BIO 331 - Structural and Functional Annotation
1/53
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
54 Terms
Genome annotation
Attaches biological information to sequences
Structural annotation
Process of identifying coding genes (an intron-exon structures) and non-coding genes (e.g. tRNAs)
Functional annotation
Attaches metadata structural annotations (e.g. which product is encoded in the gene)
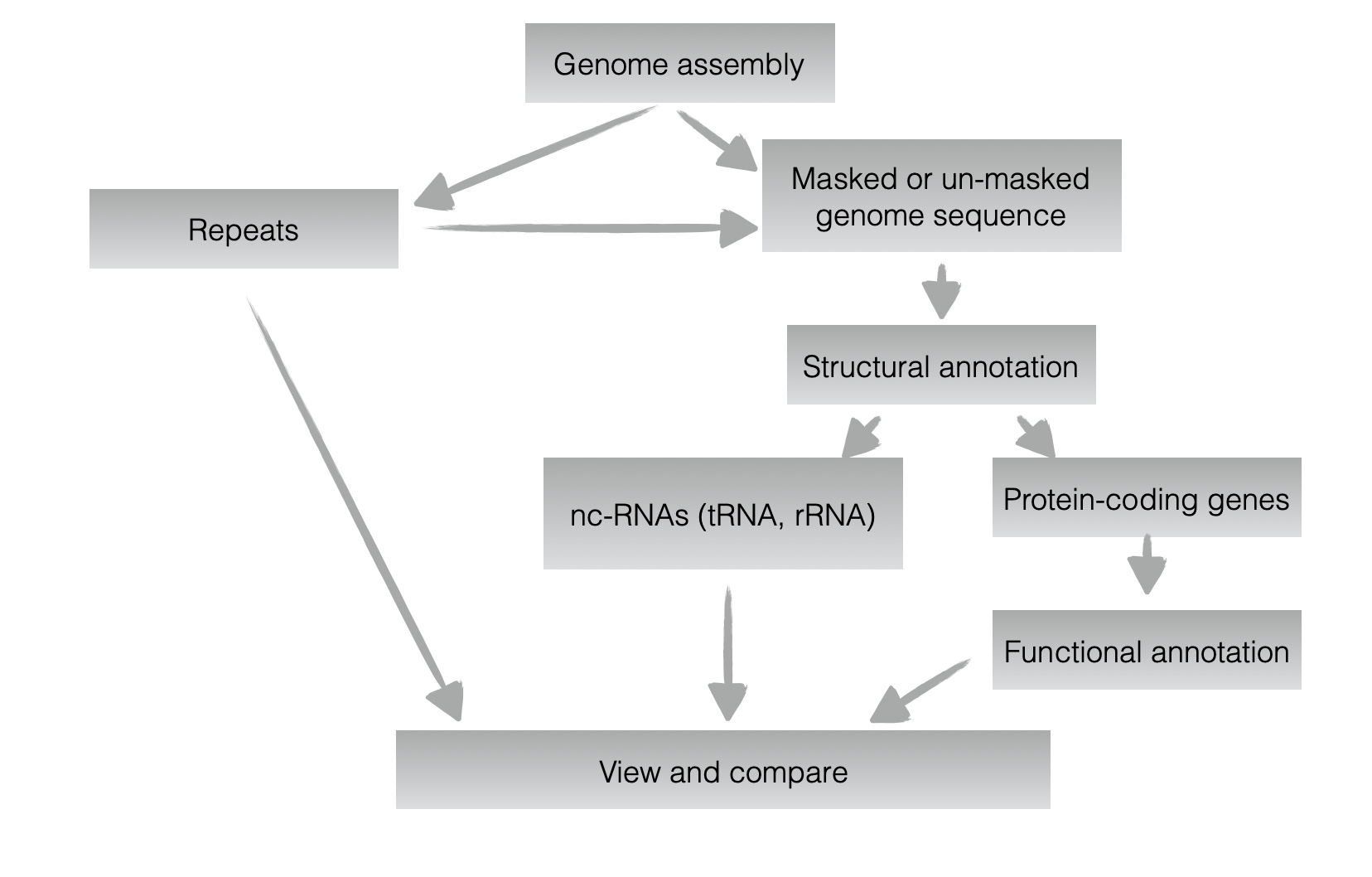
This image describes the workflow for ___
Genome annotation
Programs for finding repeats and masking/annotating them
RepeatMasker, emboss NUCLEIC REPEATS
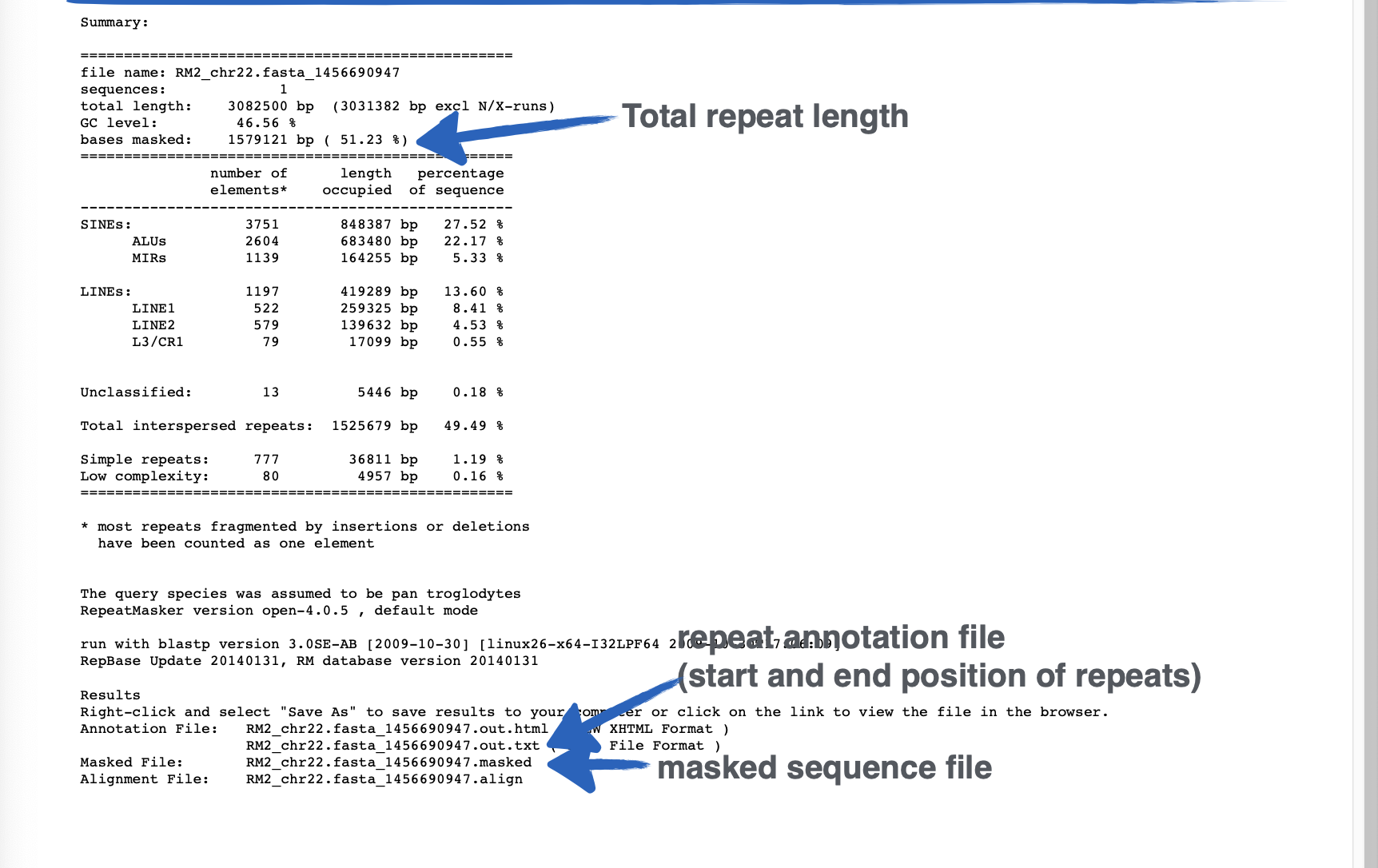
This output is from ___
RepeatMasker
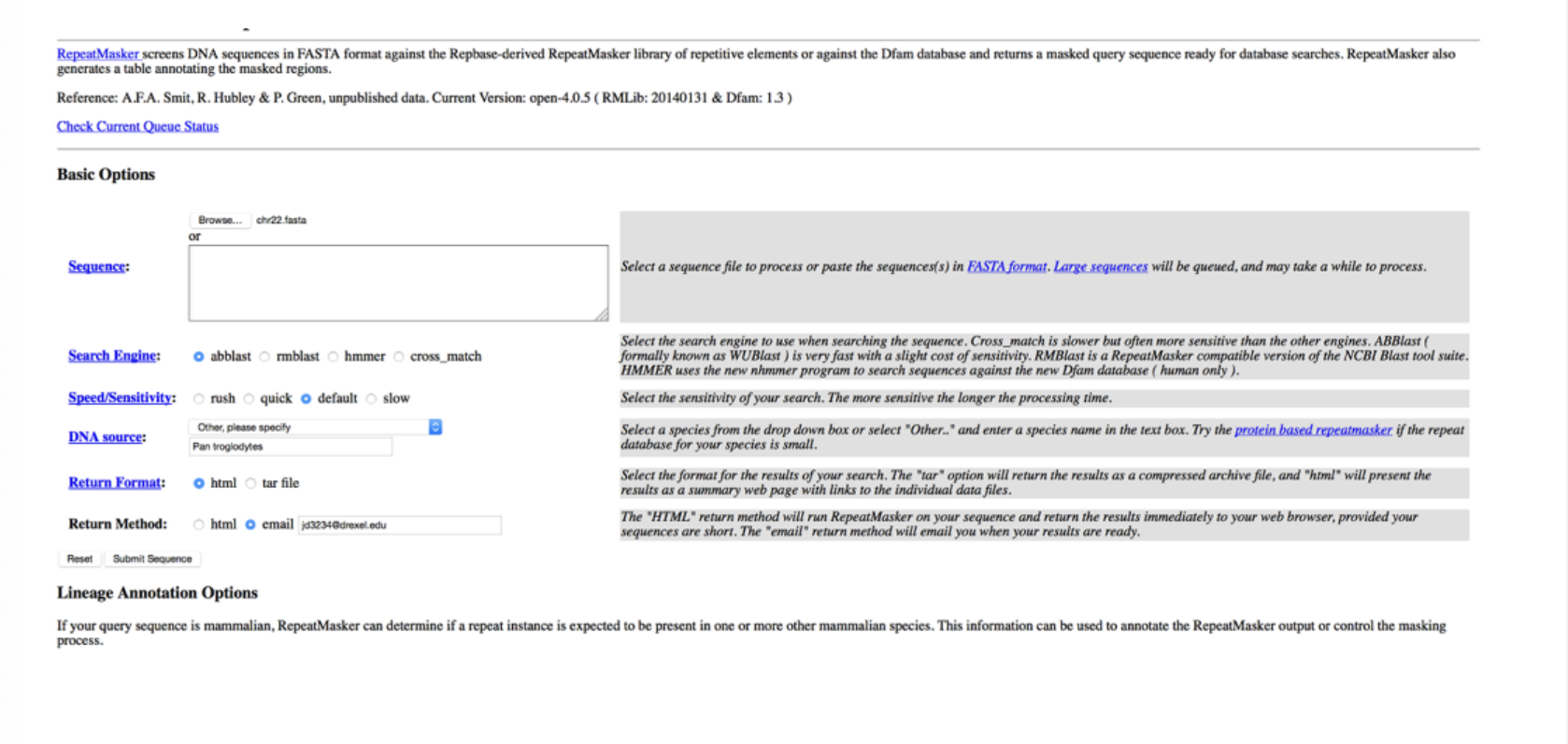
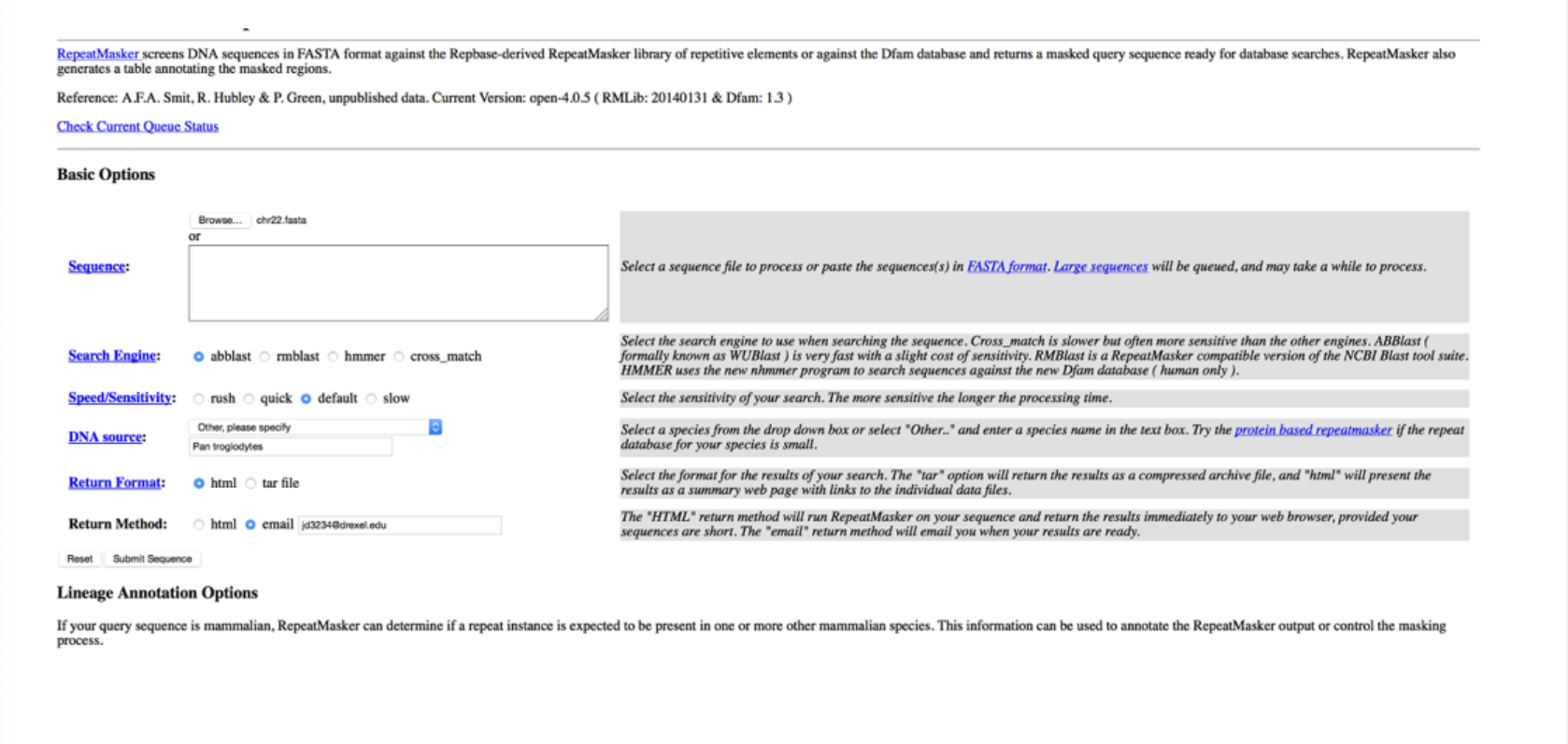
This program is ___
RepeatMasker
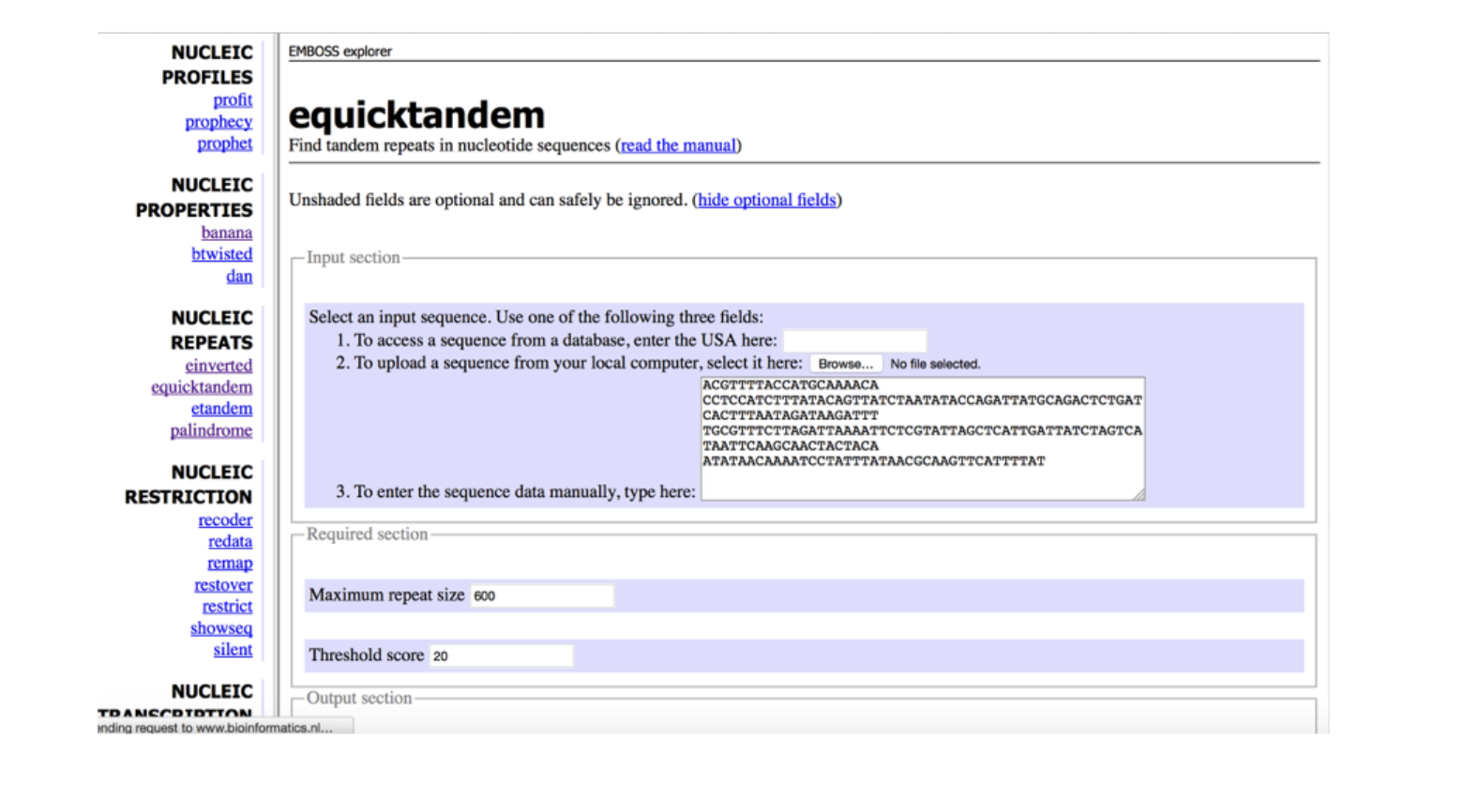
This program is ___
emboss NUCLEIC REPEATS
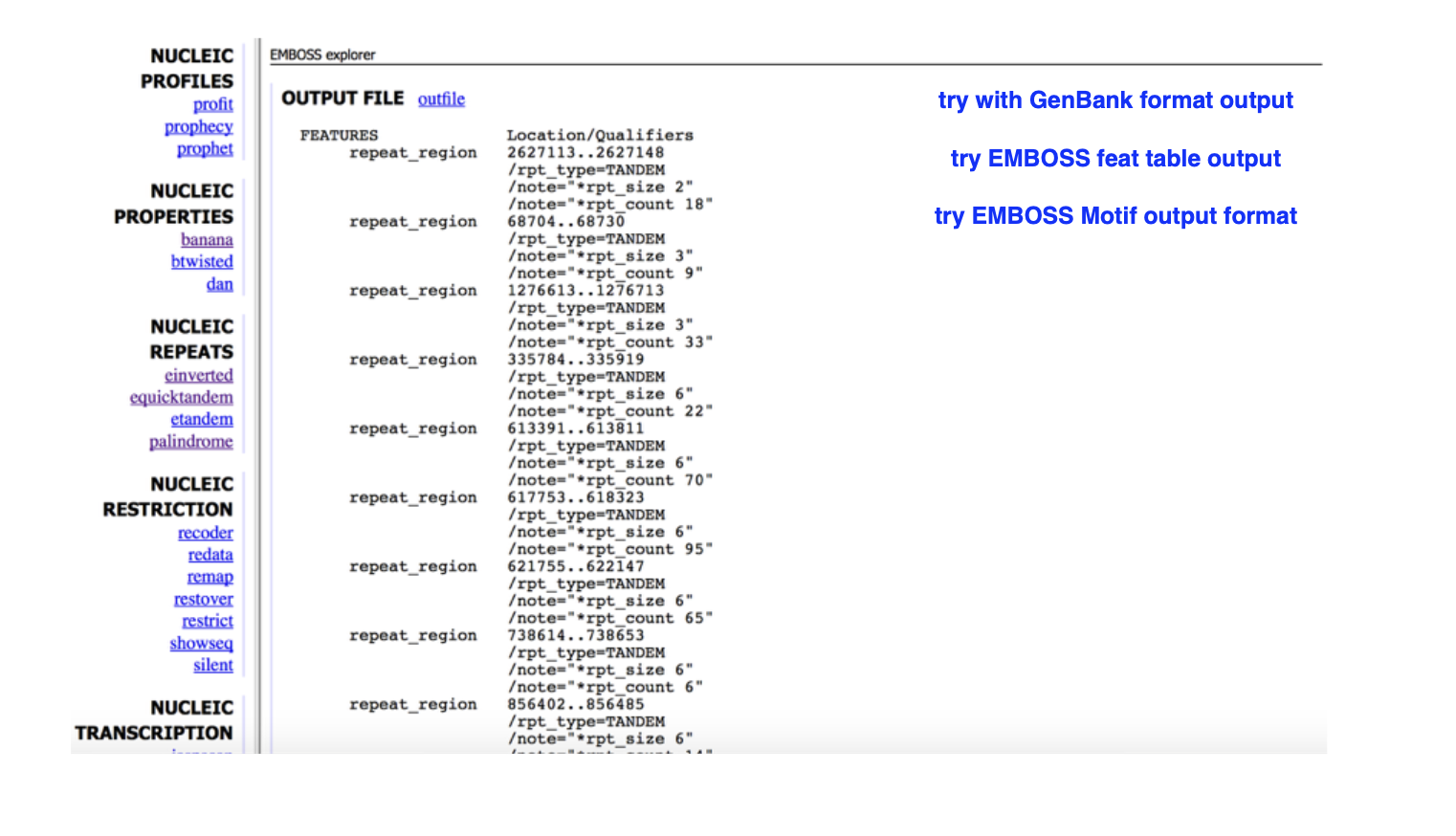
This output is from ___
emboss NUCLEIC REPEATS
Structural annotation helps identify these genomic elements
Non-coding genes, regulatory motifs/promoters, coding genes
ncRNAs
Non-coding RNAs; RNA molecule that is not translated into a protein

tRNAscan-SE
Finds tRNAs

RNAmmer
Finds rRNA
Rfam
Finds tRNA, rRNA, mtRNA, snRNA, miRNA, etc.
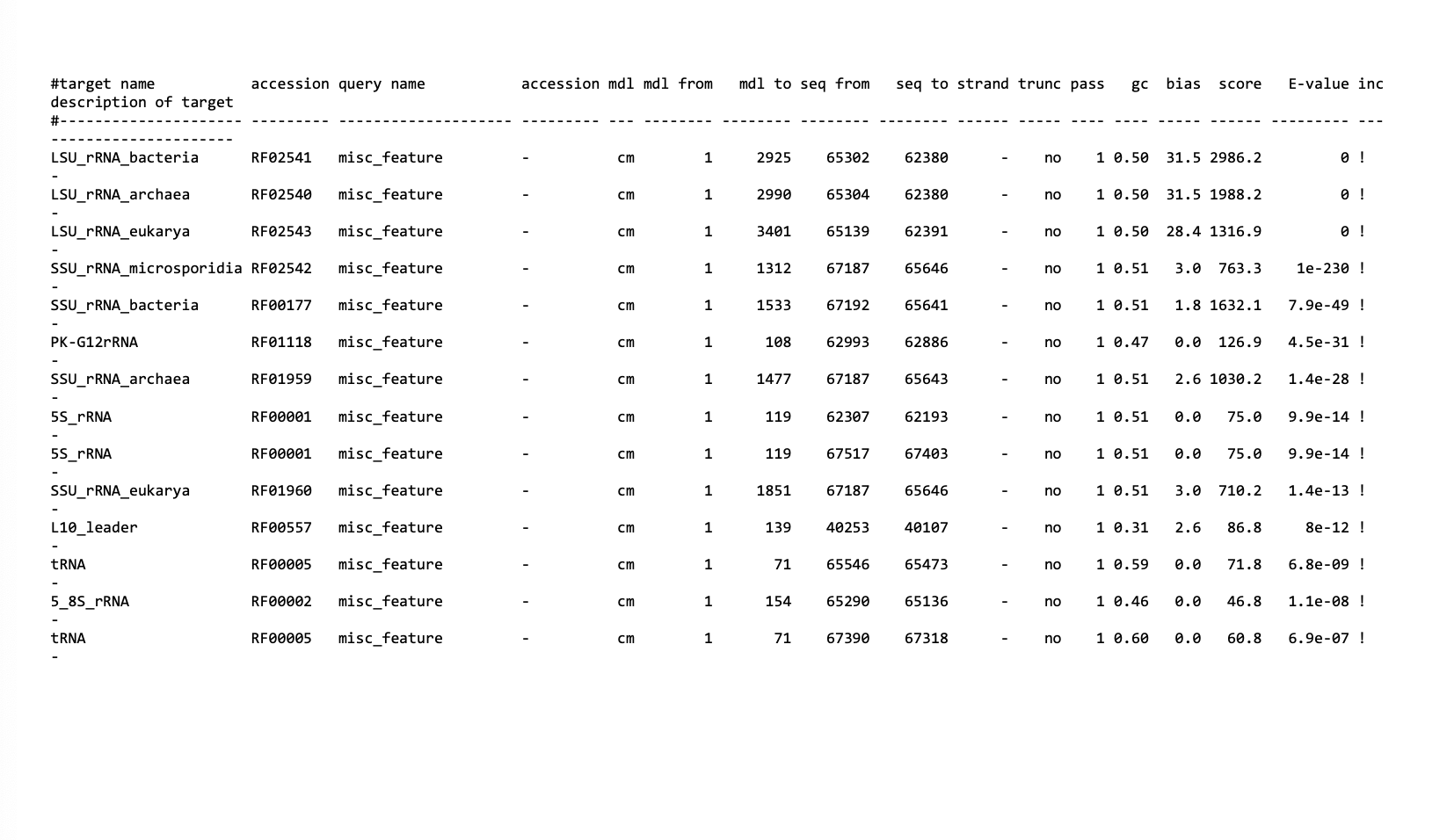
This output is from ___
Rfam
What does ab initio mean?
From the beginning
Ab initio signals
Specific sequences that indicate the presence of a gene nearby (e.g. promoter)
Ab initio content
Properties of protein-coding sequence itself (e.g. start & stop
codons)
Tools for ab initio discovery
GLIMMER, GeneMarkS, Prodigal, ORF finder
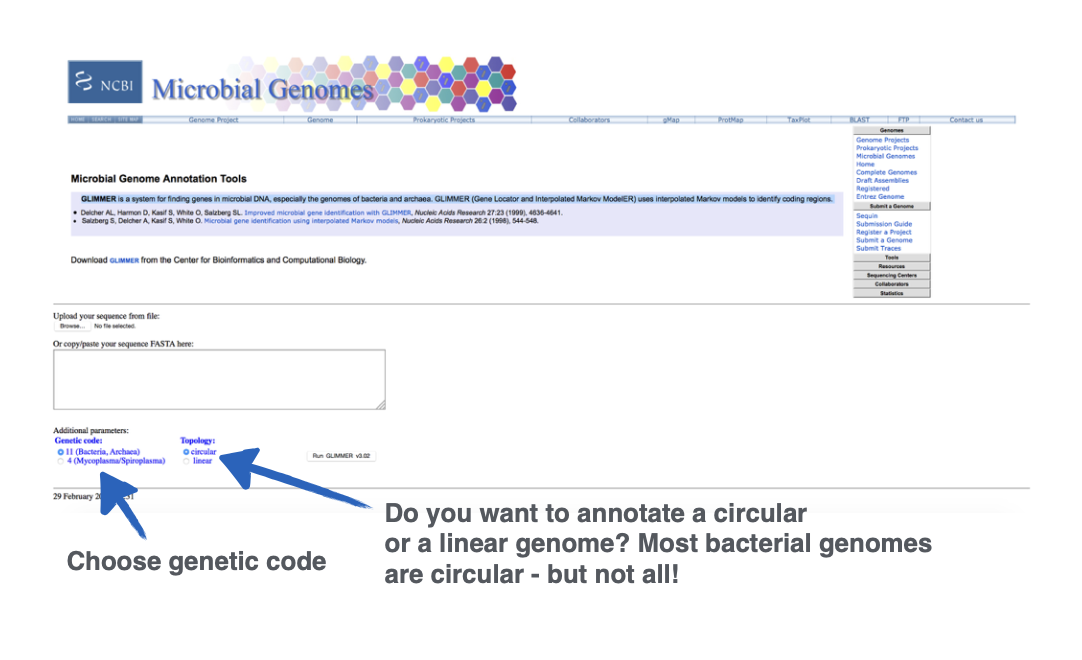
What site is this?
GLIMMER
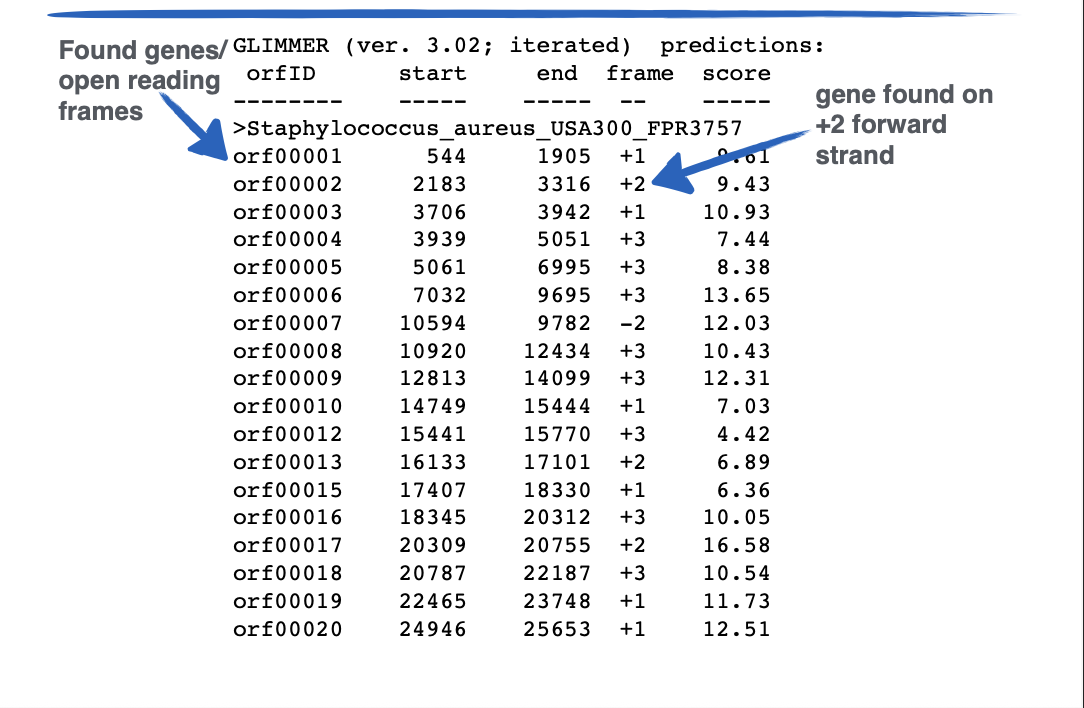
This output is from ___
GLIMMER
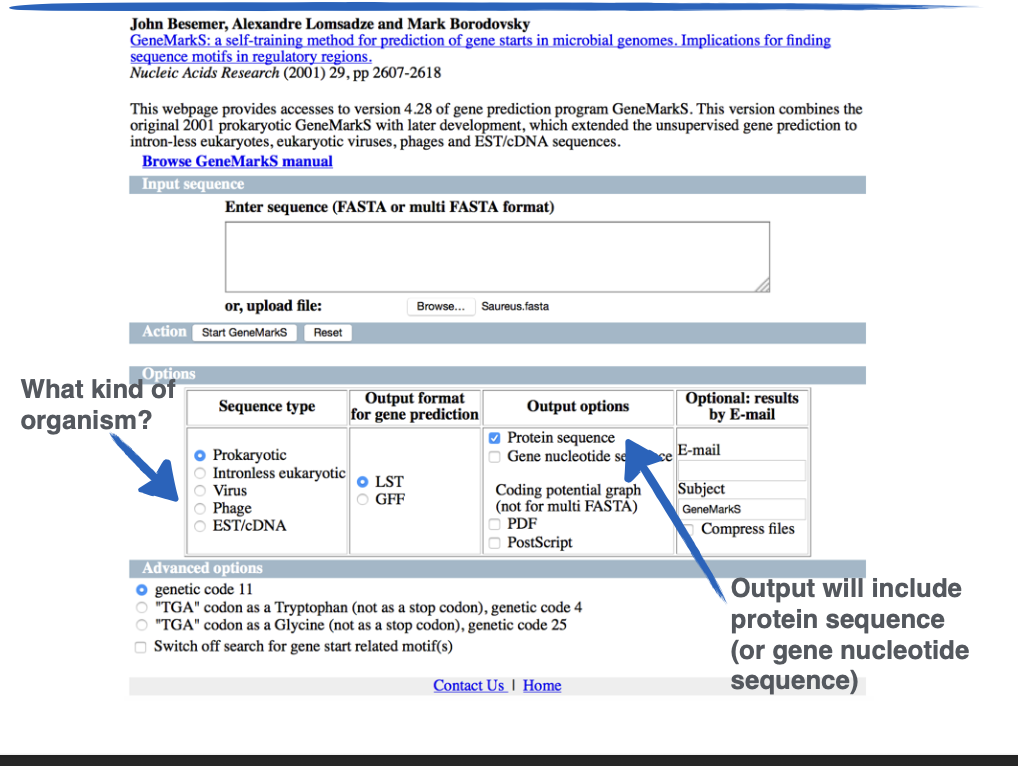
What site is this?
GeneMarkS
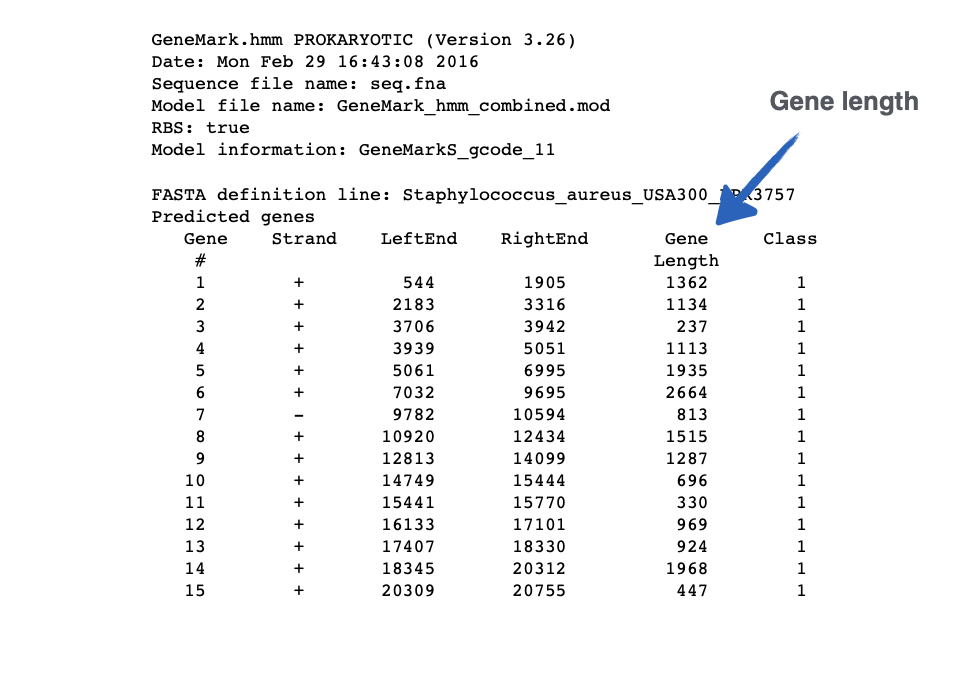
This output is from ___
GeneMarkS
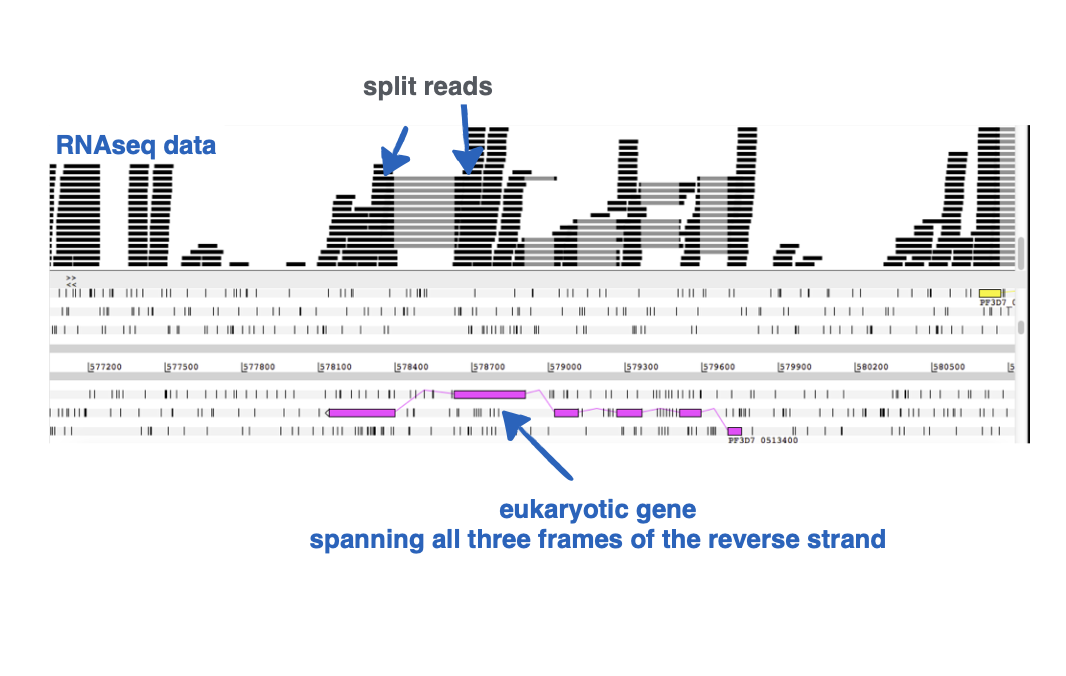
Why do split reads exist in eukaryotic RNASeq data?
Intron splicing
Average vertebrate gene is ___ kb long
30
Average vertebrate coding sequence is approx. ___ kb long
1
Average vertebrate coding region consists of ___ exons of about ___ bp each
6, 200
Exon boundaries can be defined through ___
RNASeq
FOr highest accuracy, one should use ___ data rather than screening public databases
Experimental
Databases for determining gene function
UniProt, RefSeq, Pfam/TIGERFAM, user-provided set of annotated proteins
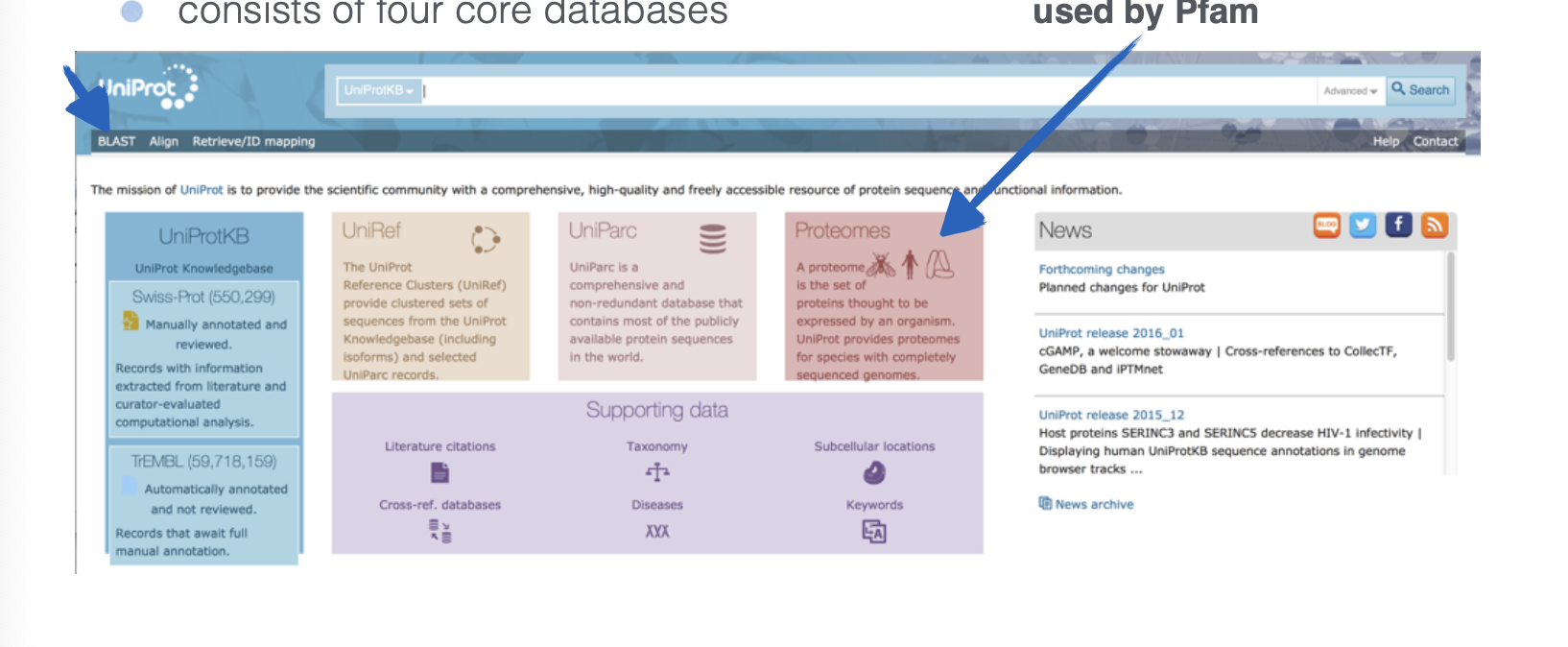
What website is this?
UniProt
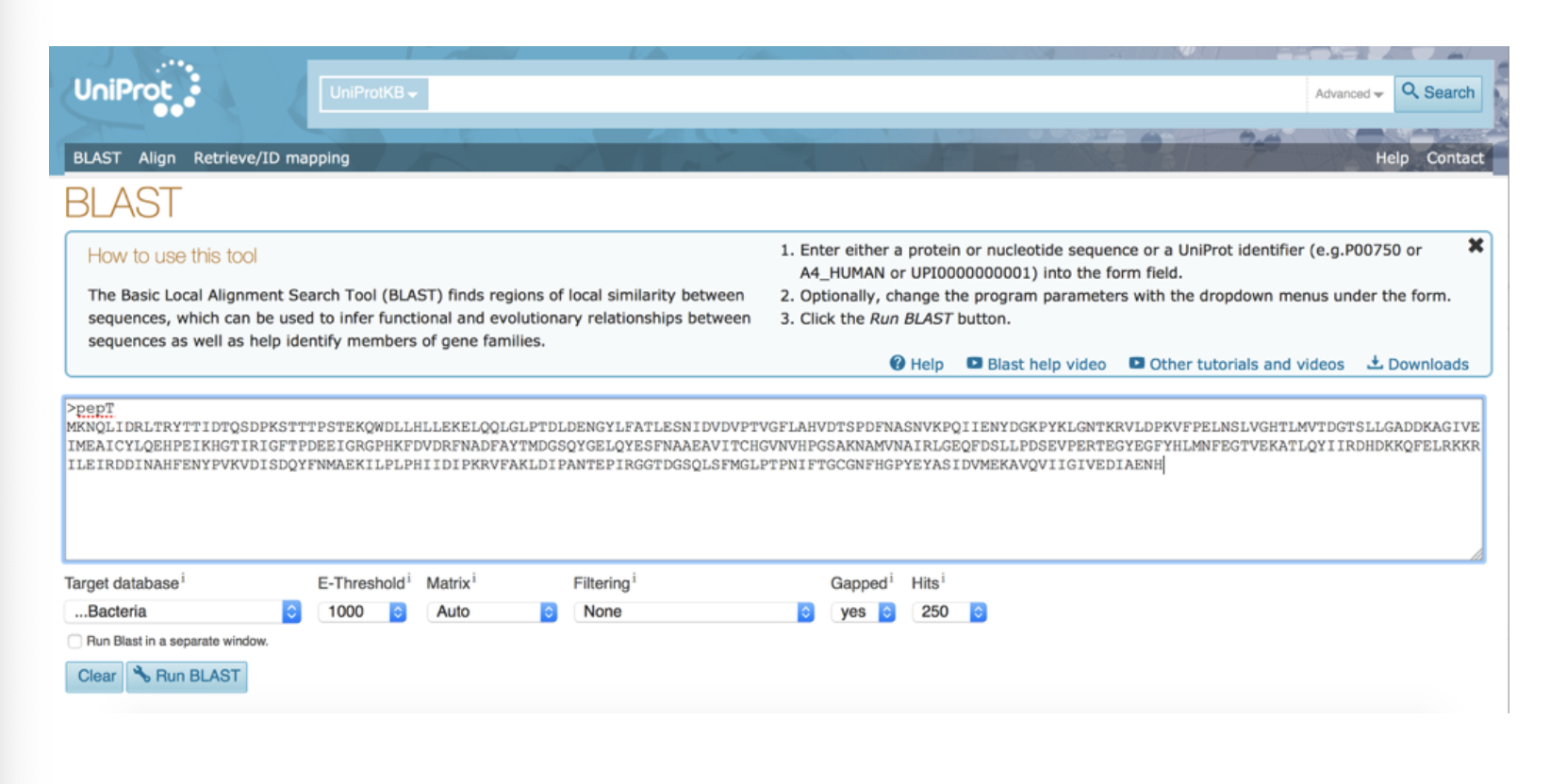
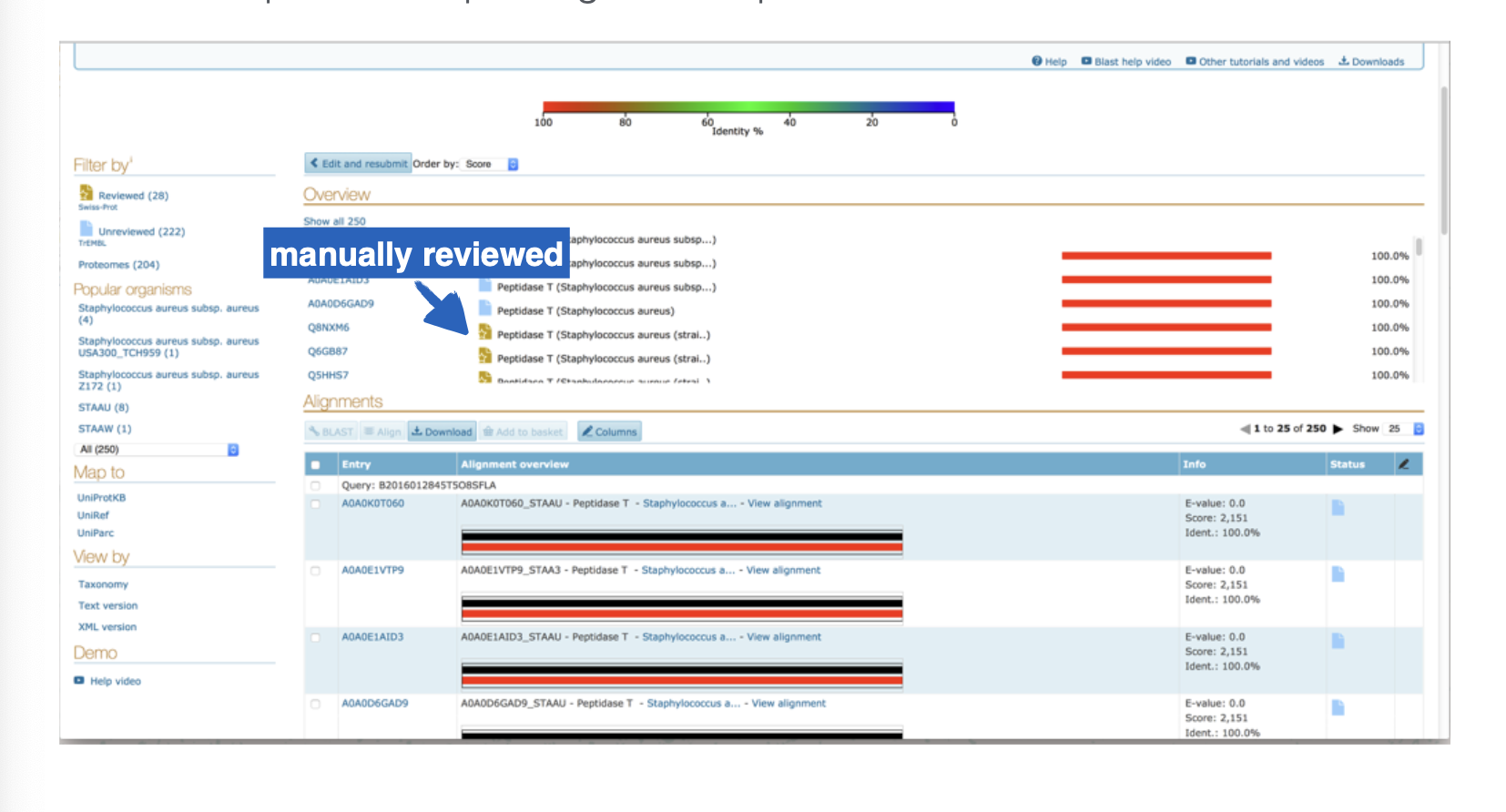
This output is from ___
UniProt
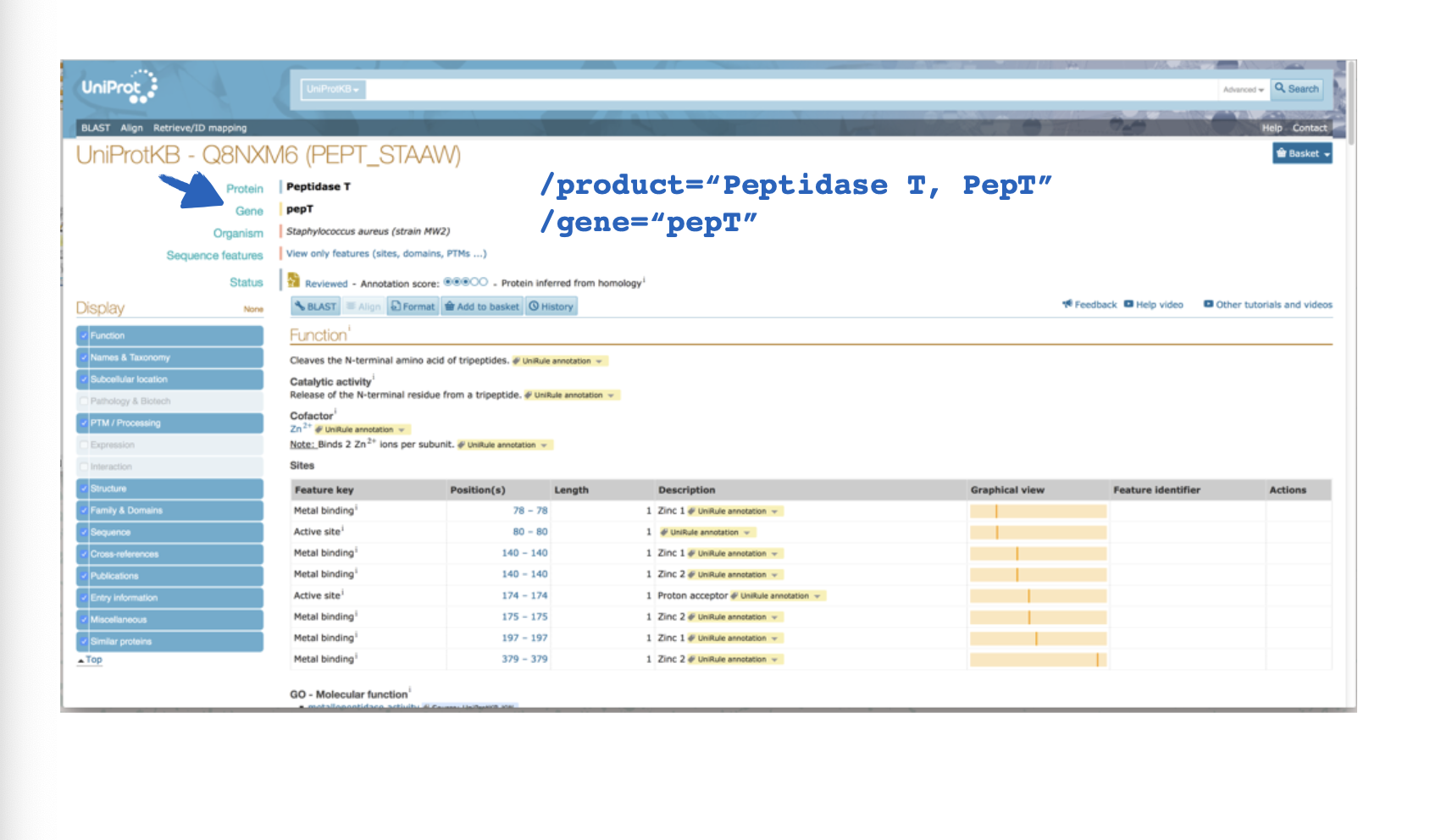
True or false, UniProt provides hits to specific proteins?
True
Pfam is used to identify the ___ your protein belongs to
Family
Superfamily vs subfamily
Superfamily: large group of distantly related proteins
Subfamily: small group of closely related proteins
Building Pfam families
Seed alignment used to build a profile HMM
Profile HMM is searched against sequence databases
All matches scoring equal to or grater than a given threshold are considered as true members of the protein family
These members are added to the seed alignment to generate the full alignment and from there a consensus HMM and a consensus sequence
HMMs
Hidden Markov Models; used in many BI applications (gene/protein prediction, phylogenetic analysis, alignments)
HMMs are systems that move from state to state with ___ probabilities. Each state produces a new possible outcome.
Finite
Issue with HMMs
States that are responsible for a possible outcome are unobservable; only final outcome is observable
Two probabilities associated with HMMs
Transition and emissions
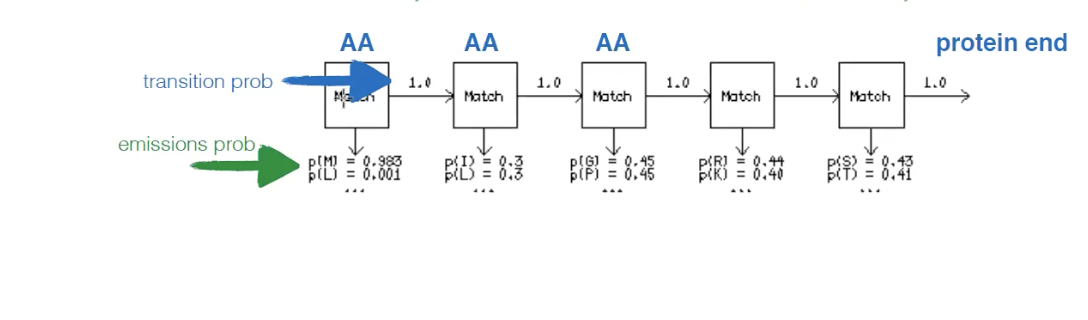
Transition probability
Probability of transitioning from one state (e.g. AA) to another
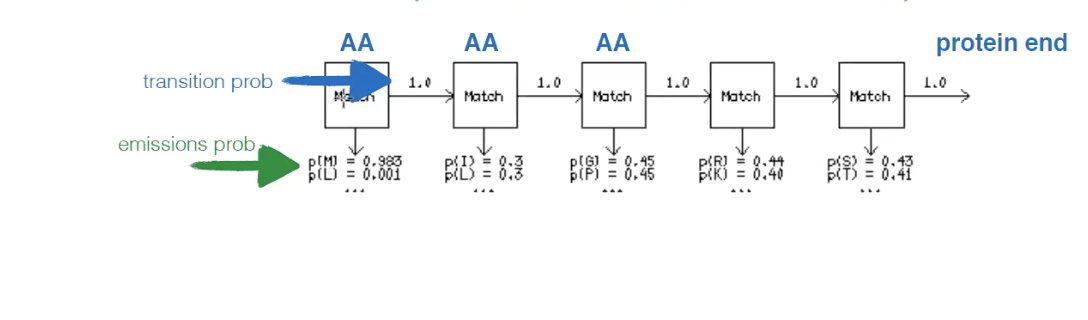
Emissions probability
Probability that a given AA exists at that position in the alignment
Comparing highly divergent protein sequences is best achieved through ___
HMMs
BLAST is based on ___ comparisons
Pairwise
Seq2EC
Takes an unknown protein and can predict the EC that would match the protein
BRENDA
Takes EC number and gives names of the protein that is usually assigned
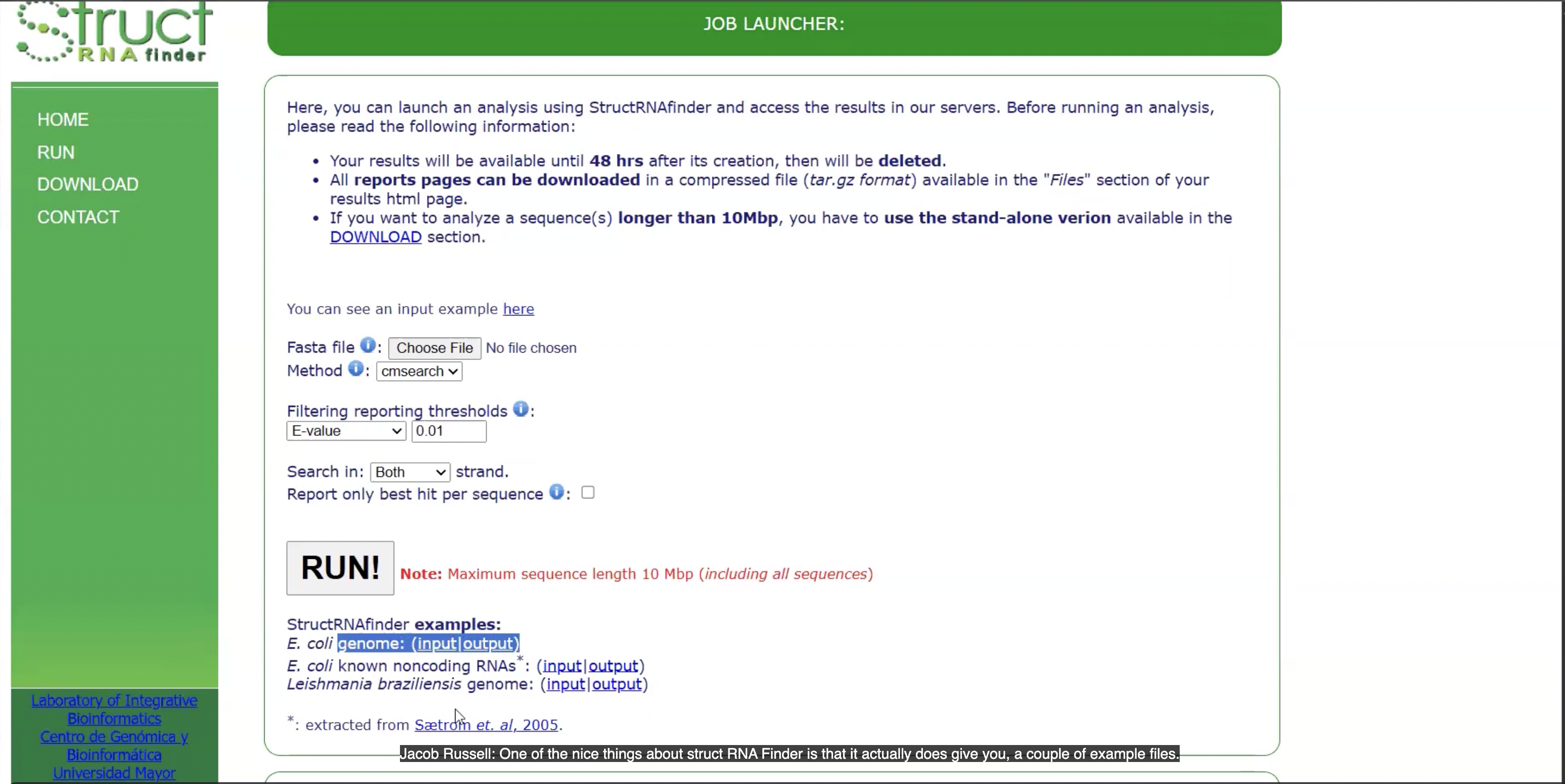
This website is ___
StructRNA finder
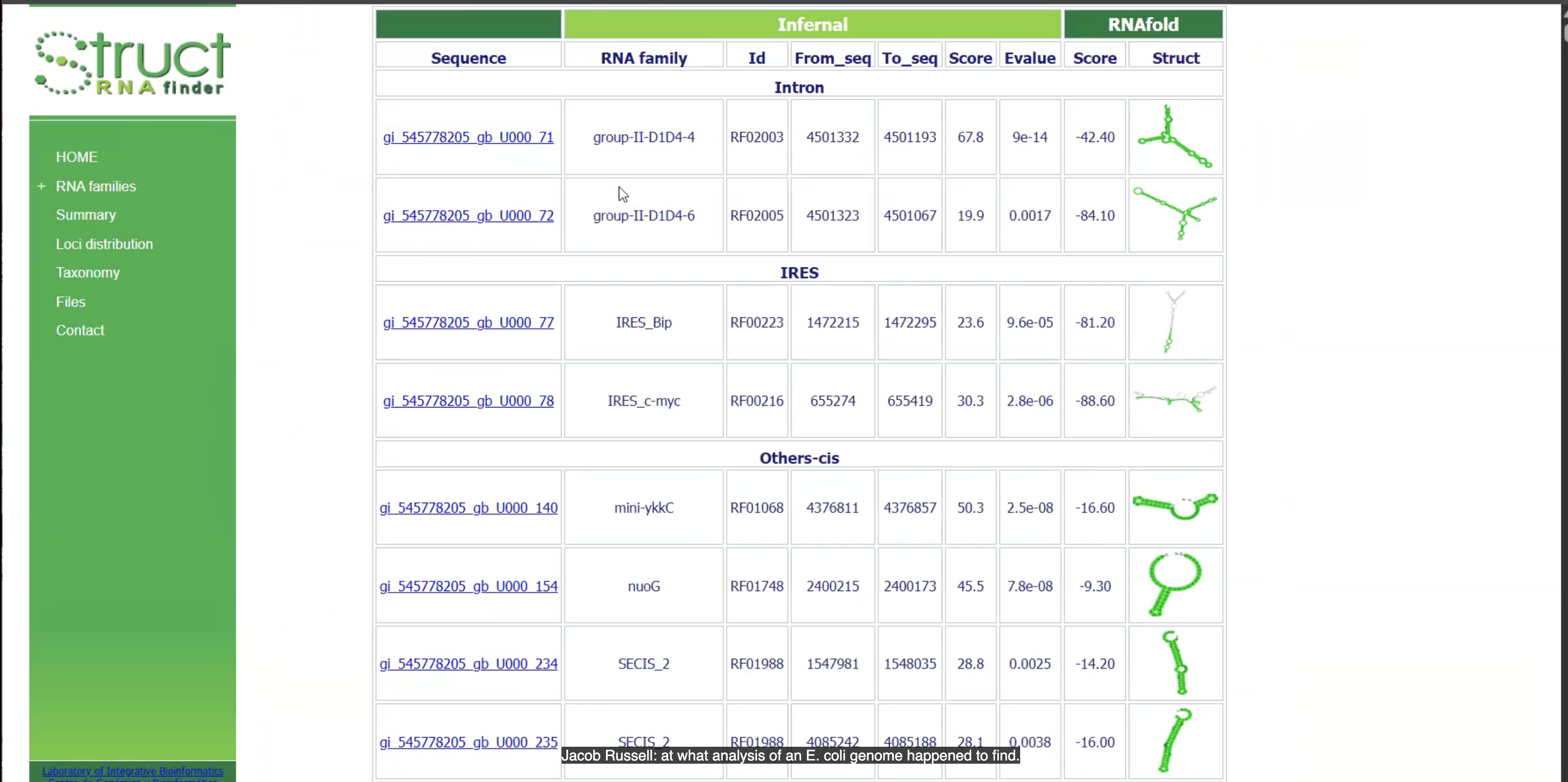
This output is from ___
StructRNA finder
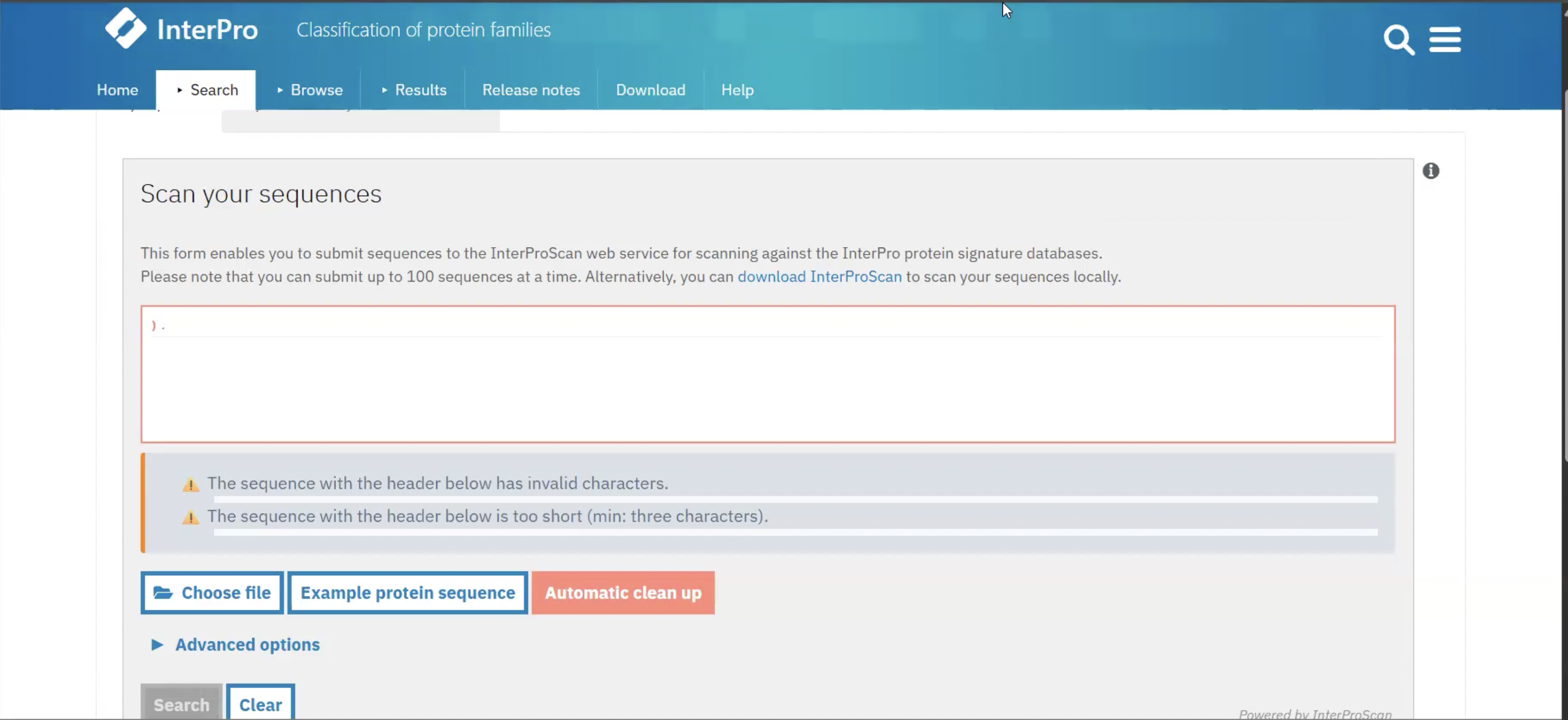
This website is ___
InterPro scan
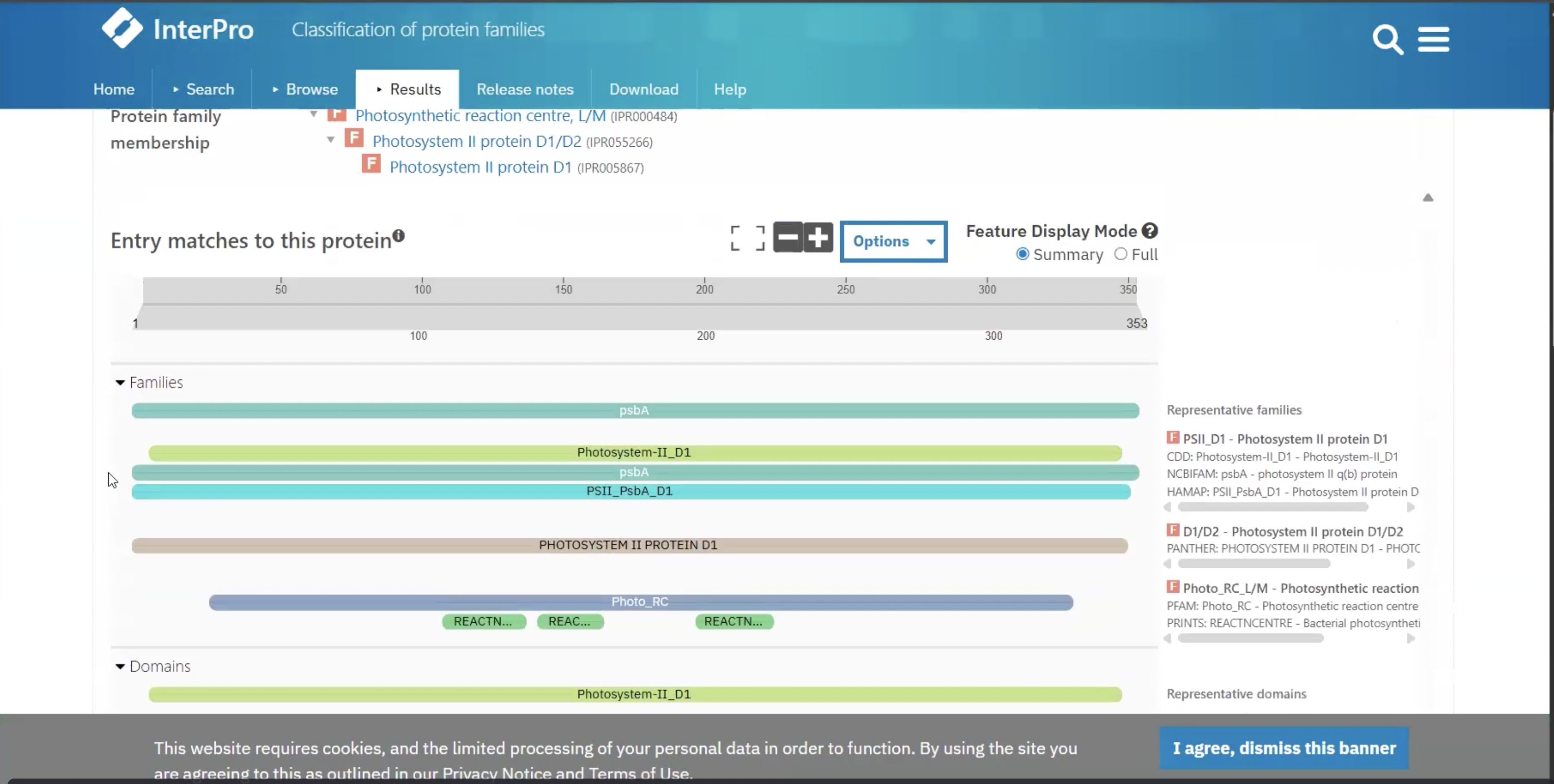
This output is from ___
InterPro scan
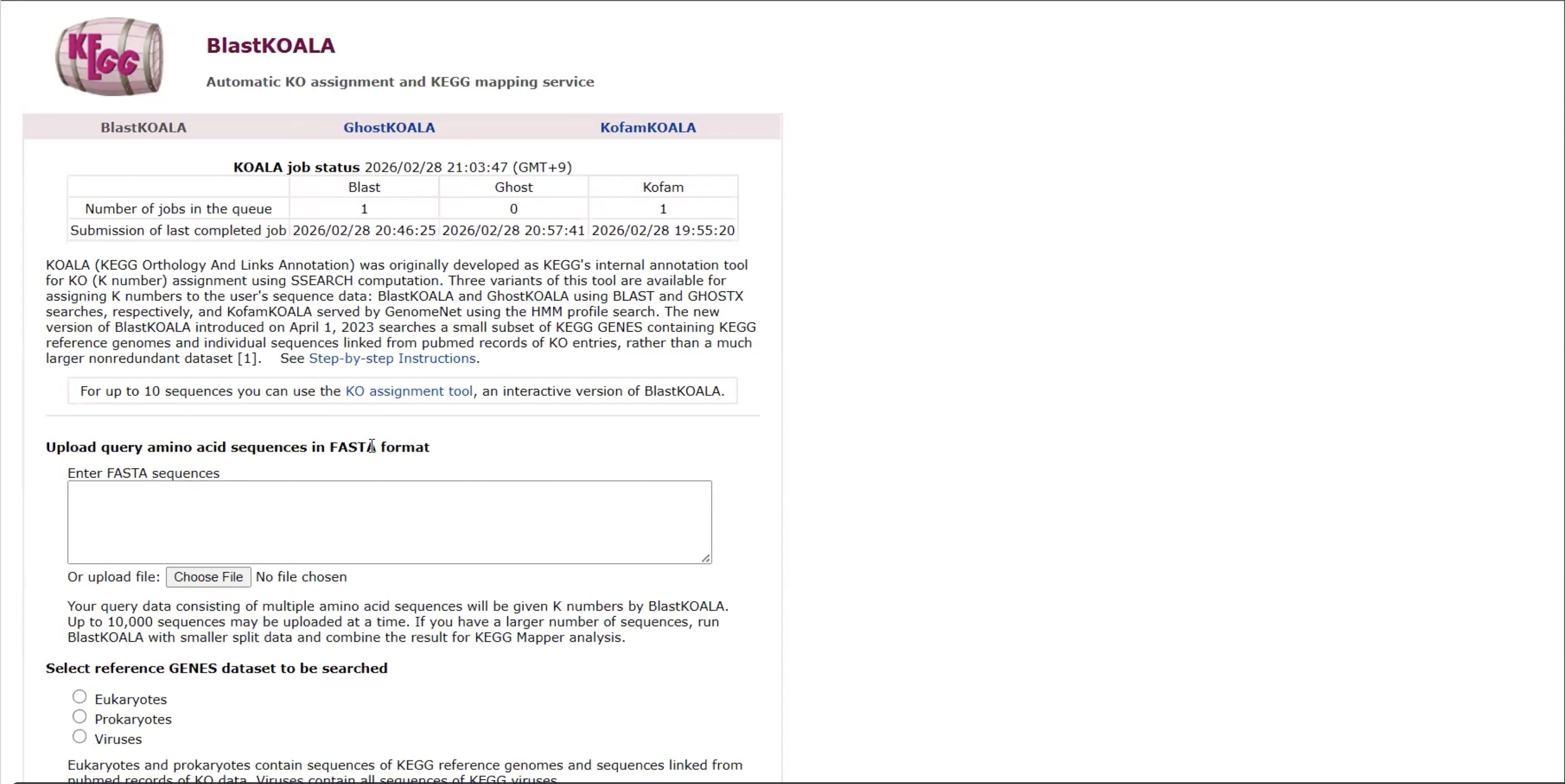
This website is ___
BlastKOALA
BlastKOALA provides information about ___
KEGG orthologs (KOs)
InterPro scan provides information about ___
Gene ontology (GO)
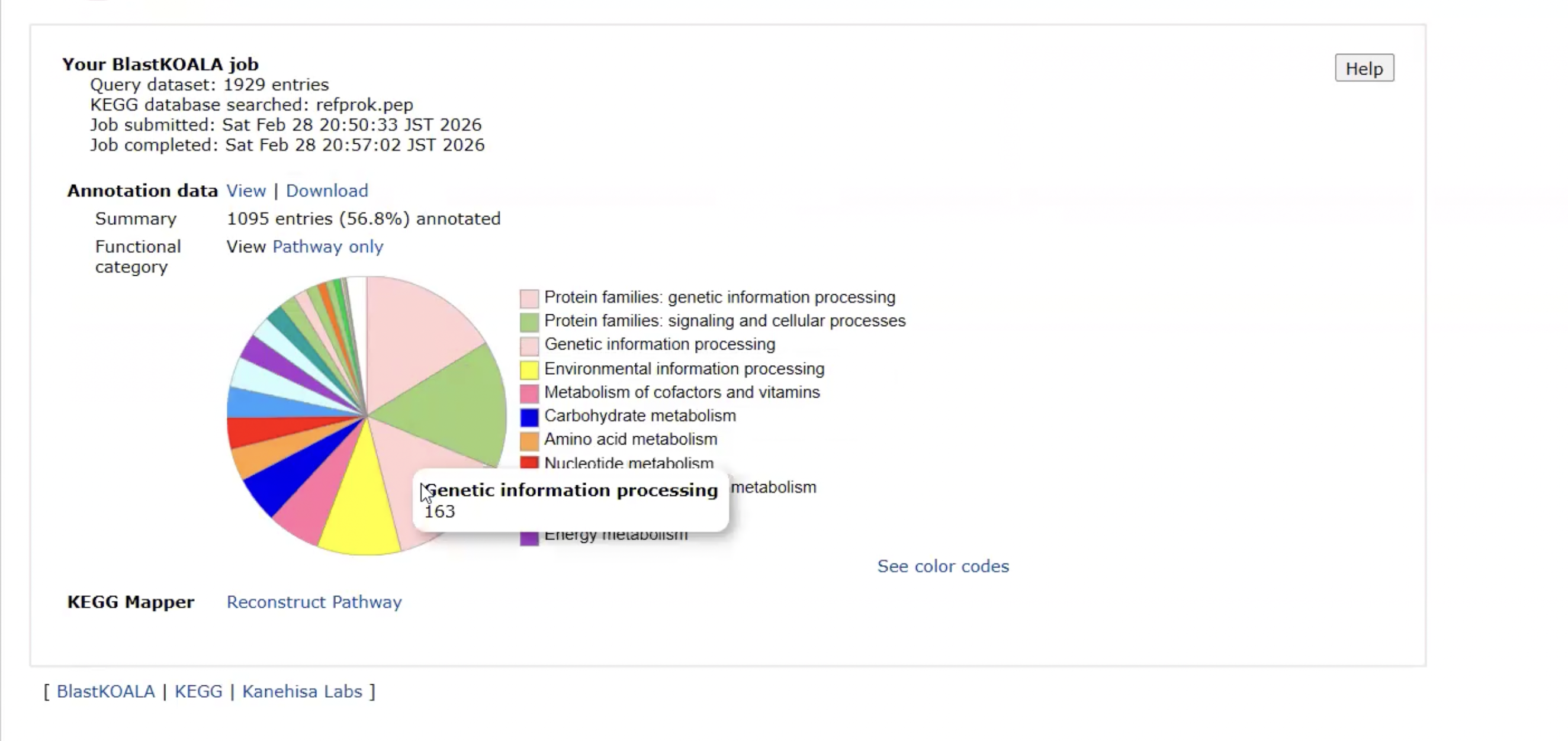
This output is from ___
BlastKOALA