CH29: RNA functions, biosynthesis, & processing (1)
1/46
Earn XP
Description and Tags
29.1-29.2
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
47 Terms
define the roles of each rna molecules in gene expression
ribosomal RNAs, transfer RNAs, messenger RNAs, small nuclear RNAs
rRNA: component of ribosomes
tRNA: deliver aa to ribosome
mRNA: carry info that ribosomes use for making specific protein seq
snRNA: guiding splicing (introns) of mRNA
other RNAs include small regulatory RNAs & long noncoding RNAs
name 6 diseases that RNA viruses are responsible for (what kind of RNA mc do they have?)
influenza, polio, mumps, Ebola, common cold, COVID-19
these viruses have either single or double-stranded RNA in single/multiple fragments
how does the “active ingredient” in an mRNA vaccine differ from that of a traditional vaccine?
instead of using a weakened or inactivated virus, mRNA vaccines use lab-generated mRNA that codes for a specific viral protein
in the context of an mRNA vaccine, what is the role of the human cell after injection?
human cell acts as production site
ribosomes translate vaccine’s mRNA into a viral protein, which is presented to immune system to trigger protective response
what is transcription and what catalyzes it?
transcription: synthesis of RNA molecules from a DNA template
RNA polymerase are large enzymes that catalyze transcription
explain what RNA polymerases do during transcription and what they are regulated by (4)
search for initiation sites called promoter seq (promoters)
unwind a short stretch of double-helical DNA to make single-stranded DNA templates
choose correct ribonucleoside triphosphate & catalyze formation of phosphodiester bond
detect termination signals that specify where transcription ends
regulated by activator & repressor proteins that interact w/ promoter
describe how RNA polymerase catalyzes formation of phosphodiester bond
RNA polymerase catalyze the nucleophilic attack of the 3’-hydroxyl group of the last nucleotide in the chain on the α-phosphoryl group of the incoming NTP, releasing a pyrophosphate.
how many metal ions do RNA polymerase require in the catalytic site? and what do they each do?
2 metal ions, normally Mg2+
one remains tightly bound
one comes in w/ the NTP and leaves w/ pyrophosphate
what is a transcription bubble
complex of double-stranded DNA that has been locally unwound in a region of ~17 base pairs where polymerization reactions occur
contains unwound DNA template & nascent RNA, where elongation takes place
newly synthesized RNA forms hybrid helix w/ template DNA strand (8 bp long or nearly one turn of a double helix)
moves w/ RNA pol during elongation
what is De Novo Synthesis? what do most newly synthesized RNA chains carry?
RNA polymerase can start from scratch, doesn’t need a pre-existing primer to get it moving unlike DNA synthesis
most newly synthesized RNA chains carry a high distinctive tag on 5’ end (pppG or pppA)
in genetic numbering, what is the designation for the first nucleotide transcribed into RNA, and what is the designation for the nucleotide right before it? (Describe upstream/downstream)
first nucleotide is +1, the one right before is -1
anything “downstream” (direction RNA grows) is positive (+2, +3)
anything “upstream” (stuff coming before start) is negative (-1, -2)
what is the template and coding strand? another name? which DNA strand has same sequence as newly synthesized DNA (w/ exception of T instead of U)
template strand: (antisense/-) has the sequence complement of RNA transcript (strand enzyme needs)
coding strand: (sense/+) has same sequence of RNA transcript instead of T in place of U)
this has same seq as newly synthesized dna
explain the 3 steps in elongation
binding: incoming ribonucleoside triphosphate binds in RNA pol active site and base pairs w. template
bond formation: Mg2+ helping orient & activate the 3’-OH of growing chain, which atks α-phosphoryl to form phosphodiester bond and release PPi
translocation: polymerase slides down DNA by one position to clear active site so next block can enter & start cycle again
what forces the separation of the RNA-DNA Hybrid?
a structure within RNA polymerase
define core enzyme
the bacterial core RNA polymerase (α2ββ’ω)
what is the σ subunit?
it helps find promoters & participates in the initiation of RNA synthesis
how is a holoenzyme formed
it is formed when the σ-subunit joins the core enzyme
what are the 2 components of the E.coli “core promoter,” and how are they identified?
-10 sequence: 6 bp long seq located ~10 nucleotides upstream from start site
-35 sequence: 6 bp long sequence located ~35 nucleotides upstream from start sitie
represented by an idealized consensus sequence
what are strong promoters and weak promoters? what kind of sequences do they tend to have?
strong promoters: promoters for genes that are transcribed frequently
tend to have -10 & -35 sequences that correspond closely to consensus sequences
weak promoters: promoters for genes that are transcribed less frequently
tend to have -10 & -35 sequences w/ multiple substitutions
what is the upstream element (UP element) and where is located, how does it increase transcription efficiency?
UP element: sequence located 40-60 nucleotides upstream of transcription start site
bound by σ subunit of RNA polymerase
increases transcription efficiency by creating an additional interaction site for polymerase
what is the primary function of the σ (sigma) subunit in the RNA polymerase holoenzyme?
the σ subunit lets the polymerase recognize & bind to specific promoter sites (at the -10 & -35 regions) rather than binding randomly to DNA
at what point does the σ subunit dissociate from the RNA polymerase, and why?
it is released once the nascent (newly synthesized) RNA chain is 9-10 nucleotides long so core enzyme can transition from initiation phase to elongation phase & move down DNA
what is the structural difference between a “closed promoter complex” and an “open promoter complex”?
in a closed complex, DNA remains a double helix
in open complex, around 17 base pairs of DNA are unwound (specifically at -10 region) to allow polymerase access to template strand
how does bacterial RNA polymerase obtain the free energy required to break DNA base pairs during initiation?
energy is from favorable interactions (binding energy) between RNA pol & DNA template
they stabilize open promoter complex & help full template strand into active site
what is the rate of elongation?
~50 nucleotides per second
what are 3 characteristics that influence gene expression in euk?
nuclear membrane allows transcription/translation to take place in diff cellular compartments
variety of promoter elements enables complex transcriptional regulation
degree of RNA processing is greater in euk
compare transcription and translation in prokaryotes vs in eukaryotes
in prok: they are closley coupled
in euk: they are spatially & temporally separate
what is chromatin?
a complex formed between DNA & set of histone proteins
compacts & organizes euk DNA
some genes & their regulatory regions are relatively accessible, whereas other genes are not
manipulation of chromatin structure is required for euk gene regulation
what is a nucleosome, and what are its 2 primary components?
nucleosome: fundamental unit of chromatin that consists of a histone octamer (protein core) & approximately 145 base pairs of DNA wrapped around it
describe the specific protein composition of the histone octamer found in the core of a nucleosome
octamer has 8 proteins: a (H3)2(H4)2 tetramer, and a pair of H2A-H2B dimers
what are the 3 RNA polymerases that catalyze eukaryotic RNA synthesis?
Type Location Cellular transcripts Effects of α-amanitin
I Nucleolus 18S, 5.8S, and 28S rRNA Insensitive
II Nucleoplasm mRNA precursors and snRNA Strongly inhibited
III Nucleoplasm tRNA and 5S rRNA Inhibited by high concentrations
what does it mean to say that the 3 euk RNA polymerases are “homologous” to eo & to prok RNA polymerase?
they share a common evolutionary ancestor & have similar structure/subunit compositions
which RNA polymerase possesses a unique carboxyl-terminal domain (CTD), and what is the function of this domain? What is the repeating heptad seq in CTD?
RNA polymerase II has the CTD
function: regulate enzyme’s activity through the phosphorylation of Ser residues, acting as signal for diff stages of transcription
seq: YSPTSPS
what is unique about the location of RNA polymerase III promoter sequences compared to pol I & pol II?
while pol I & pol II have promoters located upstream of or at start site, RNA pol III promoters are located downstream of start site, within the transcribed sequence itself
which euk RNA polymerase relies on a set of consensus sequences specifically to recruit transcription factors to the start site?
RNA polymerase II (eg uses TATA box)
Name the two key promoter elements used by RNA polymerase I to initiate transcription
ribosomal initiator element (rlnr)
upstream promoter element (UPE)
where are RNA polymerase II promoters located and what are they also called?
located on the 5’ side of the start site for transcription
AKA cis-acting elements bc they are on the same mc of DNA as the transcribed genes
where is the TATA box found? and what does it do? what happens if there’s no TATA box instead?
found upstream of promoter region between -30 and -100
helps position RNA pol for initiation
a downstream promoter element(DPE) could be present instead
define initiator element (Inr) and downstream core promoter element (DPE)
Inr: found at transcriptional start site, between -3 and +5 / defines start site
DPE: found downstream of start site, between positions +28 & +32
commonly found in conjunction w/ Inr in transcripts that lack TATA box
where are the CAAT and GC boxes typically located, and what is their general purpose?
upstream of start site, between -40 and -150
purpose: act as regulatory sequences that influence the frequency or efficiency of transcription
what are “constitutive genes,” and which specific promoter element is frequently associated with them?
housekeeping genes that are continuously expressed at a steady rate
frequently have GC boxes in their promoters
what are transcription factors? what are they for RNA polymerase II?
TF = proteins that bind to these cis-acting elements to regulate gene expression
TF for RNA pol II: TFII —> TFIIA, TFIIB, etc
what binds to the TATA box in TATA box promoters and what is its purpose?
the TATA-box-binding protein (TBP), which is a component of TFIID
binds to minor groove of TATA box to cause large conformational changes in bound DNA
shaped like a saddle, allowing it to bind to DNA & provide docking sites for other transcription factors to assembe
what is the pre-initiation complex (PIC), and what are its primary components?
PIC: assembly of proteins required to start transcription in euk
includes RNA pol II, and the general transcription factors (GTF): TFIIA, TFIIB, TFIIF, TFIIE, TFIIH
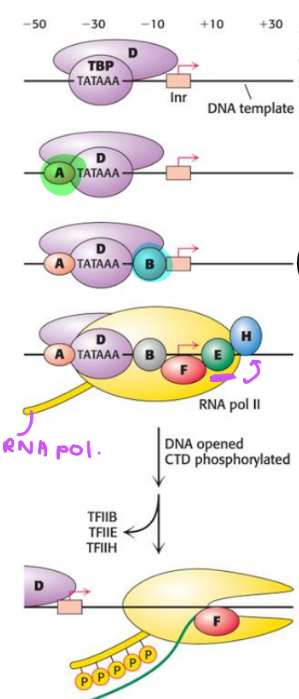
describe the roles of the following transcription factors for RNA pol II
TFIID, TFIIA, TFIB, TFIIF, TFIIE, TFIIH
TFIID: recognize core promoter elements & central to assembly process
TFIIA: aid in binding of TFIID to DNA
TFIIB: DNA-binding protein that recognizes B recognition elements near TATA box
TFIIF: aids in recruitment of polymerase II
TFIIE: brings TFIIH to complex
TFIIH: has helicase activity that unwinds DNA & kinase activity that phosphorylates CTD of polymerase II, facilitating transition to elongation
what are 2 major differences between an enhancer & a promoter?
distance: promoters are right at start site, while enhancers can be thousands of bases away (upstream, downstream, in middle of transcribed genes)
activity: promoters are required to start transcription, but enhancers have no promoter activity themselves — only stimulate existing promoter activity
enhancer is only effective in certain cells
how can an enhancer sequence function if it is located thousands of base pairs away from the gene’s start site?
DNA loops around physically, bringing distant enhancer into close proximity w/ promoter & transcription machinery (like PIC)