Ch. 13: Translation and Proteins
1/36
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
37 Terms
Translation
produces a polypeptide using the information in mRNA
Gene
-a specific sequence of nucleotides that contains in the information required to build a protein or RNA molecule
-Types of genes
Structural genes encode proteins
rDNA encode rRNA and tRNA
Regulatory RNA controls gene expression
Translation of mRNA
-the polymerization of amino acids into a polypeptide chain
-Translation occurs on ribosomes in the cytoplam
Transfer RNA (tRNA)
-the molecule that matches a triplet codon with the correct amino acid
-short molecules with 75-90 base pairs
-Transcribed by RNA polymerase III
-Each tRNA has an anticodon complementary to a codon in mRNA
-Charged tRNA have an amino acid covalently bound to the 3’ end
-Binding between the codon and anticodon of a charged tRNA directs the incorporation of amino acids into a growing polypeptide chain
Degree of folding
-all tRNA have 3 levels of structure
-Primary structure
-Secondary structure
-Tertiary structure
Primary structure folding
75-80 nucleotide sequence
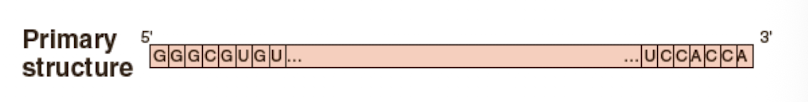
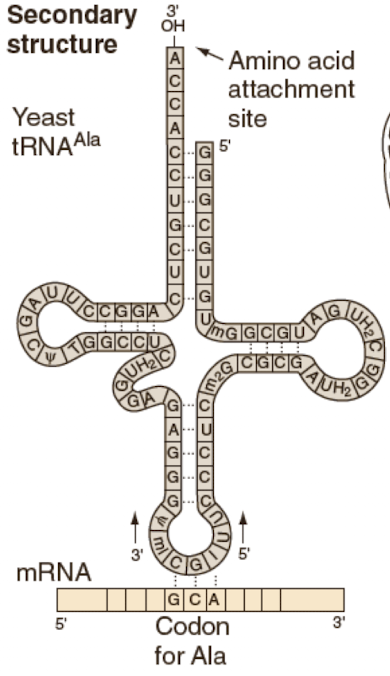
Secondary structure
-cloverleaf shape with 3 stem loops
-formed because of short complementary sequences within the tRNA
Tertiary structure
-L shaped
-formed by 3-dimensional folding
Charging
-before translation can begin, tRNA molecules must be chemically linked to their respective amino acids
-occurs under the direction of enzymes called aminoacyl tRNA synthetase
-Each synthetase recognizes ONE amino acid and the structural features of its corresponding tRNA, including its anticodon
Base pairing between an mRNA and a tRNA anticodon determines
which amino acid is added to a growing polypeptide
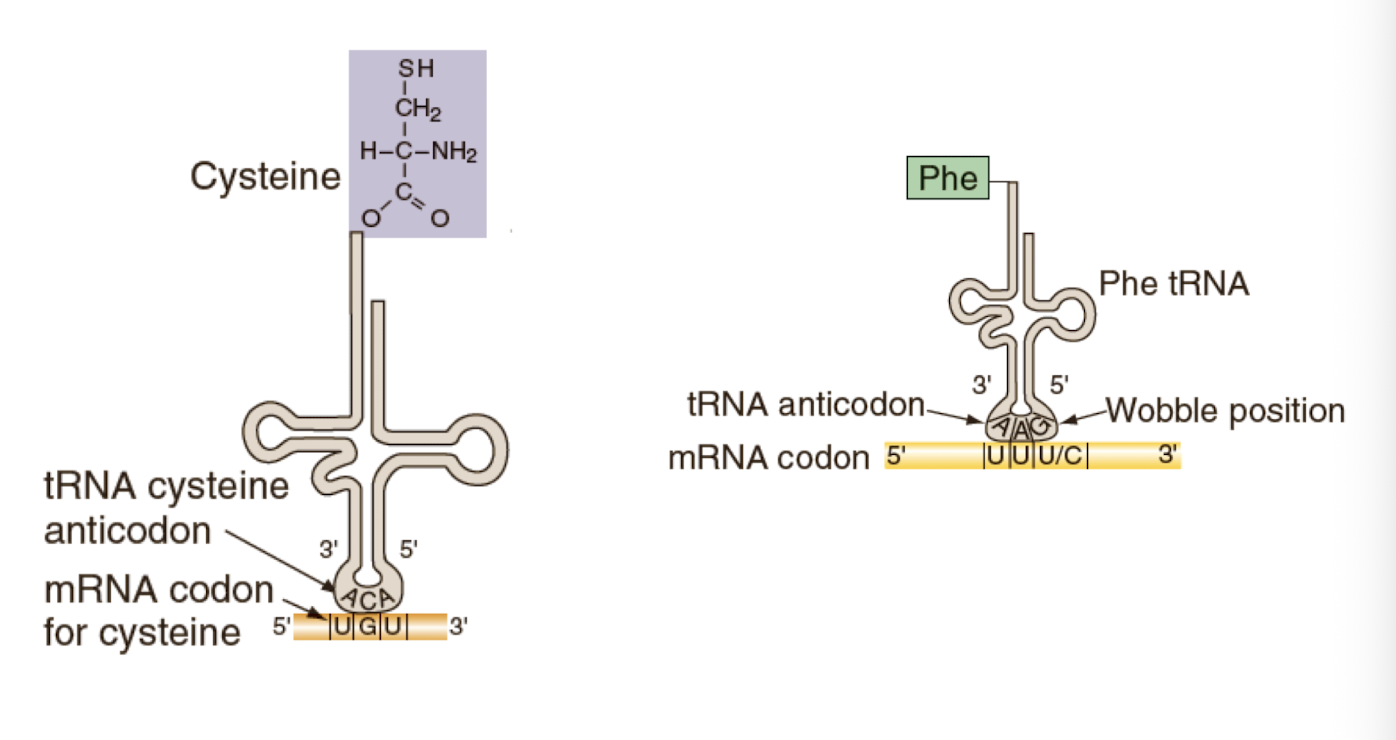
tRNA are named according…
-to the amino acid they carry
-Anticodons are complementary and antiparallel to the codon
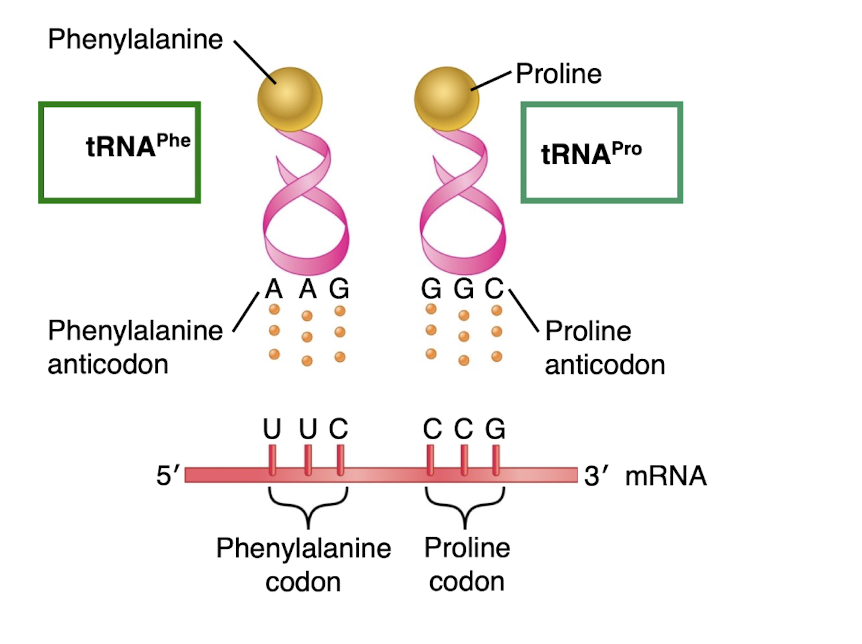
Translocation
-occurs in the cytoplasm on ribosomes
-each with 2 subunits composed of RNA and protein
RNA polymerase I transcribes…
-most rRNA
-Gene clusters (rDNA) are moderately repetitive DNA sequences near the ends of chromosomes 13, 14, 15, 21, 22
RNA polymerase III transcribes…
-5S RNA
-Gene clusters are on chromosome 1
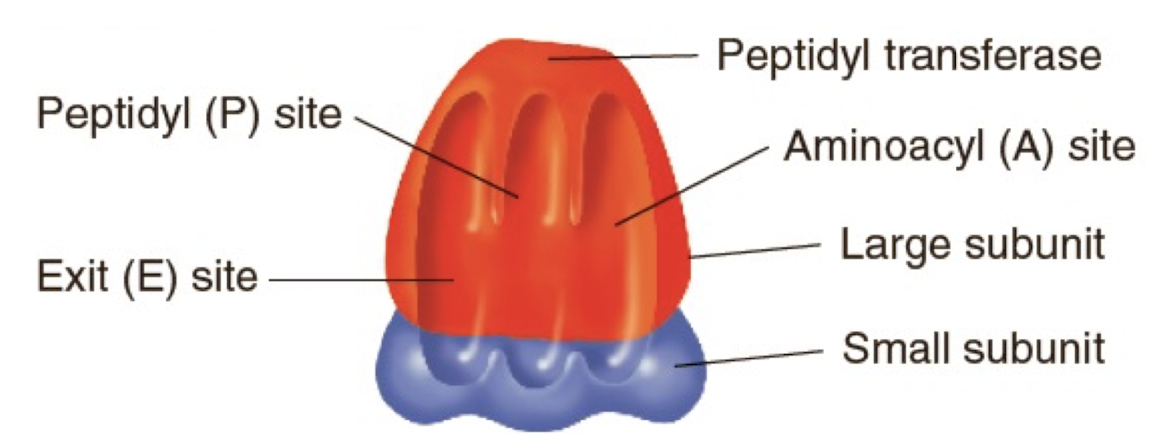
Different parts of a ribosome have different functions
-Small subunit binds to mRNA
-RNA in the large subunit has peptidyl transferase activity
It is a ribozyme that catalyzes peptide bond formation between amino acids
-3 distinct binding sites- E, P, and A sites
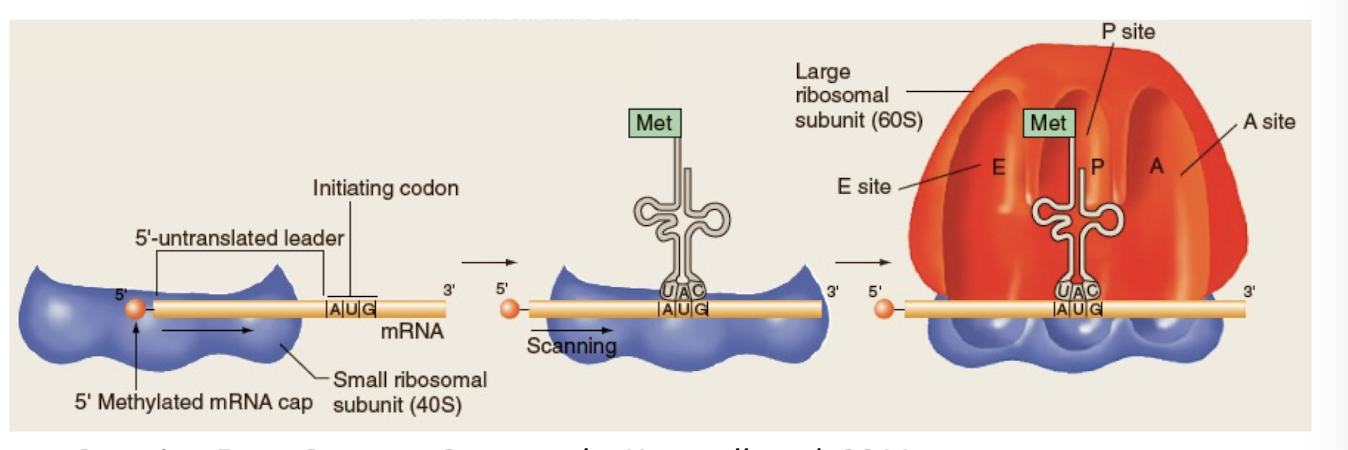
Initiation
-when translation begins in eukaryotes
-requires proteins called initiation factors (IFs)
-IFs bind to small subunit and the 5’ untranslated leader sequences position the mRNA, which is scanned for the 1st AUG codon
-tRNA with complementary anticodon loop binds mRNA
-Large subunit joins the complex, positioning tRNA at the P site
Elongation
-new tRNAs enter the A site and a peptide bond is formed between the 1st 2 amino acids
-Ribosomes move in a 5’ to 3’ direction along the mRNA
-Elongation factors (EFs) are required
-Uncharged tRNA moves to the E site and the complex shifts so the tRNA containing the polypeptide is now in the P site
Termination
-occurs when a stop codon is in the A site
-No tRNAs carry anticodons for stop codons
-Release factors bind to the stop codon and the ribosomal subunits, mRNA and the polypeptide disassociate
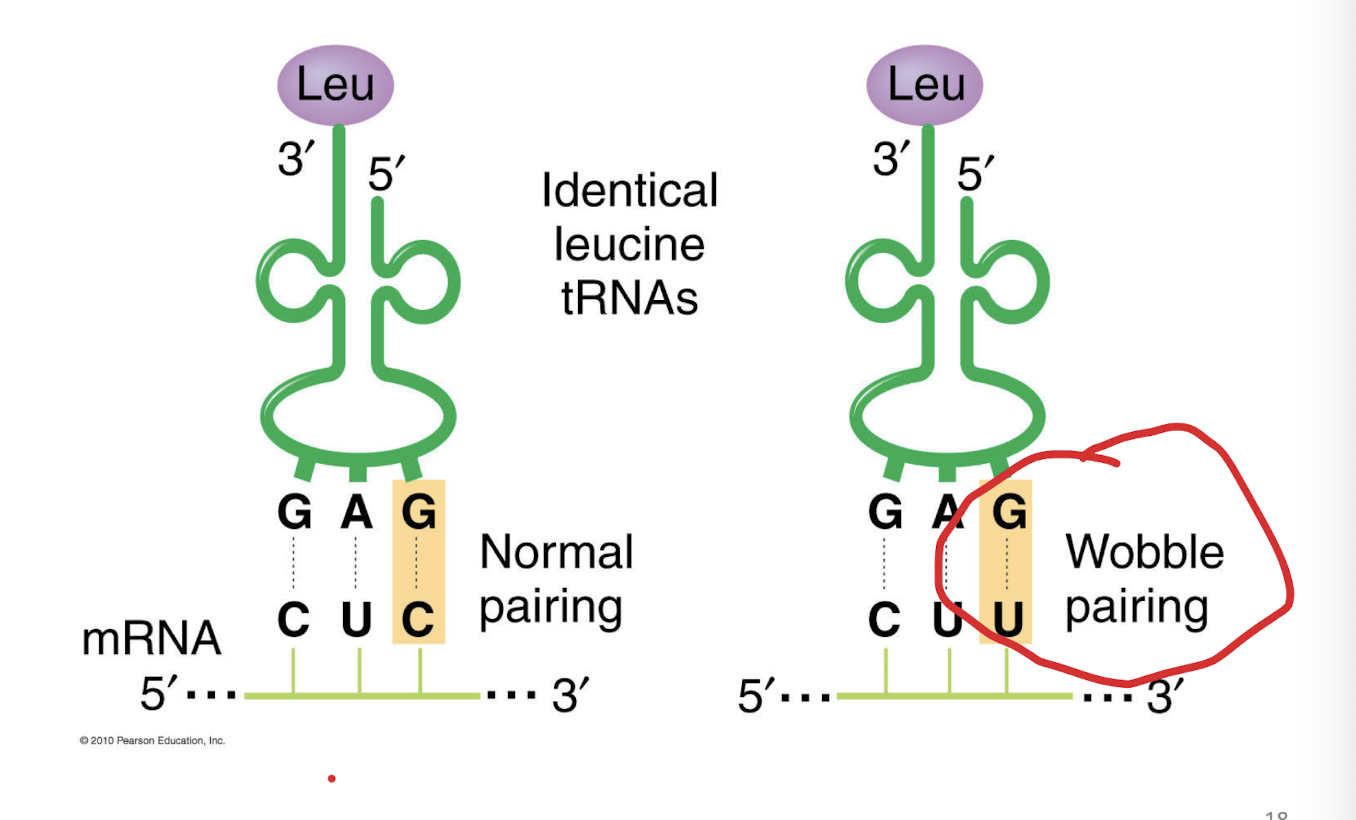
The wobble position
the third base (3' end) of an mRNA codon and the first base (5' end) of the tRNA anticodon
Third position in the triplet, when different bases makes the same amino acid, the third ring on the codon circle
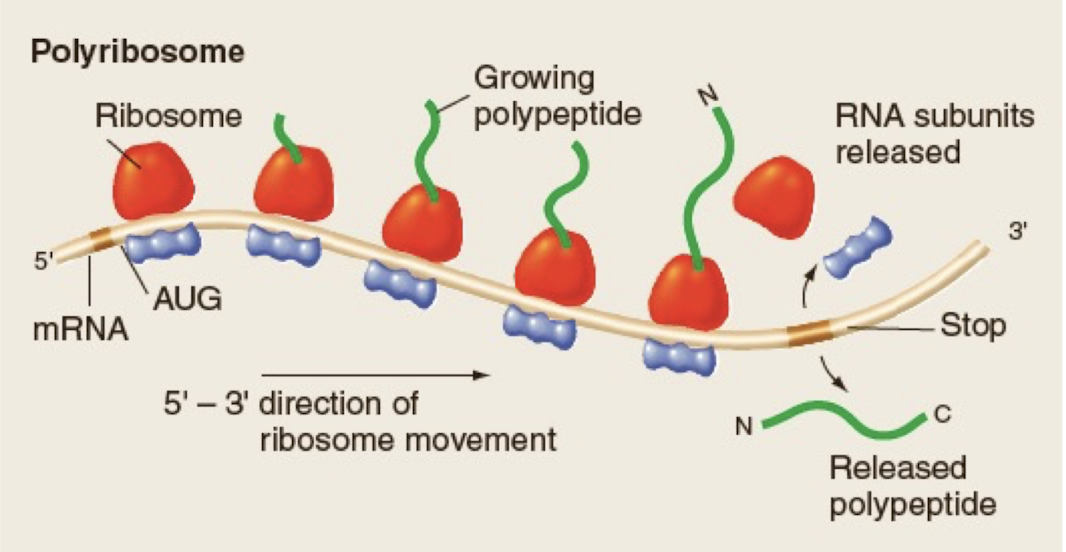
Polyribosomes
consist of multiple ribosomes translating the same mRNA
Codon-anticodon recognition
-18S rRNA that is part of the 40S subunit plays a key role
-It can detect an incorrect tRNA bound at the A site; Will prevent elongation until the mispaired tRNA is released and corrected
-Is called the decoding function of the ribosome
It is important in maintaining the high fidelity in mRNA translation
-Error rate: 1 mistake per 10,000 amino acids added
Peptidyl (P) site
Mark the following on the ribosome below: P site, A site, E site, peptidyl transferase, large subunit and small subunit
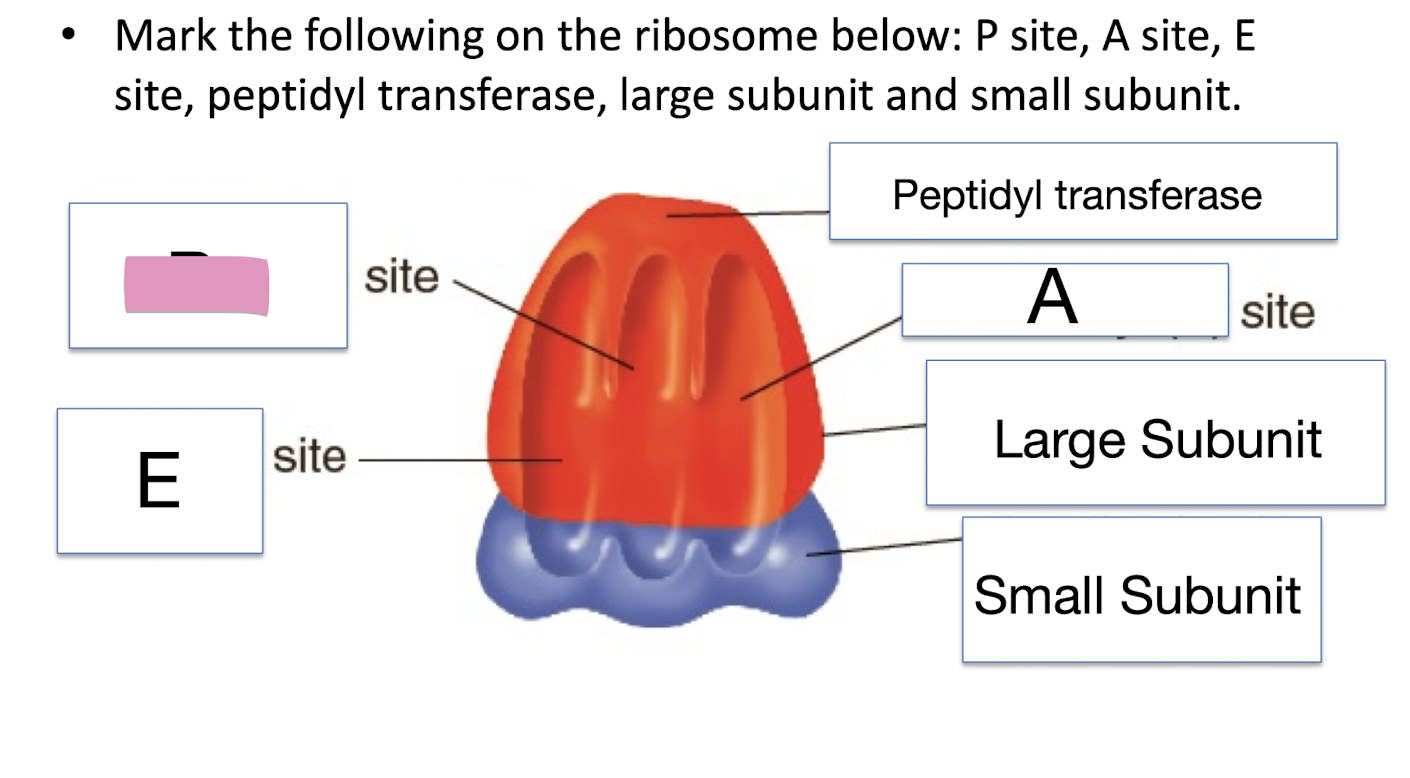
Exit (E) site
Mark the following on the ribosome below: P site, A site, E site, peptidyl transferase, large subunit and small subunit
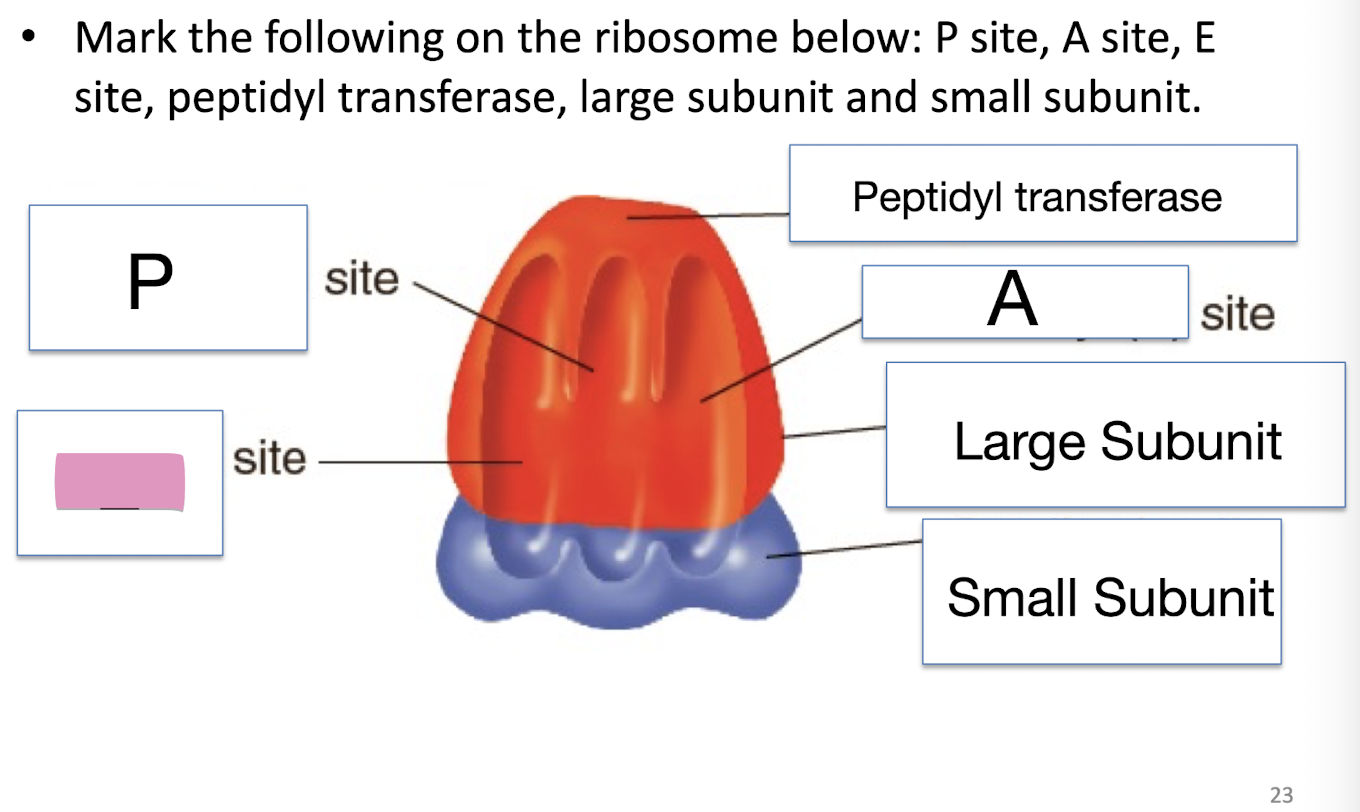
Peptudly transferase
Mark the following on the ribosome below: P site, A site, E site, peptidyl transferase, large subunit and small subunit
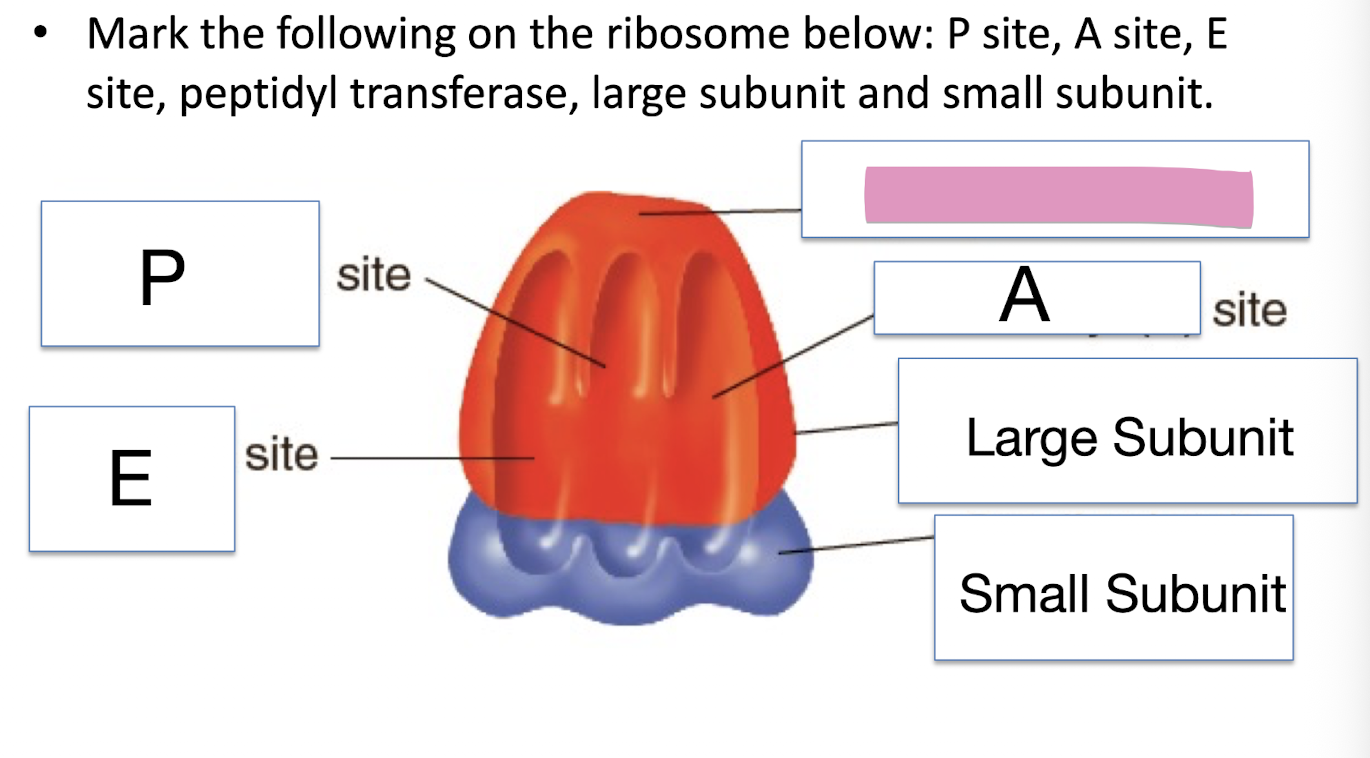
Aminoacyl (A) site
Mark the following on the ribosome below: P site, A site, E site, peptidyl transferase, large subunit and small subunit
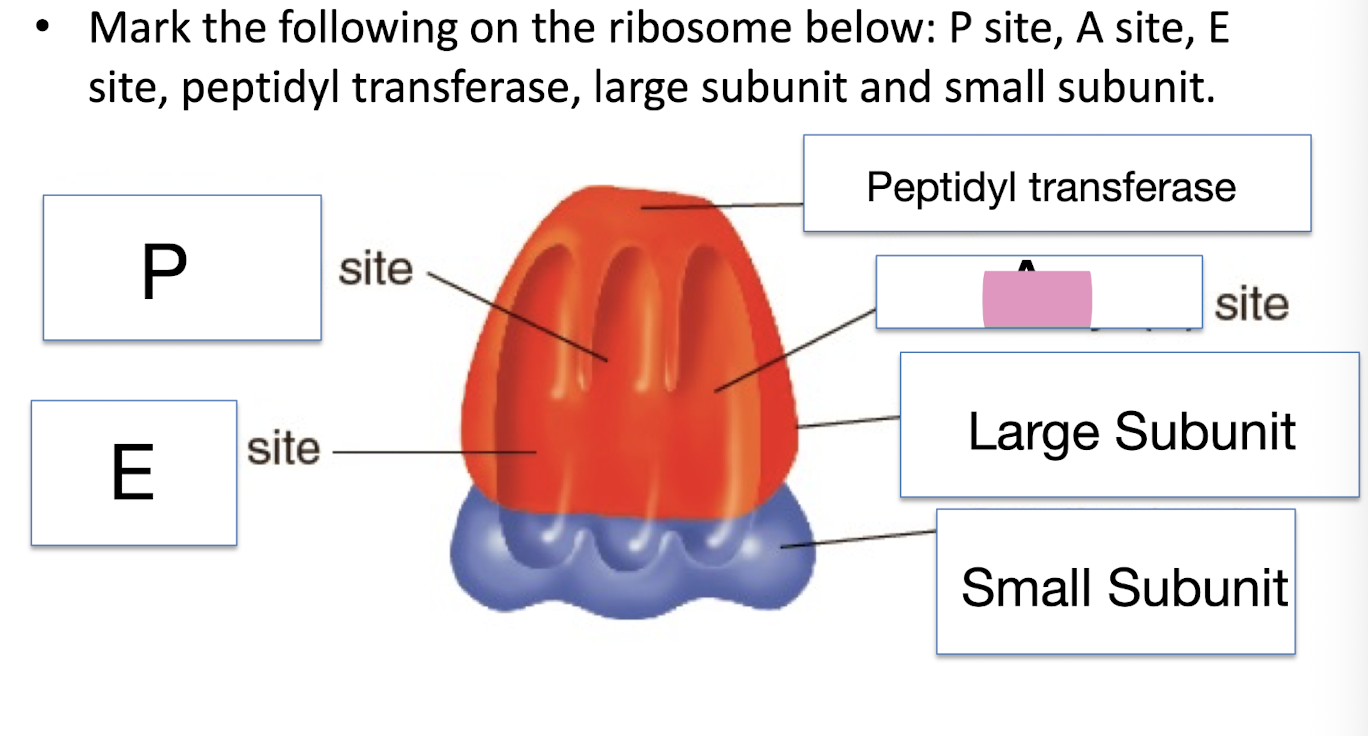
Large Subunit
Mark the following on the ribosome below: P site, A site, E site, peptidyl transferase, large subunit and small subunit
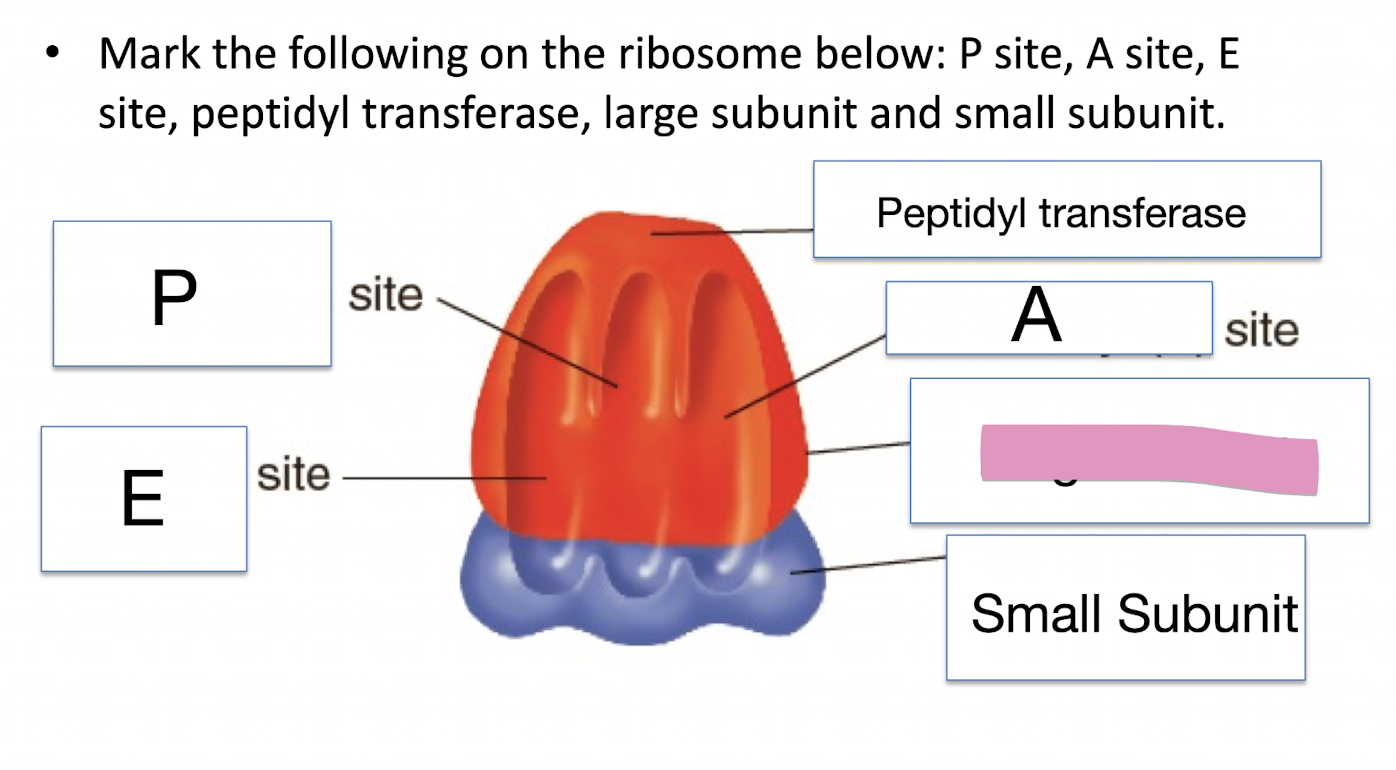
Small Subunit
Mark the following on the ribosome below: P site, A site, E site, peptidyl transferase, large subunit and small subunit
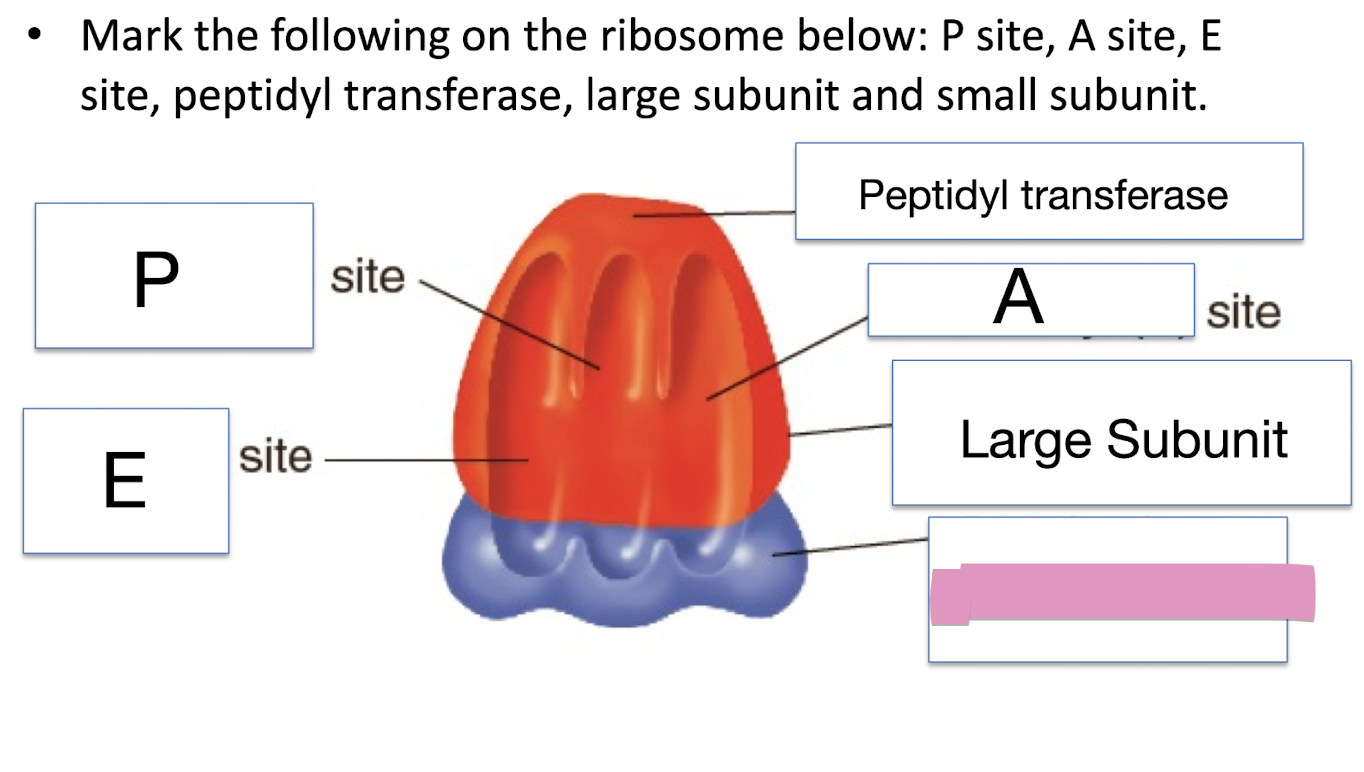
Directionality
-the polypeptide has this
-with the N terminus parallel to the 5’ end of the mRNA and the C terminus parallel to the 3’ end of mRNA
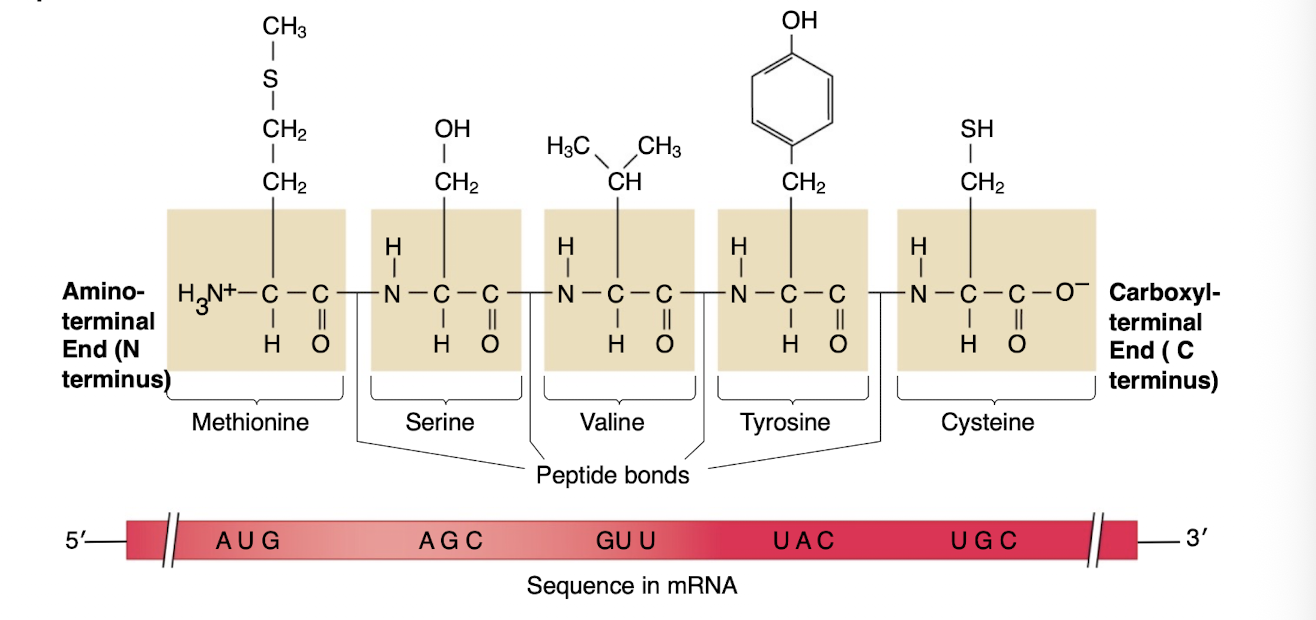
Amino acids have…
-a carboxyl group, an amino group and an R (radical group-side chain) bound to a central carbon (C) atom
R group
-gives each amino acid it’s chemical identity
-Nonpolar (hydrophobic)
-Polar (hydrophilic):
-Positively charges
-Negatively charged
Polar and charged amino acids
-hydrophilic
-more likely to be on the surface of a protein
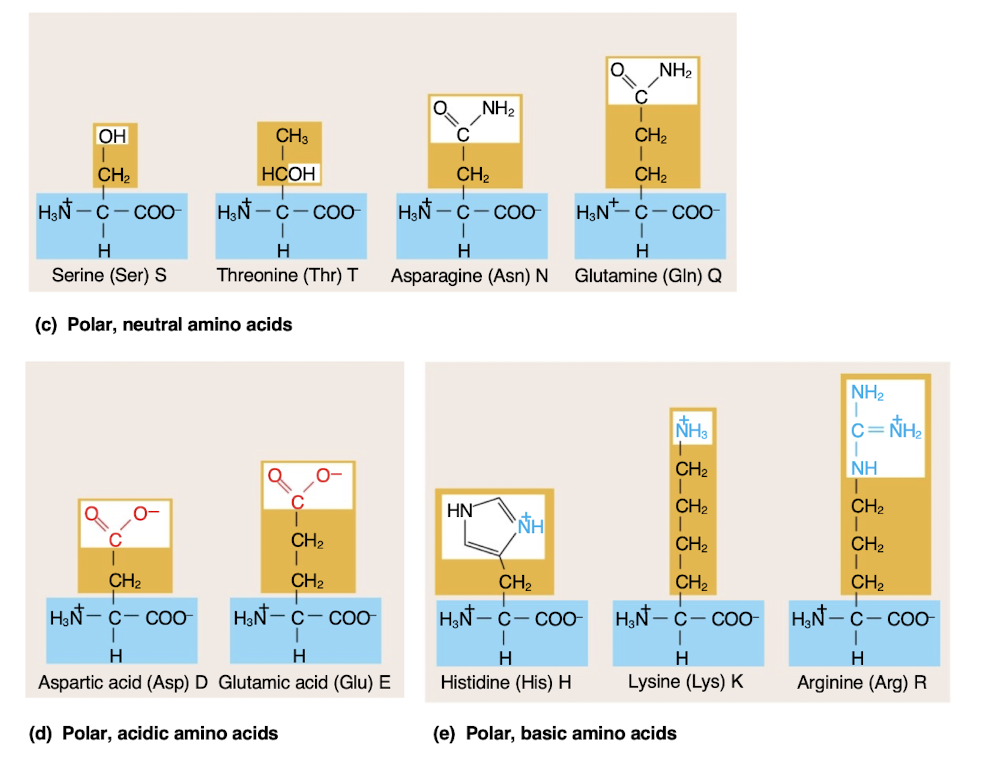
Polypeptides
-composed of a strand of amino acids, represents the protein’s primary structure
-Following translation, the polypeptide folds up and assumes a 3-dimensional structure
-Secondary is formed by hydrogen bonds between the sides of chains of amino acids that are close in proximity
alpha-helix and beta-pleated sheets are secondary structures
-Tertiary defines three dimensional conformation of entire chain in space
-Quaternary applies to those proteins comprised of more than one polypeptide chain
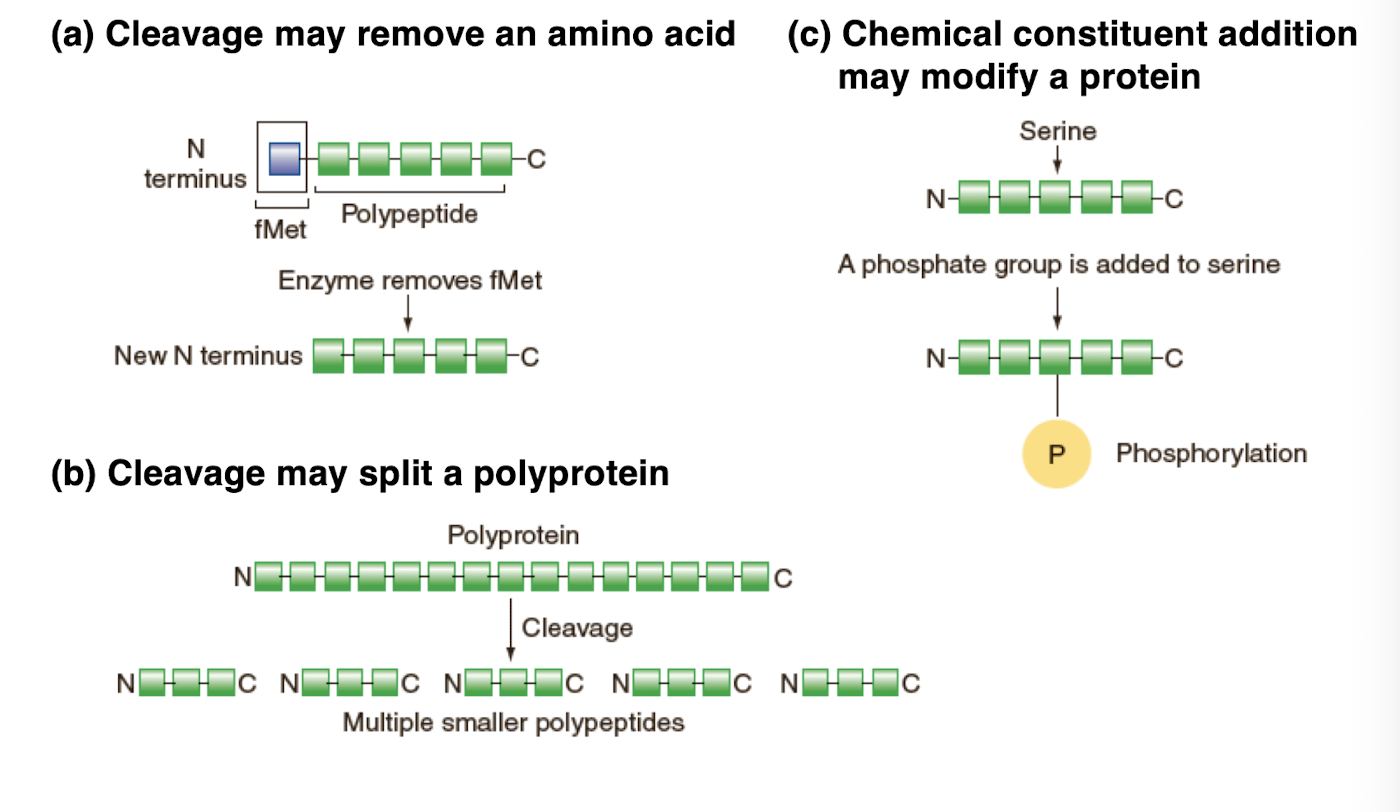
Post-translational processing
-modifies the structure of the polypeptide chain
-cleavage may remove an amino acids
-cleavage may split a polyprotein
-Chemical constituent addition may modify a protein
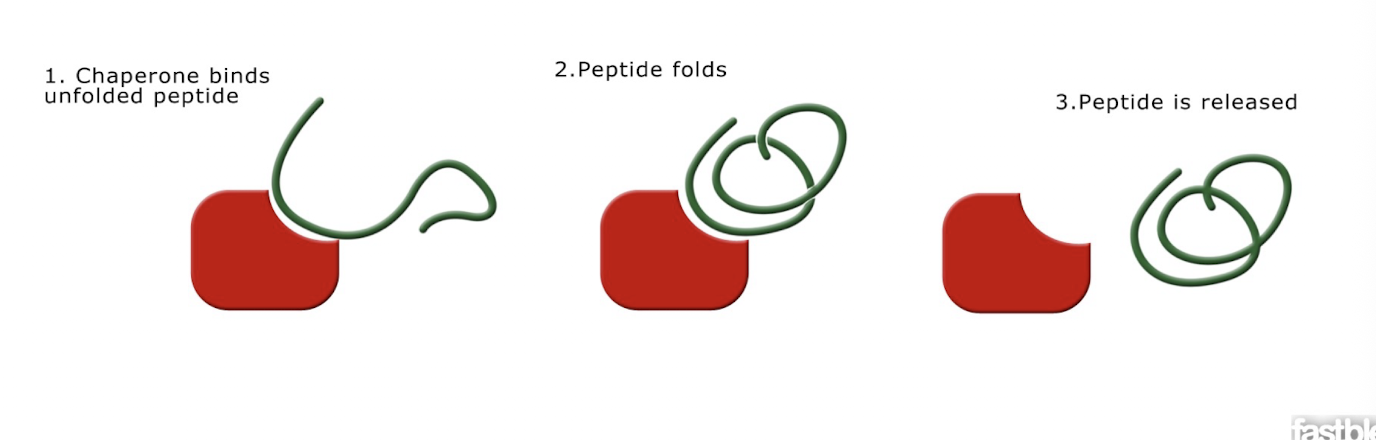
Chaperone proteins
-For many proteins, correct folding requires this
-Mediate the folding process, but do not become part of the final proteins
Ubiquitins
-Misfolded proteins are tagged by these small proteins
-Protein-ubiquitin complexes move to proteasomes, where proteins are degraded by proteases
Prions (proteinaceous infectious particles)
-cause variety of neurodegenerative diseases
-cause scrapie in sheep and bovine spongiform encephalopathy (BSE) in cows (mad cow disease): Brain tissue from affected animals resembles a sponge
-In 1993, there was an epidemic of BSE in the UK
-1st care occurred in humans in 1996 due to exposure to BSE
-Probably consumption of contaminated beef
-Called variant Creutzfeldt-Jacob disease (vCJD) due to similar clinical features as classic CJD, but at a young age
Creutzfeldt-Jacob disease (CJD)
-a very rare generative and fatal brain disorder in humans
-5-10% are inherited
-Dominant mutation in the prion gene, incomplete penetrance; average age at onset is ~60 years and 90% of cases die within 1 year NO CURE
-Most cases are sporadic and due to the spontaneous transformation of normal prion proteins
-about 1% are variant CJD and acquired; due to exposure to BSE or consumption of BSE-contaminated beef