☠️ Lecture 15: Apoptosis
1/22
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
23 Terms
Fig. 19-32: An Overview of the DNA Damage Response System
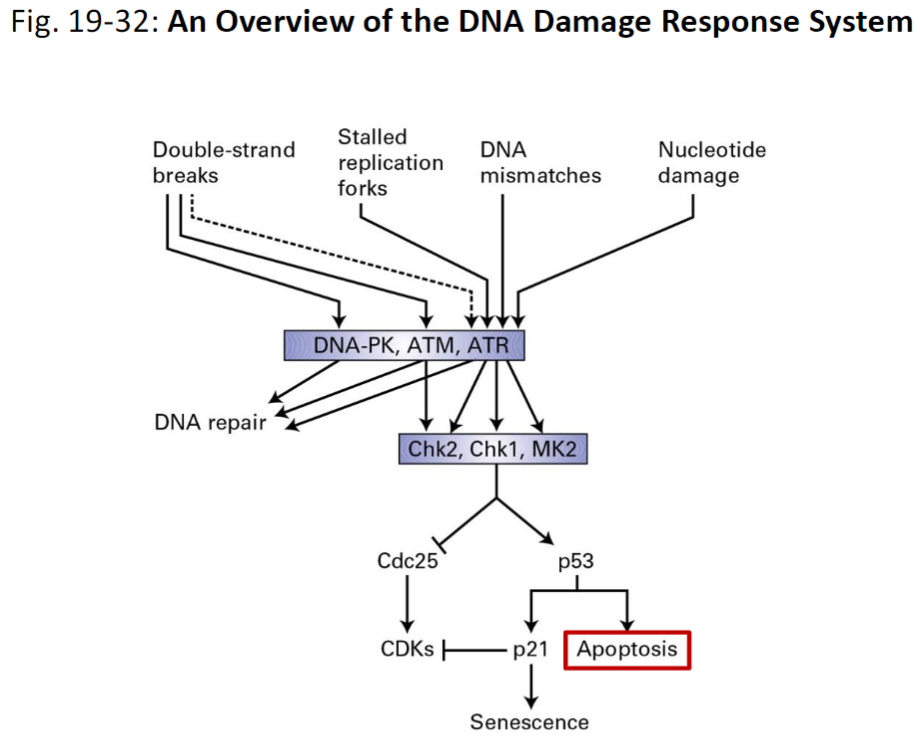
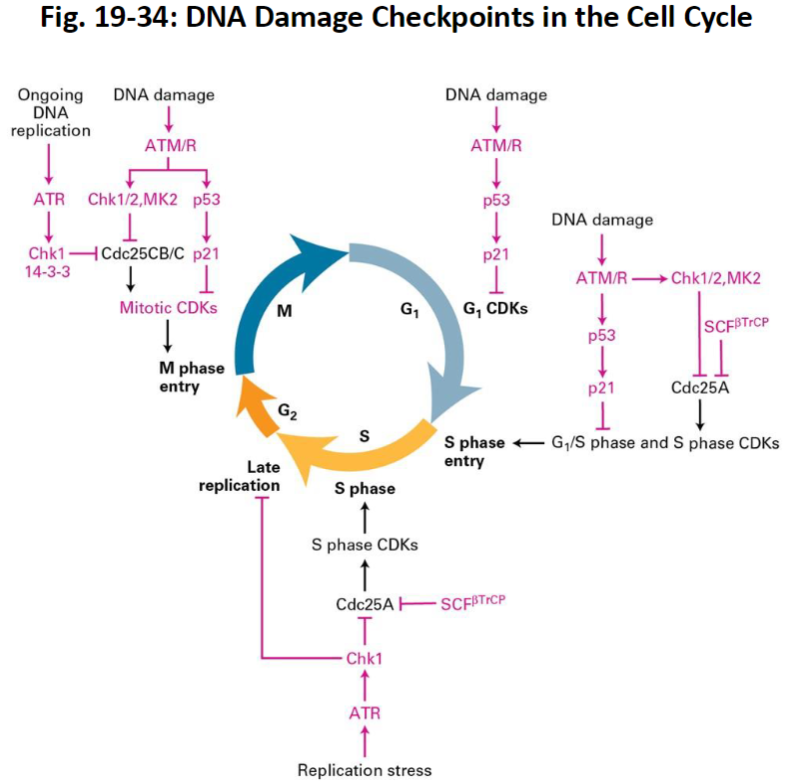
Fig. 19-34: DNA Damage Checkpoints in the Cell Cycle
Cell Death and Regulation
Apoptosis Overview
Programmed cell death that does not release cytosolic contents
Uses intrinsic pathways to safely remove cells
Caspase Activation
Caspases are proteases that degrade cellular proteins
Activated during apoptosis to break down the cell in a controlled way
Physiological Role
Essential for normal development
Removes unwanted or damaged cells
Survival Signals
Trophic factors bind to surface receptors
Repress apoptosis to allow cell survival
Triggers of Apoptosis
Deregulated cell cycle
DNA damage
Clinical Relevance
Genes regulating apoptosis are often mutated in cancers

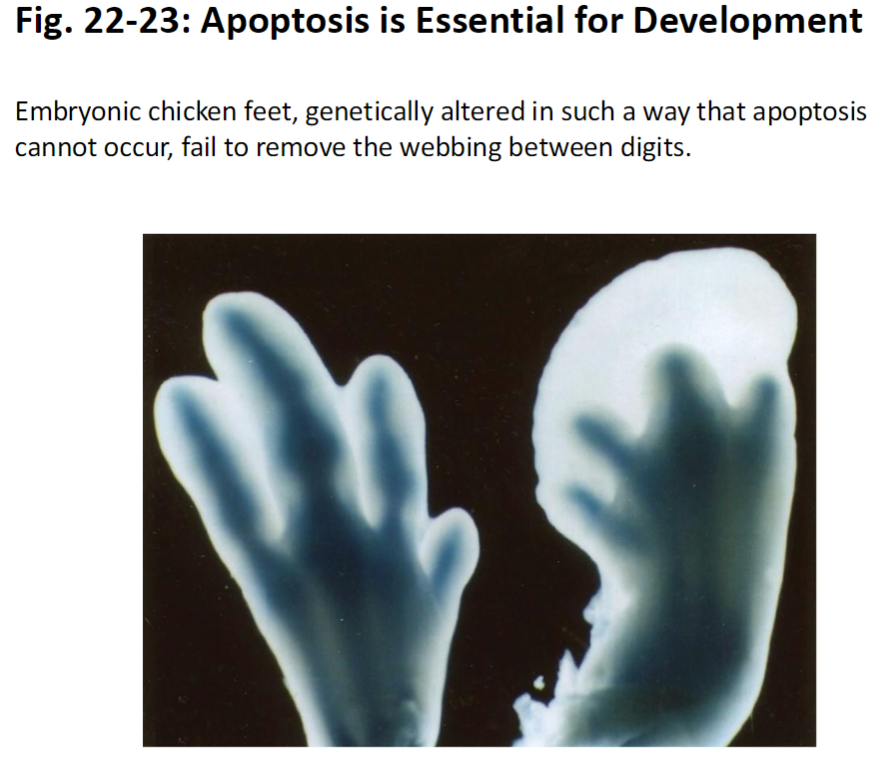
Apoptosis in Development
Example: Embryonic Chicken Feet
When apoptosis is blocked, webbing between digits is not removed
Shows that programmed cell death is essential for shaping tissues during development
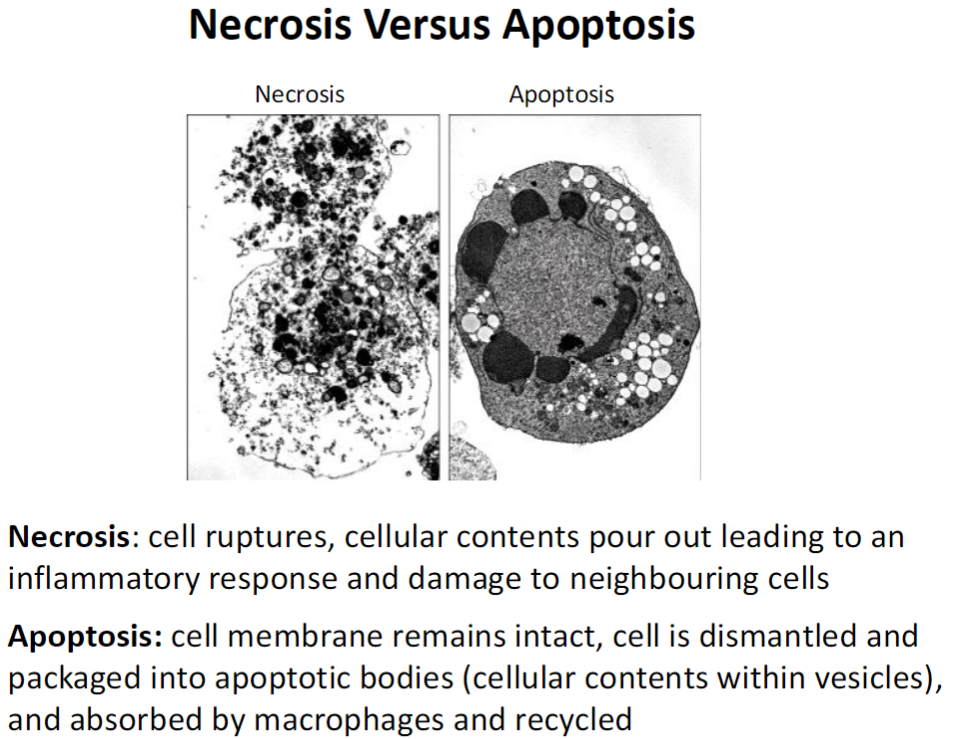
Cell Death Types – Necrosis vs Apoptosis
Necrosis
Cell ruptures and releases contents
Triggers inflammation and damages neighbouring cells
Apoptosis
Cell membrane stays intact
Cell dismantled into apoptotic bodies (vesicles containing cellular contents)
Bodies are absorbed by macrophages and recycled
No inflammation, controlled cell removal
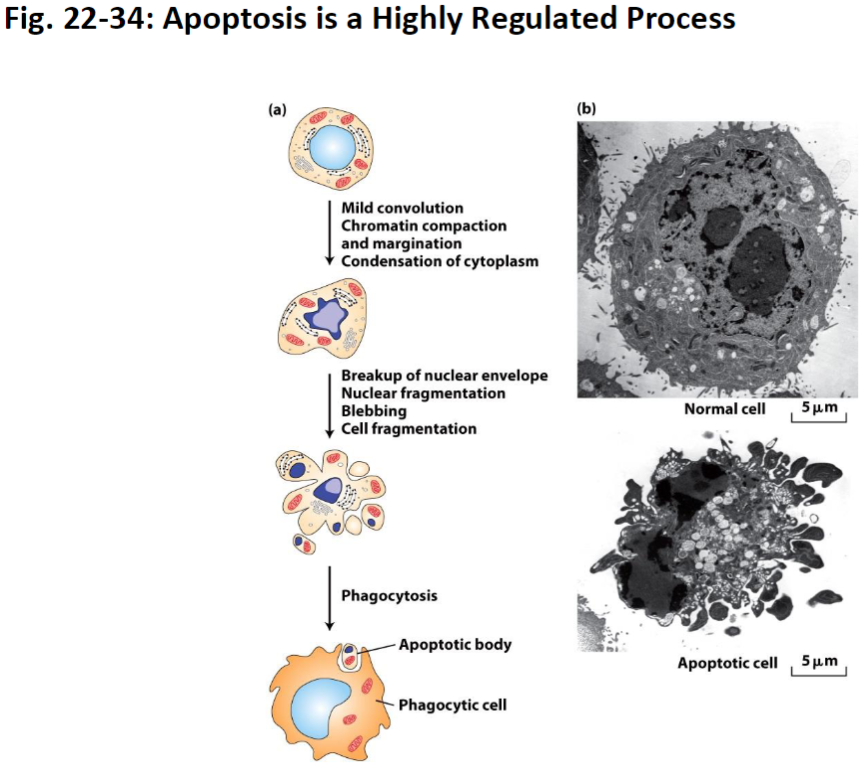
Apoptosis Process – Key Steps
Mild Convolution
Cell begins to change shape slightly
Chromatin Compaction and Margination
Chromatin (DNA + proteins) condenses and moves toward the edges of the nucleus
Cytoplasm Condensation
Cytoplasm becomes denser as cell shrinks
Nuclear Envelope Breakdown
Nuclear membrane fragments to allow controlled dismantling
Blebbing
Cell surface forms bubble-like protrusions
Phagocytosis
Apoptotic bodies are recognized and eaten by phagocytic cells for recycling
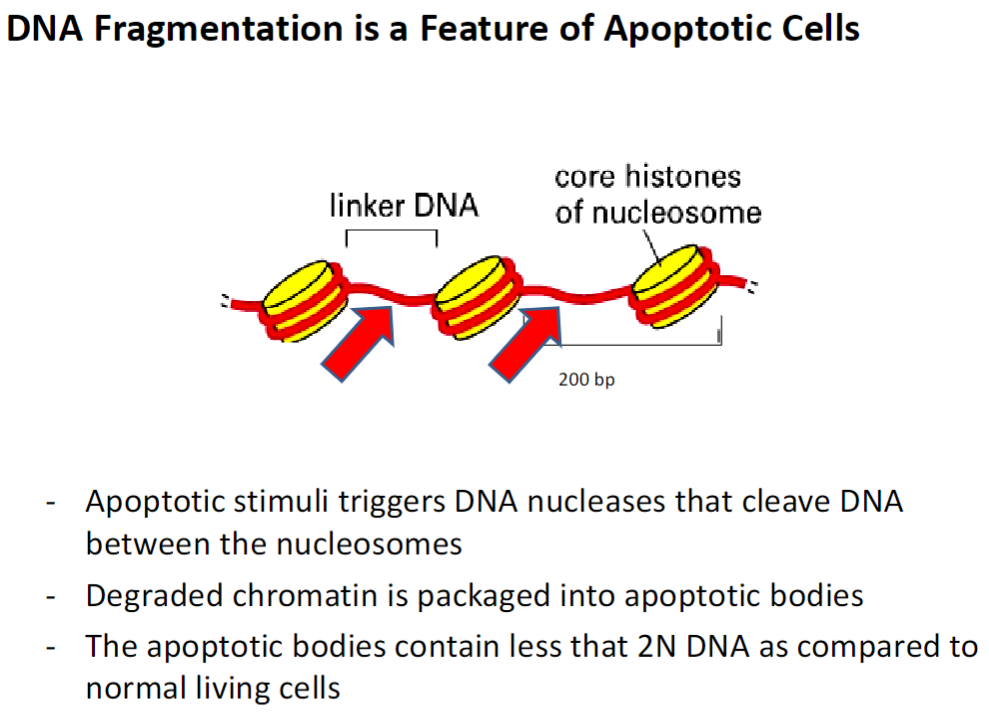
DNA Fragmentation in Apoptosis
Apoptotic Stimuli
Triggers activation of DNA nucleases that cleave DNA between nucleosomes
Chromatin Degradation
Broken-down chromatin is packaged into apoptotic bodies
DNA Content
Apoptotic bodies contain less than 2N DNA compared to normal living cells
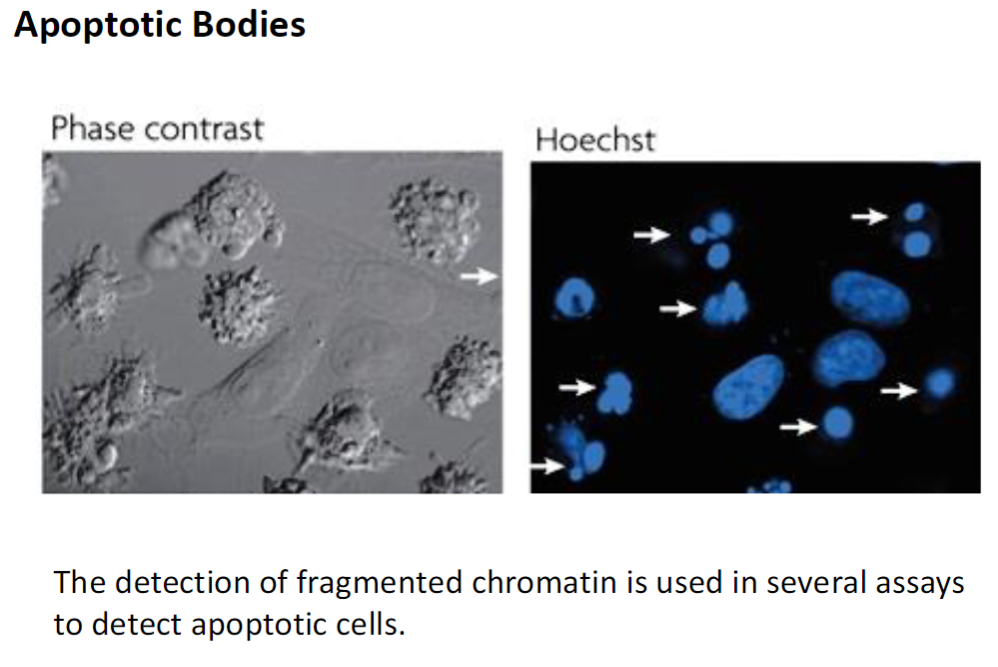
Detection of Apoptosis – Apoptotic Bodies
Assay Principle
Fragmented chromatin can be detected using various laboratory assays
Apoptotic Bodies
Small vesicles containing degraded chromatin and cellular contents
Formed during apoptosis and can be recognized and engulfed by phagocytic cells
Flow Cytometry – Detection of Apoptosis
Principle
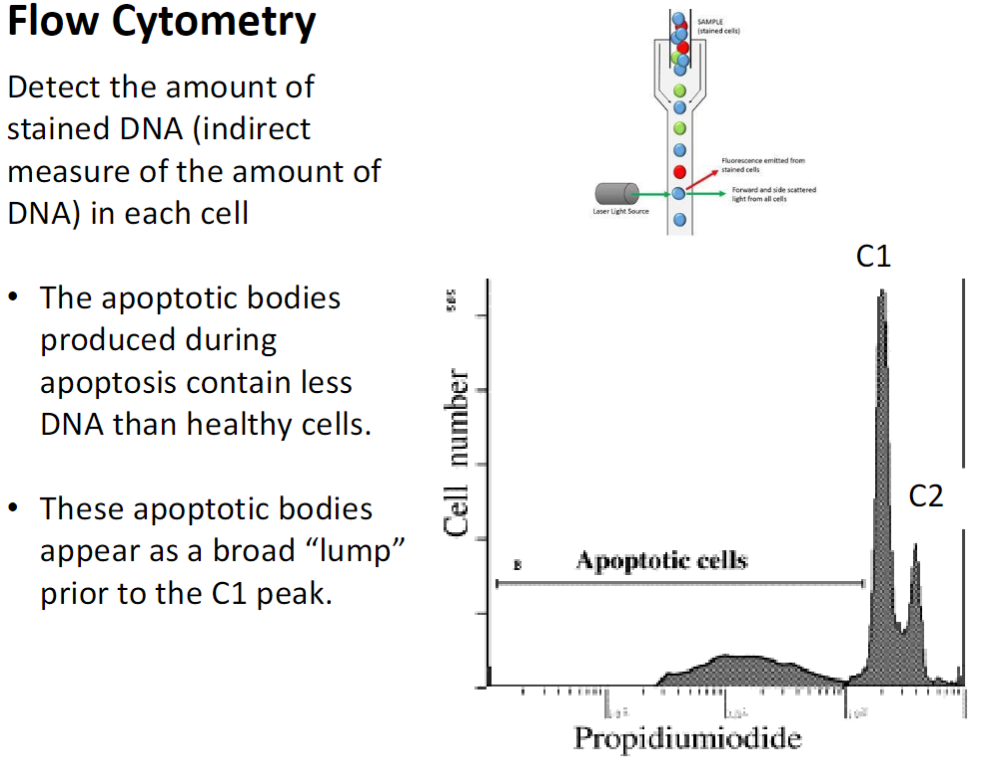
Measures the amount of stained DNA in each cell as an indirect measure of DNA content
Apoptotic Bodies
Produced during apoptosis and contain less DNA than healthy cells
Appear as a broad “lump” before the C1 peak in flow cytometry plots
Cell Cycle Peaks
C1 – cells not dividing, diploid DNA (2N)
C2 – cells that have duplicated DNA (4N) but have not divided
Between C1 and C2 – cells synthesizing DNA (S phase)
Apoptosis Detection
Apoptotic cells have DNA content less than 2N (C1)
Low number of cells in this region indicates apoptosis level
Every cell culture exhibits some baseline apoptosis
Comet Assay – Detection of DNA Fragmentation
Procedure
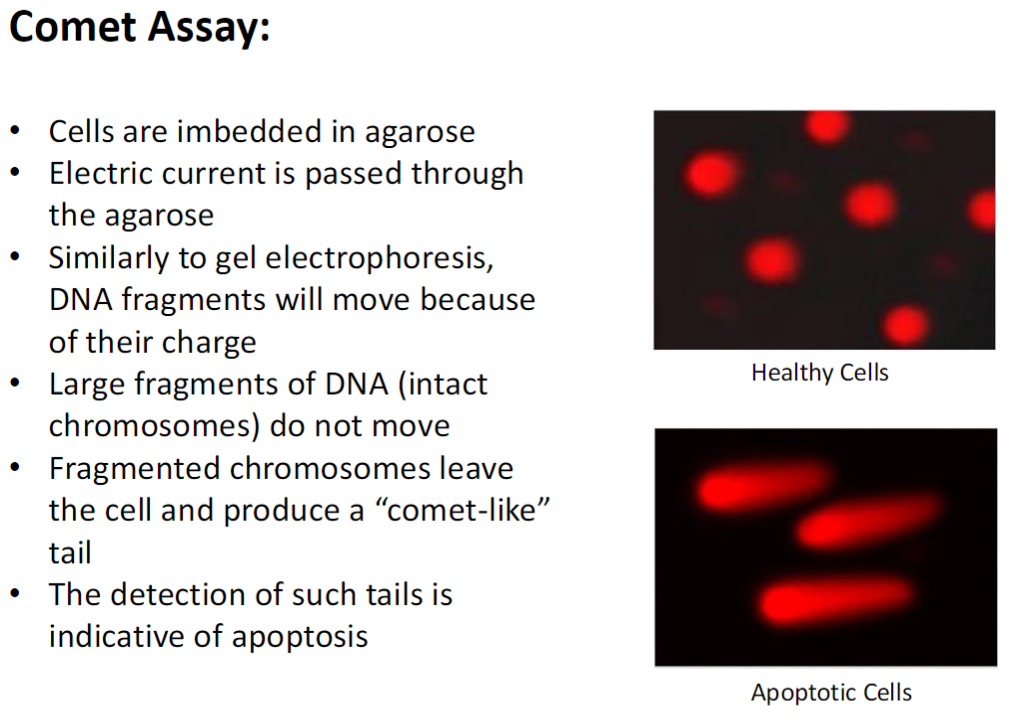
Cells embedded in agarose
Electric current passed through agarose
DNA fragments move according to charge, similar to gel electrophoresis
DNA Movement
Large DNA fragments (intact chromosomes) remain in place
Fragmented DNA leaves the cell and forms a “comet-like” tail
Interpretation
Presence of comet tails indicates apoptosis
Healthy cells show minimal or no tail, apoptotic cells show prominent tail
C. elegans – Model for Cell Death
Overview
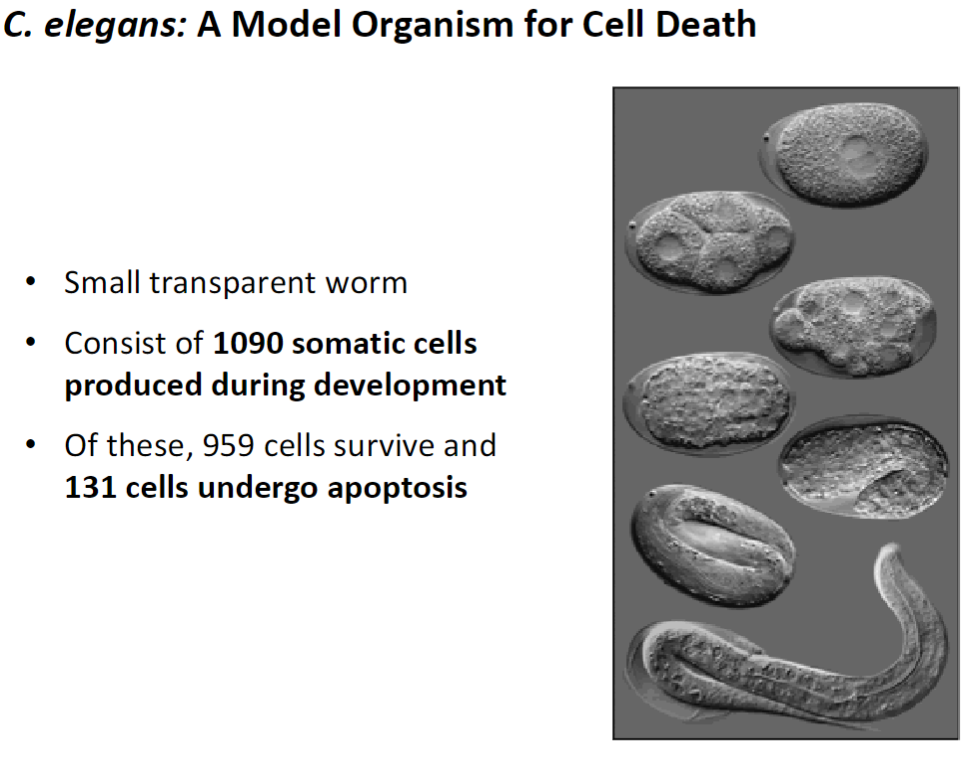
Small transparent worm used to study apoptosis
Consists of 1090 somatic cells produced during development
Cell Fate
959 cells survive
131 cells undergo programmed cell death (apoptosis)
C. elegans – Cell Lineage and Genetics
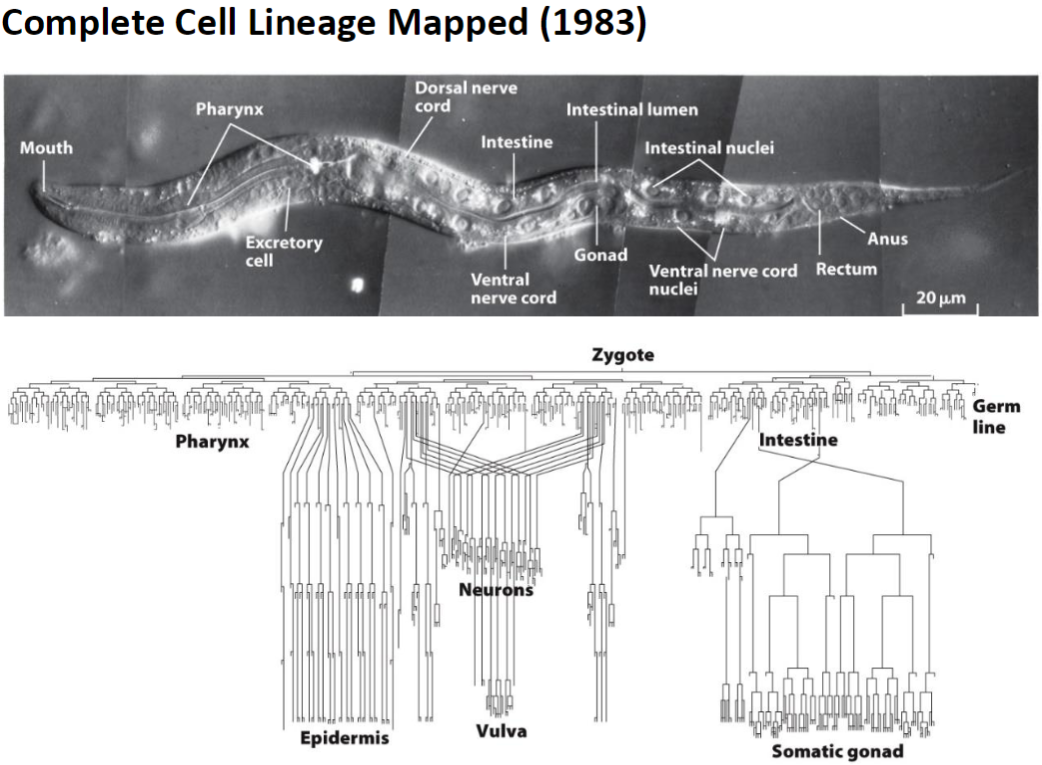
Complete Cell Lineage
Mapped in 1983, showing fate of all 1090 somatic cells
CED Genes
Cell death genes identified through genetic studies in C. elegans
Regulate which cells undergo apoptosis and which survive
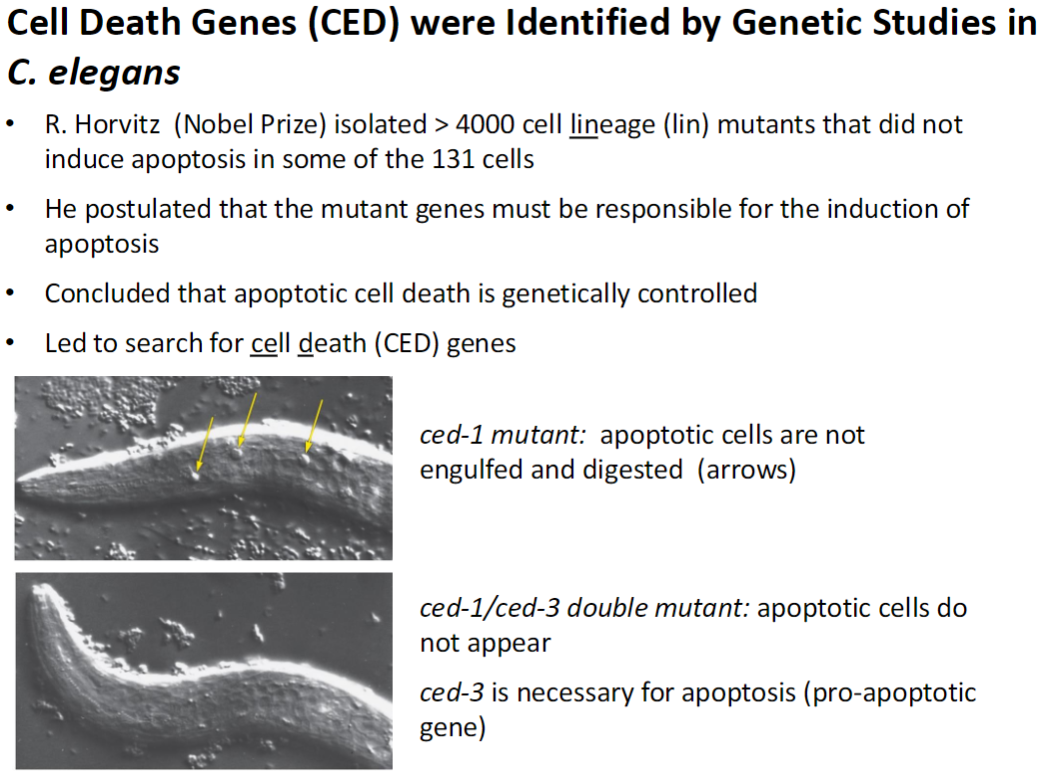
Cell Death Genes (CED) – Discovery in C. elegans
Mutant Screening
R. Horvitz (Nobel Prize) isolated >4000 cell lineage (lin) mutants
Mutants failed to induce apoptosis in some of the 131 cells
Conclusion
Apoptotic cell death is genetically controlled
Mutant genes responsible for apoptosis led to discovery of cell death (CED) genes
Key Mutants
ced-1 mutant: apoptotic cells are not engulfed or digested (visible as arrows)
ced-1/ced-3 double mutant: apoptotic cells do not appear
ced-3: necessary for apoptosis, pro-apoptotic gene
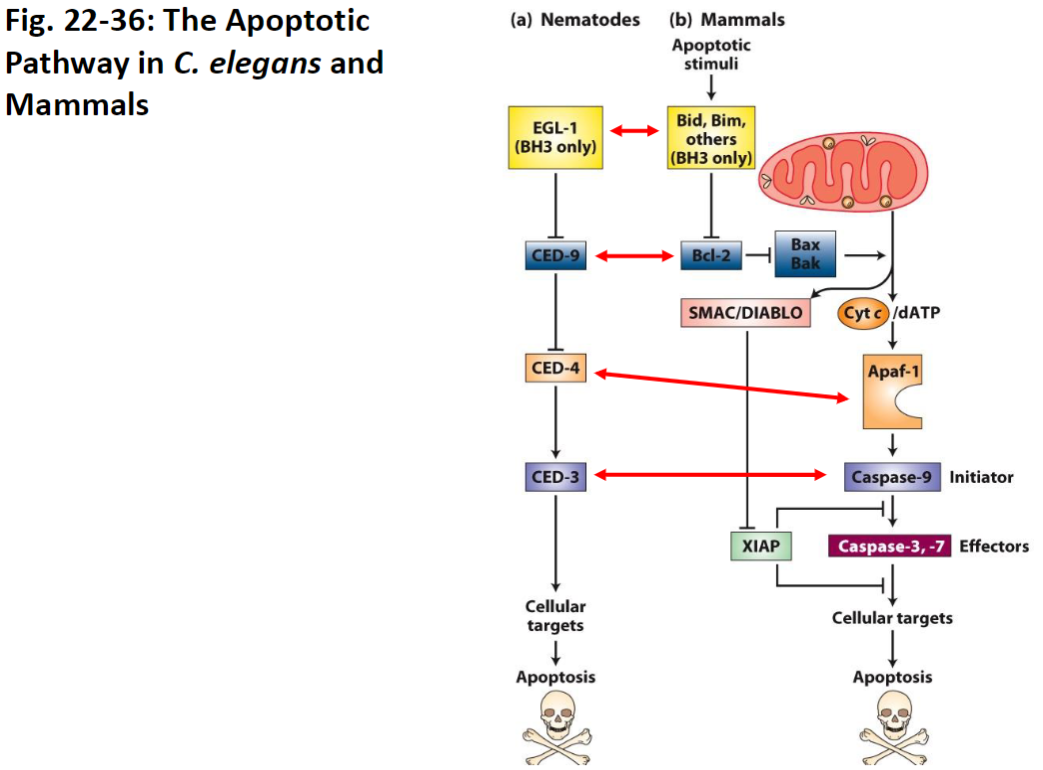
Apoptotic Pathway – C. elegans vs Mammals
Pathway Overview
Apoptosis in C. elegans is more straightforward than in mammals
All C. elegans cell death genes (CED) have analogous genes in mammals
Significance of C. elegans
Very few cells in the nematode, so cell lineage and apoptosis are easy to track
Allows precise study of genetic control of cell death
Important model organism for understanding conserved apoptotic mechanisms
Caspases – Key Classes and Functions
Definition
Caspases = Cysteinyl Aspartate-Specific Proteinases (enzymes that cleave proteins at specific sites containing aspartate)
Initiation Caspases
Process and activate effector caspases
Caspase-9: intrinsic (internal signal) pathway
Caspase-8 and Caspase-10: extrinsic (external signal) pathway
Effector (Executioner) Caspases
Cleave specific cellular proteins to dismantle the cell
Caspase-3, Caspase-6, Caspase-7
Activate nucleases that degrade DNA and chromatin
Essential for completing apoptosis

Intrinsic (Mitochondrial) Apoptosis Pathway
Activation
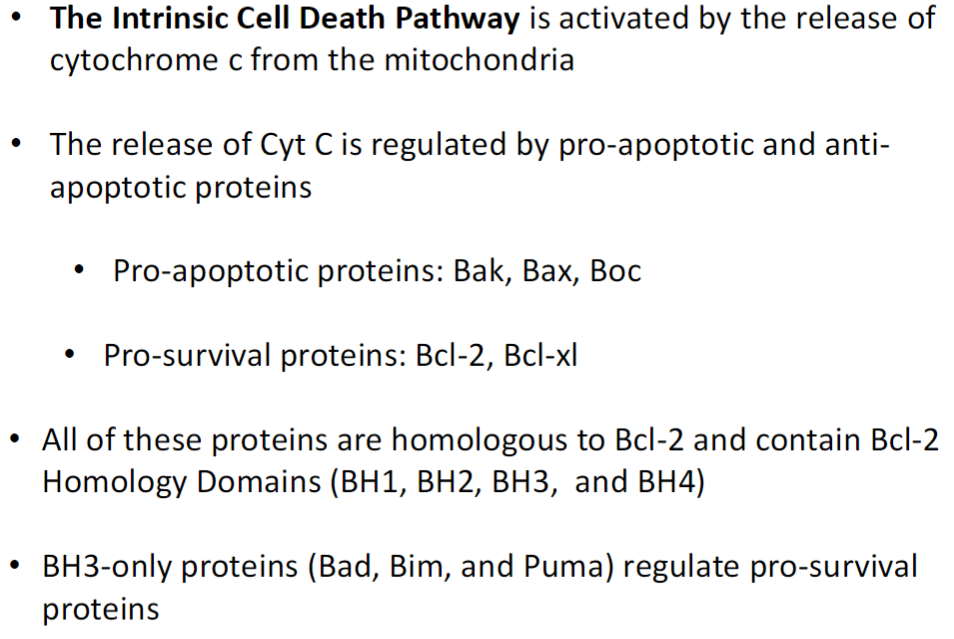
Triggered by release of cytochrome c (Cyt C) from mitochondria
Regulation by Bcl-2 Family Proteins
Pro-apoptotic proteins: Bak, Bax, Boc (promote Cyt C release)
Pro-survival proteins: Bcl-2, Bcl-xl (prevent Cyt C release)
All share Bcl-2 Homology (BH) domains: BH1, BH2, BH3, BH4
BH3-Only Proteins
Bad, Bim, Puma
Regulate pro-survival proteins by inhibiting them, tipping balance toward apoptosis

Bcl-2 Family Proteins – Mitochondrial Control
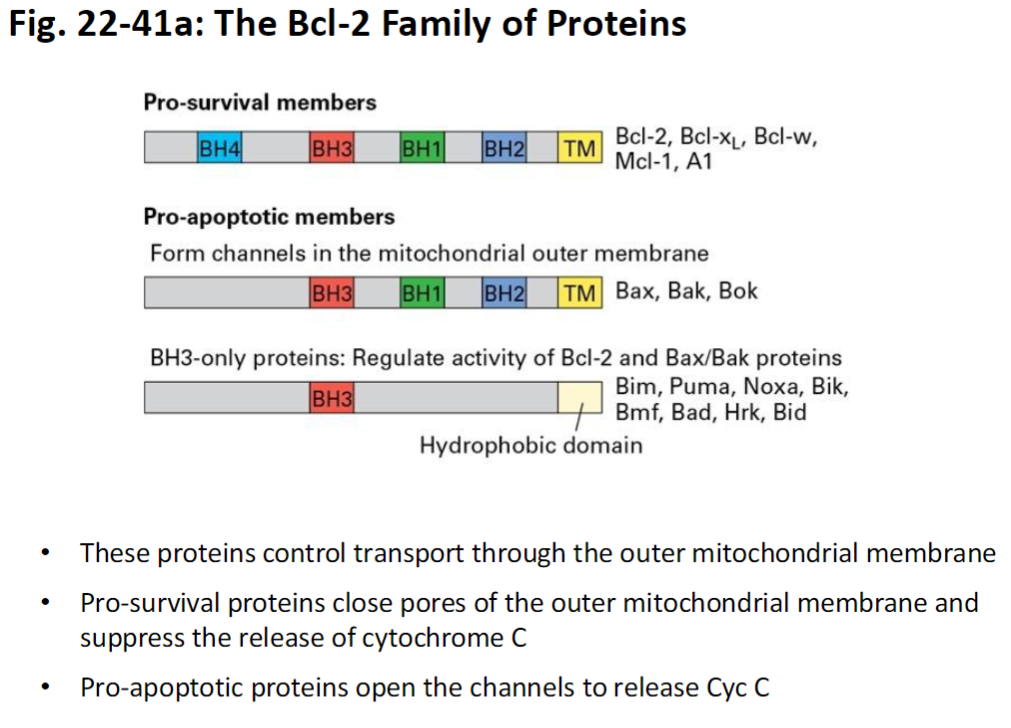
Function
Control transport through outer mitochondrial membrane
Pro-Survival Proteins
Close pores in outer mitochondrial membrane
Suppress release of cytochrome c
Pro-Apoptotic Proteins
Open channels in outer mitochondrial membrane
Promote release of cytochrome c into cytosol

Cell Death Pathways – Vertebrates
Intrinsic Pathway
Activated by mitochondrial proteins (Bak, Bax, Bcl-2, Cytochrome C)
Triggered by DNA damage, cell stress, loss of adhesion, or lack of trophic signals
Extrinsic Pathway
Activated through cell surface death receptors
Triggered by direct contact with other cells signaling apoptosis

Intrinsic Cell Death Pathway – Additional Notes
Integrins
Transmembrane proteins that link the extracellular matrix (ECM) to the intracellular cytoskeleton
Provide survival signals; loss of integrin-mediated attachment can trigger intrinsic apoptosis
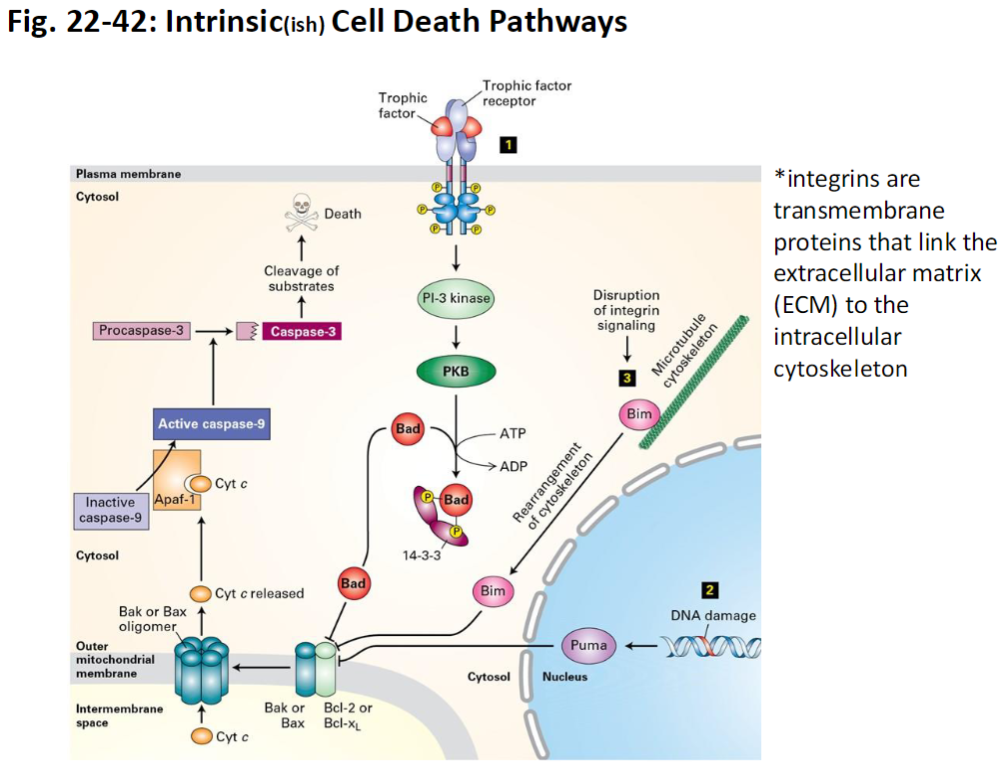
Intrinsic Death Pathway – Pro- and Anti-Apoptotic Proteins
Pro-Apoptotic Proteins
Bak and Bax can oligomerize (bind together) to form pores in the mitochondrial membrane
This allows cytochrome C release, triggering apoptosis
Pro-Survival Proteins
Bcl-2 and Bcl-XL bind to Bak or Bax and prevent pore formation
BH3-Only Proteins
Bad, Bim, and Puma displace Bcl-2 from Bak or Bax
This allows Bak/Bax to form pores and activate the intrinsic apoptosis pathway
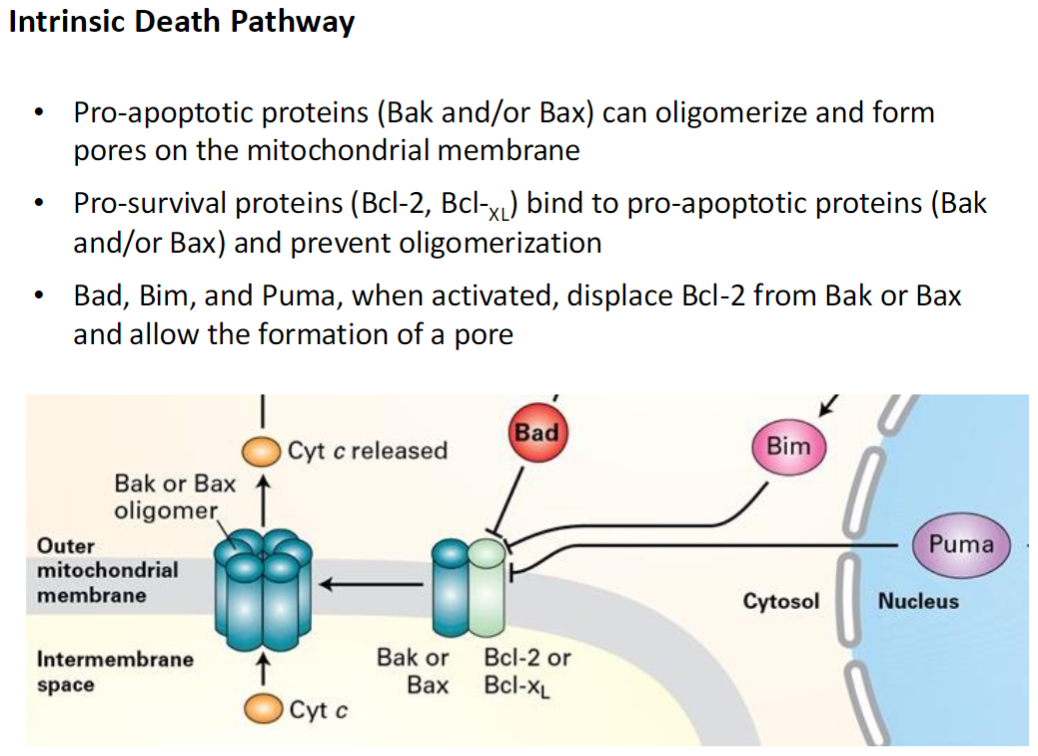
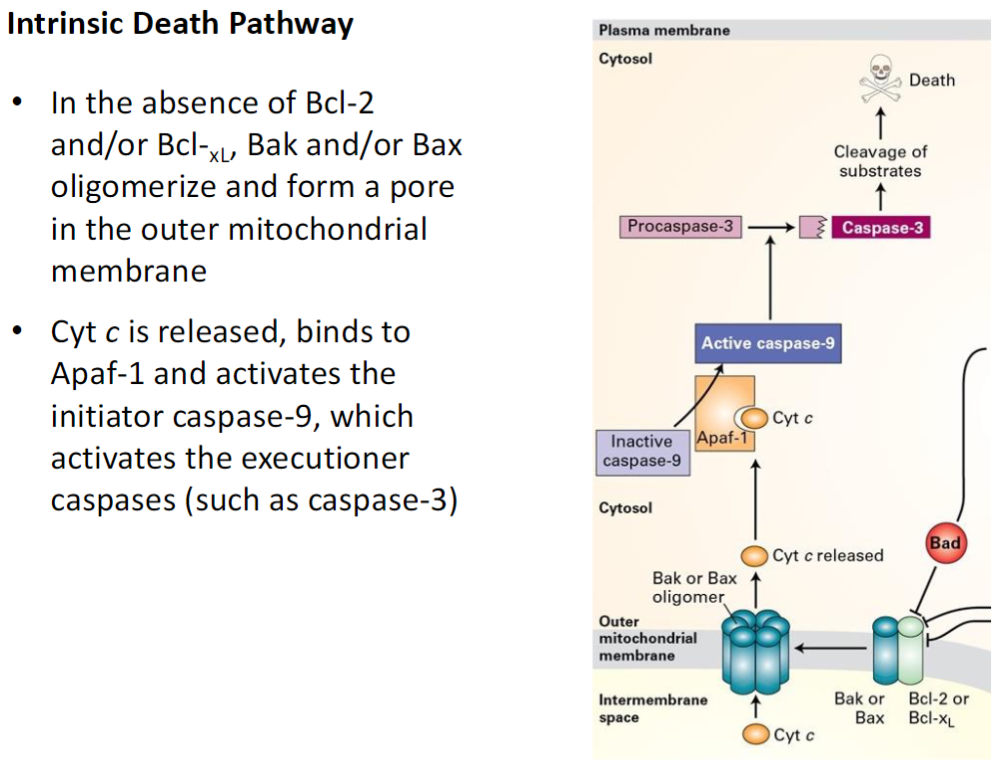
Intrinsic Death Pathway – Cyt c and Caspase Activation
Loss of Pro-Survival Proteins
Without Bcl-2 or Bcl-XL, Bak and Bax oligomerize and form pores in the outer mitochondrial membrane
Cytochrome C Release
Cytochrome C exits the mitochondria and binds Apaf-1
Caspase Activation
Apaf-1 activates initiator caspase-9
Caspase-9 activates effector (executioner) caspases such as caspase-3
Leads to systematic degradation of cellular proteins and DNA, completing apoptosis
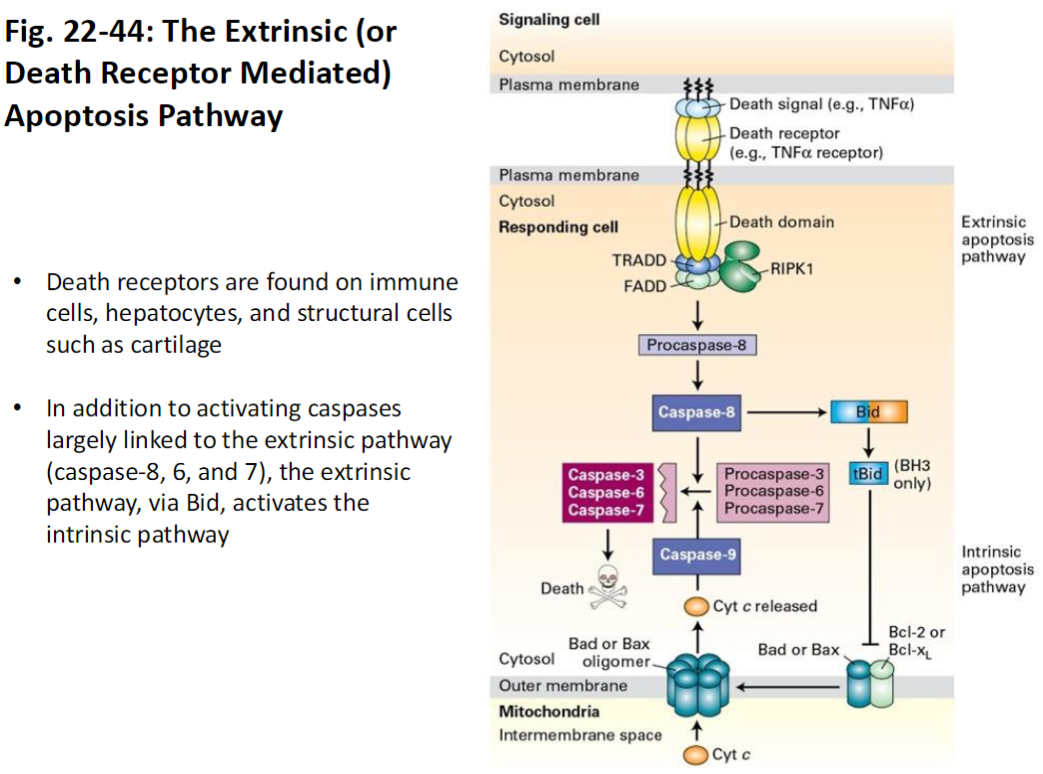
Extrinsic (Death Receptor Mediated) Apoptosis Pathway
Death Receptors
Located on immune cells, hepatocytes, and structural cells (e.g., cartilage)
Trigger apoptosis when bound by external death signals
Caspase Activation
Activates initiator caspase-8
Caspase-8 activates effector caspases 6 and 7
Leads to systematic breakdown of cellular proteins and DNA
Cross-Talk with Intrinsic Pathway
Via Bid, the extrinsic pathway can activate the intrinsic (mitochondrial) apoptosis pathway
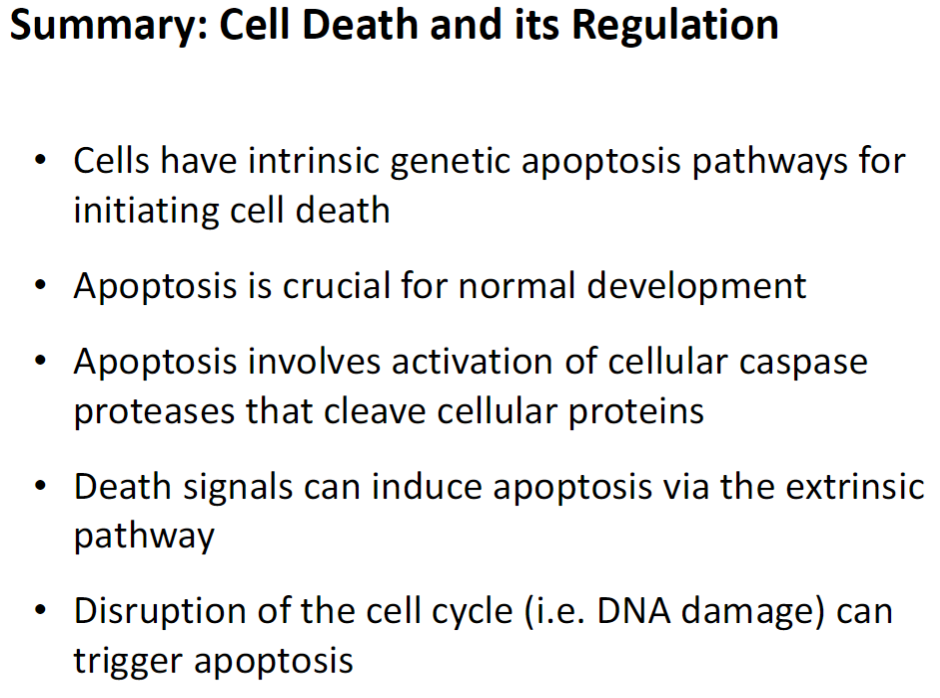
Cell Death and Its Regulation
Intrinsic Apoptosis Pathways
Genetically programmed pathways that initiate cell death from within the cell
Crucial for normal development
Caspase Activation
Cellular caspase proteases cleave cellular proteins to dismantle the cell
Extrinsic Apoptosis Pathway
Death signals from other cells can trigger apoptosis via surface receptors
Triggers
Disruption of the cell cycle, such as DNA damage, can also initiate apoptosis