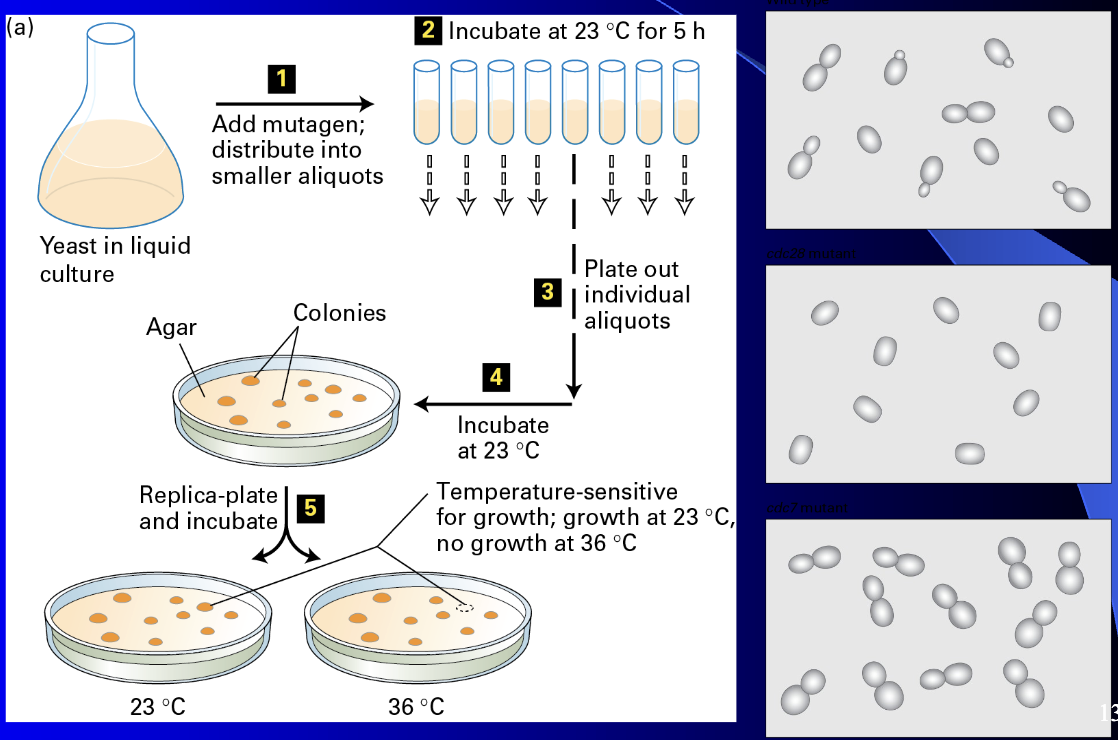
Generate temperature sensitive mutants
permissive temp at 23ºC
Non permissive temp at 36ºC (Can’t grow anymore because growth related proteins are denatured)
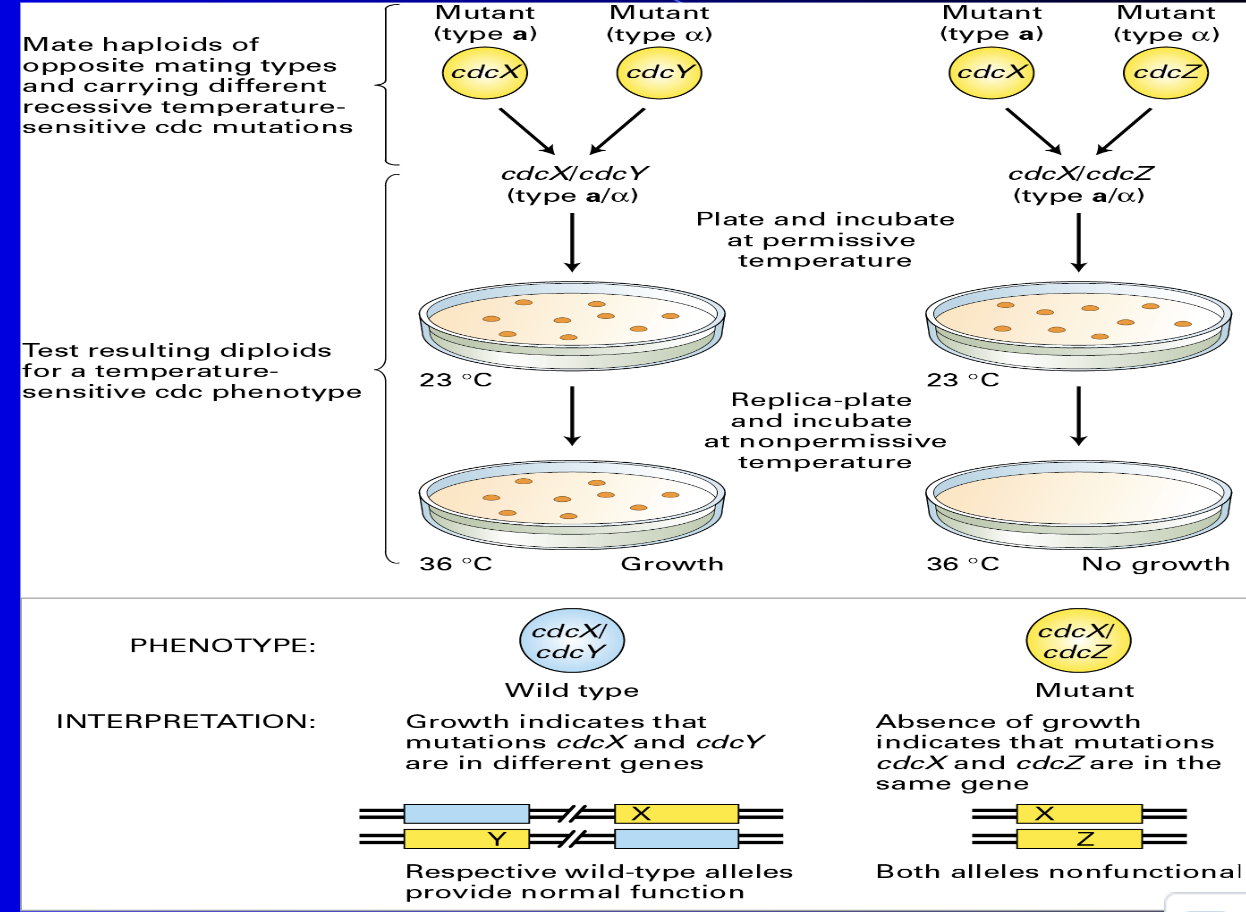
Complementation Analysis
Used to see if two mutations are on the same gene
(1) Scenario: we want to see if mutation Y or Z is found on the same gene that mutation X is on
Mutation X causes the yeast to be temp sensitive
Mutation X is recessive
Mutation Y and Z are dominant
(2) Take haploid yeast cells and grow them in 2 cultures
(a) Each culture will have yeast cells with mutation X, one culture will have mutation Y yeast cells, and another will have mutation Z
(b) Cross the two haploid yeast cells at 23ºC to make XY and XZ mutant yeast cells
(c) Isolate the diploid yeast cells and place them in a separate culture (one culture for XY and another for XZ)
(d) Grow at 36ºC
(e) If yeast grew at 36ºC then X and the other mutation are on different genes
(i) If they were on the same gene, then there would be no growth because the mutation makes them TS (temp sensitive) so there would be a dominant gene to not grow at 36ºC
C. General principles and concepts
Restriction nucleases/enzymes
Gene splicing
Nucleic Acid Hybridization
Restriction nucleases (enzymes)
Cleave DNA at select sites
Discovered in 1963 → 1970s used in laboratory
Restriction nucleases → discovered in bacteria so they can ward of invading viruses
3000 restriction nucleases known
800+ commercially available
Gene splicing
Nucleic Acid Hybridization
FISH
Antisenese RNA/DNA
Southern Blots
Northern Blots
cDNA micro arrays
Molecular Beacons
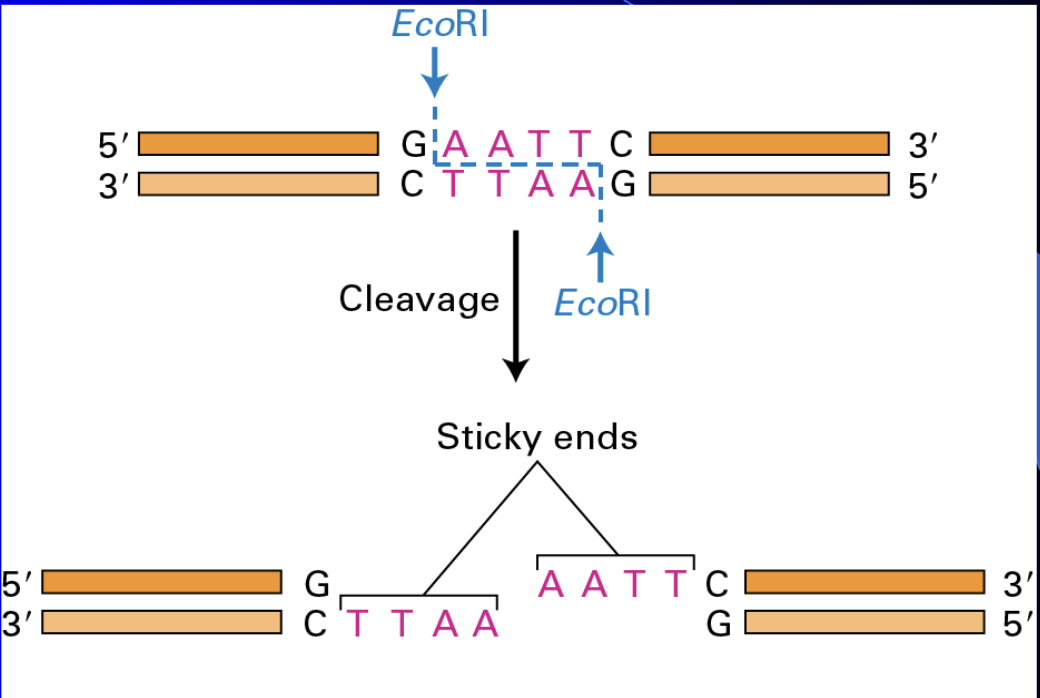
Methods - molecular bio
Plasmids
Hybridization
Plasmids
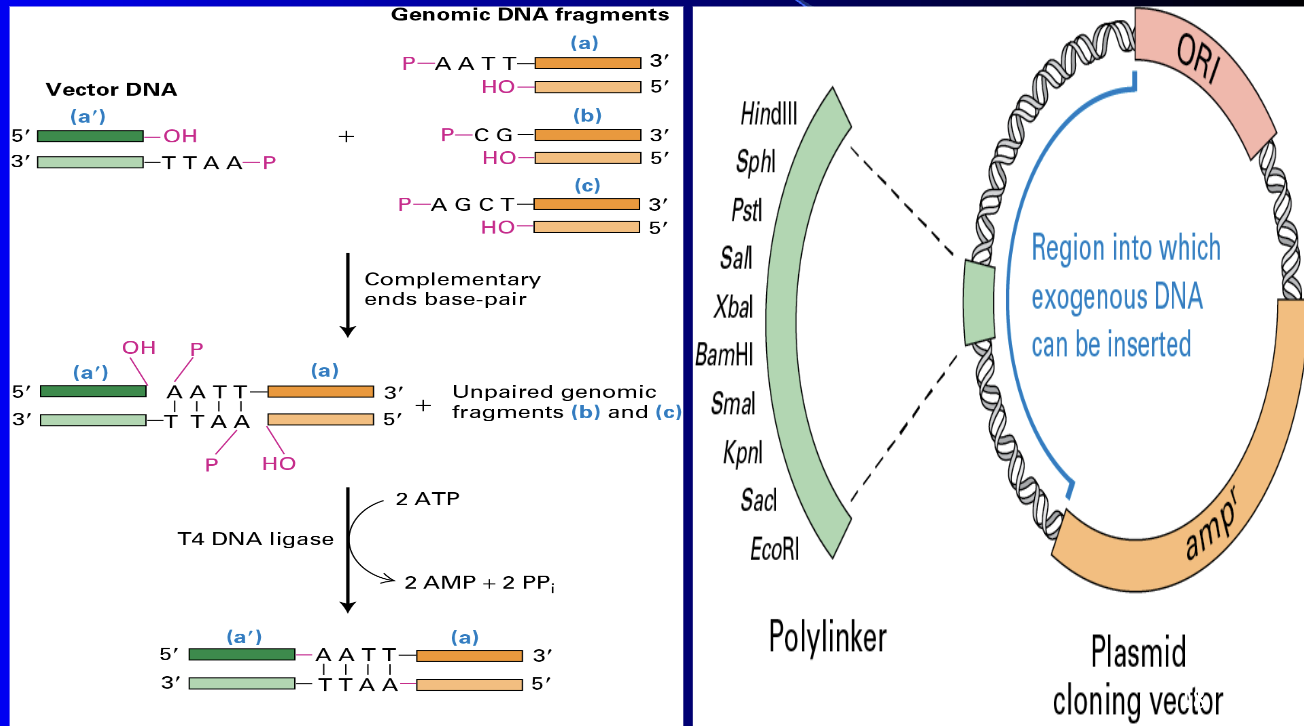
Hybridization - many applications such as anti-sense RNA/DNA
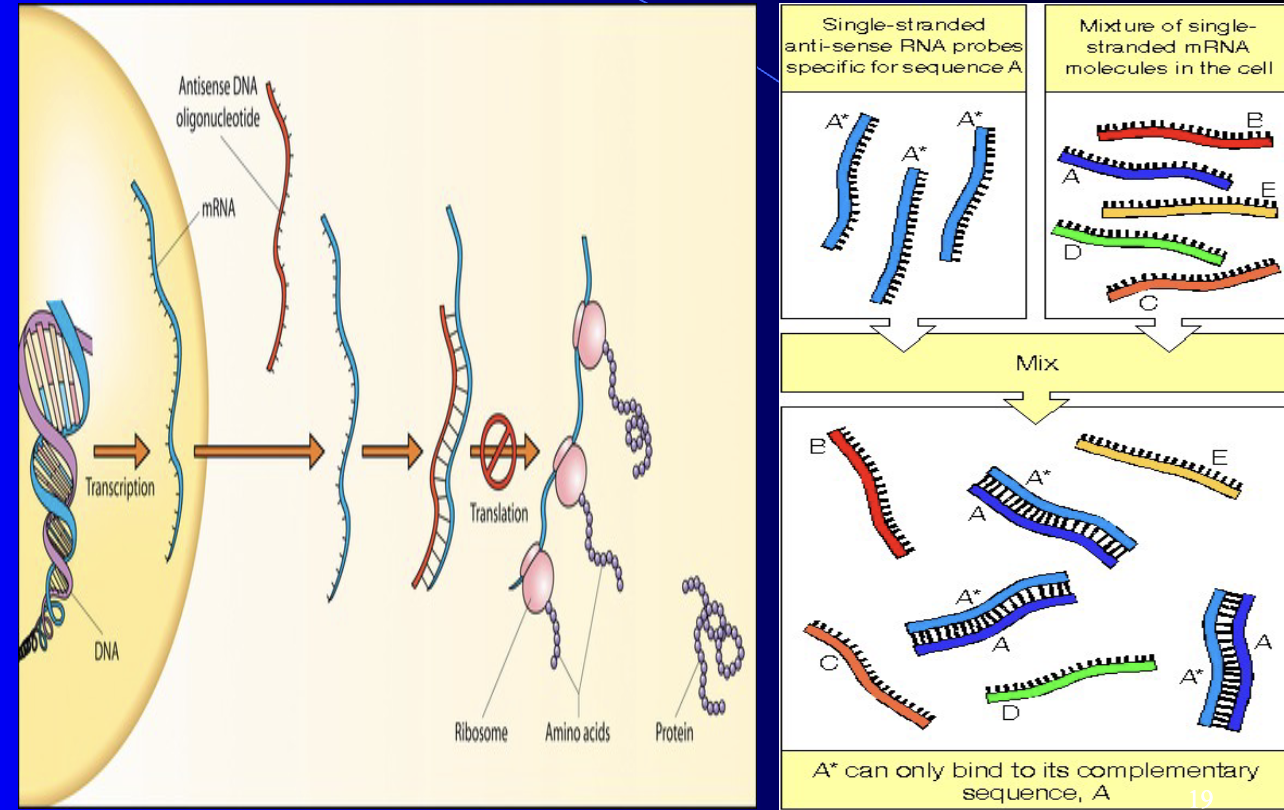
Methods - molecular bio
DNA gel electrophoresis & band gel shift assay
Can identify DNA binding proteins such as transcription factors
Evolved from SDS gel electrophoresis
Uses restriction enzymes
Uses polyacrimaline sieve for smaller DNA fragments
Uses agarose sieve for larger DNA fragments
Scenario: we want to see if our POI binds to a select gene in the genome
Band Shift Assay
DNA with gene of interest (GOI) run through one part of the gel
Other part of the gel has both GOI and POI
Compare both sides of gel - If band is higher up, POI has bound to GOI and holds it up from falling
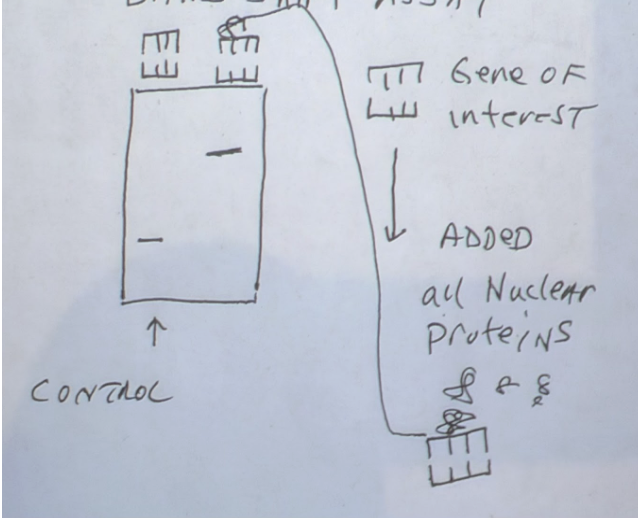
Methods - molecular bio
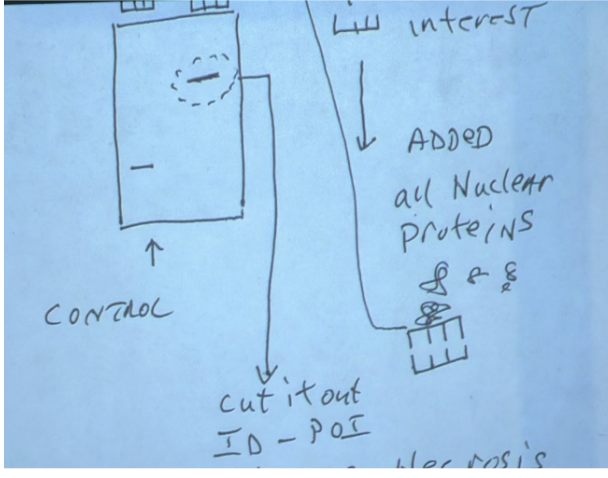
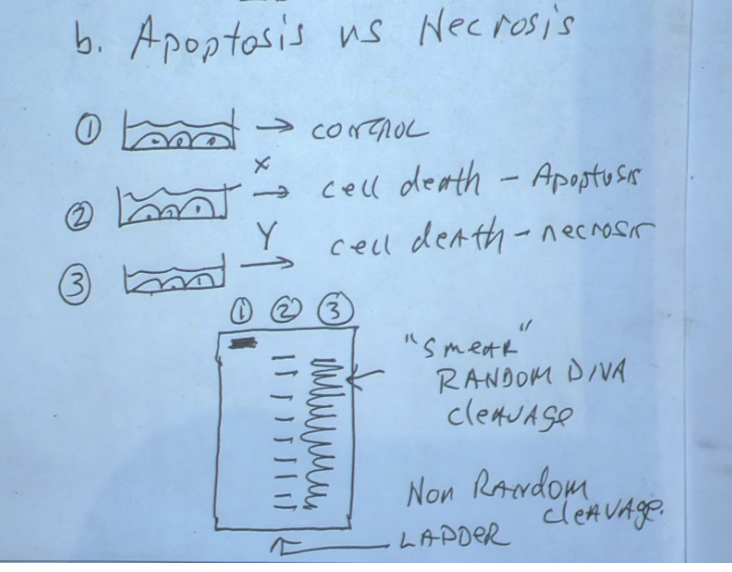
DNA gel electrophoresis can discern cell death
Can be used to see if cell undergoing late stage apoptosis or necrosis
(1) Divide gel into 3 parts
Control - DNA from healthy cells
Apoptosis
Necrosis
(2) Control DNA will stay in place at the top
(3) Apoptosis DNA will show up as a ladder
(a) Apoptosis induces non random DNA cleavage
(4) Necrosis DNA will show up as a continuous streak
(a) Necrosis induces random DNA cleavage

Methods - molecular bio
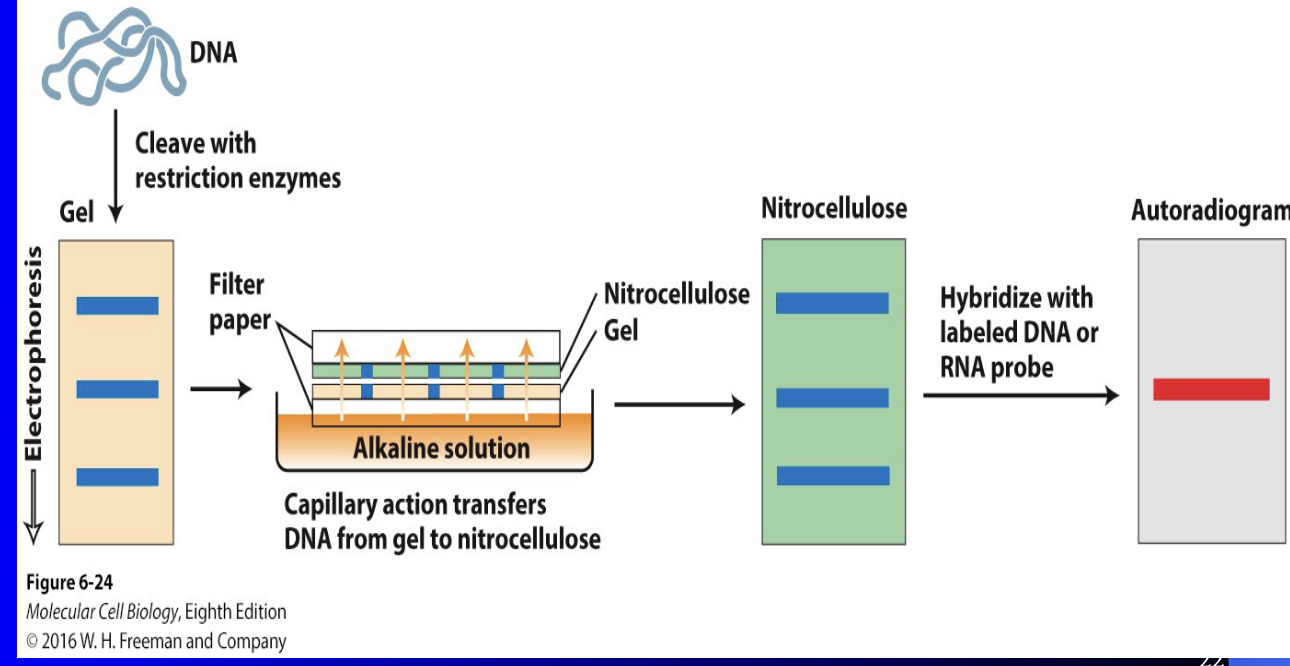
Southern Blots
Analyze DNA & genes
(1) DNA gel electrophoresis bands placed onto blotting paper (nitrocellulose membrane)
(a) Nitrocellulose membrane placed on top of gel inside alkaline solution
(b) Bands transfer from the gel to the membrane
Complementary DNA/RNA probe (generally radioactive) to GOI placed on nitrocellulose membrane
Quantitative — molecular # (relative abundance)
Qualitative — # of gene copies (molecular weight) data
The further down the gel, the lighter the molecular weight for that gene
Applications
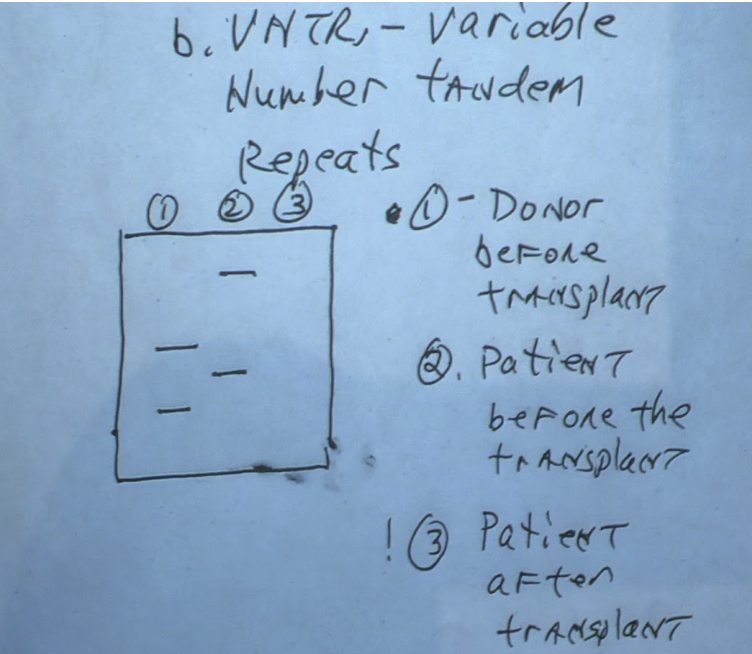
Used for bone marrow transplants to see if it was successful
Divide gel into 3 parts
Donor marrow
Receiver marrow before transplant
Receiver marrow after transplant
Bands 1 and 3 should be at the same height on the gel
Variable Number Tandem Repeats (VNTRs)
(1) Part of genome where short strand of nucleotides is repeated adjacent to each other
a) The number of times they repeat vary from person to person
b) Used in forensic medicine
Single nucleotide polymorphisms (SNPs)
single base change
(1) Used for personalized medicine. e.g. 237 SNPs in 146 genes → 86% accurate prediction of autism
(2) Looks at single nucleotide differences primarily in noncoding genes - 98-99% of your DNA are non coding so 1 base can make a big difference in the coding region
(3)Over 600 + million known SNP in the DNA genome
(4) you - 4-5 million SNPs
(5) Rapid expansion - a report estimates the global SNP genotyping market will grow from $12.9 billion in 2024 to $61.1 billion by 2035. Annual growth grate (CAGR) of 16.84%.
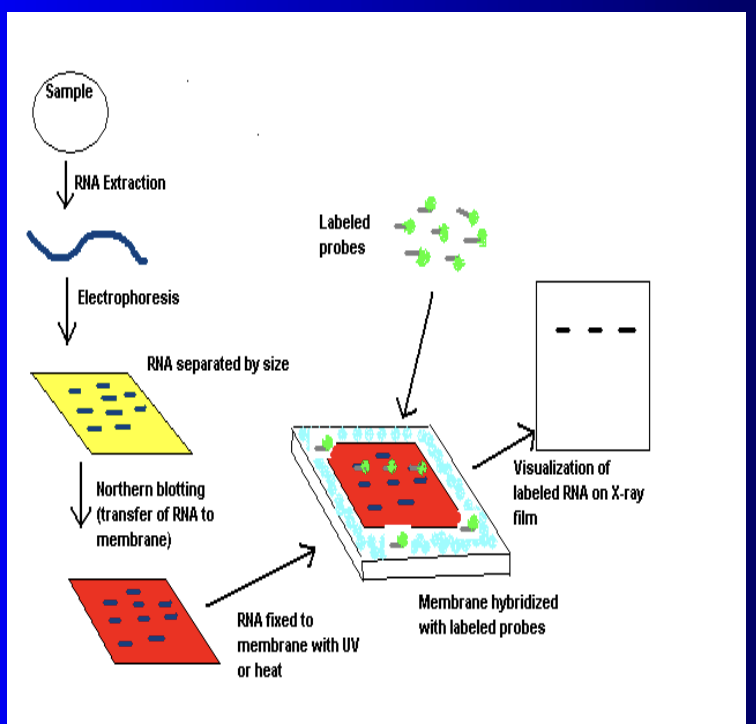
Northern Blot
(1) Same exact protocol as southern blots except tracks RNA instead of DNA
(a) Used to measure gene expression (mRNA)
(b) Can only analyze 1 mRNA transcript at a time
(i) If we look at multiple mRNA transcripts we will get mixed up on what is used to code what
summary
blots
(a) Western blots
(i) Proteins
(b) Northern blots
(i) RNA
(c) Southern blots
(i) DNA
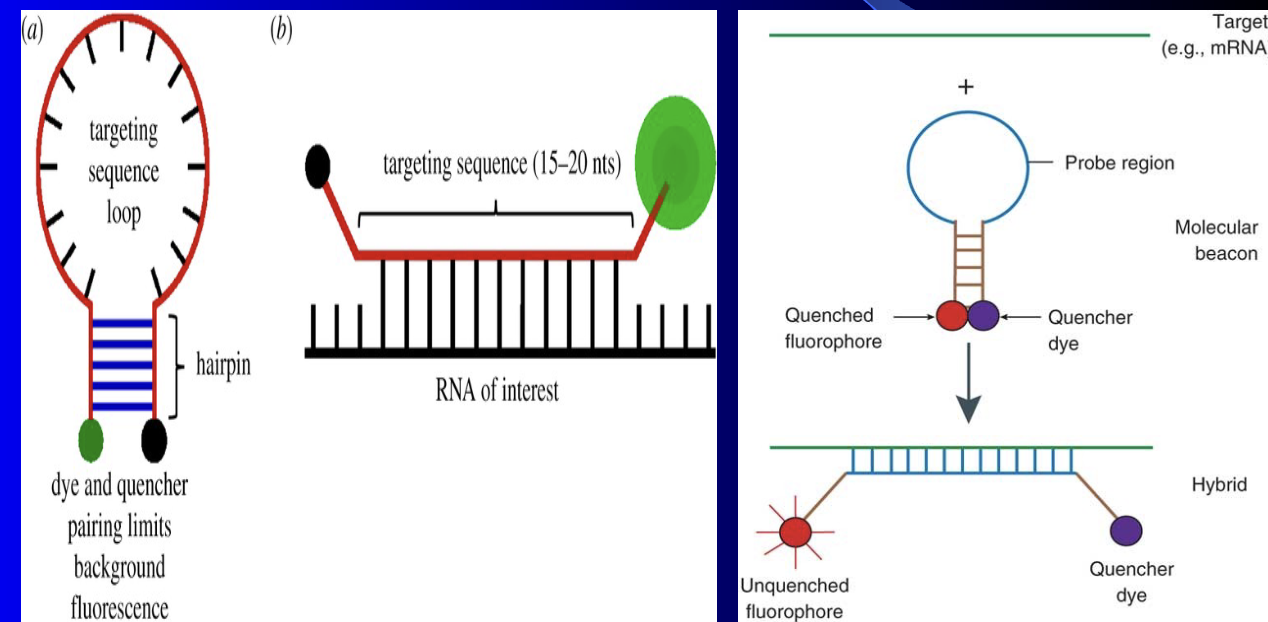
Molecular Beacon
probes can hybridize in situ in cells – better than FISH
Molecular Beacon probe’s fluorescence is quenched (not fluorescent) unless bound to target RNA or DNA strand
(1) Takes RNA or ssDNA complementary probe that has fluorophore and quencher dye on either end (
2) The probe region is circular so the fluorophore and quencher dye are next to each other
(3) Complementary probe bound to the target
(a) Fluorophore and quencher dye are farther away and the fluorophore will fluoresce
(b) Tells us the same thing as FISH but is more efficient
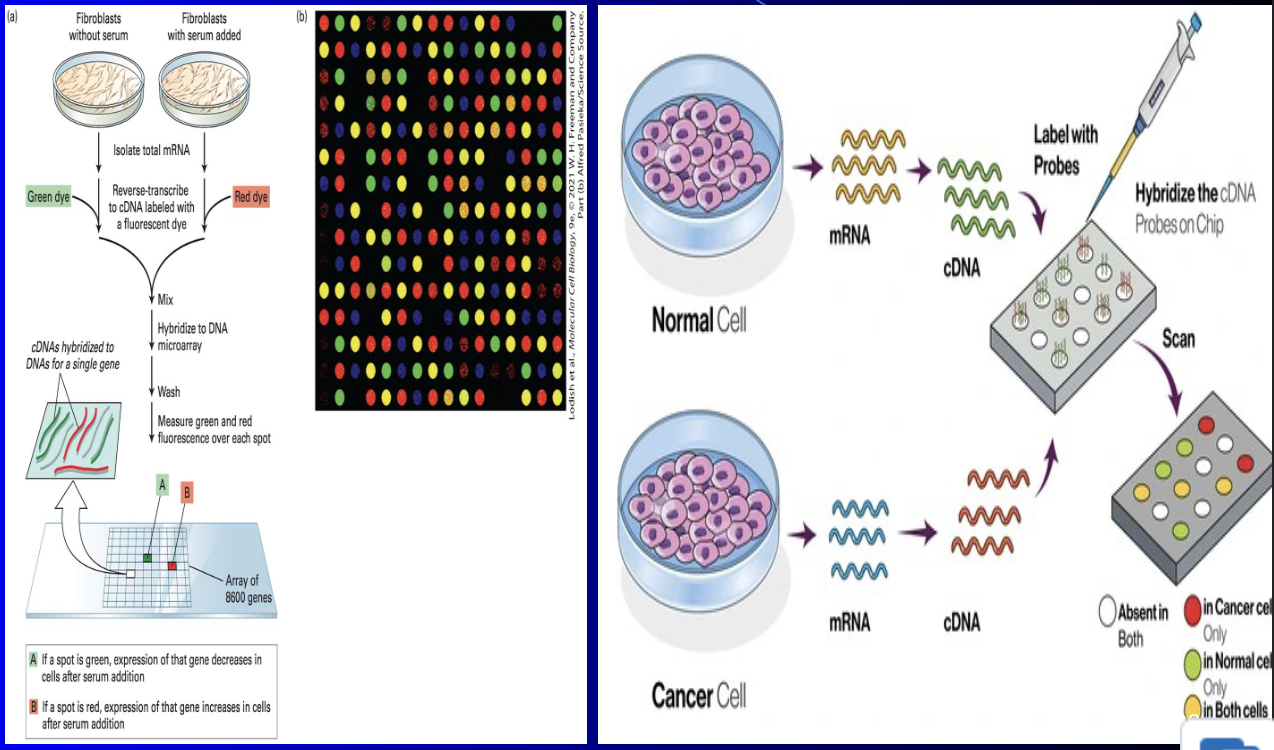
cDNA (complementary DNA) microarrays
a) Used to examine up to 8600 genes expressed
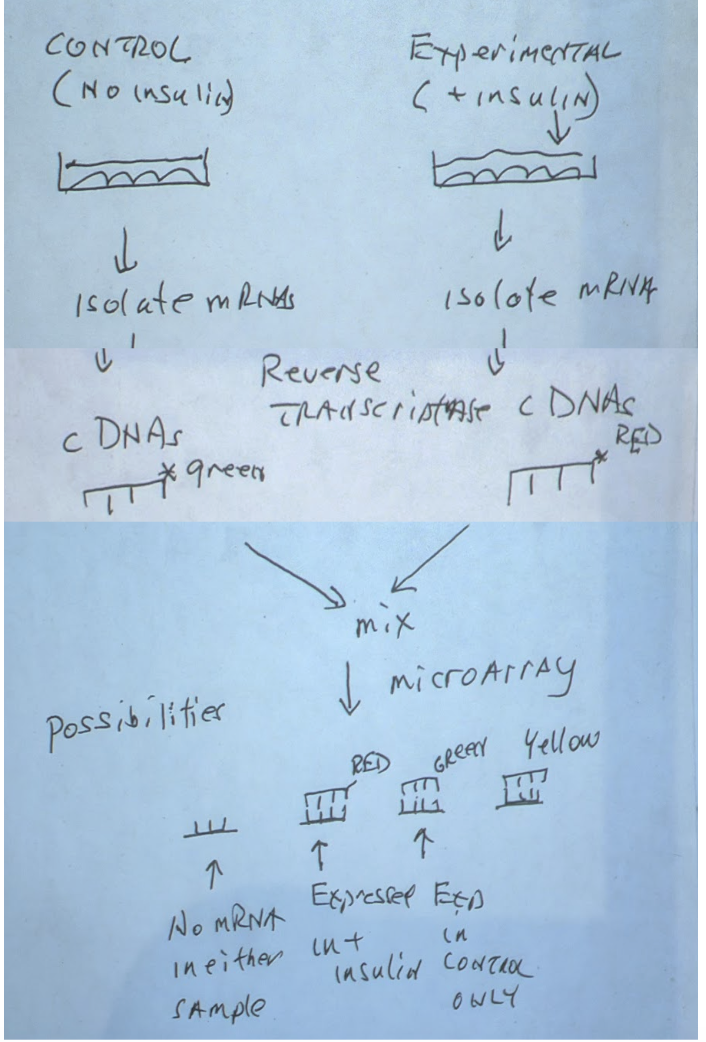
b) Scenario: we want to see what genes are being expressed when cell is introduced to insulin
Take 2 cultures
one for control group
one for insulin introduced cells
Harvest mRNA from the cells, and reverse transcriptase them to form sscDNA (single stranded complimentary DNA)
Tag the sscDNA with fluorescent dyes
Green for control group cDNA
red for insulin group cDNA
Identical oligonucleotides (short DNA or RNA sequences(in this case ssDNA)) for the testing gene placed in a well
This can be done up to 8600 genes
Identical oligonucleotides (short DNA or RNA sequences(in this case ssDNA)) for the testing gene placed in a well
If the cDNA bound to the oligonucleotide, it stays on the well and does not get washed away
Array is laser scanned
(a) if well is gray, none of the cultures express that gene
(b) if the well is green, only the control group express that gene
( c) if well is red, gene is only expressed by insulin exposed cells
If well is yellow then both samples express that gene
Microarrays uses
Gene Cluster Analysis – Patterns of Similar Expression
Subtyping human cancer cell type
Cryosurgery of the Prostate
Single Cell RNA - seq
Can identify individual cells at select points in time based on their unique RNA signatures
RNA signature - Part of mRNA that is reflective based off the state of the cell (healthy, diseased, old, etc)
Used in tissue engineering.
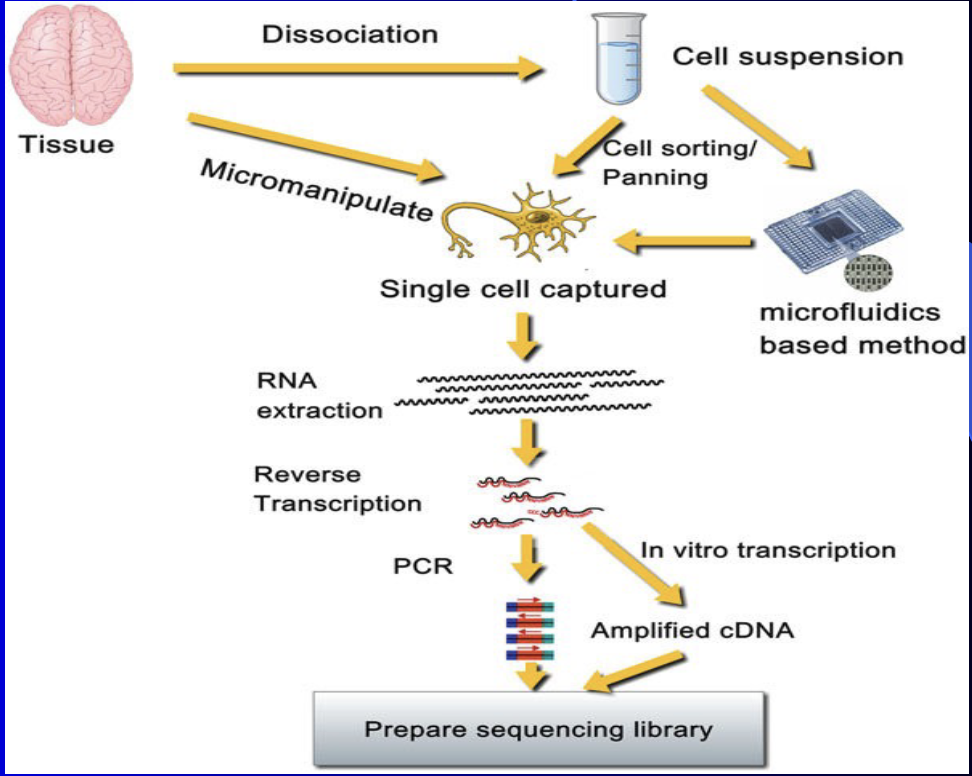
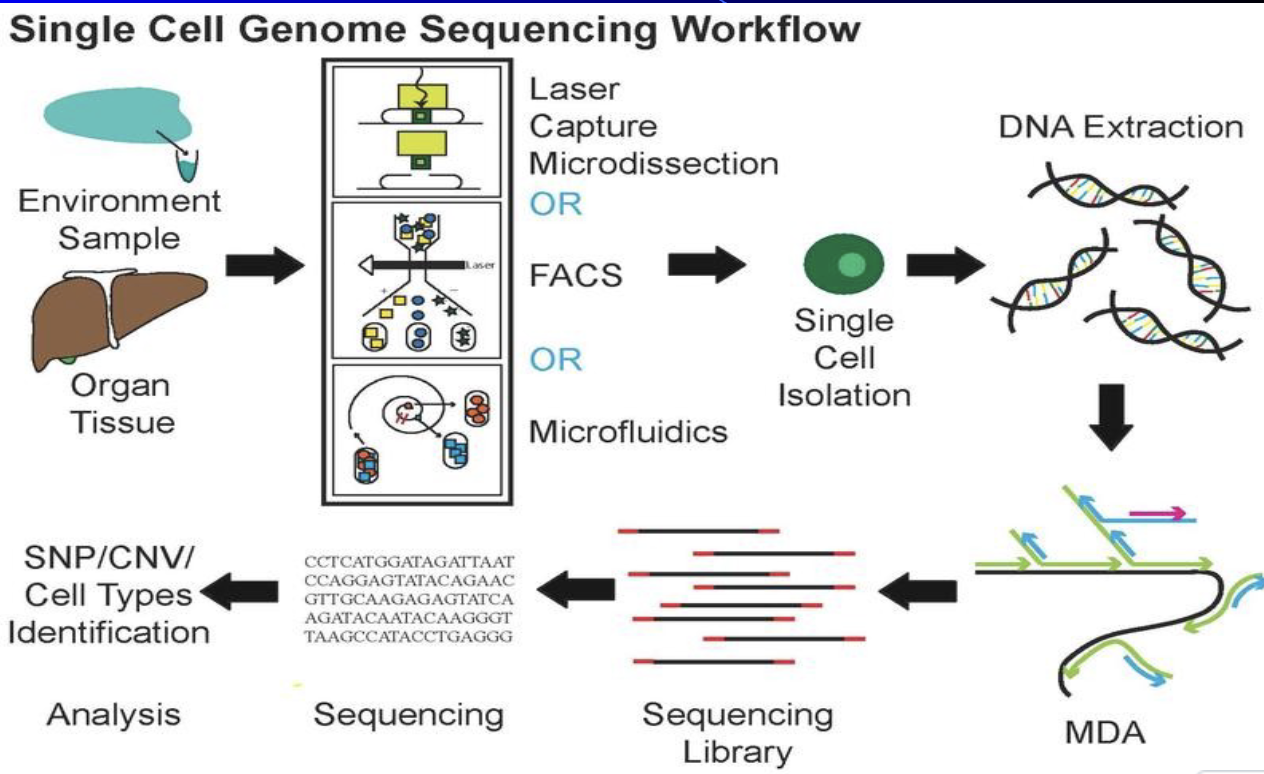
Single cell genome sequencing
a) Taking a cells genome and comparing it to sequencing library for analysis
Can tell you cell type
b) Cells from a tissue sample must be sorted before sequenced. (single cell genome sequencing requires cell sorting too
DNA cloning and cDNA cloning are key molecular biology techniques
Genomic cloning vs cDNA cloning
Genomic Cleaning
Starting material is all of the genomic DNA
Advantages: The gene of interest is present
Disadvantage: Huge library of E. coli DNA fragments to search
cDNA Cloning
Starting material is mRNA, not DNA
Advantages: Smaller library to search
Disadvantages:
Need to convert mRNA to cDNA to clone E. coli using reverse transcriptase
If gene isn’t being expressed then the gene won’t be cloned
DNA/Protein Expression Systems using E.Coli
Prokaryote gene expressions
1981-84 - Humulin - Recombinant human insulin
We can use cDNA cloning to make humulin from E.coli
Take out Lac-2 gene from lac operon, place G-CSF cDNA instead
This has been commercially available since 1982
IPTG - lactose analog doesn't break down
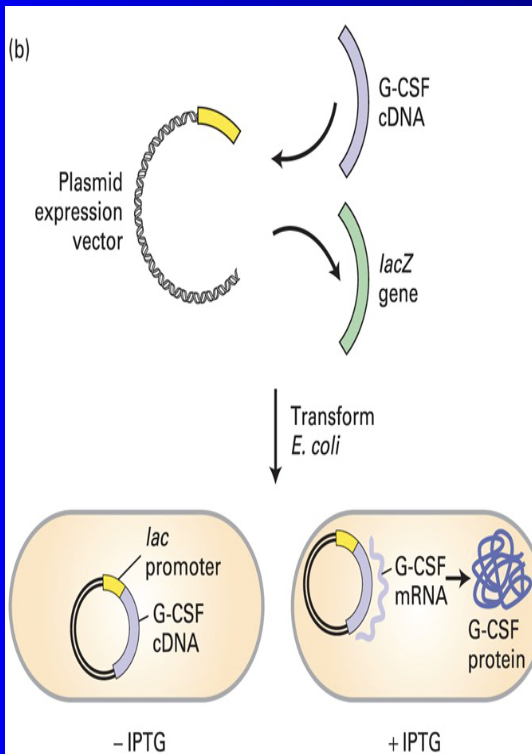
Protein Production by Bacteria
Disadvantages compared to eukaryotic cells
Bacteria tend to only produce proteins that are not hard to harvest
Protein present in inclusion bodies
Bacterial proteins are not post-translationally modified
no Glycoslation, phosphorylation, folding, etc.
Humulin works with E.Coli because of sheer luck, it should not have worked
Required for protein function