Lecture 8: QA/QC and Molecular Microbiology Part 1
1/57
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
58 Terms
Regulations and Governing Bodies:
Federal law governs implementation, usage and quality programs the surround testing of human specimens
Clinical laboratory Improvement Amendments - Regulations passed down by CMS (1988 - known as CLIA ‘88)
Joint Commision (TJC), local state laws (state DOH), and other accreditting organizations (CAP, AABB, etc)
Some states have additional requirements for testing/accredidation (CLEP - Clinical Laboratory Evaluation Program, NYS DOH)
Accrediting bodies will sometimes defer to whichever one is “stricter” regarding rules (CLEP > CAP > CLIA)
What do these “bodies” govern/say you have to do?:
EVERYTHING
Laboratory Safety/Information Systems
Test Method Validation/Verification
Standard Operating Procedures and Training
Proficiency Testing
Personnel Requirements/Competency of Staff
Quality Control and Analysis
So much, much more
Requirements for Testing/Reporting:
Must be accredited first (apply for license to do the testing)
Must be enrolled in a proficiency testing (PT) program (blinded challenges)
Must undergo periodic review by other labs/teams (biannual insepction)
Must have staff that are qualified and certified to run the tests alongside ALL THE PAPERWORK
CAP - biyearly inspection where teams come in to review lab (files, reports, licensing, etc)
The guidance documents tell all…
Requirements for Testing/Reporting: Checklists:
All Commons - All labs
Specialist section (Micro, Chemistry)
Requirements for Testing/Reporting: What’s inside?:
The thing you must do
What thy expect you to do (why)
How you have to keep track
Requirements for Testing/Reporting: This covers:
Training and competency, safety, SOPs, IT stuff, testing, QA/QC and any new technologies
A new molecular test for TIKTOKVIRUS has been made - it’s been infecting people’s brains and zombifying them: How do we do it?:
Decide what type of test to bring on (IVD v. others)
How to operate the test (SOP)
A way to show that the new assay is working correctly (blinded challenges — proficiency testing)
Data showing that the test works great in your hands (validation/verification)
A way to show that it continues to perform well and safely (QC plan)
IVD (In vitro diagnostic):
“…products are those reagents, instruments, and systems intended for use in diagnosis of disease or other conditions, including a determination of the state of health, in order to cure, mitigate, treat, or prevent disease or its sequelae. Such products are intended fro use in the collection, preparation, and examination of specimens taken from the human body
These assays are the easiest to show performance on and introduce
Everything is made by the company. submitted to FDA and they tell you how to do it, what to test on, and how the test performs
Cant deviate from manufacturer’s instructions
Pre-market approval (new diagnostic):
Goes through initial submission of safety and efficacy of new device in chosen field
510k Premarket notification:
Comparative analysis of new assay to “gold standard” (one that did PMA), showing “substanial equivalency”
Emergency Use Authorization:
Classification for emerging disease/infectious agents — limited FDA submission data (rare)
Laboratory Developed Test (LDTs):
Developed by the lab (nothing on the market)
Need to show that it works well (validation)
Vendors won’t help (no tech support)
Have to state “not FDA approved” and that “characteristics were determined by XXX laboratory”
May be cheaper than IVD (can use equipment you already have)
SOPs (Standard Operating Procedure):
Tell people how to do the test — new tests all need an SOP/manual
Must include principle of test, what specimens it can be run on, rejection criteria, TAT, detailed protocol, reagent/instruments, results (expected), troubleshooting, QA/QC and limitations
Must be signed by laboratory director and everyone who does the test
Proficiency Testing/Challenges:
Regulated analytes require annual enrollement (CLIA defines)
CMS tell us which ones are approved (PT 3× 5 samples/year)
For non-regulated analytes (those not covered by CMS), lab has to test 2x per year
There are programs that will send you challenged 2x/yr
If no PT is available, can work with other labs (peer/alt review) and show performance (blinded samples)
Alternative proficiency
Showing your test works: Validation/Verification:
IVD requires a verification (unless modified)
LDT requires a full validation
Verification:
Shows that the assay works just like the manufacturer says
FDA-cleared or approved tests only
Checks accuracy, precision, reportable range, and reference range
Compare new method to one already verified (test using same samples) — do this with ALL specimen types
Any changes require FULL validation (don’t deviate)
Validation:
Checks accuracy, precision, reportable range, and reference range
ALSO checks for interferences, sensitivity, specificity, LoD, LoQ, etc
Check against another method (IVD or LDT) that’s approved
Define: Controls, protocol, and performacne chars
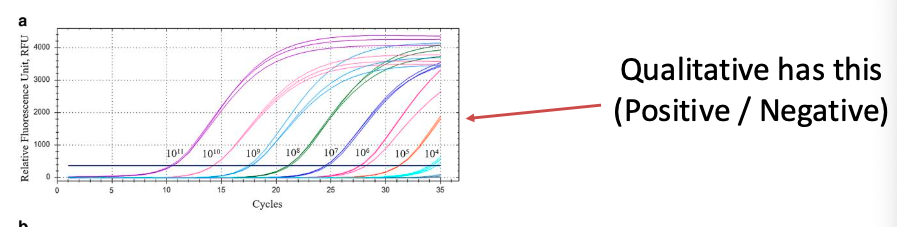
Qualitative tests:
Require (1) Positive, (2) Negative analytes and (3) Non-template control (NTC)
These show (1) assay detects what it should, (2) no false positive amplifications, and (3) no contamination
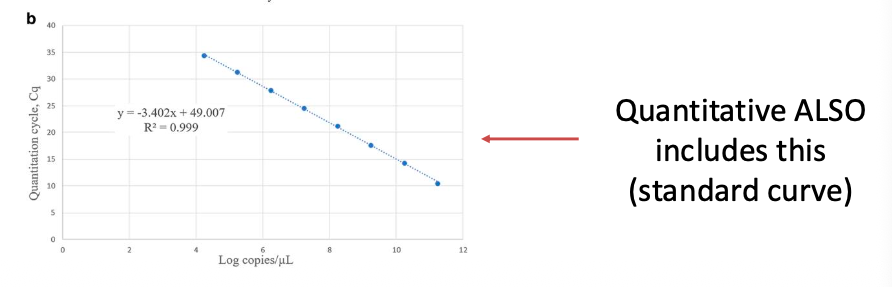
Quantitative tests:
Require (1) High and Low positive, (2) Negative, (3) Non-template control
Have a “standard curve” of known analytes and specific concentrations
External Control:
Typically bought by someone other than the test maker
Previously tested (pos/neg) samples work too
Shows assay detects targets/works
Internal control:
Typically part of the assay (checks for quality)
Can be human genes or spiked in target
beta-globulin gene in HPV endocervical samples
Bacillus spores in test for C. difficile
If does not amplify, cannot report — INVALID test
Extraction control:
Used to show that nucleic acid extraction worked
Can be same control as “internal” or “external”
Many companies will use this as a schema to save on reagent costs (fewer tubes to run and less ctrl material to use at once)
Why do we need so many controls?:
Manufactuers define conditions in IVD approval as to how many controls need to be run
Typically defined within a 24 hr window or per each “batch” of tests
If LDT, control frequency must be defined by the lab
Some tests do not have defined frequency to be run, thus the labe must work on defining this (ICQP)
All control performance must be tracked (logged and tracked for compliance)
This must be done prior to reporting ANY specimen results
IQCP: Individualized Quality Control Plan:
QC plan that allows for decreased control material utilization (weekly or monthly) IF the assay allows for it (instructions)
In NYS, no less than 1 pos sample/mon, 1 neg sample / week
Each labe must carry out risk assessment and review of performance (20-30 d of runs)
Monitored monthly following approval — signed by Lab Director
Changes/updates needed if deviates from prior approval conditions (temp change, increase in false positives, instruments non functional, etc)
Used to cut down on usage of individual cassettes / reagents (cost)
Molecular Microbiology:
Detection, identification and analysis of microorganisms
DNA or RNA
Advantages of Molecular Microbiology over traditional microbiology (culture, stains, and biochemical testing):
Rapid turnaround time
Better sensitivity and specificity
Can be quantitative
Comparison of biochemically similar organisms (epidemiology)
Why care about Molecular Microbiology?:
Molecular testing is increasingly common
Unlike genetic screening or oncology, microbiology testing is relatively widespread
EVERYONE gets sick, multiple times throughout their life
Rapid advance in technology enables us to detect previously unknown pathogens (or rarely ID’d organisms)
Legionnaire’s Disease (L. pneumophilia), COVID-19 (SARS-CoV-2)
Molecular testing can drastically impact patient care:
Improved patient care (decreased mortality, morbidity, and length of stay)
Lower costs to the hospital system
Avoid unnecessary antibiotic administration (or decrease utilization of broad spectrum antibiotics)
Lower infection rates by identifying people colonized by multidrug resistant orgnisms
Applications of Molecular Assays in Microbiology:
• Molecular assays have in almost all labs replaced viral culture:
– Rapid / high-throughput differentiation of viruses (DNA, RNA) that
cause similar syndromic conditions
• Antibiotic resistance testing for tailored antimicrobial therapy
– Change drug choices when they have a resistance marker (e.g. tetM
might not use tetracyclines)
• Quantification of viruses (or other organisms) in specimens
– HIV viral loads
• Genotyping, classification, and epidemiological studies
• Discovery of novel pathogens
– Sequencing discovery of SARS-CoV-2
• Microbiome / Virome / Mycobiome studies
– How changes in our normal flora relates to disease (more research)
Signal Amplification Methods:
Normally use a nucleic acid probe combined with some form of amplifying signal producer (enzyme):
Branched-chain DNA
Hybrid Capture
In situ hybridization
Typically considered less sensitive than NAATs
Less likely to be contaminated than NAATs (false positive results)
Fluorescent ISH:
Uses hybridized probes of DNA (or peptide, PNA) to identify intact microorganisms:
Bind to rRNA molecules on microbes (higher copy number than a single gene) and aid in ID/selection of antimicrobials
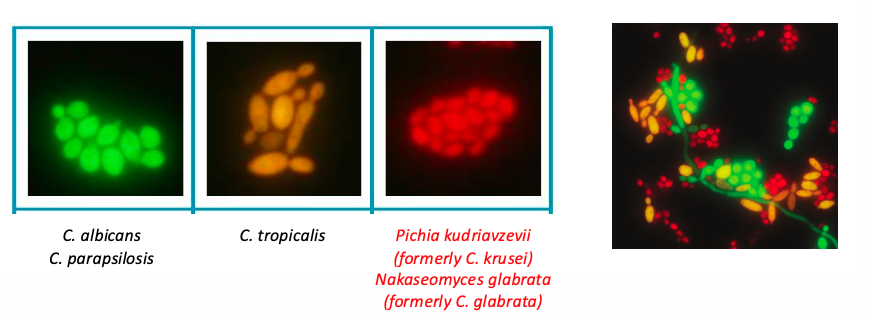
Accelerate Pheno System:
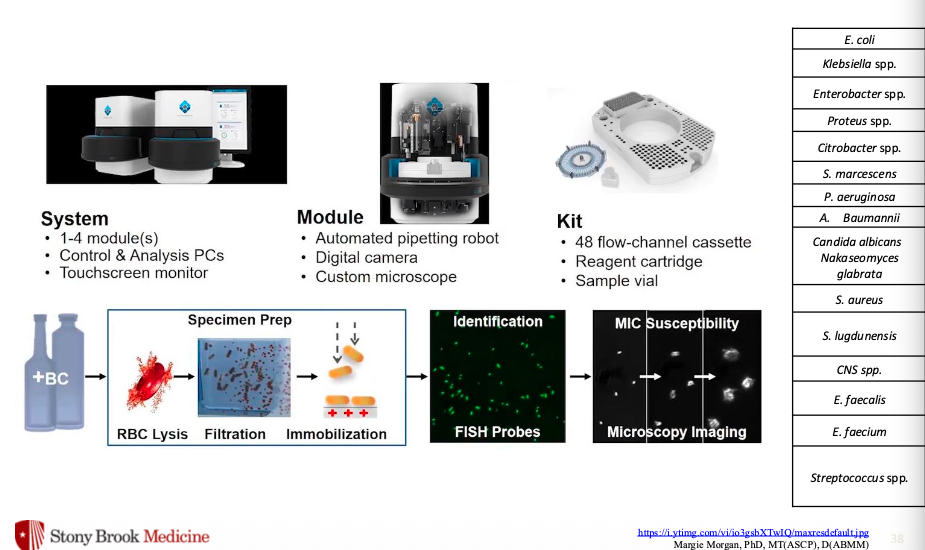
Nucleic Acid Amplification Tests (NAATs): qPCR/RT-qPCR dominates most NAATs
Predominant workhorse of most molecular diagnostic labs
End stage PCR (gel electrophoresis, PFGE, etc) has been mostly replaced by qPCR
SYBR Green:
Primers specific for a region of interest (ROI)
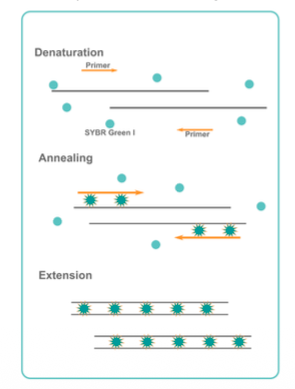
Taqman:
Primers and an ROI specific probe
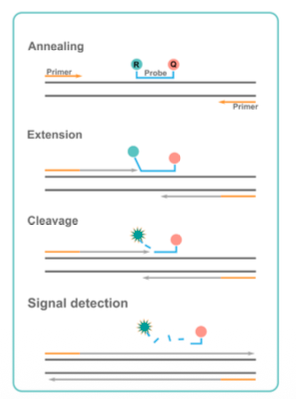
Principles of NAATs (e.g. qPCR) for pathogen detection:
DNA (or RNA) extraction → Target sequence amplification with qPCR (or RT-qPCR) → Simultaneous detection of products w/ fluorescent probes
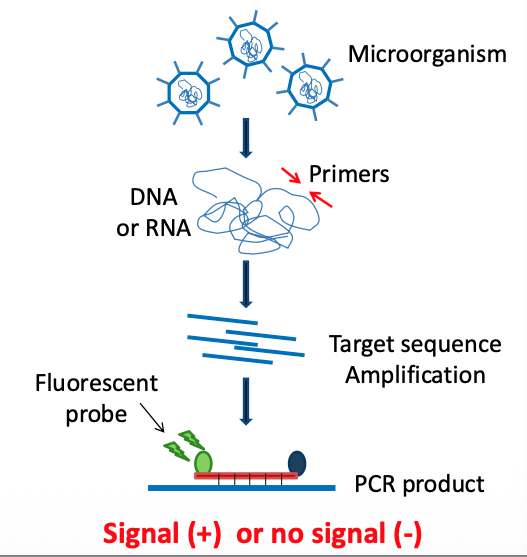
NAATs: Qualitative assays Presence or Absence of a gene/ROI:
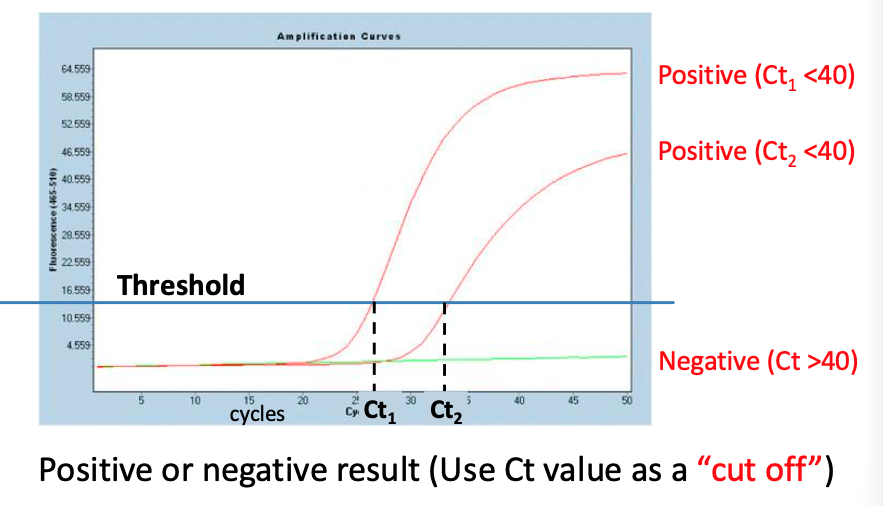
What are Qualitative Assays good for?:
Diagnosis of infections:
SARS-CoV-2
Influenza A/B
Human Papillomavirus (HPV)
HSV1, HSV2, VZV
Mpox Virus (MPXV or Ortopoxviridae)
Many others
Syndromic-based diagnostic panels:
Test for multiple pathogens in one test:
Respiratory panel
Gastrointestinal Disease Panel
Meningitis/Encephalitis Panel
Blood Culture ID Panel
Many of these are “sample-to-answer” platforms where assay complexity is highly reduced:
Nucleic acids extracted onboard
PCR carried out on cartridge
Self-sealed containers than can be discarded once used

NAATs: Quantitative assays:
Uses standard curve (serial dilution of known concentrations) run in parallel to allow for estimation of titer within a sample
Used for diagnosis/tracking of titers of organisms in bodily fluids:
namely viral titers in serum/plasma
Ex:
Human Immunodeficiency Virus-1
Hepatitis C Virus
Hepatitis B Virus
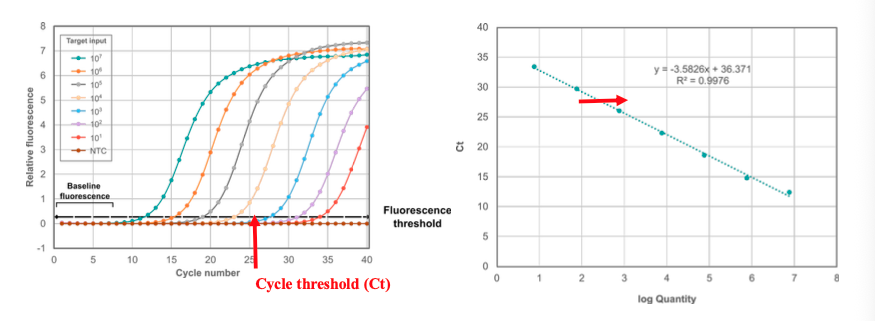
Quantifiable range vs reportable range:
Not always the same
Things can be positive < or > your limit of quantitation (LoQ), and are still positive
You cannot report outside of the LoQ (only that it’s positive)
Ct values =/= quantitation (and should not be interp that way)
NAATs: Modifications of PCR:
Different assays use modified protocols or methods within the “PCR” realm for augmenting specificity or allowing for greater number of analytes per test:
Broad range PCR
Nested PCR (Biofire plateforms use this technology)
Multiplex PCR (commercial platforms with 2+ targets use this)
Reverse Transcription PCR (RT-PCR) is used by assays with targets that originate as RNA (HIV, HCV, etc)
How do you select target sequence for identification of microorganisms?:
Genomic or plasmid DNA, genomic RNA
Unique gene/sequence for the pathogen of interest (e.g. virulence factors)
Antibiotic resistance gene
For viruses with various subtypes, use:
Sequence shared by all types for initial detection
Use type specific sequence for further typing
Major considerations of using molecular tests:
Cannot differentiate between alive/dead organisms
Remnant nucleic acid can persist for weeks or even months
Many PCRs should not be used as “test of cure” for this reason
Not all tests and targets are equal - know test limitations
Not detected =/= not present
Pathogen under detection limit, wrong location, etc
Genotype =/= Phenotype
Presence of antibiotic resistance gene does not mean the bacteria is resistant
Clostridium difficile:
Gram positive, anaerobic, spore-forming bacillus
Spores: critical for prolonged survival in the environment and ability to spread
Causes 15-25% of all antibiotic-associated pseudomembranous colitis (diarrhea)
In recent years healthcare associated diarrhea has increased in incidence and severity
C-difficile infections are associated with an increased hospital stay, morbidity, and mortality among patients
C. difficile Pathogenesis:
Antibiotic therapy → C. difficile exposure and colonization → Toxin production → Effective immune response = asymptomatic, Inadequate immune response = diarrhea
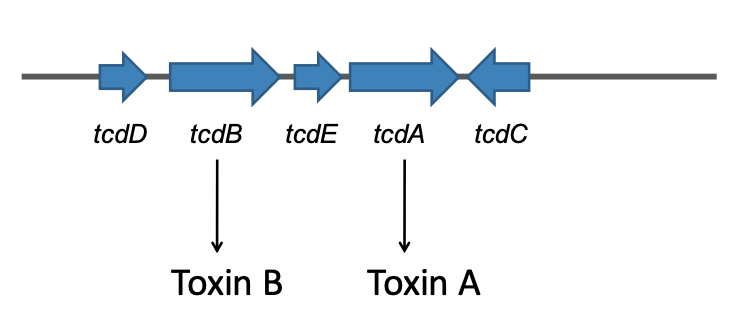
Pathogenicity locus of C.difficile: PaLoc
Glutamate dehydrogenase (GDH) EIA:
Fast and sensitive, but not specific (false-positives)
All C.difficile strains (toxic or not), other clostridial species and other bacteria produce GDH
Clinical Impact of an Ineffective Test:
Repeating the tests
Delay with prescribing an antibiotic against C.diff
Increase in severity/deterioration of the illness
Increase the length of stay
Time before isolation > exposure of other patients > transmission
Increase in number of infections
NEED A TEST FOR C.DIFFICILE DETECTION WITH MORE SENSITIVITY AND QUICK TURN AROUND TIME
C.Difficile detection by multiplex qPCR w/internal control:
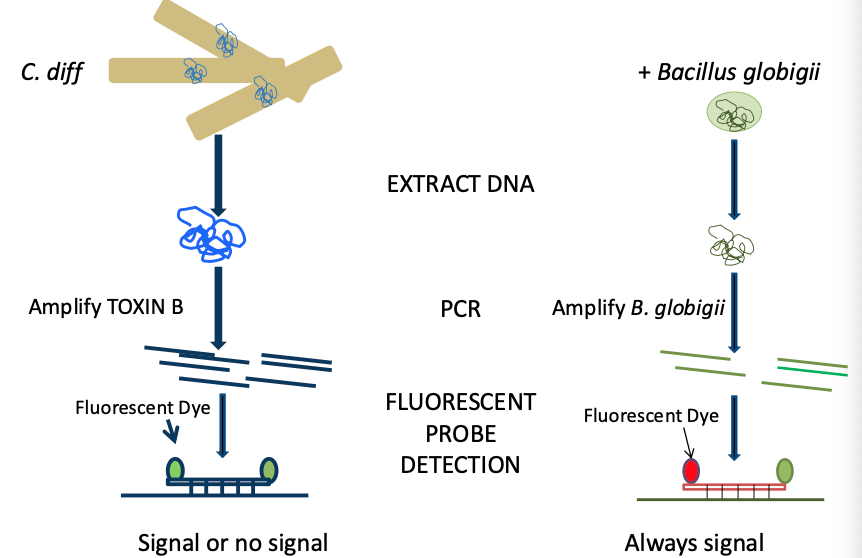
Internal Control for C.diff assay:
Add an internal control in each sample
spores of Bacillus globigii
Note: internal control tests the validity of each sample
Postive I.C
The nucleic acid extraction worked
Also checks presence of PCR inhibitors
Valid test
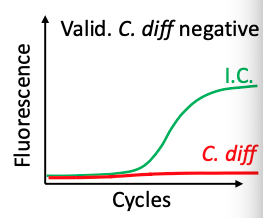
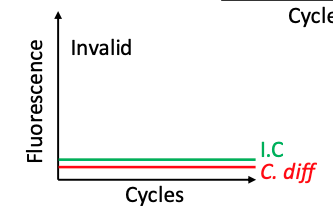
Negative I.C. C.difficile:
Invalid test
Sample needs re-extraction
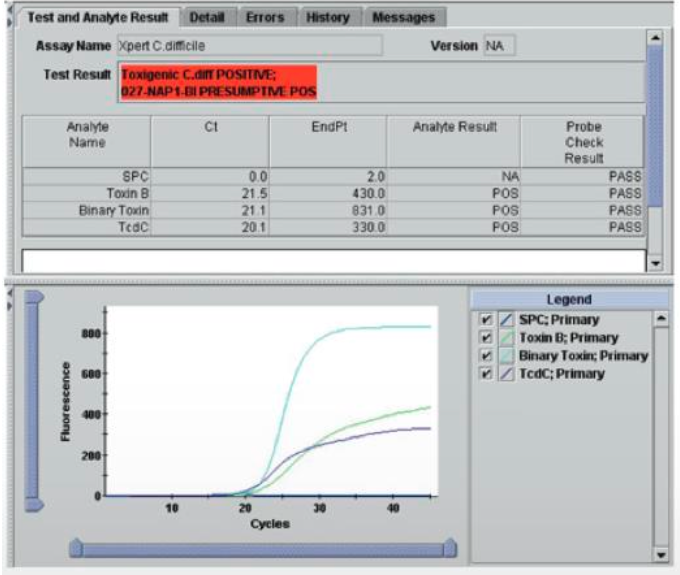
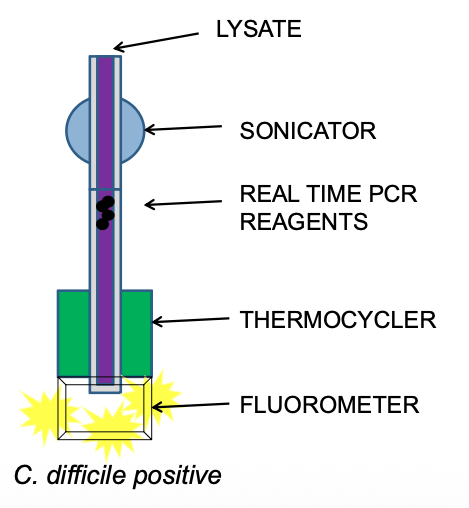
qPCR detection of C.difficile using GeneXpert (Cepheid):
Quick TAT: 50 min
Easy set up, automated (no highly trained personnel required)
Closed system (less contamination)
Detects toxB gene and internal control
High sensitivity (>94%) and specificity (>94%), NPV (98.8)
Expensive!

GeneXpert Assay: Inside the cartidge:
Closed system
Less contamination
Could be done in other sections of the lab
Result:
C.diff positive or negative
Current Epidemic Strain of C.difficile:
Increasing incidence and severity
Outbreaks of severe disease by the epidemic strain:
BI/NAP1/027, toxinotype III
Resistant to fluoroquinolones
Increased spore production
Carries extra toxin known as binary toxin (cdtB, cdtA)
Increased toxin A/B production due to a polymorphism in the regulatory gene (tcdC)
Pathogenicity Locus of C.difficile NAP-1/027:
Binary toxin is coded for by cdtA and cdtB genes outside PaLoc
1 bp deletion in tcdC inactivates repressor of tcdB
Increased tcdB transcription, i.e. Toxin B production
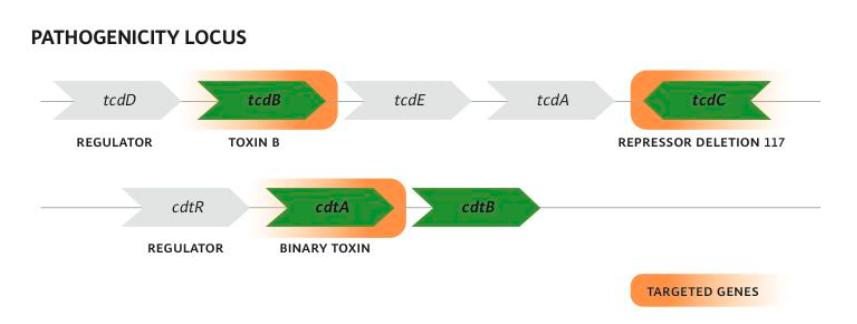
Positive Toxin B: Presumptive Positive B1/NAP1/027